Abstract
Despite their often high conservation status and international obligations to undertake regular population surveillance, the status of pine marten (Martes martes) in many countries remains poorly understood. We conducted a non-invasive survey to determine the abundance of pine marten in forested habitat located within a 360 km2 mountainous region of Northern Ireland. We deployed 126 hair tubes between June and November during which 4 sampling sessions occurred, each lasting approximately 10 days. Hair samples were collected and analysed using genetic techniques to confirm species identity, sex and provide individual identity profiles. Genotyping success rates increased with the number of hairs in a sample and were significantly greater with sample sizes of ≥10 hairs (χ 2 = 15.1, df 1, P < 0.005). Abundance estimates for the adult breeding population were 23 individuals (95 % CI 15–31). Sex ratio data were 61 % male and 39 % female, suggesting that 14 males and 9 females were present in the breeding population. The total population abundance of pine marten, including breeding adults and annual juvenile recruitment, was estimated at 32 (95 % CI 31–35). Mean pine marten density was 0.53 per km2 of forest habitat and was within the range of densities found for the species in Europe. A breeding population in the low 20’s was considered small, and further research is needed on factors that may impact on the population including habitat management, connectivity and mortality. Non-invasive surveys provided conservation relevant data within a relatively short temporal period.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The requirement to determine the conservation status of a species is often driven by government agency mandates to assess populations through various regulatory mechanisms and it can be challenging, particularly in relation to methodology and logistic constraints (Witmer 2005). Basic parameters of interest can include animal abundance, density, distribution and response(s) to specific management interventions. For medium-sized carnivores such as pine marten (Martes martes) that can be rare, elusive and have fragmented distributions, estimating abundance can be difficult. Traditionally, abundance estimates for members of the genus Martes have been assessed using basic indices per unit length of transect or unit area based on scats or tracks (Lockie 1964; Kurki et al. 1998; Baines et al. 2004; Birks et al. 2005); historical trend data derived from hunting/trapping records (Helldin 2000); and live-trapping (Miller et al. 1955; Chapin et al. 1998). Generally, capture rates and probabilities can be low leading to problematic population estimation (Otis et al. 1978; White et al. 1982).
More recently, wildlife abundance estimation has been advanced by the development of non-invasive, DNA-based survey techniques. DNA is extracted from remotely collected samples and tested using microsatellite analysis to identify individuals (Mowat and Strobeck 2000). Non-invasive survey methods are increasingly being applied to pine marten populations (Balestrieri et al. 2010; Mullins et al. 2010; Ruiz-González et al. 2013; Sheehy et al. 2014). Primarily, these studies involve obtaining hair or scat samples, creating ‘capture histories’ for individuals based on microsatellite analysis and then applying a range of statistical techniques based primarily on capture–mark–recapture concepts (Poole et al. 2001; Boulanger et al. 2004; Frantz et al. 2004). This framework has recently been extended to the use of camera traps to estimate pine marten population density (Manzo et al. 2012).
Whilst non-invasive techniques offer exciting opportunities for wildlife studies in terms of costs, logistics and reduced disturbance to populations of interest, they are not a panacea having potential drawbacks that include shadow effects and genotyping errors that could cause negative bias in estimation (Mills et al. 2000; Lampa et al. 2013). There are also issues related to sampling design, study area delineation and accounting for the ecology of the target species (Lukacs and Burnham 2005; Kelly et al. 2012). Despite these potential drawbacks, non-invasive genetic sampling techniques offer an increasingly cost effective monitoring strategy for wildlife populations as they can provide a range of data on animal distribution, individual identity, population parameters, landscape genetics and occupancy, which are of paramount importance for conservation management (Mullins et al. 2010; Wasserman et al. 2010; Pauli et al. 2011; O’Mahony et al. 2012).
Hair samples, in particular, may offer an especially valuable tool for abundance estimation in non-invasive surveys as they typically yield higher quality DNA (Kelly et al. 2012) than other sources. Hair sampling devices can be placed to optimise survey design and can be used by inexperienced surveyors. Samples can be fresh and hair samples are simple to handle in terms of health and safety. This is not always the case with scats that are often opportunistically obtained, although recent advances in molecular techniques in relation to scats make them an increasingly valuable resource for species management (Ruiz-González et al. 2013).
After major reductions in range and distribution during the nineteenth and twentieth centuries (O’Sullivan 1983), over recent decades pine marten distribution has been increasing in Ireland in response to increased afforestation rates and reduced persecution (O’Mahony et al. 2012).
Preliminary population abundance estimates for Ireland in the period 2005–2007 suggest an overall breeding population of approximately 3,000 individuals (O’Mahony et al. 2012).
Pine marten are a species of high conservation interest in Ireland and receive full statutory protection at all times of the year. The population is in a phase of natural range expansion and may face increasing human–wildlife conflict scenarios into the future, however, in the historical context, current pine marten distribution remains diminished (O’Mahony et al. 2012). Whilst data on pine marten density is available from much of Europe (see Zalewski and Jędrzejewski 2006), apart from a few recent studies (Mullins et al. 2010; Sheehy et al. 2014), there is generally a lack of Irish data on pine marten abundance at a local and regional level, which is an important knowledge gap in terms of species management.
In the current study, we used non-invasive genetic sampling techniques coupled with abundance estimation statistics to provide estimates of the population size and density of pine marten in commercial forest plantations in a mountainous region of Northern Ireland. We also provide data on average movement distances of pine marten between inter-session centres of activity. To our knowledge, this study represents the largest scale deployment of non-invasive hair snagging devices to estimate pine marten abundance across the species entire geographic range.
Materials and methods
Study area
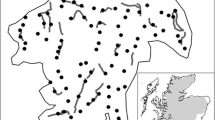
The Mourne Mountains are located in the southeast of Northern Ireland (Fig. 1). The study area was approximately 360 km2 in size with altitude ranging from 0 to 852 m. The region has a largely temperate climate with average annual rainfall from 1,300 to 2,000 mm and temperature ranges from 0 to 25 °C.
Forest habitat (3,350 ha) occurred in several discrete management units that ranged in size from 56 to 1,325 ha (Fig. 1). The majority of forest land cover was either owned or managed by the Forest Service of Northern Ireland with conifer (1,800 ha), mixed conifer (500 ha) and mixed conifer/broadleaf (150 ha) stands occupying the largest land area (approximately 72 % of forest area). There were also significant areas of open moorland habitat within forests (approximately 900 ha; 26 % of area), which was also the main habitat available between forest units. This consisted mainly of tussock grasses (Molinia spp), some heather (Calluna spp.) and rushes (Juncus spp.). There was little habitat connectivity between forest management units as hedgerows were rare; stone walls were the only linear boundary feature, where such features existed.
It was known that pine marten occurred in the Mourne Mountains and that the population was located approximately 35 km from pine marten core area (O’Mahony et al. 2012). The species was thought to have been extirpated in the Mournes by 1870, but individuals were live-trapped in the late 1950s (Rogers 1959) suggesting a possible mid-twentieth century re-colonisation event(s), although it could represent a relict population.
Non-invasive surveys
We deployed hair tubes, which consisted of 110 mm width drainpipe cut into 400 mm sections, throughout forest habitat in a systematic grid pattern (Fig. 1). Hair tubes were secured to trees at breast height at approximate spacing intervals of 500 m. This distance represented the approximate average home-range diameter of pine marten in Irish conifer forests (O’Mahony 2014). Two sticky patches were attached to the bottom of each hair tube to allow sample collection and the bait was placed internally, approximately 100 mm above patches. In total, 126 hair tubes were deployed throughout the duration of the study, an approximate density of 1 tube per 26 ha of forest habitat.
Hair tubes were deployed continuously between June and November 2011, during which four sampling sessions occurred, with each session 4–6 weeks apart (Table 1). During each sampling session, tubes were baited with chicken and checked approximately 10 days later, with all patches removed and stored at −30 °C until subsequent DNA analysis of hairs. Between sampling sessions hair tubes were left in situ without bait.
Hair tubes were potentially available for use by multiple individuals during sampling sessions as tubes did not have a closing mechanism after first use. In the current study, mixed genotypes were not found (see the ‘Results’ section) indicating that multiple individuals did not enter tubes. Multiple entries by different individuals have not been reported from other studies that have used the same, or similar, sampling device as in the current study (e.g. Mullins et al. 2010; Sheehy et al. 2014). Observations from camera traps show that pine marten do not re-enter tubes once bait has been removed indicating a possible behavioural response that may act as a proxy for a tube closure mechanism.
DNA analysis
Genomic DNA was isolated from hair samples using the ZR Genomic DNA™-Tissue MicroPrep (D3041) kit (ZYMO Research, CA, USA) using the protocol for hair DNA extraction (D3040). Real-time quantitative polymerase chain reaction (qPCR) assays for species and sex identification were carried out as described in Mullins et al. (2010). Two PCR replicates were carried out for molecular sexing (Lynch and Brown 2006). Females were identified through the amplification of ZFX only, while a signal from both ZFX and ZFY probes indicated male DNA was amplified. The ZFX allele, therefore, acted as an internal amplification control for the assay. Samples were selected for genotyping based on the results of the qPCR sex-typing assay with samples with a ZFX cycle threshold (CT) value <34 selected for genotyping.
Microsatellite analysis to identify individual pine marten was carried out using 9 microsatellite markers. These were: Gg7; Ggu234; Ma2; Ma8; Mel1; Mer041; Mvi1341, Mvi1354 and Mvis075 (see Mullins et al. 2010). Each sample was analysed in duplicate and only samples giving identical results in the replicates were scored. PCR reactions were carried out in two multiplex reactions. Multiplex 1 contained Gg7, Mer041, Mvi1341, Mvi1354 and Mvis075 primers and Multiplex 2 contained Ggu234, Ma2, Ma8 and Mel1 primers. Each PCR reaction contained 5 μl GoTaq Hotstart master mix (Promega Inc.), 1 μl multiplex primer mix (200 nM each primer) and 4 μl DNA extract. The PCR protocol was 95 °C initial denaturation for 5 min, followed by 40 cycles of 94 °C for 30 s, 58 °C for 1 min and 72 °C for 30 s, with a final extension time of 30 min at 72 °C. Fragment analysis was carried out on an ABI PRISM 310 genetic analyser under standard run conditions with 4 % polyacrylamide. Alleles were scored against a GS500 LIZ™ size standard using GeneMapper software version 3.7 (Applied Biosystems). Genotype data were analysed for probability of identity (PI and PIsibs), observed (Ho) and expected (He) heterozygosity and allele frequencies using GENALEX version 6 (Peakall and Smouse 2006). As only identical replicate genotypes were accepted no false alleles or allele drop-out were observed.
Microsatellite genotyping success rates were assessed for hair sample sizes between 1–5, 6–9 and ≥10. Genotyping success rates for individual samples that had ≤9 hairs and ≥10 hairs were analysed using a chi-square analysis (with Yates correction factor) test.
Abundance and density estimation
Dependent on the estimation method used, data were used to determine either (a) a pine marten breeding population estimate (data from sampling sessions 1 and 2, i.e. June to August) and/or (b) a post-breeding population estimate (data from sampling sessions 3 and 4, i.e. September to November), which included juvenile recruitment that year. Juveniles would have been unlikely to be detected in hair tubes during June–August as they tend to remain close to natal den sites (Helldin and Lindström 1995). Minimum number alive (MNA) estimates were calculated for the breeding and post-breeding pine marten population and represented the minimum and cumulative number of unique individuals known to be alive in each sampling period. MNA estimates were also used to identify forests where breeding was likely to have occurred as inferred by the presence of males and females within forest units during June–August sampling sessions. MNA estimates are particularly useful for rare and cryptic species in determining if multiple individuals are present within an area (Kendall and McKelvey 2008).
Capture-mark-recapture analysis was undertaken on microsatellite derived individual pine marten identity data obtained from non-invasive surveys, using CAPWIRE (Miller et al. 2005) and Chapman’s modified Lincoln–Petersen estimator (Chapman 1951). CAPWIRE allowed for sampling with replacement, which is approximated in DNA-based mark recapture studies as animals can visit multiple locations within a survey session (Miller et al. 2005). A likelihood-ratio test between two competing models (even capture model or two innate rates model) of capture frequency was used to choose the most appropriate model for abundance estimation. Confidence intervals were estimated using a parametric bootstrap procedure. This analysis was based on grouping all data into a single sampling session as each individual sampling session and sampling period (i.e. breeding and post-breeding) had relatively low counts of individual pine marten (≤20), which can lead to inaccurate estimates that have high confidence intervals (Miller et al. 2005). Therefore, to avoid biased estimates of low precision, CAPWIRE was applied to all multisession data only and produced a total abundance estimate combining adult and juvenile individuals identified during the study period. The Chapman modification of the Lincoln–Petersen estimator is a two-sample closed population model and was applied to provide breeding and post-breeding population estimates of the pine marten population, with 95 % confidence intervals. The assumption of population closure was likely met given the short temporal period between sampling sessions for breeding and post-breeding estimates (i.e. 4–5 weeks).
To estimate pine marten density a maximum likelihood approach using a spatially explicit capture–recapture (SECR) model, which incorporate animal capture location information to develop individual centre of activities, was implemented in DENSITY 4.4 (Efford 2004). For SECR analysis, the detector type chosen was ‘proximity’ to allow for multiple individuals to be captured at the same location; a Poisson distribution of home range centres was specified; and probability density functions were modelled using half-normal and hazard detection functions with a constant model for capture probability (g0) and spatial scale parameter (σ). Overall model selection was based on the lowest Akaike Information Criterion value, corrected for small samples sizes (AICc). A habitat mask that included only forested habitat within the study area was used with a specified buffer width of 1,000 m (approximately two home range centres (σ); from O’Mahony 2014). Data from each sampling session were used to produce within session density estimates, and an overall mean density estimate across all sessions was also determined. Density was expressed as number of pine marten per km2 of forest habitat.
Individual movement analysis
Average individual pine marten movement distances were calculated between consecutive pairs of sampling sessions (i.e. 1–2, 2–3 and 3–4), hereafter, termed inter-session. This analysis only included individuals that were ‘recaptured’ during at least two inter-session periods (n = 11). A mean centre of activity (i.e. point location) was determined for each individual pine marten within each sampling session, using the mean centre of spatial locations of hair tubes visited by the same individual determined in ArcMap 10 (ESRI Systems, USA). Inter-session straight line movement distances for individual pine marten were calculated using ArcMap 10. Inter-session movement distance analysis provided preliminary data on the minimum distance travelled over the time elapsed between sampling sessions and was used to assess home-range stability.
Results
In total, 228 hair samples were obtained from hair tubes of which 96 % were identified as pine marten using qPCR (Table 1). Other species that visited tubes were either rat (Rattus norvegicus) or domestic dog (Canis familiaris). The percentage of hair tubes with hair present increased over the duration of the study (Fig. 2). Based on CT values a total of 123 hair samples were analysed, with 110 giving full genotypes at the 9 microsatellite markers (success rate 48.2 %). All 9 microsatellite loci were polymorphic with a maximum of 4 alleles (Gg7). The PI using all nine loci was PI = 1.0 × 10−4 and PIsibs = 0.012. Only one loci, Mer041, showed a significant deviation from Hardy–Weinberg expectations (P = 0.033) although this may not be significant due to the small sample size. Genotyping success rates increased from 26.7, 42.1 and 63.5 %, respectively, for hair sample sizes of 1–5, 6–9 and ≥10 hairs. Significantly more genotype data were determined from hair samples with ≥10 hairs (χ 2 = 15.1, df 1, P < 0.005).
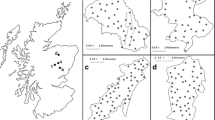
Minimum number alive estimates for the breeding population were 18 unique individuals (11 male and 7 female) and for post-breeding were 13 unique animals (6 male and 7 female), with a cumulative of 31 individual genotypes identified throughout the study (Fig. 3). The maximum number of detections for an individual pine marten across all sampling sessions was 18, whereas 10 individuals were detected only once during the survey period. Data from sampling session 1 and 2 indicated that breeding may have occurred in 3 of the larger (>500 ha) forest units (Castlewellan, Rostrevor and Tollymore; see Fig. 1) only. Automatic model selection in CAPWIRE selected the TIRM model to estimate a total population abundance estimate of 32 (95 % CI 31–35) pine marten within the study area. The Lincoln–Petersen model was used to estimate the breeding and post-breeding population, which were 23 (95 % CI 15–31) and 29 individuals (95 % CI 22–37), respectively. Applying sex ratio data to Lincoln–Petersen estimates for the breeding population indicated that there were 14 males (61 %) and 9 females (39 %) present. Pine marten mean density estimates across sampling session ranged from 0.40 to 0.70 marten per km2 of forest habitat, with a mean density of 0.53 (Table 2). Inter-session movement distances by individual pine marten ranged from 0 to 6,324 m (mean = 881 ± 303 m), with mean inter-session movement distances across all individuals between sampling sessions less than 1,100 m.
Discussion
We deployed a systematic non-invasive survey to a previously unstudied population of pine marten and obtained data that suggested the population had a low abundance, low density and that there were short-range movements between pine marten inter-session centre of activities, indicating general home range stability. As all available forest habitats within the study area were sampled, the research functioned as a population census and indicated that a small breeding population of 23 pine marten existed, with concordance across estimation methods, of which only 9 may have been female. Preliminary data also suggested that breeding may only have occurred within 3 forest units within the study area. Small and isolated populations are more susceptible to stochastic events (Shaffer 1981) and genetic considerations such as inbreeding that could lead to local extinction events. The available evidence suggests that the pine marten population in the study area relied on a very small number of breeding individuals, particularly females.
Breeding pine marten density estimates have been reviewed throughout eastern and northern Europe and can range from 0.01 to 1.75 per km2 with strong biogeographic limiting factors found to be winter severity and prey availability (Zalewski and Jędrzejewski 2006). Pine marten density estimates obtained from live-trapping and non-invasive studies vary from 0.36 to 0.76 per km2 in Poland (Zalewski and Jędrzejewski 2006), 0.46 to 4.42 per km2 in Ireland (Lynch et al. 2006; Mullins et al. 2010; O’Mahony 2014; Sheehy et al. 2014) and 0.34 per km2 in Italy (Manzo et al. 2012). Although differences in methodological and data analysis procedures constrain comparisons between studies, there is clear variation across even small geographic scales that could be influenced by habitat type, prey availability and levels of fragmentation.
In the current study, pine marten density was at the lower scale of that which has been previously reported for Ireland. However, this study took place at the largest geographical scale, in primarily commercial conifer forests encompassing a complete census of all available forest units within the landscape. Other Irish studies have focussed on either a single site (Lynch et al. 2006) or several relatively small sites (60 to 250 ha) consisting of deciduous or mixed conifer habitat, within much more highly fragmented landscapes than the current study (Mullins et al. 2010; Sheehy et al. 2014). The highest densities reported in Ireland have been achieved in deciduous woodland (Sheehy et al. 2014), which is a very rare habitat due to previous deforestation events. In such deciduous woodlands, pine marten can reach high densities and occupy small home ranges (Zalewski and Jędrzejewski 2006), which can be exacerbated in fragmented habitat (Mergey et al. 2011; Caryl et al. 2012).
Additionally, in deciduous habitat, pine marten have a wide dietary niche breadth (Lynch and McCann 2007). A mixture of differences in habitat types, spatial scales of investigation, food resource availability, density estimation methods and Ireland’s lack of seasonality, including mild winters (Sheehy et al. 2014), may influence the wide variation in estimated densities. Our estimates may be more representative of the range of densities pine marten can achieve in multi-scale commercial conifer plantations, which are the dominant habitat resources for the species throughout Ireland.
Within the study region, density estimates were derived only for forest land cover. There was little habitat connectivity in terms of hedgerows or riparian woodland between forest units, with open moorland habitat only available. As the habitat outside forest was more similar to inhospitable matrices (i.e. moorland) suggested by Caryl et al. (2012), rather than the fragmented yet highly connected habitats in Mergey et al. (2011), density estimates were unlikely to have been biased.
Implementing research and monitoring programmes for pine marten is a priority for the species across their range to address acknowledged gaps in the management of the species (Proulx et al. 2005). Non-invasive methodologies may provide a cost effect mechanism through which data can be obtained on species. In the current study, there was a clear increase in the number of tubes with hair samples as the study progressed. This could have been related to several factors including pine marten showing a positive response to tubes as a food reward was offered (an equivalent of individuals that become ‘trap happy’ in live-capture studies), increased marten abundance through juvenile recruitment over the study period or it could have reflected natural changes in the behaviour and spatial ecology of pine marten according to time of year (Zalewski and Jędrzejewski 2006; O’Mahony 2014).
In terms of the deployment of the method, a relatively large number of hair samples in the current study did not provide individual identity information due to poor quality DNA, with larger samples of ≥10 hairs yielding improved results. In similar studies carried out on carnivores, genotyping success rates using hair samples can vary from 45.5 to 76.6 % (Mowat and Paetkau 2002; Balestrieri et al. 2010; Kopatz et al. 2012; Davoli et al. 2013; Sheehy et al. 2014), which are generally higher than for scats (43 to 56 %; Frantz et al. 2006; Vergara et al. 2014). Hair sample DNA quality can be influenced by many factors including moisture, UV radiation, temperature and age (Piggott 2004). Practically, in terms of designing non-invasive sampling devices, all of these potential threats to DNA quality should be addressed and snagging mechanisms should preferentially collect large samples. The PIsib value (0.012) was found to be slightly above the recommended <0.01 threshold necessary to prevent the shadow effect. In future studies, the loci number used should be increased to potentially avoid the presence of different individuals with the same multilocus genotype. In the current study, a 10-day interval between tube placement and sampling collection was necessary in logistical terms. This could have led to some degradation of DNA and also provided for unsuitable weather conditions that may have affected the effectiveness of the sticky patches in collecting samples. Hair samples should preferentially be collected within short intervals, but not so short that they could impact on hair capture success rates (Mowat and Paetkau 2002).
Within a relatively short period of time, the current study has provided conservation relevant data on this elusive carnivore from a landscape that included difficult survey terrain. The method has potential as a national surveillance technique in that not only does it provide basic data on presence but also key demographic and population structure data that is critical for conservation management. Given that our study indicated low population abundance for pine marten in the Mourne Mountains, it is important to consider local management issues that may impact on the population. These include road traffic collisions, habitat fragmentation, population connectivity, potential illegal persecution and forest management. Perhaps of the most immediate interest is forest management as recent forest disease outbreaks (Phytophthora ramorum and Phytophthora lateralis) have led to an increase in clear felling rates, along with normal commercial harvesting operations. Forest disease management strategies may influence habitat quality, abundance and viability of red squirrel populations (Shuttleworth et al. 2012) and such concerns are equally relevant to forest dwelling species such as pine marten.
References
Baines D, Moss R, Dugan D (2004) Capercaillie breeding success in relation to forest habitat and predator abundance. J Appl Ecol 41:59–71
Balestrieri A, Remonti L, Frantz AC, Capelli E, Zenato M, Dettori EE, Guidali F, Prigioni C (2010) Efficacy of passive hair-traps for the genetic sampling of a low-density badger population. Hystrix 21:137–146
Birks J, Messenger J, Braithwaite T, Davison A, Brookes R, Strachan C (2005) Are Scat Surveys a Reliable Method for Assessing Distribution and Population Status of Pine Martens? In: Harrison DJ, Fuller AK, Proulx G (eds) Martens and fishers (Martes) in human-altered environments. Springer, USA, pp 235–252
Boulanger J, Himmer S, Swan C (2004) Monitoring of grizzly bear population trends and demography using DNA mark-recapture methods in the Owikeno Lake area of British Columbia. Can J Zool 82:1267–1277
Caryl FM, Quine CP, Park KJ (2012) Martens in the matrix: the importance of nonforested habitats for forest carnivores in fragmented landscapes. J Mammal 93:464–474
Chapin TG, Harrison DJ, Katnik DD (1998) Influence of landscape pattern on habitat use by American marten in an industrial forest. Conserv Biol 12:1327–1337
Chapman DG (1951) Some properties of the hypergeometric distribution with applications to zoological sample censuses. Univ Calif Publ Stat 1:131–160
Davoli F, Schmidt K, Kowalczyk R, Randi E (2013) Hair snaring and molecular genetic identification for reconstructing the spatial structure of Eurasian lynx populations. Mamm Biol 78:118–126
Efford M (2004) Density estimation in live-trapping studies. Oikos 106:598–610
Frantz AC, Schaul M, Pope LC, Fack F, Schley L, Muller CP, Roper TJ (2004) Estimating population size by genotyping remotely plucked hair: the Eurasian badger. J Appl Ecol 41:985–995
Frantz AC, Fack F, Muller CP, Roper TJ (2006) Faecal DNA typing as a tool for investigating territorial behaviour of badgers (Meles meles). Eur J Wildl Res 52:138–141
Helldin J (2000) Population trends and harvest management of pine marten Martes martes in Scandinavia. Wildl Biol 6:111–120
Helldin J, Lindström ER (1995) Late winter social activity in pine marten (Martes martes)—false heat or dispersal? Ann Zool Fenn 32:145–149
Kelly MJ, Betsch J, Wultsch C, Mesa B, Mills LS (2012) Noninvasive sampling for carnivores. In: Boitani L, Powell RA (eds) Carnivore ecology and conservation: a handbook of techniques. Oxford University Press, New York, pp 47–69
Kendall KC, McKelvey KS (2008) Hair collection. In: Long RA, Mackay P, Zielinski WJ, Ray JC (eds) Noninvasive survey methods for carnivores. Island Press, Washington DC, pp 141–182
Kopatz A, Eiken H, Hagen S, Ruokonen M, Esparza-Salas R, Schregel J, Kojola I, Smith M, Wartiainen I, Aspholm P, Wikan S, Rykov A, Makarova O, Polikarpova N, Tirronen K, Danilov P, Aspi J (2012) Connectivity and population subdivision at the fringe of a large brown bear (Ursus arctos) population in North Western Europe. Conserv Genet 13:681–692
Kurki S, Nikula A, Helle P, Linden H (1998) Abundances of red fox and pine marten in relation to the composition of boreal forest landscapes. J Anim Ecol 67:874–886
Lampa S, Henle K, Klenke R, Hoehn M, Gruber B (2013) How to overcome genotyping errors in non-invasive genetic mark-recapture population size estimation—a review of available methods illustrated by a case study. J Wild Manag 77:1490–1511
Lockie J (1964) Distribution and fluctuations of the pine marten, Martes martes (L.), in Scotland. J Anim Ecol 33:349–356
Lukacs PM, Burnham KP (2005) Review of capture–recapture methods applicable to noninvasive genetic sampling. Mol Ecol 14:3909–3919
Lynch Á, Brown M (2006) Molecular sexing of pine marten (Martes martes): how many replicates. Mol Ecol 6:631–633
Lynch ÁB, McCann Y (2007) The diet of the pine marten (Martes martes) in Killarney National Park. Biol Environ 107B:67–76
Lynch Á, Brown M, Rochford J (2006) Fur snagging as a method of evaluating the presence and abundance of a small carnivore, the pine marten (Martes martes). J Zool 270:330–339
Manzo E, Bartolommei P, Rowcliffe JM, Cozzolino R (2012) Estimation of population density of European pine marten in central Italy using camera trapping. Acta Theriol 57:165–172
Mergey M, Helder R, Roeder JJ (2011) Effect of forest fragmentation on space-use patterns in the European pine marten (Martes martes). J Mammal 92:328–335
Miller R, Ritcey R, Edwards R (1955) Live-trapping marten in British Columbia. Murrelet 36:1–8
Miller CR, Joyce P, Waits LP (2005) A new method for estimating the size of small populations from genetic mark–recapture data. Mol Ecol 14:1991–2005
Mills LS, Citta JJ, Lair KP, Schwartz MK, Tallmon DA (2000) Estimating animal abundance using noninvasive DNA sampling: promise and pitfalls. Ecol Appl 10:283–294
Mowat G, Paetkau D (2002) Estimating marten Martes americana population size using hair capture and genetic tagging. Wildl Biol 8:201–209
Mowat G, Strobeck C (2000) Estimating population size of grizzly bears using hair capture, DNA profiling, and mark-recapture analysis. J Wild Manag 64:183–193
Mullins J, Statham MJ, Roche T, Turner PD, O’Reilly C (2010) Remotely plucked hair genotyping: a reliable and non-invasive method for censusing pine marten (Martes martes, L. 1758) populations. Eur J Wildl Res 56:443–453
O’Mahony DT (2014) Socio-spatial ecology of pine marten (Martes martes) in conifer forests, Ireland. Acta Theriol 59:251–256
O’Mahony D, O’Reilly C, Turner P (2012) Pine marten (Martes martes) distribution and abundance in Ireland: a cross-jurisdictional analysis using non-invasive genetic survey techniques. Mamm Biol 77:351–357
O'Sullivan P (1983) The distribution of the Pine marten (Martes martes) in the Republic of Ireland. Mamm Rev 13:39–44
Otis DL, Burnham KP, White GC, Anderson DR (1978) Statistical inference from capture data on closed animal populations. Wildl Monogr 62:1–135
Pauli JN, Whiteman JP, Marcot BG, McClean TM, Ben-David M (2011) DNA-based approach to aging martens (Martes americana and M. caurina). J Mammal 92:500–510
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol 6:288–295
Piggott MP (2004) Effect of sample age and season of collection on the reliability of microsatellite genotyping of faecal DNA. Wildl Res 31:485–493
Poole KG, Mowat G, Fear DA (2001) DNA-based population estimate for grizzly bears Ursus arctos in northeastern British Columbia, Canada. Wildl Biol 7:105–115
Proulx G, Aubry K, Birks JDS, Buskirk S, Fortin C, Frost H, Krohn W, Mayo L, Monakhov V, Payer D, Saeki M, Santos-Reis M, Weir R, Zielinski W (2005) World Distribution and Status of the Genus Martes in 2000. In: Harrison DJ, Fuller AK, Proulx G (eds) Martens and fishers (Martes) in human-altered environments. Springer, USA, pp 21–76
Rogers A (1959) Pine marten, Martes martes (L.) in Co. Down woods. Ir Nat J 13:42
Ruiz-González A, Madeira MJ, Randi E, Urra F, Gómez-Moliner BJ (2013) Non-invasive genetic sampling of sympatric marten species (Martes martes and Martes foina): assessing species and individual identification success rates on faecal DNA genotyping. Eur J Wildl Res 59:371–386
Shaffer ML (1981) Minimum population sizes for species conservation. Bioscience 31:131–134
Sheehy E, O’Meara DB, O’Reilly C, Smart A, Lawton C (2014) A non-invasive approach to determining pine marten abundance and predation. Eur J Wildl Res 60:223–236
Shuttleworth CM, Lurz PWW, Geddes N, Browne J (2012) Integrating red squirrel (Sciurus vulgaris) habitat requirements with the management of pathogenic tree disease in commercial forests in the UK. For Ecol Manag 279:167–175
Vergara M, Ruiz-González A, López de Luzuriaga J, Gómez-Moliner BJ (2014) Individual identification and distribution assessment of otters (Lutra lutra) through non-invasive genetic sampling: recovery of an endangered species in the Basque Country (Northern Spain). Mamm Biol 79:259–267
Wasserman TN, Cushman SA, Schwartz MK, Wallin DO (2010) Spatial scaling and multi-model inference in landscape genetics: Martes americana in northern Idaho. Landsc Ecol 25:1601–1612
White GC, Anderson DR, Burnham KP, Otis DL (1982) Capture-recapture and removal methods for sampling closed populations. Los Alamos Natl Lab USA 1–230
Witmer GW (2005) Wildlife population monitoring: some practical considerations. Wildl Res 32:259–263
Zalewski A, Jędrzejewski W (2006) Spatial organisation and dynamics of the pine marten Martes martes population in Białowieza Forest (E Poland) compared with other European woodlands. Ecography 29:31–43
Acknowledgments
This study was funded by a grant from the Peoples Trust for Endangered Species and the Northern Ireland Environment Agency. The Forest Service of Northern Ireland provided considerable logistical support to the project including a permit to work in their forests. Graham Cherry (AFBI) undertook substantial field work for the project and his contribution is gratefully acknowledged. Two anonymous referees improved the manuscript and their contribution is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Gortázar
Rights and permissions
About this article
Cite this article
O’Mahony, D.T., Turner, P. & O’Reilly, C. Pine marten (Martes martes) abundance in an insular mountainous region using non-invasive techniques. Eur J Wildl Res 61, 103–110 (2015). https://doi.org/10.1007/s10344-014-0878-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10344-014-0878-0