Abstract
The larvae of many hoverflies (Diptera: Syrphidae) are important polyphagous predators used in integrated pest management programs. Because the accurate identification of preimaginal stages by morphological characters is difficult, we have developed a multiplex PCR to identify the immature and/or adult stages of the most common syrphid species in Mediterranean vegetable crops: Episyrphus balteatus, Scaeva pyrastri, Eupeodes corollae, Meliscaeva auricollis, Sphaerophoria scripta, and Sphaerophoria rueppellii. The latter two species were amplified by the same primer pair due to the high similarity of their cytochrome oxidase subunit I sequences. Additionally, the assay included a primer pair targeting Diplazon laetatorius, a common koinobiont ichneumonid endoparasitoid of predatory syrphid larvae. The multiplex PCR assay proved to be highly specific and sensitive, and it was used to study the assemblage of hoverfly species in larval stage in two Mediterranean lettuce crops in two consecutive years. The molecular analysis revealed that Eu. corollae, Ep. balteatus, and Sph. scripta/Sph. rueppellii were the species present in the investigated fields. Species composition differed depending on sampling date and whether the larvae were collected on the plants or on the ground. The parasitoid D. laetatorius was not detected in any of the analyzed hoverfly larvae, suggesting low-parasitism pressure in the studied syrphid populations. The wide distribution of most of these syrphid species makes this multiplex PCR assay an ideal tool to deepen our knowledge on the ecology of these polyphagous hoverfly species in preimaginal stages and to improve the use of hoverflies to control insect pests.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hoverflies (Diptera: Syrphidae) are an abundant group of insects present in natural and agriculture related ecosystems. With about 750 species recorded in Europe (Speight 2011), at least 355 species are recorded from Spain (Marcos-García et al. 2002). Their adults provide crucial ecosystem services as important pollinators, obtaining their energy requirements by feeding on nectar and pollen (Haslett 1983; Branquart and Hemptinne 2000; Jauker et al. 2012). The larvae of about 35 % of the species of the family of syrphids are polyphagous predators of a broad range of soft-bodied arthropods, including coleopteran and lepidopteran larvae, mites, thrips, and hemipterans (e.g., coccids, psyllids, whiteflies and particularly aphids) being the preferred prey (Rojo et al. 2003; Rotheray and Gilbert 2011). Predatory larvae of many syrphid species hide under bark or underneath soil particles during the day and are mostly active at dawn and dusk. This behavior makes them less conspicuous than other natural enemies (Hagen et al. 1999). In Spain, 124 syrphid species with predaceous larvae have been reported by Marcos-García et al. (1998), most of them commonly found in the Mediterranean basin (Speight 2011). Some predatory hoverflies are abundant in different agroecosystems such as fruit orchards, woodlands, grasslands, scrublands, as well as in arable and vegetable crops (Ghahari et al. 2008; Haenke et al. 2009; Hopper et al. 2011). Less than 30 syrphid predaceous species had been related to herbaceous plants at the Iberian Peninsula (Rojo and Marcos-García 1998; Rojo et al. 2003). Six of these hoverfly species are commonly found in Mediterranean vegetable crops such as lettuce (Rojo 1995; Pascual-Villalobos et al. 2006; Morales et al. 2007): Episyrphus balteatus (De Geer), Scaeva pyrastri (Linnaeus), Eupeodes corollae (Fabricius), Meliscaeva auricollis (Meigen), Sphaerophoria scripta (Linnaeus), and Sphaerophoria rueppellii (Wiedemann).
Syrphid larvae may be attacked by a wide range of hymenopteran parasitoids belonging to the families Ichneumonidae, Encyrtidae, Pteromalidae, Megaspilidae, and Figitidae (Scott 1939; Rotheray and Gilbert 2011). However, the most common endoparasitoids of predatory species belong to the family Ichneumonidae and the subfamily Diplazontinae (Bordera et al. 2000, 2001). Particularly Diplazon laetatorius (Fabricius) has been reported as the most important natural enemy of hoverflies in terms of abundance of the taxon around the world (Greco 1997; Jankowska 2004). This species is a koinobiont endoparasitoid that oviposits into the syrphid eggs or first instars larvae, with the imago emerging from the syrphid puparium (Mayadunnage et al. 2009).
Although the larvae of many hoverfly species are important biocontrol agents (Rojo et al. 2003; Hopper et al. 2011), it is quite difficult to obtain accurate identification of preimaginal stages, particularly in the first larval stages, using exclusively morphological characters (Bastian 1986; Laska et al. 2006). Moreover, larvae of many species are unknown and the color pattern of common species it is not retained after preserving them in ethanol (Rotheray 1993). Rearing field-collected larvae to the adult stage is recommended for a correct morphological identification (Gilbert 1993), however, this is a time-consuming process which can be accompanied by a high mortality (Jankowska 2004). At the same time, it is also difficult to discriminate between parasitized and nonparasitized hoverfly larvae to assess how parasitoids may impact hoverfly larval populations and their biocontrol success (Hazell et al. 2005). Hence, an alternative technique is needed which allows identifying hoverflies in their larval stage and to detect parasitism by D. laetatorius.
To date, two DNA-based approaches are most widely used for species identification: (i) DNA barcoding (Hebert et al. 2003), where species-specific sequences are generated and identified via a reference database (e.g., Mengual et al. 2008; Stahls et al. 2009; Benefer et al. 2013), or (ii) diagnostic PCR where species-specific primers may be used either individually in one PCR amplifying just one target species (singleplex PCR) or simultaneously in a multiplex PCR which enables the parallel identification of several species (King et al. 2011; Staudacher et al. 2011). While the former approach can be limited by the sequence barcode information available in databases such as GenBank or Bold to identify the sequence, the latter technique, is particularly useful once species-specific primers have been developed and when large sample numbers have to be screened because it is cost-effective and quick. Results obtained by multiplex PCR are usually not corrupted by the presence of endoparasitoid DNA, which can be a problem when using the barcoding approach because the mixture of different sequences may foil species identification (Traugott et al. 2013). On the other hand, multiplex PCR can only identify those taxa for which primers have been developed, which means that this approach needs to be carefully checked for cross-reactivity to ensure accurate results.
The aims of this study were: (1) to design species-specific primers for the six most common hoverfly species found in Mediterranean vegetable crops, as well as the parasitoid D. laetatorius; (2) to embed these primers in a multiplex PCR assay to easily and rapidly identify these syrphid species including the detection of parasitoid DNA; and (3) to use this molecular tool to identify which of these hoverflies species are present in larval stage in two Mediterranean lettuce crops in two consecutive years as well as to assess the levels of parasitism by D. laetatorius.
Materials and methods
Insects
Twenty hoverfly species commonly present in European agricultural environments (Table 1) were used for designing species-specific primers targeting the most common hoverfly species found in Mediterranean vegetable crops: Ep. balteatus, Sc. pyrastri, Eu. corollae, M. auricollis, Sph. scripta, and Sph. rueppellii. These specimens were collected in several locations of Spain and Germany (Table 1).
Sequencing and primer design
A nondestructive DNA extraction method was used to avoid morphological damage to the adult syrphid samples (Staudacher et al. 2011), and a minimum of one adult specimen per species was sequenced. The adult hoverflies were incubated overnight at 58 °C with 180 μl of buffer ATL and 20 μl of Proteinase K (10 mg ml−1, AppliChem, Darmstadt, Germany). DNA was extracted from this solution using the DNeasy Tissue Kit (Qiagen, Hilden, Germany; protocol for animal tissues) following the manufacturer’s protocol and stored at −20 °C. One negative extraction control was included in each batch of 30 samples. All syrphids were amplified using the universal primers LC01490/HC02198 described in Folmer et al. (1994), obtaining fragments of the cytochrome c oxidase subunit I (COI) gene of approximately 700 bp in length. Each 10 μl PCR contained 1.5 μl of DNA extract, 5 μl of 2× Multiplex PCR Master Mix (Qiagen), 1 μM of each primer, and 1.5 μl of PCR-grade RNase-free water (Qiagen). Thermocycling was done using Mastercycler Gradient PCR machines (Eppendorf, Hamburg, Germany); the thermocycling program consisted of an initial denaturation step of 15 min at 95 °C, followed by 35 cycles of 20 s at 94 °C, 30 s at 52 °C, 45 s at 72 °C, and a 3 min final extension at 72 °C. PCR products were electrophoresed on 1.5 % agarose gels stained with GelRed™ (Biotium, Hayward, USA) and visualized under UV light. PCR products were purified with ExoSAP®-IT (GE Helthcare, Little Chalfont, UK) following the manufacturer’s recommendation and sequenced according to the dideoxychain-termination method. Sequences were aligned and edited manually using Bioedit Sequence Alignment Editor v. 7.0.9.0 (Hall 1999). The obtained sequences were submitted to GenBank database (see Table s1 for accession numbers). These sequences were also aligned with other sequences from the GenBank database (Table s1) using CLUSTALW2 (www.ebi.ac.uk/Tools/msa/clustalw2) and checked for species-specific primer-binding sites. All primer pairs (five for the six hoverfly target species and one for the parasitoid D. laetatorius) were designed using Primer Premier 5 (Premier Biosoft International, CA, USA).
Multiplex PCR and specificity assay
All field-collected larval syrphid specimens tested by multiplex PCR were also DNA extracted using the DNeasy Tissue Kit (QIAGEN; protocol for animal tissues). Total DNA was eluted in 100 μl of AE buffer provided by the manufacturer and stored at −20 °C. Two negative extraction controls were added to each set of 28 samples. Multiplex PCR was optimized testing different concentrations of primers and thermocycling conditions. The final reaction volumes (10 μl) contained 1.5 μl of DNA extract, 5 μl of 2× Multiplex PCR Master Mix (Qiagen), 1 μl of 10× primer mix, 1 μl of 5× Q-solution, and 1.5 μl of PCR-grade RNase-free water (Qiagen). Primer concentrations in the primer mix were different depending on the species (see Table 2). In a 2720 thermocycler (Applied Biosystems, CA, USA), the DNA extracts were subjected to 95 °C for 15 min, followed by 35 cycles of 94 °C for 30 s, 64 °C for 90 s, and 72 °C for 60 s and a final extension of 72 °C for 10 min. Target DNA and water were always included as positive and negative controls, respectively. PCR products were separated by electrophoresis in 3.6 % agarose gels stained with ethidium bromide and visualized under UV light.
Primer specificity was evaluated not only by testing the six target hoverfly species, but also the other 14 nontarget hoverfly species (1–4 individuals/species) used for primer design. Additionally, nine potential hoverfly prey species which are commonly found in Mediterranean lettuce crops, including aphids, thrips, and collembolans, as well as the hoverfly parasitoid D. laetatorius, were tested (3 individuals/species) (Table 1).
Different concentrations of D. laetatorius DNA were analyzed to characterize the sensitivity of the primer pair targeting the parasitoid. The initial DNA concentration tested with the multiplex PCR protocol described above was 0.4 ng/μl which was twofold diluted down to 2.5 pg/μl. DNA concentrations were measured in a Qubit Fluorometer (Invitrogen, CA, USA) using the Quant-iT™ dsDNA HS assay kit (Invitrogen).
Analysis of field-collected hoverfly larvae
Two lettuce fields (var. Maravilla) located in El Maresme area (Barcelona, Spain) were sampled. One was an experimental field at IRTA (41°31′4.33″N, 2°22′37.87″E) and the other one was a commercial field in 50-km distance (41°28′26.07″N, 1°57′34.52″E).
In the experimental field, two consecutive lettuce crops were planted: one from beginning of April until end of May and another from beginning of June until beginning of August, both in 2009 and 2010. Twenty to thirty lettuces were collected on May18th and 19th 2009; July 7th 2009; and May 11th, 18th, and 25th 2010. All lettuces were brought individually in plastic bags to the laboratory, where all syrphid larvae were collected. On May 12th 2009, the experimental field was also manually sampled once for syrphid larvae found on the ground. In the commercial field, also twenty to thirty lettuces were sampled once on April 22th 2009. All collected larvae were stored at −20 °C until molecular analyses.
All syrphid larvae were individually analyzed by multiplex PCR to study parasitism by D. laetatorius and the syrphid larval species composition depending on the sampled season (spring and summer), year (2009 and 2010) and substrate (lettuce or ground). Species percentages were calculated and compared in order to determine whether they were influenced by the season, year, substrate and sample location.
Results
Multiplex PCR and specificity assay
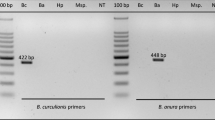
COI sequences of 21 hoverfly species were generated and submitted to GenBank (accession numbers are shown in Table s1). Six specific primer pairs were designed for the hoverflies Ep. balteatus, Eu. corollae, M. auricollis, Sc. pyrastri, Sph. scripta/Sph. rueppellii and the parasitoid D. laetatorius (Table 2). Sphaerophoria scripta and Sph. rueppellii were covered by one primer pair as their sequences were very similar (97.6 % sequence identity for a 570 bp long stretch of COI sequence). The hoverfly primers generated DNA fragments ranging from 96 to 754 bp depending on the species (Fig. 1; Table 2). The parasitoid D. laetatorius was also detected with the parasitoid primers, amplifying a specific 220 bp fragment. Detection of the parasitoid was possible down to a DNA concentration of 0.4 pg/μl PCR.
DNA fragments obtained by multiplex PCR amplification using the specific syrphid- and parasitoid-specific primers. Lane 1 DNA size marker (50 bp ladder), L2: Meliscaeva auricollis (96 bp), L3 Sphaerophoria scripta (165 bp), L4 Sphaerophoria rueppellii (165 bp), L5 Scaeva pyrastri (314 bp), L6 Eupeodes corollae (395 bp), L7 Episyrphus balteatus (754 bp), L8 Diplazon laetatorius (220 bp), and L9 negative control
When these primers were tested in the multiplex PCR for cross-amplification against the other hoverfly species and potential prey of hoverfly larvae (Table 1), all nontarget samples were negative, demonstrating the specificity of the assay.
Analysis of field-collected hoverfly larvae
Diagnostic PCR allowed identifying 169 field-collected syrphid larvae from both fields and years. Only three taxa (Eu. corollae, Ep. balteatus and Sph. scripta/Sph. rueppellii) were found. Overall, Eu. corollae dominated the catches (74 % of all collected larvae), followed by Sph. scripta/Sph. rueppellii (14 %) and Ep. balteatus (12 %). On the lettuce plants in spring 2009, the species assemblage in the experimental field was very similar to that in the commercial field (Fig. 2). On the ground however, Eu. corollae was found almost exclusively when searching for hoverfly larvae on the soil surface. In spring 2010, only Eu. corollae and Ep. balteatus were captured while in summer 2009, Sph. scripta/Sph. rueppellii were the taxa with the highest representation, followed by Eu. corollae, whereas, Ep. balteatus was only occasionally found (Fig. 2). None of the syrphid larvae tested positive for DNA of the parasitoid D. laetatorius.
Discussion
The multiplex PCR assay developed in this study allows unambiguous identification of the five most common predatory hoverfly taxa present in Mediterranean vegetable crops. Moreover, the assay includes a primer pair for the parasitoid D. laetatorius, a common ichneumon-parasitoid of hoverfly larvae. A primer pair for the parasitoid has been included in the assay because parasitoid eggs and larvae are easily missed when inspecting the hoverfly larvae under a dissecting microscope which can lead to an underestimation of the real parasitism rate (Moreno-Ripoll et al. 2012). Compared to an identification of the larvae via a DNA barcode (Jinbo et al. 2011), the current approach has the advantage that whole body DNA extracts which might also contain DNA of prey and/or parasitoids can be tested. This nonsyrphid DNA would cause problems for DNA barcoding if general invertebrate/metazoan primers are used to generate the COI fragment used as the barcode DNA region. Using Sanger sequencing, sequence-based identification of one type of DNA in the sample is preferable. A mixture of syrphid, prey and/or parasitoid DNAs can lead to unreadable sequences or preferential amplification of parasitoid DNA (Lee and Lee 2012) and prohibit species identification. This could be avoided by using Next Generation Sequencing technologies, which have been also recently used to identify a wide range of insect prey items present in the gut of predaceous syrphid larvae (author's unpublished results).
When the designed primers were tested for specificity, none of the other syrphid species potentially present in Mediterranean vegetable crops nor any other potential prey species yielded false positives. The latter were tested because of the possibility of amplifying prey remains from the gut content of the hoverflies. The lack of amplification demonstrated that the PCR products were exclusive from the syrphid taxa. The assay developed here can be used to identify all developmental stages, and even parts or remains of the targeted species, which makes its possibility of application manifold. In the case of the primers that produce a band smaller than 400 bp [i. e., M. auricollis (96 bp), Sphaerophoria spp. (165 bp), Sc. pyrastri (314 bp), and Eu. corollae (395 bp)], they could also be used to test other predators for consumption of these hoverfly species.
When the multiplex PCR assay designed here was used to study the composition of hoverfly larvae communities in Mediterranean lettuce fields, only three syrphid taxa were found: Eu. corollae, Ep. balteatus, and Sph. scripta/Sph. rueppellii. Previous studies conducted also in lettuce crops in Spain confirm these results (Pascual-Villalobos et al. 2006; Morales et al. 2007), being also the main syrphid species found. Other species, such as M. auricollis have also been observed in lettuce crops in Spain, but in much less proportion (Rojo and Marcos-García 1998).
The multiplex PCR assay also detected temporal differences in the hoverfly species assemblages. Eupeodes corollae and Ep. balteatus were more abundant in spring whereas Sph. scripta/Sph. rueppellii densities peaked in summer. The same temporal pattern (Eu. corollae/Ep. balteatus/Sph. rueppellii) was found in a previous study on aphidophagous syrphid population dynamics in pepper greenhouses in the southeast Spain (Pineda and Marcos-García 2008). Eupeodes corollae, Ep. balteatus, and Sph. scripta are highly migratory species (Speight 2011) that move to Central Europe during summer and the mated females returning to South Europe in autumn (Rotheray and Gilbert 2011). On the other hand, Sph. rueppellii is a resident Mediterranean species which is well adapted to high-ambient temperatures (Pineda and Marcos-García 2008; Amorós-Jiménez et al. 2012). In relation with these biological traits, larvae of both species of the genus Sphaerophoria were found in Spanish lettuce crops during spring, but only Sph. scripta was found in autumn (Morales et al. 2007). For this reason, those syrphid larvae which were collected in summer 2009 in this study and which were assigned by the multiplex PCR approach to the two molecularly indistinguishable species Sph. scripta/Sph. rueppellii probably belong to Sph. rueppellii.
When analyzing the syrphid larvae collected on the ground, we did not find a complex of syrphid species like on the lettuce plants. Instead, Eu. corollae was the most abundant species. Episyrphus balteatus and Sph. scripta/Sph. rueppellii were hardly and not found on the ground, respectively. This behavior is also related with the preference of these hoverfly species (like most Syrphinae) to pupate on the plant on which their prey occur. However, according to Dusek and Laska (1961), Eu. corollae overwinters as pupa, which is unusual for aphidophagous hoverflies (Stubbs and Falk 1983).
From all syrphid larvae analyzed here, none was found to be parasitized by D. laetatorius, suggesting that the syrphid populations in the investigated fields did not experience top-down pressure by this endoparasitoid. Note, however, that the current result could also be explained by the comparably low number of syrphid larvae analyzed, as parasitism rates are usually not very high in hoverfly larvae. For example, in lettuce crops, Smith and Chaney (2007) found less than 5 % of parasitism by D. laetatorius after analyzing 1,087 syrphid larvae collected in Californian crops. Krawczyk et al. (2011) reported that 3 % of the syrphid pupae inspected (n = 538) were parasitized in maize fields in Poland, where the dominant syrphid parasitoid was Pachyneuron grande (Hymemoptera: Pteromalidae). In cabbage fields, also in Poland, parasitism by D. laetatorius was found as high as 22 % when 410 syrphid larvae and pupae were analyzed (Jankowska 2004). Lacking parasitoid DNA detection in diagnostic PCRs could also be ascribed to a low sensitivity of the assay (Traugott and Symondson 2008).The sensitivity of the current multiplex PCR for detecting parasitoid DNA, however, is highly comparable to previous assays which allowed detection of eggs and early instar larvae of parasitoids (e.g., Traugott et al. 2006). Therefore, we think that the current results are not due to a methodological artifact but represent a nonexisting/very low level of parasitism of these hoverfly larvae by D. laetatorius.
The multiplex PCR approach described here is an efficient tool for the rapid identification of the main hoverfly species present in Mediterranean vegetable crops. Because the larvae of these hoverfly species are known to be important predators of several insect pests, and the species studied in the present study have been identified in other agroecosystems (Jansen 2000; Marshall and West 2007; Sajjad et al. 2008) or forest ecosystems (Kehlmaier and Martínez de Murguía 2004), this molecular method will be particularly useful for further studies on population dynamics, distribution, and abundances of these syrphid species. A molecular tool for detecting D. laetatorius parasitism within syrphid larvae has also been described here, allowing to further examine which effect this parasitoid has on syrphid populations and their ability to control pest populations. A better understanding of the identity of the predators and their feeding activities would allow to better conserve key predators in conservation biological programs in vegetable crops.
References
Amorós-Jiménez R, Pineda A, Fereres A, Marcos-García MA (2012) Prey availability and abiotic requirements of immature stages of the aphid predator Sphaerophoria rueppellii. Biol Control 63:17–24
Bastian O (1986) Schwebfliegen (Syrphidae). Die Neue Brehm-Bücherei, Bd. 576. 168 S, 352 Abb (z T farb)—A Ziemsen Verlag, Wittenberg
Benefer CM, van Herk WG, Ellis JS, Blackshaw RP, Vernon RS, Knight ME (2013) The molecular identification and genetic diversity of economically important wireworm species (Coleoptera: Elateridae) in Canada. J Pest Sci 86:19–27
Bordera S, Agulló P, Rojo S (2000) Nuevos Diplazontinae (Hymenoptera, Ichneumonidae) para la entomofauna iberbalear y potenciales sírfidos hospedadores (Diptera, Syrphidae). Bol Asoc Esp Ent 24(1–2):131–139
Bordera S, Agulló P, Rojo S (2001) Catálogo de los Diplazontinae iberobaleares (Hymenoptera, Ichneumonidae) y potenciales sírfidos hospedadores (Diptera, Syrphidae). Bol Asoc Esp Ent 25(1–2):153–174
Branquart E, Hemptinne JL (2000) Selectivity in the exploitation of floral resources by hoverflies (Diptera: Syrphinae). Ecography 23:732–742
Dusek J, Laska P (1961) Beitragzur Kenntnis der Schwebfliegen-Larven III (Syrphidae, Diptera). Prirod cas slezsky 22:513–541
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3:294–299
Ghahari H, Hayat R, Tabari M, Ostovan H (2008) Hoverflies (Diptera: Syrphidae) from rice fields and around grasslands of norther Iran. Munis Entomol Zool 3:275–284
Gilbert FS (1993) Hoverflies. Richmond Publishing Co. Ltd., Slough
Greco CF (1997) Specificity and instar preference of Diplazon laetatorius (Hym.: Ichneumonidae) parasitizing aphidophagous syrphids (Dipt: Syrphidae). Entomophaga 42:315–318
Haenke S, Scheid B, Schaefer M, Tscharntke T, Thies C (2009) Increasing syrphid fly diversity and density in sown flower strips within simple vs. complex landscapes. J Appl Ecol 46:1106–1114
Hagen KS, Mills NJ, Gordh G, McMurtry JA (1999) Terrestrial arthropod predators of insect and mite pests. In: Bellows TS, Fisher TW (eds) Handbook of biological control. Academic Press, San Diego, pp 383–503
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Haslett JR (1983) A photographic account of pollen digestion by adult hoverflies. Physiol Entomol 8:167–171
Hazell SP, Wenlock C, Bachel S, Fellowes MDE (2005) The costs and consequences of parasitoid attack for the predatory hoverfly, Episyrphus balteatus. Evol Ecol Res 7:669–679
Hebert PDN, Cywinska A, Ball SL, DeWaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc A 270:313–321
Hopper JV, Nelson EH, Daane KM, Mills NJ (2011) Growth, development and consumption by four syrphid species associated with the lettuce aphid, Nasonovia ribisnigri, in California. Biol Control 58:271–276
Jankowska B (2004) Parasitoids of aphidophagous Syrphidae occurring in cabbage aphid (Brevicoryne brassicae L.) colonies on cabbage vegetables. J Plant Prot Res 44:299–305
Jansen JP (2000) A three-year field study on the short-term effects of insecticides used to control cereal aphids on plant-dwelling aphid predators in winter wheat. Pest Manag Sci 56:533–539
Jauker F, Bondarenko B, Becker HC, Steffan-Dewenter I (2012) Pollination efficiency of wild bees and hoverflies provided to oilseed rape. Agric Forest Entomol 14:81–87
Jinbo U, Kato T, Ito M (2011) Current progress in DNA barcoding and future implications for entomology. Entomol Sci 14:107–124
Kehlmaier C, Martínez de Murguía L (2004) Syrphidae recorded in a heterogenous forest ecosystem at the “Finca de Artikutza” (Navarre, Northern Spain) (Diptera). Fragm Entomol 36:253–274
King RA, Moreno-Ripoll R, Agustí N, Shayler SP, Bell JR, Bohan DA, Symondson WOC (2011) Multiplex reactions for the molecular detection of predation on pest and nonpest invertebrates in agroecosystems. Mol Ecol Resour 11:370–373
Krawczyk A, Hurej M, Jackowski J (2011) Syrphids and their parasitoids from maize crop. J Plant Prot Res 51:93–97
Laska P, Perez-Bañón C, Mazanek L, Rojo S, Ståhls G, Marcos-García MA, Bicik V, Dusek J (2006) Taxonomy of the genera Scaeva, Simosyrphus and Ischiodon (Diptera: Syrphidae): Descriptions of immature stages and status of taxa. Eur J Entomol 103:637–655
Lee W, Lee S (2012) Unexpected problem in aphid DNA barcoding by universal primers. Entomol Sci 15:121–126
Marcos-García MA, Isidro PM, Rojo S, Pérez-Bañón C (1998) Catálogo y distribución geográfica de las especies de Syrphidae (Diptera) en la Península Ibérica. I.—Syrphinae y Microdontinae. Bol Asoc Esp Entomol 22:37–61
Marcos-García MA, Rojo S, Pérez-Bañón C (2002) Family Syrphidae. In: Catálogo de los dípteros de España, Portugal y Andorra (Insecta), vol 8. Monografías SEA, Zaragoza, pp 132–136
Marshall EJP, West TM (2007) Impacts of field margins, landscape and crop on the distributions of Syrphidae on an arable farm. Asp Appl Biol 81:91–100
Mayadunnage S, Wijayagunasekara HNP, Hemachandra KS, Nugaliyadde L (2009) Occurrence of aphidophagous syrphids in apphid colonies on cabbage (Brassica oleracea) and their parasitoids. J Trop Agri Res 21:99–109
Mengual X, Stahls G, Rojo S (2008) First phylogeny of predatory flower flies (Diptera, Syrphidae, Syrphinae) using mitochondrial COI and nuclear 28S rRNA genes: conflict and congruence with the current tribal classification. Cladistics 24:543–562
Morales I, Díaz BM, Nebreda M, López-Lastra C, Goldarazena A, Sánchez JA, Pineda A, Marcos-García MA, Fereres A (2007) Principales agentes de biocontrol en cultivos de lechuga en la zona centro de España. Revista Horticultura 49:46–49
Moreno-Ripoll R, Agustí N, Berruezo R, Gabarra R (2012) Conspecific and heterospecific interactions between two omnivorous predators on tomato. Biol Control 62:189–196
Pascual-Villalobos MJ, La casa A, González A, Varo P, García MJ (2006) Effect of flowering plant strips on aphid and syrphid populations in lettuce. Eur J Agron 24:182–185
Pineda A, Marcos-García MA (2008) Seasonal abundance of aphidophagous hoverflies (Diptera : Syrphidae) and their population levels in and outside Mediterranean sweet pepper greenhouses. Ann Entomol Soc Am 101:384–391
Rojo S (1995) Biología de los sírfidos afidófagos (Diptera, Syrphidae), presentes en cultivos hortofrutícolas mediterráneos. Implicaciones en el control biológico de pulgones (Homoptera, Aphididae). PhD dissertation, University of Alicante, Alicante, Spain
Rojo S, Marcos-García MA (1998) Catálogo de los sírfidos (Diptera, Syrphidae) afidófagos (Homoptera, Aphididae) presentes en cultivos y plantas herbáceas de España y Portugal. Boll Zool Agrar Bachic 30:39–54
Rojo S, Gilbert FS, Marcos-García MA, Nieto JM, Mier MP (2003) A world review of predatory hoverflies (Diptera, Syrphidae, Syrphinae) and their prey. CIBIO (Centro Iberoamericano de la Biodiversidad) ediciones, Alicante
Rotheray GE (1993) Colour guide to hoverfly larvae (Diptera, Syrphidae) in Britain and Europe. Dipterists Digest 9:156
Rotheray GE, Gilbert F (2011) The natural history of hoverflies. Forrest, Tresaith
Sajjad A, Saeed S, Masood A (2008) Pollinator community of onion (Allium cepa L.) and its role in crop reproductive success. Pak J Zool 40:451–456
Scott EI (1939) An account of the developmental stages of some aphidophagous Syrphidae (Dipt) and their parasites (Hymenopt). Ann Appl Biol 26:509–532
Smith HA, Chaney WE (2007) A survey of syrphid predators of Nasonovia ribisnigri in organic lettuce on the central coast of California. J Econ Entomol 100:39–48
Speight MCD (2011) Species accounts of European Syrphidae (Diptera). Syrph the Net, the database of European Syrphidae, vol 695. Syrph the Net publications, Dublin
Stahls G, Vujic A, Pérez-Bañón C, Radenkovic S, Rojo S, Petanidou T (2009) COI barcodes for identification of Merodon hoverflies (Diptera, Syrphidae) of Lesvos Island, Greece. Mol Ecol Resour 9:1431–1438
Staudacher K, Pitterl P, Furlan L, Cate PC, Traugott M (2011) PCR-based species identification of Agriotes larvae. Bull Entomol Res 101:201–210
Stubbs AE, Falk SJ (1983) British hoverflies: an illustrated identification guide. British Entomology and Natural History Society, London
Traugott M, Symondson WOC (2008) Molecular analysis of predation on parasitized hosts. Bull Entomol Res 98:223–231
Traugott M, Zangerl P, Juen A, Schallhart N, Pfiffner L (2006) Detecting key parasitoids of lepidopteran pests by multiplex PCR. Biol Control 39:39–46
Traugott M, Kamenova S, Ruess L, Seeber J, Plantegenest M (2013) Empirically characterising trophic networks: what emerging DNA-based methods, stable isotope and fatty acid analyses can offer. Adv Ecol Res 49:177–224
Acknowledgments
The authors thank Thaïs Aznar and Vanesa Vega for their technical support. The authors also thank Drs. R. Meyhöfer and P. Hondelmann (Leibniz University, Hannover, Germany) for providing syrphid samples, as well as Dr. A. Fereres (Institute of Agricultural Sciences-Spanish National Research Council (ICA-CSIC), Madrid, Spain) for providing specimens of several aphid species. This study has been funded by the Spanish Ministry of Economy and Competitiveness (MINECO) (Projects AGL2008-00546 and AGL2011-24349). P. Gomez-Polo was supported by a FPI Grant funded by the MINECO.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Dr. Donald C. Weber.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gomez-Polo, P., Traugott, M., Alomar, O. et al. Identification of the most common predatory hoverflies of Mediterranean vegetable crops and their parasitism using multiplex PCR. J Pest Sci 87, 371–378 (2014). https://doi.org/10.1007/s10340-013-0550-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10340-013-0550-6