Abstract
The genes involved in l-arginine biosynthesis in Corynebacterium crenatum are organized as the argCJBDFRGH cluster like in Corynebacterium glutamicum. However, the argC~H cluster of the C. crenatum SYPA 5-5, which is an industrialized l-arginine producer, had a lethal mutation occurring in the ArgR repressor encoding gene. The argC~H cluster with an inactive argR was overexpressed in E. coli and C. crenatum. In the recombinant E. coli JM109 enzyme activities were increased, and more l-arginine was found in the supernatants from l-glutamine. When the argC~H cluster was overexpressed in C. crenatum under its native promoter Parg, l-arginine production was increased by 24.9%, but the presence of the recombinant plasmid pJC-9039 had a negative effect on cell growth. Surprisingly, the DO value of the recombinant strain dropped gently and stayed at a lower level from 24 h to the end of fermentation. The results demonstrated an increasing utilization of oxygen and the distinct enhancement of unit cell l-arginine yields with the cluster argC~H-bearing in C. crenatum SYPA-9039. This study provides a kind of Corynebacteria with improved l-arginine-producing ability and an efficient elevation for producing amino acid. Moreover, the promoter Parg would be used as a valid promoter to express objective genes for metabolic engineering in Corynebacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
l-Arginine is a semi-essential amino acid involved in numerous areas of human biochemistry, including ammonia detoxification, hormone secretion, and the immune system. Arginine is also well known as a precursor to nitric oxide, which is a key component of endothelial-derived relaxing factor [8]. So far, most l-arginine has been produced by the direct fermentation method from natural carbon sources. l-Glutamine is an important metabolic precursor for l-arginine biosynthesis; therefore, l-glutamine producers have always been used as starting strains for l-arginine production [6, 7].
A traditional method to develop strains has been random mutation followed by selection of strains showing the desired phenotypes. Strategies for developing amino acid producers are now in transition toward systems metabolic engineering from random mutagenesis. Targeted metabolic engineering of several genes and pathways has become a standard method of strain improvement. After the genome sequence of the C. glutamicum wild-type strain, ATCC 13032, was determined [12], the genome-wide analyses contributed to the understanding of genetic regulation and became integral parts of C. glutamicum systems biology [28]. Recently, several amino acid producers have been successfully developed by systems metabolic engineering [22]. Due to the important role of Corynebacterium species as amino acid producers, several genes from these strains involved in amino acid biosynthesis have been isolated and characterized, and some of these genes have been used to design engineered strains with improved amino acid production [4].
The genes within the cluster of genes involved in arginine biosynthesis are organized as two separate parts, which are argCJBDFR and argGH operons in C. glutamicum [9]. The gene cluster encodes all of the enzymes required to convert glutamate to arginine via ornithine [23, 32]. l-Arginine biosynthesis starts with the acetylation of the amino group of glutamate, mediated by N-acetylglutamate synthase (in linear pathway) or N-acetyltransferase (in cyclic pathway). The cyclic acetyl pathway directs the l-arginine flow in procaryotic organisms such as C. glutamicum, Pseudomonas aeruginosa [10], Bacillus subtilis [15], Streptomyces coelicolor [1], and Thermus thermophilus [21].
In this article, we studied the corresponding genes for l-arginine biosynthesis from glutamate in C. crenatum SYPA5-5. Because the similarity of the 16SrDNA sequence and the homology of the functional DNA sequence between C. crenatum and C. glutamicum are highly homogeneous, we investigated the evolution of the genes involved in the l-arginine biosynthesis of C. crenatum with the genomic data of C. glutamicum [12]. The argCJBDFRJH (argC~H) cluster involved in l-arginine biosynthesis was overexpressed with the aim to improve l-arginine production in C. crenatum. The overexpression of arginine cluster was set up as follows: firstly, we reported our results with the overexpression of the complete argC~H cluster (argC–argH genes) in E. coli. Then, motivated by the results obtained from this work, the simultaneous overexpression of the genes involved together in arginine biosynthesis was performed in C. crenatum. All the data obtained are presented in this article.
Materials and methods
Strains and plasmids
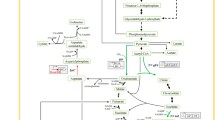
Corynebacterium crenatum SYA is an aerobic, gram-positive, non-sporulating coryneform bacterium, and its UV&EMS-mutant strain C. crenatum SYPA5-5 (CGMCC no. 0890) could produce l-arginine (30 g/l) under optimal culture conditions [30]. Firstly, the argC~H was cloned on the pMD-18T vector (AmpR, TaKaRa, Japan) and transformed into E. coli JM109 (Invitrogen) to form JM109-9039. The C. glutamicum/E. coli shuttle vector pJC1 (KmR) had been demonstrated to replicate in C. crenatum, too [5, 31]. Then the argC~H cluster (9,039 bp) was subcloned on the plasmid pJC1 and introduced into C. crenatum SYPA5-5. Recombinant plasmids were constructed (Fig. 1).
Biosynthetic pathway of l-arginine in Corynebacteria, the configuration of argC~H cluster, and the maps of recombinant plasmids constructed in this study. argJ: ornithine acetyltransferase; argB: acetylglutamate kinase; argC: acetylglutamate semialdehyde dehydrogenase; argD: acetylornithine transaminase; argF: ornithine transcarbamylase; argG: argininosuccinate synthase; argH: arginosuccinase
Cloning and expression in E. coli of argC~H cluster
Standard protocols were used for transformation of E. coli and for isolation, digestion, and ligation of DNA [24]. ArgC~H cluster was amplified from templates genomic DNA from C. crenatum SYPA 5-5. The PCR amplification was carried out in a 50-μl reaction mixture containing 5 ng of template, 200 μM dNTPs, 20 μM primers, and 2 U LATaq™ DNA polymerase (TaKaRa, Japan). Then the PCR products were purified by Gel Extraction Kit and cloned into pMD-18T vector. The primers (P1: 5′-CGCTCTAGAAAATTCATGCTTTTACCCACTTGCAGTTTTAGC-3′; P2: 5′-CGCTCTAGAGAATTGATAATAAATGGCCTGTGCAACTCCCG-3′) were designed to introduce the XbaI site at the promoter of the argC~H and the XbaI site at the end of the amplified region argC~H cluster. The PCR products, digested with XbaI–XbaI, were inserted into the same site's digestion plasmid pJC1. The ligation mixture was used for transforming into E. coli JM109 by the calcium chloride method [2]. When necessary, ampicillin or kanamycin was added at the proper final concentration for the transformant selection.
The resulting plasmids, pT-9039 and pJC-9039, isolated from the recombinant E. coli JM109 cells grown in Luria-Bertani (LB) medium containing 100 μg/ml ampicillin or 50 μg/ml kanamycin, were tested. After growth of the cells at 37°C in liquid LB medium supplemented with ampicillin or kanamycin, the subsequent steps were carried out at 4°C. The cells were harvested by centrifugation, suspended in 20% of the original culture volume of 10 mM Tris-HCl (pH 7.4), sonicated, and the resulting extract was centrifuged at 15,000 rpm for 30 min. The supernatant was used for sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and for the activity assays of seven enzymes involved in the argC~H cluster.
Cloning and expression in C. crenatum of argC~H cluster
Corynebacterium crenatum was cultivated at 31°C in LBG medium, LB containing (in g/l) glucose 5 and competent cell medium, and LB including (in g/l) glycine 30 and Tween 1. Then the recombinant plasmid pJC-9039 was transformed into C. crenatum SYPA5-5 to form engineered SYPA-9039 using the electroporation methods described by Tauch et al. [26], and transformants were selected on LBG agar containing 50 μg/ml kanamycin.
Corynebacterium crenatum strains were grown in 200 ml LBG media and harvested when OD562 reached 25.0. Cell pellets from 200 ml cell culture were resuspended in 5 ml Tris-HCl buffer (100 mM, pH 7.4) to a concentration of 100 g/l wet cells. Then the cells were sonicated and centrifuged at 15,000 rpm for 30 min. The resulting cell-free extracts were used for SDS-PAGE analysis, with 40 μg total proteins loaded on each lane.
Enzyme assays
The same cell-free extracts were used to measure the activity of the seven enzymes involved in the cluster. N-Acetylglutamate 5-semialdehyde dehydrogenase (EC. 1.2.1.38), N-acetylornithine aminotransferase (EC. 2.6.1.11), N-acetylglutamate kinase (EC. 2.7.2.8.), ornithine N-acetyltransferase (EC. 2.3.1.35), orinithine transcarbamoylase (EC. 2.1.3.3), argininosuccinate synthetase (EC. 6.3.4.5), and argininosuccinate lyase (EC. 4.3.2.1) activities were assayed in different E. coli transformants and C. crenatum according to the methods of Takahara et al. [25], Martin et al. [17], Wolf et al. [29], Billhemier et al. [3], Mountain et al. [18], Kumar et al. [14], and Troshina et al. [27], respectively.
Growth medium and conditions for l-arginine production
Corynebacterium crenatum SYPA 5-5 and its transformants are auxotrophic for l-histidine. A stock culture was maintained on agar slants containing (in g/l) peptone 10, beef extract 10, yeast extract 5, NaCl 5, and agar 20. The seed culture medium (in g/l) consisted of glucose 30, corn steep liquor 20, (NH4)2SO4 20, KH2PO4 1, MgSO4·7H2O 0.5, and urea 1.5 (pH 7.0). The seed was inoculated from agar slants and cultured at 31°C for 14–16 h in a shake flask. The shake flask culture was then transferred into a 5-l bioreactor (BIOTECH-5BG, Baoxing Co., China) containing 3 l fermentation medium A (in g/l) consisting of glucose 150, corn steep liquor 40, (NH4)2SO4 20, KH2PO4 1.5, MgSO4·7H2O 0.5, FeSO4·7H2O 0.02, MnSO4·H2O 0.02, biotin 8 × 10−5, and l-histidine 5 × 10−4 (pH 7.0). The batch fermentation was performed at 31°C for 108 h until steady strain growth. The aeration rate was controlled at 1 vvm for all the experiments. Studies in the bioreactor were carried out at the agitation rate of 600 rpm. The dissolved oxygen concentration (DOC, %) under the operation conditions was expressed in terms of dissolved oxygen (DO) saturation level, while 100% DO saturation level corresponded to an actual DO concentration of about 7.3 mg/l at 31°C, 1.0 atm. l-Arginine was also produced by culturing E. coli, which was studied in this article. The E. coli recombinants had the ability to produce l-arginine in a medium containing glucose with l-glutamate added to produce and accumulate l-arginine in a culture. The E. coli fermentation medium B (in g/l) contained glucose 30, l-glutamate 5, corn steep liquor 20, (NH4)2SO4 25, KH2PO4 2, MgSO4·7H2O 1, and chalk 15 (pH 7.0). Glucose/l-glutamate and chalk were sterilized separately. Then 2 ml of the medium was placed into the test flask, and the cultivation was carried out at 37°C for 60 h with shaking at 150 rpm.
Assays of cell concentration, glucose, and l-arginine
The cell concentration was first monitored at 562 nm, and the dry cell weight (DCW) was determined by a pre-calibrated relationship (1 OD = 0.375 g/l DCW). Glucose concentration in the media was measured by using the anthrone method [11]. Concentrations of amino acids were measured by an Agilent 1100 HPLC under the following conditions: column Hypersil ODS-C18 4 × 125 mm, temperature 40°C, flow rate 1.0 ml/min, detection fluorescence detector, Ex 340 nm Em 450 nm, eluent A 20 mM Na-acetate, and eluent B 20 mM of Na-acetate: methanol: acetonitrile = 1:2:2 (v/v). All of the measurements, particularly the most important state variables, such as the concentration of cells, l-arginine, and glucose, were measured in three parallels.
Plasmid stability test
Samples from fermentation were spread on selective (50 μg/ml kanamycin) and nonselective LBG agar plates, respectively, after proper dilution. Then, the plates were incubated at 31°C for 24–36 h. The plasmid stability was shown as the ratio in percentage of colonies on the antibiotic agar plates over those on the plates without kanamycin.
Results
Cloning and sequence analysis of the C. crenatum argC~H cluster gene
The argC~H cluster on a single 9,039-bp chromosomal fragment encoding the seven successive enzymes of the arginine biosynthetic pathway was isolated from C. crenatum SYPA 5-5. Analysis of the 9.0-kb nucleotide sequence revealed eight ORFs, and the ORFs were numbered as indicated in Fig. 1. The arginine biosynthetic gene cluster of C. crenatum contains eight genes (argC, argJ, argB, argD, argF, argR, argG, and argH). Accordingly, the major arginine biosynthetic cluster involves eight genes of C. crenatum SYA and SYPA5-5 showing 99.23 and 99.22%, respectively, homologous to the argC~H cluster of C.glutamicum ATCC13032 (GenBank accession no. BX927147). In fact, there was only 1-bp difference in the ORF6 (argR) strain SYPA5-5 compared to the argR of its wild-type C. crenatum SYA. But the ORF6 (argR) had an early termination codon, i.e., TAG. Therefore, the argR repressor could not carry out the feedback repression with arginine in its arginine biosynthesis of UV&EMS-mutant strain C. crenatum SYPA5-5. The nucleotide sequence data in this study were shown in the GenBank Nucleotide Databases under accession no. HQ602711.
Construction of recombinant plasmids harboring argC~H cluster
The recombinant plasmid pT-9039 was constructed as described in “Materials and methods.” The recombinant plasmid harboring argC~H cluster was digested with XbaI; this recombinant plasmid was divided by XbaI into two major fragments with sizes of 9.0 and 2.7 kb, respectively, which are consistent with the map of pT-9039. The resulting E. coli JM109 containing plasmid pT-9039 was designated JM109-9039.
To further increase the arginine production in C. crenatum, we aimed to overexpress argC~H cluster in the strain SYPA5-5. Similarly, the recombinant plasmid pJC-9039 was constructed. The recombinant plasmid was digested with XbaI and NotI, and the recombinant plasmid was divided into two major fragments with sizes of 9.0 and 6.1 kb, respectively, which are consistent with the map of pJC-9039. There were only two fragments in the digestion: one had the same size as the empty plasmid, pJC1 vector; the other had the same size as the argC~H cluster. The Not I digested product had the same 15.1-kb size as the fragment of argC~H cluster and pJC1.
Expression of argC~H cluster in E. coli
It is supposed that there is a promoter in the upstream sequence of the cluster. It was verified that the argC~H cluster had been cloned and could express in E. coli, that is to say, the native promoter (Parg) had an evident transcription activity in E. coli. The seven enzyme activities involved in arginine biosynthesis were determined by the methods described in Materials and methods. The data in Table 1 demonstrate only low enzyme activities in E. coli; they were apparently increased over the control, demonstrating the apparent use of the specific promoter of the argC~H cluster from C. crenatum. The six enzyme activities except for OAT were increased 1.6–6.2 times in the engineered JM109-9039. The specific activity, OAT, could be detected with 1.14 U/mg; however it was very low because of the lack of OAT in the control E. coli JM109. Consequently, the argC~H cluster was expressed with a certain amount of protein in E. coli JM109 under the promoter Parg. When analyzed by using SDS-PAGE, some protein bands in the JM109-9039 cell were different from those in JM109 (Fig. 2a). However, no specific bands corresponded to the argC~H cluster visible in the cell extract of JM109-9039. The presumable reason is that the quantity of the overexpressed protein was below the detection limit of Coomassie brilliant blue (CBB) R250 because of the low transcription efficiency of the promoter Parg.
Coomassie blue-stained SDS-PAGE of cell-free extracts of the recombinant E. coli and C. crenatum. About 40 μg of total proteins was loaded on each lane. a Lane 1, protein molecular weight marker; lane 2, sample from E. coli JM109; lane 3, sample from JM109-9039. b Lane 1, protein molecular weight marker; lane 2, sample from C. crenatum SYPA5-5; lane 3, sample from SYPA-9039
Enhanced l-arginine production from glutamate by engineered JM109-9039
The engineered JM109-9039 was cultured for 60 h in the medium containing glucose (and l-glutamate or not), and the concentration of l-arginine in the fermentation culture was measured. The effect of argC~H expression on cell growth and l-arginine accumulation by E. coli JM109 and JM109-9039 was determined. It was proved that the engineered JM109-9039 could harvest more arginine in the supernatants when the medium contained l-glutamate, whereas no more arginine was produced if it only contained glucose. It seems that the overexpression of argC~H could only further boost the metabolic flux from l-glutamate to l-arginine in the pathway of E. coli JM109 (Fig. 3). The immediate reason for no more arginine being harvested from glucose is the lower production of l-glutamate in E. coli, that is to say, the expression of argC~H cluster in E. coli does little to boost the l-glutamate productivity.
Expression of argC~H cluster in C. crenatum SYPA
The recombinant plasmid pJC-9039 was transformed into C. crenatum SYPA 5-5. The expression of argC~H cluster by using the plasmid pJC1 under its native promoter Parg was first checked using SDS-PAGE. The expected seven distinct proteins (from NAGSD to AL) corresponding to the overexpressed argC~H cluster could not be observed, although a few more bands appeared in the samples of SYPA-9039 (Fig. 2b). Then the specific enzyme activities in the recombinant SYPA-9039 were evaluated with the control activities in SYPA5-5. As a result, it was found that the activities from the cell-free extract of SYPA5-5 were much higher than those from E. coli. Furthermore, specific enzyme activities in SYPA-9039 displayed 1.5–2.7 times more than those in the original strain SYPA 5-5 (data shown in Table 1). Additionally, the results indicate that the native Parg promoter is a functional promoter; however, its transcription efficiency in C. crenatum was not high enough compared to that in E. coli JM109.
Improved l-arginine production by recombinant C. crenatum carrying argC~H cluster
The argC~H cluster was introduced to overexpress NAGSD~AL in C. crenatum under its native promoter Parg with the aim to increase l-arginine production. The C. crenatum recombinants were first fermented to produce l-arginine in a 30/250-ml shake flask test, and the recombinant SYPA-9039 was screened for its highest production in a 250-ml flask. Then one of the SYPA-9039 recombinants was fermented in the 5-l fermentor. The effects of the overexpression of the argC~H cluster on cell growth, glucose consumption, and l-arginine accumulation of the recombinant C. crenatum were determined. Starting from the 100% air saturation point, the fermentation curves, including concentrations of cells, glucose, and l-arginine, as well as the l-arginine production rate of wild-type strain and SYPA-9039 compared in the same media for proper aeration fermentation, are described in Fig. 4. The results illustrated that the presence of the recombinant plasmid pJC-9039 could enhance l-arginine production, despite the fact that the expression of the long gene cluster had a negative effect on cell growth from the beginning of the fermentation. However, the final biomass reached the same level as it did in the wild-type strain (Fig. 4a). Meanwhile, the glucose consumption rate in the cluster-bearing strain remained faster than the control (Fig. 4b). Finally, the l-arginine production of recombinant and wild-type C. crenatum reached concentrations of 45.30 and 36.27 g/l under the same fermentation conditions, respectively, that is to say, l-arginine production was increased 24.9% (Fig. 4c). Thus, the distinct enhancement of unit cell arginine yields with the cluster argC~H-bearing in C. crenatum SYPA-9039 could be observed. At the same time, the l-arginine production rate data also indicated the differences between the two strains, and hyperproduction rates in the recombinant were examined (Fig. 4d). The time changing profiles of the DO of the two strains are depicted in Fig. 4e. During the growth phase of the two strains, DO of wild C. crenatum dropped rapidly and got to the lowest levels of 41.9% of the saturation point at 15 h and then began to rebound, while the DO of the recombinant strain dropped gently and was kept at a lower level from 20 h to the end. These results illustrated a distinct increasing utilization of oxygen with the cluster argC~H-bearing in C. crenatum SYPA.
Comparison of l-arginine production between C. crenatum SYPA-9039 and C. crenatum SYPA5-5. a Cell concentration; b glucose concentration; c l-arginine concentration; d l-arginine production rate. Open triangle/thick solid line: SYPA-9039 recombinant, open square/thin solid line: C. crenatum SYPA5-5; e the changing dissolved oxygen (DO) patterns under proper oxygen supply conditions. Thick solid line: SYPA-9039 recombinant, thin solid line: C. crenatum SYPA5-5
To check the plasmid stability of pJC-9039, the aforementioned assays were performed. The ratio in percentage of colonies on the antibiotic agar plates over those on the plates without antibiotics was 99.14%. At the same time, all the assays showed similar specific activities and the fermentation characteristics with samples taken from several continuous fermentations of SYPA-9039, suggesting the stable fermentation stability of SYPA-9039.
Discussion
In this study, an early termination codon in the argR was found to be involved in the argC~H cluster from strain SYPA5-5. Therefore, we supposed that the ArgR repressor could not act in the feedback repression of arginine biosynthesis in UV&NTG-mutated strain C. crenatum SYPA5-5. Consequently, the argC~H cluster was introduced to overexpress in C. crenatum and E. coli under its native promoter Parg, respectively. In E. coli JM109, the enzymatic activities of the arginine biosynthesis enzymes exhibited higher levels with the functional promoter Parg in JM109-9039. Regardless, this situation raised up the problem of the efficiency and effectiveness of Parg in E. coli amd C. crenatum. The comprehensive study of promoter Parg has been undertaken.
The recombinant C. crenatum SYPA-9039 had a better l-arginine-producing ability because of the enhancement of the enzyme activities in l-arginine biosynthesis than the wild-type, wherein the expression of intracellular enzymes was enhanced by increasing the copy number of the argC~H cluster involved in arginine production in C. crenatum SYPA. Simultaneously, the results in this study demonstrated an obvious increasing utilization of oxygen in the cluster argC~H-bearing C. crenatum. Maghnouj et al. reported that the dissolved oxygen (DO) levels largely determined the interactions between the metabolic reactions and genetic regulatory mechanism, as well as the formations of products and by-products in the production of l-arginine [13, 16]. Our previous study demonstrated that the l-arginine production would be increased in a way enhancing the flux distribution ratio directed to glutamate synthesis in the TCA cycle during late fermentation phases throughout the fermentation at the higher oxygen supply condition [30, 31]. Therefore, the effects of the improved DO with the overexpression of argC~H could lead to more l-arginine production as well in the recombinant SYPA-9039. Consequently, it could clearly be considered that the high load pressure of the expression of the gene cluster 9,039-bp resulted in slow growth, but more oxygen utilization and glucose consumption, which resulted in an enhancement of l-arginine production. Thus, the most significant characteristic of recombinant C. crenatum was the increased unit cell arginine yields.
Recently, there have been several reports on the use of omics analysis for the development of strains for amino acid production [19, 20]. On the basis of more protein bands being shown in SYPA-9039 using SDS-PAGE, further metabolic engineering should be carried out. Proteomics analysis is performed to examine the effect on the other protein expression of the cluster argC~H in the engineering strain. Then the data and information obtained would be used for further improvement of the strain. In addition, these results indicate that argC~H cluster was expressed at a level under its own promoter in the upstream region of argC gene ORF. Consequently, Parg could be used as the valid promoter to express objective genes in Corynebacterium species, and moreover, this Parg promoter could be conveniently site-directed and improved for great efficiency to develop the Corynebacteria vector systems for metabolic engineering in Corynebacterium species. The present invention provides a coryneform bacterium with improved l-arginine-producing ability and an efficient method for producing l-arginine.
References
Auchter M, Cramer A, Hüser A, Rückert C, Emer D, Schwarz P, Arndt A, Lange C, Kalinowski J, Wendisch VF (2010) RamA and RamB are global transcriptional regulators in Corynebacterium glutamicum and control genes for enzymes of the central metabolism. J Biotechnol. doi:10.1016/j.jbiotec.2010.07.001
Ausubel F, Brent R, Kingston R, Moore D, Seidman J, Smith J, Struhl K (1987) Current protocols in molecular biology, vol 1 and 2. Green Publishing Associates and Wiley Inter-science, New York, NY 12:1C12.12
Billhemier JT, Carnevale HN, Leisinger T, Eckhardt T, Jones EE (1976) Ornithine delta-transaminase activity in Escherichia coli: its identity with acetylornithine delta-transaminase. J Bacteriol 127(3):1315–1323
Blombach B, Hans S, Bathe B, Eikmanns BJ (2008) Acetohydroxyacid synthase, a novel target for improvement of l-lysine production by Corynebacterium glutamicum. Appl Envir Microbiol 75(2):419–427. doi:10.1128/aem.01844-08
Cremer J, Eggeling L, Sahm H (1990) Cloning the dapA dapB cluster of the lysine-secreting bacterium Corynebacterium glutamicum. Mol Gen Genet 220(3):478–480. doi:10.1007/bf00391757
Faurie R, Thommel J, Bathe B, Debabov V, Huebner S, Ikeda M, Kimura E, Marx A, Möckel B, Mueller U, Pfefferle W, Ikeda M (2003) Amino acid production processes. In: Microbial production of l-amino acids, vol 79. Adv Biochem Eng/Biotechnol, Springer Berlin/Heidelberg, pp 1–35. doi:10.1007/3-540-45989-8_1
Glansdorff N, Xu Y (2007) Microbial arginine biosynthesis: pathway, regulation and industrial production. Microbiol Monogr 5:219–257. doi:10.1007/7171_2006_061
Grillo MA, Colombatto S (2004) Arginine revisited: minireview article. Amino Acids 26(4):345–351. doi:10.1007/s00726-004-0081-9
Hänßler E, Müller T, Jeßberger N, Völzke A, Plassmeier J, Kalinowski J, Krämer R, Burkovski A (2007) FarR, a putative regulator of amino acid metabolism in Corynebacterium glutamicum. Appl Microbiol Biot 76(3):625–632. doi:10.1007/s00253-007-0929-5
Ikeda M, Mitsuhashi S, Tanaka K, Hayashi M (2009) Reengineering of a Corynebacterium glutamicum l-arginine and l-citrulline producer. Appl Environ Microbiol 75(6):1635–1641. doi:10.1128/aem.02027-08
Jermyn M (1975) Increasing the sensitivity of the anthrone method for carbohydrate. Anal Biochem 68(1):332–335. doi:10.1016/0003-2697(75)90713-7
Kalinowski J, Bathe B, Bartels D, Bischoff N, Bott M, Burkovski A, Dusch N, Eggeling L, Eikmanns BJ, Gaigalat L, Goesmann A, Hartmann M, Huthmacher K, Kramer R, Linke B, McHardy AC, Meyer F, Mockel B, Pfefferle W, Puhler A, Rey DA, Ruckert C, Rupp O, Sahm H, Wendisch VF, Wiegrabe I, Tauch A (2003) The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of l-aspartate-derived amino acids and vitamins. J Biotechnol 104(1–3):5–25. doi:S0168165603001548
Koffas M, Roberge C, Lee K, Stephanopoulos G (1999) Metabolic engineering. Annu Rev Biomed Eng 1:535–557. doi:10.1146/annurev.bioeng.1.1.535
Kumar A, Vij N, Randhawa GS (2003) Isolation and symbiotic characterization of transposon Tn5-induced arginine auxotrophs of Sinorhizobium meliloti. Indian J Exp Biol 41(10):1198–1204
Larsen R (2005) Interaction between ArgR and AhrC controls regulation of arginine metabolism in Lactococcus lactis. J Biol Chem 280(19):19319–19330. doi:10.1074/jbc.M413983200
Maghnouj A, Abu-Bakr AAW, Baumberg S, Stalon V, Van der Wauven C (2000) Regulation of anaerobic arginine catabolism in Bacillus licheniformis by a protein of the Crp/Fnr family, vol 191. Blackwell Publishing Ltd. doi:10.1111/j.1574-6968.2000.tb09344.x
Martin PR, Mulks MH (1992) Sequence analysis and complementation studies of the argJ gene encoding ornithine acetyltransferase from Neisseria gonorrhoeae. J Bacteriol 174(8):2694–2701. doi:0021-9193/92/082694-08$02.00/0
Mountain A, McChesney J, Smith MC, Baumberg S (1986) Gene sequence encoding early enzymes of arginine synthesis within a cluster in Bacillus subtilis, as revealed by cloning in Escherichia coli. J Bacteriol 165(3):1026–1028. doi:0021-9193/86/031026-03$02.00/0
Ohnishi J (2008) Characterization of mutations induced by N-methyl-N′-nitro-N-nitrosoguanidine in an industrial Corynebacterium glutamicum strain. Mut Res/Gen Tox Environ Mutag 649(1–2):239–244. doi:10.1016/j.mrgentox.2007.10.003
Ohnishi J, Katahira R, Mitsuhashi S, Kakita S, Ikeda M (2005) A novel gnd mutation leading to increased l-lysine production in Corynebacterium glutamicum. FEMS Microbiol Lett 242(2):265–274. doi:10.1016/j.femsle.2004.11.014
Panhorst M, Sorger-Herrmann U, Wendisch VF (2010) The pstSCAB operon for phosphate uptake is regulated by the global regulator GlxR in Corynebacterium glutamicum. J Biotechnol. doi:10.1016/j.jbiotec.2010.07.015
Park JH, Lee SY (2008) Towards systems metabolic engineering of microorganisms for amino acid production. Curr Opin Biotechnol 19(5):454–460. doi:10.1016/j.copbio.2008.08.007
Sakanyan V, Petrosyan P, Lecocq M, Boyen A, Legrain C, Demarez M, Hallet JN, Glansdorff N (1996) Genes and enzymes of the acetyl cycle of arginine biosynthesis in Corynebacterium glutamicum: enzyme evolution in the early steps of the arginine pathway. Microbiol 142(Pt 1):99–108. doi:10.1099/13500872-142-1-99
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York
Takahara K, Akashi K, Yokota A (2007) Continuous spectrophotometric assays for three regulatory enzymes of the arginine biosynthetic pathway. Anal Biochem 368(2):138–147. doi:10.1016/j.ab.2007.06.032
Tauch A, Kirchner O, Löffler B, Götker S, Pühler A, Kalinowski J (2002) Efficient electrotransformation of Corynebacterium diphtheriae with a mini-replicon derived from the Corynebacterium glutamicum plasmid pGA1. Curr Microbiol 45(5):362–367. doi:10.1007/s00284-002-3728-3
Troshina O, Hansel A, Lindblad P (2001) Cloning, characterization, and functional expression in Escherichia coli of argH encoding argininosuccinate lyase in the cyanobacterium Nostoc sp. strain PCC 73102. Curr Microbiol 43(4):260–264. doi:10.1007/s002840010298
Wendisch VF (2006) Genetic regulation of Corynebacterium glutamicum metabolism. J Microbiol Biotechnol 16(7):999–1009
Wolf E, Weiss R (1980) Acetylglutamate kinase. A mitochondrial feedback-sensitive enzyme of arginine biosynthesis in Neurospora crassa. J Biol Chem 255(19):9189
Xu H, Dou WF, Xu HY, Zhang XM, Rao ZM, Shi ZP, Xu ZH (2009) A two-stage oxygen supply strategy for enhanced l-arginine production by Corynebacterium crenatum based on metabolic fluxes analysis. Biochem Eng J 43(1):41–51. doi:10.1016/j.bej.2008.08.007
Xu MJ, Rao ZM, Xu H, Lan CY, Dou WF, Zhang XM, Xu HY, Jin JA, Xu ZH (2011) Enhanced production of l-arginine by expression of Vitreoscilla hemoglobin using a novel expression system in Corynebacterium crenatum. Appl Biochem Biotechnol 163(6):707–719. doi:10.1007/s12010-010-9076-z
Xu Y, Labedan B, Glansdorff N (2007) Surprising arginine biosynthesis: a reappraisal of the enzymology and evolution of the pathway in microorganisms. Microbiol Mol Biol Rev 71(1):36–47. doi:10.1128/mmbr.00032-06
Acknowledgments
We are grateful to Dr. Andreas Burkovski (Universität zu Köln, Germany) for the generous donation of plasmid pJC1. This work was supported by the High-tech Research and Development Programs of China (2007AA02Z207), the National Basic Research Program of China (2007CB707804), the National Natural Science Foundation of China (30970056), and the Program for New Century Excellent Talents in the University (NCET-07-0380, NCET-10-0459), the Fundamental Research Funds for the Central Universities (JUSRP31001), the Program of Introducing Talents of Discipline to Universities (111-2-06), and a Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Xu, M., Rao, Z., Yang, J. et al. Heterologous and homologous expression of the arginine biosynthetic argC~H cluster from Corynebacterium crenatum for improvement of l-arginine production. J Ind Microbiol Biotechnol 39, 495–502 (2012). https://doi.org/10.1007/s10295-011-1042-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-011-1042-4