Abstract
Yeasts that ferment both hexose and pentose are important for cost-effective ethanol production. We found that the soil yeast strain NY7122 isolated from a blueberry field in Tsukuba (East Japan) could ferment both hexose and pentose (d-xylose and l-arabinose). NY7122 was closely related to Candida subhashii on the basis of the results of molecular identification using the sequence in the D1/D2 domains of 26S rDNA and 5.8S-internal transcribed spacer region. NY7122 produced at least 7.40 and 3.86 g l−1 ethanol from 20 g l−1 d-xylose and l-arabinose within 24 h. NY7122 could produce ethanol from pentose and hexose sugars at 37°C. The highest ethanol productivity of NY7122 was achieved under a low pH condition (pH 3.5). Fermentation of mixed sugars (50 g l−1 glucose, 20 g l−1 d-xylose, and 10 g l−1 l-arabinose) resulted in a maximum ethanol concentration of 27.3 g l−1 for the NY7122 strain versus 25.1 g l−1 for Scheffersomyces stipitis. This is the first study to report that Candida sp. NY7122 from a soil environment could produce ethanol from both d-xylose and l-arabinose.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ethanol production from plant biomass such as rice straw, waste wood, and agricultural residues is noncompetitive with food sources for animals and humans. Plant cell walls are composed of three major polymers, cellulose, hemicellulose, and lignin. Hemicellulose, which accounts for approximately 15–35% of lignocellulose, is a complex polymer containing a mixture of hexose (d-galactose, d-glucose, and d-mannose) and pentose (d-xylose and l-arabinose) sugars [11]. d-Xylose and l-arabinose are a major component of hemicellulosic hydrolysate derived from agricultural crops and hardwoods [24, 26]. One of the barriers that have prevented large-scale ethanol production from lignocellulose is the cost of production. Fermentation of all sugars present in lignocellulose material, including d-glucose, d-mannose, d-galactose, d-xylose, and l-arabinose, would be essential for cost-effective ethanol production. Thus one of the most critical features that an industrial ethanol producer from lignocellulose must have is its ability to effectively ferment pentose sugars.
Saccharomyces cerevisiae is currently used for conversion of starch and sucrose based crops into ethanol. However, S. cerevisiae lacks the ability to ferment pentose sugars. There are only a few natural xylose-fermenting yeasts, including Scheffersomyces stipitis, formerly known as Pichia stipitis [18], Candida shehatae, and Pachysolen tannophilus [8, 28]. Among these yeasts, S. stipitis is commonly reported as the best native xylose-fermenting yeast in terms of product yield and productivity. Although many yeasts can assimilate l-arabinose [15, 22], only a few have been reported to ferment l-arabinose to ethanol [6, 16]. The highest ethanol yield has been reported for Candida arabinofermentans, but it only produced 1.9–3.4 g l−1 ethanol from 80–100 g l−1 l-arabinose in 12–14 days [16].
Most research has centered on engineering S. cerevisiae to ferment pentoses with—in the case of xylose—increasing success [21, 29, 31]. S. cerevisiae has also been engineered to ferment arabinose to ethanol but with less success. Discovering further pentose fermenting yeasts is desirable because it furthers our knowledge of pentose metabolism in yeasts in general and serves as an additional source of genes. Use of a nonrecombinant yeast strain might be desirable as well in some cases.
In the present study, a massive screening (over 1,600 strains) was undertaken to identify yeasts that ferment arabinose and xylose. This work describes the isolation of a yeast capable of fermenting both sugars, its initial characterization, and comparison with S. stipitis in parallel fermentations.
Materials and methods
Yeast strains
Yeast strains (1,699 strains) used for screening of pentose-fermenting yeasts were obtained from the Microbiological Bank of the National Food Research Institute (NFRI) and other culture collections, and from natural sources such as fermented foods. Candida arabinofermentans JCM 10727, Saccharomyces cerevisiae NBRC 0224, and Scheffersomyces stipitis NBRC 1687 were used as control strains. All strains were maintained on YPD agar plates (10 g l−1 yeast extract [Difco Laboratory, Detroit, MI], 20 g l−1 peptone [Difco], 20 g l−1 glucose, 20 g l−1 agar).
Screening of pentose-fermenting yeast
Yeast strains were inoculated into 100 μl of YPD medium in microtiter plates (Corning Inc., Corning, NY) and incubated at 30°C for 48 h (preculture). An aliquot (1.2 μl) of the preculture was transferred into 100 μl of YNBX media containing 6.7 g l−1 yeast nitrogen base without amino acid and 20 g l−1 d-xylose, and then the cultures were incubated at 30°C for 48 h. The optical density at 630 nm (OD630) of the cultures was measured using a microtiter plate reader (Elx800; BioTek Instruments, Winooski, VT). The selected xylose-assimilating yeasts were further screened to identify the strains with an ability to assimilate l-arabinose; for this search, the same method was used as for the selection of xylose-assimilating yeasts, with the exception that a different sugar was used.
Taxonomic identification and phylogenetic analysis
The selected yeast strain was taxonomically identified by determining the sequences of the D1/D2 domain of the 26S rDNA and 5.8S-internal transcribed spacer (ITS) regions of the rDNA. The partial 26S rDNA of the yeast was amplified by PCR using the primers NL-1 and NL-4 [17], and the ITS rDNA region was amplified with primers BMBC-R [19] and ITS4 [30]. Then PCR products were purified by using the QIAquick PCR purification kit (Qiagen, Valencia, CA). Cycle sequencing was done for the D1/D2 domain of the 26S rDNA with the primers NL-1 or NL-4, and for the complete 5.8S-ITS region with primers BMBC-R [19], ITS1, ITS2, ITS3, and ITS4 [30]. The sequences of the D1/D2 domain of the 26S rDNA and 5.8S-ITS regions including portions of the 3′ end of the 18S and 5′ start of the 28S were deposited in DNA Data Bank of Japan (DDBJ) under accession no. AB619618 and AB663086, respectively. The comparison of the sequence with known DNA sequences in the DDBJ was conducted using the BLAST system. The determined sequence of the D1/D2 domain of the 26S rDNA was aligned with the sequences of xylose-fermenting yeasts retrieved from DDBJ using CLUSTAL_X 2.0 [20]. Kimura’s two-parameter correction [14] was applied for the calculation of evolutionary distance, and a phylogenetic tree was constructed using the neighbor-joining method [25]. Bootstrapping analysis was performed with 1,000 replicates [10] and the resulting tree was plotted using the NJplot software program [23].
Morphological and physiological characterization
The macroscopic morphology of the yeast was observed by using a phase-contrast microscope (Leica DMLB 100; Leica, Solms, Germany). The cells grown on YPD agar or cornmeal agar for 1 or 5 days were used for microscopic analysis. Sugar fermentation tests were carried out using d-cellobiose, d-galactose, d-glucose, lactose, maltose, d-mannose, d-raffinose, sucrose, d-trehalose, d-xylose, and l-arabinose. To determine the sugar fermentation ability, gas and acid generation were analyzed in a fermentation basal medium consisting of 4.5 g l−1 yeast extract, 7.5 g l−1 peptone, 0.02 g l−1 bromocresol purple, and 20 g l−1 sugar (40 g l−1 for raffinose) with a Durham tube. Incubation was carried out at 30°C for 7 days. To detect slow fermentation, high-performance liquid chromatography (HPLC) analysis was also conducted to measure the ethanol concentration. The assimilation abilities of sugars were assessed by using an API 20C AUX system (BioMerieux, Tokyo, Japan), and ascospores were observed after growing the strains on McClary’s acetate agar (1 g l−1 glucose, 1.8 g l−1 potassium chloride, 8.2 g l−1 sodium acetate trihydrate, 2.5 g l−1 yeast extract, and 15 g l−1 agar) for 7 days at 25°C. Growth at different temperatures was determined by incubation on YPD agar plates for 48 h.
Characterization of growth and ethanol production
Single sugar fermentation tests were carried out to characterize the time-dependent changes of cell growth, sugar consumption, and ethanol production of the yeasts. NY7122 grew rarely under anaerobic conditions using a gas pack (Anaeropack; Mitsubishi Gas Chemical Company Inc., Tokyo, Japan) in an anaerobic jar (static culture), and grew poorly under low-oxygen conditions using a shake flask with a silicon rubber plug at a low agitation speed (data not shown). Thus cultures were incubated under aerobic conditions at an agitation speed of 120 rpm. One-hundred-millilter cotton-plugged Erlenmeyer flasks were used with a working volume of 25 ml using d-glucose, d-xylose, or l-arabinose as a carbon source. The culture media contained 10 g l−1 yeast extract, 20 g l−1 peptone, and 20 g l−1 of the desired sugar (approximate pH 6.2). The flasks were incubated at 30°C or 37°C on a rotary shaker at 120 rpm for 72 h. In control experiments, the dissolved oxygen level in the medium was increased from 35.8 (static flask condition) to 79.6% (shake flask condition) in 2 min; loss of ethanol from the 30 g l−1 ethanol media as a result of evaporation was about 3.3 g l−1 after 5 days (data not shown).
To assess the potential ability of NY7122 for fermentation of lignocellulosic hydrolysates, mixed sugar fermentation tests were performed. Because lignocellulosic hydrolysate is generally not nutrition rich [13], a synthetic minimal medium was chosen in this experiment over yeast extract peptone (YP) medium to simulate lignocellulosic hydrolysates as closely as possible. The volume of fermentation medium was increased from 25 ml in single sugar fermentation tests to 50 ml in order to suppress ethanol reassimilation due to excess aeration [27]. One-hundred-milliliter Erlenmeyer flasks containing 50 ml medium comprising 1.7 g l−1 yeast nitrogen base without ammonium sulfate and amino acids, 3 g l−1 urea, 50 g l−1 of d-glucose, 20 g l−1 of d-xylose, and 10 g l−1 of l-arabinose were used and the pH was adjusted with sulfuric acid (H2SO4) to 3.5. We also examined the effects of nutrient supply on ethanol production by adding 5 g l−1 yeast extract to the medium, because addition of nutrients to the hydrolysate improved the fermentation in some cases [3, 7, 12]. The cultures were incubated at 30°C on a rotary shaker at 120 rpm for 120 h. In control experiments, the dissolved oxygen level in the medium was increased from 27.1 (static flask condition) to 70.0% (shake flask condition) in 2 min; loss of ethanol from the 30 g l−1 ethanol media as a result of evaporation was about 2.0 g l−1 after 5 days (data not shown). For preparing the inocula for both single and mixed sugar fermentation experiments, cells were cultured in YPD overnight at 30°C on a rotary shaker (150 rpm), harvested by centrifuging, and then washed twice with 9 g l−1 NaCl. Washed cells were resuspended in mixed sugar medium at a cell concentration giving 0.5 g dry cell weight per liter.
Effect of initial pH on ethanol production by NY7122
The effect of the initial pH of the fermentation medium was studied by using pH values of 3.5, 4.5, 5.0, 6.5, and 9.5. The pH was adjusted with H2SO4 or sodium hydroxide (NaOH). The overnight YPD culture was inoculated into 25 ml of medium containing 10 g l−1 yeast extract, 20 g l−1 peptone, and 20 g l−1 of the desired sugar described above, and incubated at 30°C at 120 rpm for 48 h. Precultured cells were inoculated as described above.
Measurement of cell density, ethanol content, and residual sugar concentration
The growth was followed by measuring the optical density at 600 nm using a spectrophotometer. One unit of OD600 of screened strain used in this study corresponded to 0.25 g l−1 dry cell weight (dried at 105°C to a constant weight). One unit of OD600 of C. arabinofermentans cells, S. cerevisiae cells, and S. stipitis cells was about 0.36, 0.36, and 0.23 g l−1 of dry cells, respectively. Ethanol contents in the culture medium were measured using a Prominence series HPLC system (Shimadzu, Kyoto, Japan) equipped with a fermentation-monitoring column (Bio-Rad Laboratories, Hercules, CA). Samples were eluted with 5 mM H2SO4 as the mobile phase at a flow rate of 0.6 ml min−1 and temperature of 30°C. The concentrations of d-glucose, d-xylose, l-arabinose, xylitol, and l-arabitol were measured with HPLC using a Shodex Sugar SC1011 ion exchange column (Showa Denko, Tokyo, Japan) at 70°C with water as the mobile phase at a flow rate of 0.6 ml min−1. A refractive index detector (RID-6A; Shimadzu) was used for quantification in all HPLC analyses. The ethanol yield was calculated as grams of ethanol produced per grams of total sugar consumed or added.
Results and discussion
Screening of pentose-fermenting yeast
To obtain pentose-fermenting yeasts, we first assessed the growth abilities of 1,699 yeast strains in a medium with d-xylose as the sole carbon source. We obtained 580 strains that assimilate d-xylose. Among them, 36 strains could produce ethanol from d-xylose. Then we investigated the ability of the selected xylose-fermenting yeasts to assimilate l-arabinose. Finally, we obtained three strains that assimilate both d-xylose and l-arabinose and produce a detectable amount of ethanol from l-arabinose (data not shown). These three strains were obtained from a neighbor soil sample and our preliminary data obtained from a genetical identification approach based on the D1/D2 domain of 26S rDNA sequences suggested that these three strains belong to the same species. Among the three strains, NY7122, which was obtained from a soil sample collected from a blueberry field in Tsukuba, grew rapidly in the medium containing l-arabinose and produced the most ethanol (data not shown). On the basis of these results, we decided to investigate the strain NY7122 in further analyses.
Taxonomic identification and phylogenetic analysis
The sequence of the D1/D2 domain of the 26S rDNA and 5.8S-ITS regions in NY7122 was compared with sequences in the DDBJ nucleotide database. The sequence of the NY7122 gene in the D1/D2 domain of the 26S rDNA (AB619618) and 5.8S-ITS regions (AB663086) showed that the strain was most similar to Candida subhashii CBS 10753 (EU836708 and EU836707), which was the first reported type strain of C. subhashii [1]. The difference between NY7122 and C. subhashii CBS 10753 was a single nucleotide substitution out of 514 nucleotides (99.8% identical) in the D1/D2 domain of 26S rDNA. Kurtzman and Robnett [17] reported that ascomycetous yeast strains showing nucleotide substitutions constituting less than 1% of the D1/D2 domain of 26S rDNA usually represent the same species. On the other hand, the comparison of the 5.8S-ITS rDNA between NY7122 and C. subhashii CBS 10753 showed higher variability than that of the D1/D2 domain of 26S rDNA. The nucleotide length of the complete 5.8S-ITS region was 475 bp. The sequence was composed of the complete ITS1 sequence of 134 bp, complete 5.8S rDNA sequence of 158 bp, and complete ITS2 sequence of 183 bp. The whole 5.8S-ITS rDNA sequence of NY7122 differed from those of C. subhashii CBS 10753 by seven substitutions and one indel, and the sequence differences were 3.7% for the ITS1 region, 1.6% for ITS2, and 1.3% for the entire 5.8S-ITS region. There was no nucleotide variation in the 5.8S gene. Judging by the results of the sequencing, NY7122 was a closely related species of C. subhashii. We compared the sequence of NY7122 with that of other pentose-fermenting yeasts (Fig. 1). The phylogenetic tree based on neighbor-joining analysis of the D1/D2 domain revealed that NY7122 was phylogenetically distinct from other pentose-fermenting yeast species.
Neighbor-joining analysis based on the D1/D2 region of 26S rDNA sequence data, depicting the relationship between NY7122 and pentose-assimilating yeasts. Numbers on the branches represent bootstrap values based on 1,000 replicates and the values below 50% are not shown. The scale bar represents a 0.01 genetic distance (in nucleotide substitutions per site)
Morphological and physiological characterization
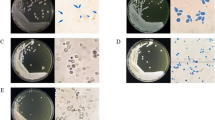
The colony morphology of NY7122 grown on YPD agar medium was observed. The strain produced a colony with convex, smooth, cream-colored, and faintly glistening surfaces during the early stage of colony growth. After 2–3 days of incubation, these colonies became dry and ring-shaped, and finally became irregular and wrinkled. The appearance of colonies of NY7122 was different from that of the colonies of C. subhashii CBS 10753 described by Adam et al. [1] in regard to surface texture, because CBS 10753 produced only smooth colonies at 7 days of incubation. NY7122 showed a dimorphic behavior and mycelia morphology. Microscopic observation of NY7122 grown on YPD broth indicated that the yeast showed budding proliferation and produced oval or round cells and a chain of elongated cells (Fig. 2a). However, pseudohyphae were formed abundantly on cornmeal agar after 5 days (Fig. 2b). Table 1 summarizes the physiological characteristics of NY7122. No ascospores were observed in NY7122, which was the same as the result for C. subhashii CBS 10753. The sugar assimilation pattern of NY7122 almost agreed with that of C. subhashii CBS 10753, except for the culture on pentose sugars. C. subhashii CBS 10753 assimilated these pentose sugars weakly, in contrast to NY7122, which strongly assimilated l-arabinose and d-xylose. The sugar fermentation profile of NY7122 was also different from that of C. subhashii CBS 10753, which was not able to ferment any sugars. NY7122 was able to ferment several sugars (Table 1). NY7122 grew at 30 and 37°C, and weakly at 40°C, but not at 42°C. The profiles of high temperature tolerance were similar to that of C. subhashii CBS 10753. The physiological characteristics of NY7122 suggested that NY7122 was a different species from C. subhashii.
Characterization of growth and ethanol production in NY7122
The characteristics of the growth and ethanol production of NY7122 were investigated on 25 ml of YP medium (pH 6.2) containing d-glucose, d-xylose, or l-arabinose as a sole carbon source. The time course analyses of cell growth at 30 or 37°C are shown in Fig. 3. For comparison, we also examined the ethanol production ability of the C. arabinofermentans JCM 10727, S. stipitis NBRC 1687, and S. cerevisiae NBRC 0224 under the same conditions. NY7122 had the most rapid growth, regardless of sugar and temperature. The ethanol accumulation, yield, residual sugar, and by-products formation are summarized in Table 2. The results showed that NY7122 produced more ethanol from pentose than control strains, but less ethanol from glucose. It is noteworthy that NY7122 could produce 1.9 g l−1 ethanol from 20 g l−1 l-arabinose at 30°C (pH 6.2) within 24 h. C. arabinofermentans JCM 10727, which is known as one of the best fermenters of l-arabinose [6, 16], produced a small amount of ethanol (0.3 g l−1) from l-arabinose in this experiment. S. stipitis was slightly able to consume l-arabinose, but as expected did not produce ethanol.
Time course analyses of growth on YP medium containing d-glucose (a), d-xylose (b) and l-arabinose (c) by NY7122 (circles), C. arabinofermentans JCM 10727 (triangles), S. stipitis NBRC 1687 (squares), and S. cerevisiae NBRC 0224 (diamonds). Yeasts are grown at 30°C (open symbols) or 37°C (closed symbols). Results shown are the means of three independent experiments
We found that NY7122 was able to grow well even at 37°C in any of the carbon sources tested. NY7122 had the ability to produce ethanol from pentose at 37°C at a higher rate than the other pentose-fermenting yeasts used in the present study as well as several previously reported strains [9]. Among the tested strains, C. arabinofermentans did not grow well at 37°C. S. stipitis was able to grow and to ferment glucose and xylose at 37°C, but its fermentation abilities were significantly depressed compared with those at 30°C. NY7122 produced glycerol in d-glucose medium to the extent of 1.08 and 1.12 g l−1 at 30 and 37°C, respectively, and in d-xylose medium, 1.07 and 1.37 g l−1 at 30 and 37°C, respectively. NY7122 produced glycerol in more minor amounts in l-arabinose medium to the extent of 0.92 and 0.97 g l−1 at 30 and 37°C, respectively. The ability to grow at 37°C is advantageous because simultaneous saccharification and fermentations in the presence of cellulases are typically operated at 35°C. Therefore, the cooling costs could be reduced. However, the production of the by-product sugar alcohols, including xylitol and l-arabitol, at 37°C was higher than that at 30°C. This could be a problem, because an increase of sugar alcohols would reduce the yield of ethanol. The increased production of sugar alcohols by relatively high temperature has also been reported in other yeast [5, 32]. Sugar alcohols are known as compatible solutes that protect organisms from stresses [4]. The increase of sugar alcohol in NY7122 at 37°C may thus be a metabolic response for adaptation to stresses.
Effect of initial pH on ethanol production
To determine the optimum pH for fermentation of NY7122, the cells were cultured in the medium at various pH values (ranging from 3.5 to 9.5) for 48 h. The effects of the initial pH of the medium on the ethanol production from d-glucose, d-xylose, and l-arabinose and their ethanol yields are shown in Table 3. Samples were collected at 6, 12, 24, and 48 h for measuring the concentration of products by HPLC. NY7122 produced the maximum amount of ethanol from pentose at pH 3.5 with low concentrations of sugar alcohols. When l-arabinose was used as a substrate, the concentrations of ethanol in the medium at pH 3.5–5.0 were more than twice as high as that at pH 6.5. These results indicated that an acidified medium might be favorable for ethanol production of NY7122.
Mixed sugar fermentation by NY7122
The results of ethanol production, by-product formation, sugar consumption, and cell growth of NY7122 in a mixed sugar medium at 30°C and pH 3.5 are shown in Fig. 4a. We also examined S. stipitis under the same conditions as used for the control (Fig. 4b). The maximum ethanol concentration obtained from mixed sugar fermentation was 26.2 g l−1 for the NY7122 and 22.3 g l−1 for the S. stipitis culture. The ethanol yields were 0.33 (grams of ethanol per gram of sugar added) for NY7122 and 0.28 for S. stipitis. NY7122 accumulated 3.19 g l−1 xylitol and 5.79 g l−1 l-arabitol. NY7122 completely consumed all sugars, whereas S. stipitis culture still contained 5.5 g l−1 of d-xylose after 120-h cultivation.
Time course analyses of sugar consumption, products formation, and cell growth in mixed sugar fermentation using NY7122 (a and c) and S. stipitis (b and d). The batch fermentation experiments were performed with medium containing 50 g l−1 d-glucose, 20 g l−1 d-xylose, and 10 g l−1 l-arabinose at pH 3.5 at 30°C for 120 h with agitation rate of 120 rpm. Yeast extract was added as nutrient supplement (c and d). The concentrations of ethanol (closed circles), d-glucose (open circles), d-xylose (open triangles), l-arabinose (open squares), xylitol (closed triangles), l-arabitol (closed squares), and OD600 of culture (open diamonds) were monitored. Results shown are the means of three independent experiments
The effects of nutrient supplement on ethanol production were examined. Both the ethanol production and sugar consumption were improved by addition of yeast extract to the medium (Figs. 4c and d). In the case of yeast extract addition, the maximum ethanol concentration was 27.3 g l−1 for the NY7122 and 25.1 g l−1 for the S. stipitis culture. The ethanol yields were 0.34 (grams of ethanol per gram of sugar added) for NY7122 and 0.31 for S. stipitis. Both NY7122 and S. stipitis completely consumed the glucose within 24 h. d-Xylose utilization in both yeasts began at a point when almost all of the glucose was consumed, and d-xylose was completely consumed within 72 h. After consumption of d-xylose, NY7122 began to utilize l-arabinose. The ethanol concentration in the culture medium of NY7122 was increased during l-arabinose fermentation (48–96 h). NY7122 accumulated more xylitol (1.43 g l−1) and l-arabitol (4.93 g l−1) compared with S. stipitis. S. stipitis used 2.12 g l−1 of l-arabinose, which seemed to be almost entirely excreted as l-arabitol. These results indicated that NY7122 preferentially utilized d-glucose before d-xylose, and then finally utilized l-arabinose.
Conclusions
We successfully isolated a novel strain that fermented both d-xylose and l-arabinose. The results of taxonomical and physiological analyses revealed that NY7122 is a closely related species or subspecies of C. subhashii. To our knowledge, this is the first report on ethanol production by a species closely related to C. subhashii. NY7122 has good potential for use in ethanol production from lignocellulose-related sugars. Moreover, even at high temperature (37°C), NY7122 produces ethanol from hexose and pentose. NY7122 thus shows promise as a candidate yeast for bioethanol production under circumstances when pentose-rich hydrolysate is used as a fermentation substrate and the temperature cannot be tightly controlled. During the industrial-scale ethanol production from lignocellulosic hydrolysates, there are many environmental stresses such as ethanol, osmotic pressure, and weak organic acid. Most stress factors are synergistic [2]. Therefore, such stress tolerance of NY7122 should be clear. Furthermore, the metabolic profile of NY7122 in regard to pentose metabolism might provide a source of genes for improving pentose fermentation.
References
Adam H, Groenewald M, Mohan S, Richardson S, Bunn U, Gibas CF, Poutanen S, Sigler L (2009) Identification of a new species, Candida subhashii, as a cause of peritonitis. Med Mycol 47:305–311
Aldiguier AS, Alfenore S, Cameleyre X, Goma G, Uribelarrea JL, Guillouet SE, Molina-Jouve C (2004) Synergistic temperature and ethanol effect on Saccharomyces cerevisiae dynamic behaviour in ethanol bio-fuel production. Bioprocess Biosyst Eng 26:217–222
Amartey S, Jeffries TW (1994) Comparison of corn steep liquor with other nutrients in the fermentation of d-xylose by Pichia stipitis CBS 6054. Biotechnol Lett 19:211–214
Brown AD (1978) Compatible solutes and extreme water stress in eukaryotic micro-organisms. Adv Microb Physiol 17:181–242
Converti A, Dominguez JM (2001) Influence of temperature and pH on xylitol production from xylose by Debaryomyces hansenii. Biotechnol Bioeng 75:39–45
Dien BS, Kurtzman CP, Saha BC, Bothast RJ (1996) Screening for l-arabinose fermenting yeasts. Appl Biochem Biotechnol 57–58:233–242
Du Preez JC, Prior BA, Monteiro AMT (1984) The effect of aeration on xylose fermentation by Candida shehatae and Pachysolen tannophilus. Appl Microbiol Biotechnol 19:261–266
Du Preeze JC, Bosch M, Prior BA (1986) The fermentation of hexose and pentose sugars by Candida shehatae and Pichia stipitis. Appl Biochem Biotechnol 23:228–233
Du Preeze JC, Bosch M, Prior BA (1987) Temperature profiles of growth and ethanol tolerance of the xylose-fermenting yeasts Candida shehatae and Pichia stipitis. Appl Microbiol Biotechnol 25:521–525
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Girio FM, Fonseca C, Carvalheiro F, Duarte LC, Marques S, Bogel-Lukasik R (2010) Hemicelluloses for fuel ethanol: a review. Bioresour Technol 101:4775–4800
Grootjen DRJ, van der Lans RGJM, Luyben AM (1990) Effects of the aeration rate on the fermentation of glucose and xylose by Pichia stipitis CBS 5773. Enzyme Microb Technol 12:20–23
Kadam KL, Newman MM (1997) Development of a low-cost fermentation medium for ethanol production from biomass. Appl Microbiol Biotechnol 47:625–629
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Knoshaug EP, Franden MA, Stambuk BU, Zhang M, Singh A (2009) Utilization and transport of l-arabinose by non-Saccharomyces yeasts. Cellulose 16:729–741
Kurtzman CP, Dien BS (1998) Candida arabinofermentans, a new l-arabinose fermenting yeast. Antonie Van Leeuwenhoek 74:237–243
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331–371
Kurtzman CP, Suzuki M (2010) Phylogenetic analysis of the ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces and Scheffersomyces. Mycoscience 51:2–14
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A 82:6955–6959
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and clustal X version 2.0. Bioinformatics 23:2947–2948
Matsushika A, Inoue H, Kodaki T, Sawayama S (2009) Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives. Appl Microbiol Biotechnol 84:37–53
McMillan JD, Boynton BL (1994) Arabinose utilization by xylose-fermenting yeasts and fungi. Appl Biochem Biotechnol 45–46:569–584
Perriere G, Gouy M (1996) WWW-query: an on-line retrieval system for biological sequence banks. Biochimie 78:364–369
Saha BC (2003) Hemicellulose bioconversion. J Ind Microbiol Biotechnol 30:279–291
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schell DJ, Farmer J, Newman M, McMillan JD (2003) Dilute-sulfuric acid pretreatment of corn stover in pilot-scale reactor: investigation of yields, kinetics, and enzymatic digestibilities of solids. Appl Biochem Biotechnol 105–108:69–85
Skoog K, Hahn-Hagerdal B, Degn H, Jacobsen JP, Jacobsen HS (1992) Ethanol reassimilation and ethanol tolerance in Pichia stipitis CBS 6054 as studied by C nuclear magnetic resonance spectroscopy. Appl Environ Microbiol 58:2552–2558
Toivola A, Yarrow D, van den Bosch E, van Dijken JP, Scheffers WA (1984) Alcoholic fermentation of d-xylose by yeasts. Appl Environ Microbiol 47:1221–1223
Van Vleet JH, Jeffries TW (2009) Yeast metabolic engineering for hemicellulosic ethanol production. Curr Opin Biotechnol 20:300–306
White TJ (1990) Amplification and direct sequencing of fungal ribosomal DNA for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, CA, pp 315–322
Wisselink HW, Toirkens MJ, Wu Q, Pronk JT, van Maris AJ (2009) Novel evolutionary engineering approach for accelerated utilization of glucose, xylose, and arabinose mixtures by engineered Saccharomyces cerevisiae strains. Appl Environ Microbiol 75:907–914
Zhu HY, Xu H, Dai XY, Zhang Y, Ying HJ, Ouyang PK (2010) Production of d-arabitol by a newly isolated Kodamaea ohmeri. Bioprocess Biosyst Eng 33:565–571
Acknowledgments
This work was supported by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Rural Biomass Research Project, BEC-BC050).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Watanabe, I., Ando, A. & Nakamura, T. Characterization of Candida sp. NY7122, a novel pentose-fermenting soil yeast. J Ind Microbiol Biotechnol 39, 307–315 (2012). https://doi.org/10.1007/s10295-011-1033-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-011-1033-5