Abstract
The composition of the cellulase system in the cellulosome-producing bacterium, Clostridium thermocellum, has been reported to change in response to growth on different carbon sources. Recently, an extensive carbohydrate-sensing mechanism, purported to regulate the activation of genes coding for polysaccharide-degrading enzymes, was suggested. In this system, CBM modules, comprising extracellular components of RsgI-like anti-σ factors, were proposed to function as carbohydrate sensors, through which a set of cellulose utilization genes are activated by the associated σI-like factors. An extracellular module of one of these RsgI-like proteins (Cthe_2119) was annotated as a family 10 glycoside hydrolase, RsgI6-GH10, and a second putative anti-σ factor (Cthe_1471), related in sequence to Rsi24, was found to contain a module that resembles a family 5 glycoside hydrolase (termed herein Rsi24C-GH5). The present study examines the relevance of these two glycoside hydrolases as sensors in this signal-transmission system. The RsgI6-GH10 was found to bind xylan matrices but exhibited low enzymatic activity on this substrate. In addition, this glycoside hydrolase module was shown to interact with crystalline cellulose although no hydrolytic activity was detected on cellulosic substrates. Bioinformatic analysis of the Rsi24C-GH5 showed a glutamate-to-glutamine substitution that would presumably preclude catalytic activity. Indeed, the recombinant module was shown to bind to cellulose, but showed no hydrolytic activity. These observations suggest that these two glycoside hydrolases underwent an evolutionary adaptation to function as polysaccharide binding agents rather than enzymatic components and thus serve in the capacity of extracellular carbohydrate sensors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clostridium thermocellum is a cellulolytic Gram-positive bacterium that serves as a model organism for plant cell wall degradation by virtue of its multi-enzyme cellulosome complex [1–3]. The central component of the cellulosome is the nonhydrolytic “scaffoldin” subunit that integrates the various catalytic subunits into the complex via high-affinity protein–protein interactions between the multiple copies of scaffoldin-borne “cohesin” modules and a complementary “dockerin” module borne by each catalytic subunit. The C. thermocellum scaffoldin can thus integrate nine different catalytic subunits per complex, although the genome encodes for more than 70 different dockerin-containing components. The targeting of the cellulosome to its cellulosic substrate is mediated by a carbohydrate-binding module (CBM) that comprises part of the scaffoldin. Carbohydrate degradation by this bacterium may also be carried out by free or cell-bound carbohydrate-degrading enzymes that lack dockerins, although the great majority of plant cell wall degrading enzymes found in this bacterium are clearly cellulosomal in nature.
Plant cell walls consist of several intertwined heterogeneous polymers, primarily cellulose, hemicellulose (substituted xylans, mannans, etc.), pectin, and lignin. Therefore, the action of several enzymes with diverse catalytic activities is needed in order to unravel and break down this inherently intricate polymer network in an efficient manner. Indeed, previous studies have demonstrated that the enzyme composition in the cellulosome changes in response to growth on different carbon sources [4–6]. The latter studies indicate that decomposition of lignocellulosic substrates is governed by a coordinated substrate-specific regulation of cellulosomal subunit composition in C. thermocellum.
Recently, systematic analysis of the C. thermocellum genome revealed multiple copies of putative σI- and RsgI-like proteins, some of which were predicted to be involved in regulatory mechanisms that control carbohydrate degradation processes including formation and function of the cellulosome complex [7]. Nine of the deduced proteins share strong similarity in their N-terminal sequences to the Bacillus subtilis membrane-integrated anti-σI factor RsgI. Eight of them appeared to form bicistronic operons downstream of genes, which encode proteins that bear strong similarity to the B. subtilis σI factor.
The RsgI-like proteins have an atypical domain organization. In five proteins, the C-terminal regions, presumed to be located outside the cell membrane, contain putative CBMs, three of which were classified as family 3b (CBM3b), which are known to bind crystalline cellulose. One (Cthe_1273) contains a putative arabinose-binding domain classified as CBM family 42 (CBM42), and another (Cthe_0316) includes two tandem PA14 motifs (pfam07691, smart00758), which together bind to pectin and related polysaccharides [7].
Of the remaining RsgI-like proteins, one of them, Cthe_2119 (RsgI6), contains a family-10 glycoside hydrolase (GH10) catalytic module (Fig. 1), known to hydrolyze mainly xylans (http://www.cazy.org/GH10.html) [8]. Upon additional mining of the C. thermocellum genome, we also discovered an unusual family-5 glycoside hydrolase (GH5) module associated with a deduced protein sequence (Cthe_1471), similar to a B. subtilis membrane-associated anti-σ factor. The prediction of such carbohydrate-active GH modules as a component of putative anti-σ factors may suggest that they also function as carbohydrate sensors that detect extracellular polysaccharide biomass components. In the present communication, we examine the carbohydrate-binding and enzymatic performance of these two GH modules and discuss their relevance as components of biomass sensors in the signal-transmission system of C. thermocellum.
Putative GH-containing biomass-sensing genes in C. thermocellum. a Genomic organization of sigI6-rsgI6 and sig24C-rsi24C operons. Numbers in parentheses indicate the size of the gene product in amino acid residues. b Comparison of modular structures of putative anti-sigma factors RsgI6 and Rsi24C. Each protein harbors an RsgI- or Rsi24-like domain (dark gray) containing an N-terminal transmembrane helix (black). The GH10 and GH5 modular components are indicated; UNK refers to a divergent domain of unknown function. The Rsi24-like domain contains a zinc-binding anti-sigma region (ZAS). Ruler indicates the length of the proteins (number of amino acids)
Materials and methods
Materials
Pectin (from apple), oat-spelt xylan, chitin (from crab shells), carboxymethyl cellulose, and polygalacturonic acid (from orange) were purchased from Sigma Chemical Co. (St. Louis, MO). Wheat arabinoxylan medium viscosity (L-arabinose, 38%, D-xylose, 62%) was purchased from Megazyme International, Ltd. (Wicklow, Ireland). Microcrystalline cellulose (Avicel) was purchased from FMC BioPolymer (Philadelphia, PA). Phosphoric acid-swollen cellulose (PASC) was prepared according to Lamed et al. [9]. GFP-CBM3a was prepared according to Demishtein et al. [10].
Bacterial strains
Clostridium thermocellum ATCC 27405 is referred to as a type and genome-sequenced strain. Alternatively, strain YS was also used in some experiments. Escherichia coli strain XL1-Blue (Stratagene, La Jolla, CA) was used for plasmid constructions, and strain BL21(DE3) (Novagen, Madison, WI) was used for protein expression via the T7 RNA polymerase system.
DNA manipulation and cloning procedures
DNA sequences encoding GH10 (Cthe_2119, residues 378–760) and GH5 (Cthe_1471, residues 114–561) regions were amplified by PCR using C. thermocellum ATCC 27405 genomic DNA as template, Phusion™ high-fidelity DNA polymerase (Finnzymes Oy, Espoo, Finland) and the following primers. For GH10 (Cthe_2119): 5′-AATTCCATGGGCATTCAATGGATTGACCAGGC-3′ (NcoI site is underlined) and 5′-AATTCTCGAGGGGAATTCTGTAAGTAGTCTGCAATG-3′ (XhoI site is underlined), and For GH5 (Cthe_1471): 5′-AATTCCATGGGTTTTGACAATCTCGGCAACTGGATTG-3′ (NcoI site is underlined) and 5′-AATTCTCGAGTTTCCAAAAATTTTGCAATGTTTCAAG-3′ (XhoI site is underlined). The genes of the desired modules were then ligated into NcoI-XhoI linearized-pET28a vectors, thus encoding an attached His-tag for subsequent purification of the gene product. The DNA plasmids were transformed into E. coli XL1-Blue, purified using Qiaprep spin Miniprep Kit (Qiagen, Alameda, CA) and verified by DNA sequencing.
Protein expression and purification
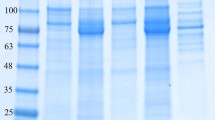
Cultures of E. coli BL21(DE3) cells containing the above plasmids were grown at 37°C in LB medium, supplemented with kanamycin (50 μg/ml) to an A600 = 0.5–0.6. Then, isopropyl β-D-1-thiogalactopyranoside (IPTG) was added to a final concentration of 0.1 mM and the culture was subjected to additional incubation at 16°C, overnight. The cells were harvested, resuspended in Tris-buffered saline (TBS) (137 mM NaCl, 2.7 mM KCl, and 25 mM Tris-HCl; pH 7.4) and sonicated in the presence of 1 mM phenylmethylsulfonyl fluoride (PMSF) and 10 μg/ml DNase. The sonicate was heated for 30 min at 60°C and then centrifuged. The supernatant fluids were mixed with 5 ml packed Ni-nitrilotriacetic acid (NTA) beads supplemented with 5 mM imidazole for 1 h at 4°C (batch purification system). The beads were then loaded on a 20-ml Econo-pack column, washed by using gravity flow with 50 to 100 ml wash buffer (TBS with 15 mM imidazole), and eluted with TBS containing 100 mM imidazole and then 250 mM imidazole. The purity of the recombinant proteins was tested by SDS-PAGE on 12% or 15% acrylamide gels. Highly purified protein fractions were merged, dialyzed against TBS buffer, and concentrated using Millipore 3,000 MW cutoff concentrators (Millipore, Billerica, MA). Protein concentrations were estimated by using the absorbance at 280 nm. The extinction coefficient was determined based on the known amino acid composition of each protein using the ProtParam tool on the ExPASy server (http://www.expasy.org/tools/protparam.html). Proteins were stored in 50% (vol/vol) glycerol at −20°C.
Enzymatic activity
Xylanase activity was determined quantitatively by measuring the reducing sugars released from soluble arabinoxylan and insoluble oat-spelt xylan by the dinitrosalicyclic acid (DNS) method [11]. A typical assay mixture consisted of 1 μM enzyme in reaction buffer (50 mM sodium acetate pH 5.5, 12 mM CaCl2) and 1% soluble arabinoxylan in a final volume of 200 μl. The reaction was performed in triplicate for 20 min at 50°C and was terminated by transferring the tubes to ice. An aliquot (100 μl) of the reaction mixture was then added to 150 μl of DNS reagent, and the tubes were boiled for 10 min, after which absorbance was measured at 540 nm. Thermobifida fusca Xyn10A (kind gift of Sarah Moraïs, Weizmann Institute) was used as a positive control for these experiments. Michaelis constant (K m) and maximal rate (V max) were determined using arabinoxylan concentrations of 0.2, 0.5, 0.8, 1, and 1.5% (w/v) and 1 μM enzyme. These kinetic values were calculated from a plot of the initial reaction rates vs. substrate concentration. Values of K m and V max were determined from a Lineweaver-Burk plot of the data [12]. The results were expressed in nkat, where 1 nkat corresponds to the amount of enzyme necessary to release 1 nmol of reducing ends per second.
Xylanase activity on 1% insoluble oat-spelt xylan was measured as above at different enzyme concentrations using a xylose standard curve.
Carbohydrate binding assay
Qualitative assessment of binding to the insoluble polysaccharides was achieved using Avicel, oat-spelt xylan, chitin (from crab shells), pectin and polygalacturonic acid. Insoluble xylan was pretreated as described before [13]. Pectin and polygalacturonic acid were immersed in buffer containing 7 mM CaCl2 in order to precipitate the polysaccharides. Each assay mixture consisted of 3 μg of the purified protein in binding buffer (50 mM Tris-HCl pH 8.0, 300 mM NaCl, 0.05% Tween 20) and 5 mg polysaccharide in a final volume of 200 μl. The mixture was incubated at 4°C for 30 min with gentle agitation and then centrifuged at 13,000 rpm for 5 min. The supernatant containing the unbound protein was separated and kept on ice. The pellet was washed three times with 800 μl binding buffer for 10 min to reduce nonspecific binding. Protein sample buffer (60 μl, containing SDS and DTT) was added to the polysaccharide pellet and then boiled for 10 min to dissociate any bound protein. Bound and unbound fractions were analyzed by SDS-PAGE using a 12% or 15% SDS polyacrylamide gel.
Affinity gel electrophoresis
Qualitative assessment of binding to soluble arabinoxylan was performed by loading 7 μg of the recombinant proteins onto a 7.5% polyacrylamide gel containing 0.1% arabinoxylan and comparing the mobility of each protein to a native gel without arabinoxylan. Non-denaturing conditions (absence of SDS) were used throughout the procedure.
Bioinformatic analysis
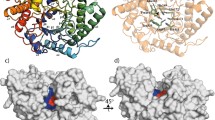
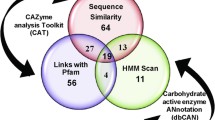
Sequence analysis of the GH modules in putative C. thermocellum anti-sigma factors Cthe_1471 and Cthe_2119 was based on their similarity to protein families established at both the CAZy [8] (http://www.cazy.org) and Pfam [14] (http://pfam.sanger.ac.uk) databases. The SWISS-MODEL Repository (http://swissmodel.expasy.org/repository/) was used to more precisely map amino acid sequences resembling either GH5 or GH10 domains in all available 3D structures presented in PDB (http://www.pdb.org/pdb/home/home.do). Amino acid residues 424–685 of Cthe_2119 (RsgI6) were shown to be strongly related (identity >30%) to certain PDB entries representing the 3D structures of GH10 proteins (1TA3, 1ISZ, 1IT0, 1XYF, 1ISV etc). A C-terminal module of Cthe_1471 (Rsi24C), residues 134–536, was shown to be related to the PDB entry 1EQC (the Candida albicans family 5 glycoside hydrolase). The Pfam and InterPro [15] (http://www.ebi.ac.uk/interpro/) databases annotated a C-terminal module of Cthe_1471 as PF00150 Cellulase (residues 158–484) and as IPR001547 Glyco_hydro_5 (residues 190–480), respectively. Multiple sequence alignments were performed with the ClustalW program [16].
The following GenBank accession numbers were used to construct the alignments: for GH5 proteins—YP_001037225, ACM60954, BAA25878, YP_001037893, AAA23230, BAA14354, AAG45159, AAA23231, BAA12826, YP_001036965, BAA32286, AAC09379, AAD36302 and BAE44526; and for GH10 proteins—YP_001038519, ABN53181, YP_001037339, YP_001038983, CAA84631, AAK76861, AAQ83581, AAT37531, AAZ56824, ACE84815, CBA13561 and AAA56791.
Results
Binding of RsgI6-GH10 to xylan
According to the CAZy Web site and additional resources (e.g., Pfam, InterPro, and PROSITE), the extracellular C-terminal region of Cthe_2119 (RsgI6) is annotated as a glycoside hydrolase family 10 [7], which contains enzymes that primarily exhibit xylanase activity [8, 17, 18]. Hence, the interaction of the purified 283-residue recombinant GH10 module (RsgI6-GH10, residues 378–760) was examined with xylan matrices. For this purpose, we first employed a gel-retardation binding assay system, using soluble arabinoxylan as a target substrate. As revealed from the affinity gel in Fig. 2a, in the presence of the substrate, the electrophoretic mobility of the GH10 module was retarded in the non-denaturing gel relative to its mobility in the absence of target polysaccharide. This shift in protein migration on the gel indicates that the module binds soluble arabinoxylan. Similar binding of RsgI6-GH10 to insoluble oat-spelt xylan was demonstrated using a centrifugation-based assay system (Fig. 2b). The protein also bound to Avicel, but not to chitin or pectin.
Binding of RsgI6-GH10 to xylan matrices. a Interaction of RsgI6-GH10 with a soluble xylan derivative. The protein was subjected to non-denaturing gel electrophoresis in the presence or absence of 0.1% arabinoxylan (±AX). BSA was used as a negative control. Note the reduced mobility of the protein band (asterisks) in the presence of substrate. b Interaction of RsgI6-GH10 with an insoluble oat-spelt xylan matrix. The recombinant RsgI6-GH10 was incubated with the indicated insoluble polysaccharides, centrifuged, and the bound (+) and unbound (–) fractions were analyzed by SDS-PAGE
Enzymatic activity of RsgI6-GH10
Since RsgI6-GH10 exhibited an interaction with xylan matrices as was anticipated, we were further interested to test its enzymatic capabilities on this substrate. GH10 xylanases are retaining enzymes, liberating the product with an overall retention in the anomeric configuration. Retaining glycosidases utilize a two-step double-displacement mechanism involving two key active site acidic residues, one functioning as the nucleophile and the other as the acid-base catalyst. In the case of xylanases, only glutamic acid residues have been identified as the catalytic amino acids that promote the hydrolysis [19–21]. Therefore, we examined the Cthe_2119 sequence for the presence of these two catalytic residues, using multiple sequence alignment analysis of the RsgI6-GH10 module with the 502 available bacterial sequences of the GH10 family. The two conserved glutamate residues were found at positions 538 and 635 (Fig. 3), as the acid-base catalyst and catalytic nucleophile, respectively [22, 23]. Consequently, RsgI6-GH10 would be expected to be functional.
Sequence conservation of two distinctive portions of the GH10 modules from various representative enzymes. Putative catalytic residues are marked in bold font. GenBank accession numbers used to construct the alignment are given in the Materials and methods section
The enzymatic activity of RsgI6-GH10 was tested on soluble arabinoxylan and on insoluble oat-spelt xylan and was found to be active on both substrates. The activity of RsgI6-GH10 was examined on different concentrations of soluble substrate, which enabled calculation of the kinetic constants. The K m was calculated to be 2.9 mg/ml and the V max was 43.5 nkat/mg protein. The RsgI6-GH10 module showed no detectable activity on carboxymethyl cellulose, phosphoric acid-swollen cellulose or microcrystalline cellulose (Avicel).
Identification and genomic organization of the sig24 and rsi24 genes in C. thermocellum
Previous bioinformatic analysis by Kahel-Raifer and coworkers [7] revealed for the first time the unique set of σI- and RsgI-like proteins in C. thermocellum, and their presence in the genome may implicate a major regulatory role in the degradation of plant cell wall biomass. More extensive analysis of the C. thermocellum genome databases revealed several additional predicted operons that encode pairs of putative alternative sigma factors together with cognate transmembrane anti-sigma-like factors. These alternative sigma factors are weakly similar to σW, a subfamily of the B. subtilis extracytoplasmic function (ECF) sigma factors, and to E. coli σ24. Consequently, we hereby assign the common prefix sig24 for C. thermocellum genes encoding putative ECF sigma factors, and the prefix rsi24 for their cognate anti-sigma factors. However, only one of these operons contains a gene, Cthe_1471, which encodes a protein, herein termed Rsi24C, that includes a recognizable carbohydrate-active module, which shares similarity to GH family 5 (Fig. 1). All of the other rsi24 genes encode proteins with C-terminal regions that have no significant similarity to other sequences in the known databases.
Binding of Rsi24C-GH5 to insoluble polysaccharides
Glycoside hydrolase family 5 (GH5) is one of the largest and most diverse families of glycoside hydrolases [24, 25]. GH5 enzymes are known to hydrolyze various substrates, including cellulose, mannan, xylan, chitin, glucan, and lichenin [17]. In order to predict the substrate specificity of Rsi24C-GH5, its amino acid sequence was compared with other members of the GH5 family using Clustal analysis. This protein showed low sequence similarity with other GH5 enzymes but did not cluster into one of the known sub-groups of this family (e.g., endoglucanase, xylanase, mannanase). Interestingly, in the CAZy database, this Cthe_1471 module is not annotated as a glycoside hydrolase, in contrast to both Pfam and InterPro databases (PF00150: Cellulase and IPR001547: Glyco_hydro_5, respectively).
The 448-residue recombinant GH5 module (114–561) was cloned and expressed, and its binding properties were tested against various insoluble carbohydrates. The results shown in Fig. 4 demonstrate relatively weak but reproducible interaction with crystalline cellulose but shows little (polygalactauronic acid) or no interaction with any of the other examined polysaccharides (oat-spelt xylan, pectin, and chitin). The binding of Rsi24C-GH5 to cellulosic substrates was weak, compared to a CBM3-containing chimeric protein (GFP-CBM3a) used as a positive control (data not shown).
Enzymatic performance of Rsi24C-GH5
Family 5 glycoside hydrolases catalyze the hydrolysis of different target substrates (cellulose, xylan, mannan) with net retention of configuration at the anomeric center [19, 26]. Both the acid/base catalyst and the stabilizing anion/nucleophile residues that catalyze the hydrolysis of the glycosidic bond have been identified experimentally within enzymes of this family. An Asn-Glu-Pro motif distinguishes the acid catalyst, while a Glu-X-Gly sequence, where X is typically an aromatic amino acid residue, characterizes the Glu participating as the stabilizing nucleophile.
Before performing an enzymatic test to investigate the function of the Rsi24C-GH5 module, we analyzed the prospective position of the conserved glutamic acid residues in its sequence. Multiple sequence alignment analysis of the Rsi24C-GH5 sequence with 1357 Pfam available bacterial sequences of the GH family 5 revealed the catalytic motifs depicted above as Asn-Gln-Pro in positions 314–316 and Glu-Tyr-Gly in positions 443–445 (Fig. 5). This extremely conserved pattern of catalytic residues, although largely maintained in our GH5 module, contains a curious Glu-to-Gln replacement (position 315) of the general acid/base catalyst Glu (position 315) into Gln. This switch in the conserved Glu is the only one found throughout the alignment among the entire collection of GH5 sequences that maintained this pattern of catalytic site. This observation implies that the putative enzyme may have lost its hydrolytic function.
Sequence conservation of the regions containing the catalytic residues of GH5 modules from various representative enzymes. Putative catalytic residues are marked in bold font. Note the E315Q substitution in the position of the catalytic acid of Rsi24C-GH5 (arrow). GenBank accession numbers used to construct the alignment are given in the Materials and methods section
In order to clarify this issue, the enzymatic activity of the purified recombinant GH5 module was tested on various polysaccharides. The degradation of crystalline cellulose (Avicel), amorphous cellulose, carboxymethyl cellulose, oat-spelt xylan, chitin, pectin, and polygalactauronic acid was determined after 20 min at 50°C using the reducing sugar method. Under these conditions, no degradation of any of these carbohydrates was detected, even for cellulose to which this putative enzyme was shown to bind (Fig. 4).
Discussion
The process of plant cell wall degradation either by free cellulolytic enzymes or by cellulosomes begins with a binding interaction to the component polysaccharides via appropriate CBMs [27]. Based on their amino acid similarity, CBMs are currently grouped into 59 families that show notable variation in substrate specificity [8]. For example, CBMs can bind specifically to cellulose, xylan, chitin, pectin, β-glucans, starch and many others polysaccharides, such as mannan, arabinan, and polygalacturonic acid. The most recognized function of these domains is to bind polysaccharides, bringing the biocatalyst into close and prolonged proximity with its substrate, thereby allowing efficient carbohydrate hydrolysis. However, CBMs are also important in events related to metabolism, pathogen defense, polysaccharide biosynthesis, virulence, plant development, etc., as reviewed in Guillén et al. [28].
Recently, a new role for CBMs as extracellular carbohydrate sensors was suggested by Kahel-Raifer et al. [7]. In this case, CBM-annotated modules were found at the C-terminal end of several membrane-associated anti-σI factors, predicted to be localized and to act outside the cell membrane. The N-terminal portion of these proteins was expected to be on the cytoplasmic side of the membrane and to interact with σI-like factors (i.e., alternative σ factors that control specialized regulon activation), such that the activated genes would encode proteins involved in carbohydrate utilization. Accordingly, a novel carbohydrate-sensing mechanism was proposed, whereby the presence of polysaccharide biomass components is detected by a corresponding RsgI-borne CBM extracellularly and the signal is transferred intracellularly by which the σI is released from the RsgI-like subdomain and, together with RNA polymerase, transcribes the target genes.
One of the nine putative RsgI-like proteins, Cthe_2119 (RsgI6), contains a predicted glycoside hydrolase family 10 instead of a CBM. This GH10 module was studied here for its relevance in this system, since its major function is to catalyze the hydrolysis of xylan. Indeed, RsgI6-GH10 both binds to and hydrolyzes insoluble and soluble xylan substrates. Although the K m was similar to previously published values for GH10 enzymes on arabinoxylan [29–31], the V max value was very low, exhibiting between 0.1–10% of the activity of other characterized xylanases, from C. thermocellum [31, 32], as well as from Aspergillus aculeatus, Bacillus subtilis, Geobacillus stearothermophilus, and Themobacillus xylanilyticus [29, 30, 33]. This indicates that the affinity for the soluble xylan substrate was similar to those of a typical GH10 enzyme, but the hydrolysis of the substrate was limited. Moreover, RsgI6-GH10 also binds to cellulose, although it fails to hydrolyze cellulosic substrates. Another glycoside hydrolase from family 5 was revealed at the C-terminal region of Cthe_1471, which is similar in its N-terminal sequence to a B. subtilis membrane-associated anti-σ factor (RsiW, a negative regulator of the ECF family 24 sigma factor σw). This prospective enzymatic module, however, completely lost its catalytic function, presumably due to replacement of the catalytic acid, which is essential for the hydrolysis. Nevertheless, the binding function to microcrystalline cellulose of this module was preserved.
These observations suggest an evolutionary adaptation of the above-described glycoside hydrolases to function as CBMs in polysaccharide binding, due to their extracellular localization in proteins that play a role in regulating enzyme expression for polysaccharide utilization by the parent cells. In this context, the hydrolytic function appears not to be crucial. These modified glycoside hydrolases, together with the other CBMs in this extensive biosensing system, promote various binding specificities for different plant cell wall polysaccharides, and the specificity is maintained in the cognate σ-like factors, which together serve to activate different sets of carbohydrate-active enzymes, located at various loci on the C. thermocellum genome.
References
Lamed R, Setter E, Bayer EA (1983) Characterization of a cellulose-binding, cellulase-containing complex in Clostridium thermocellum. J Bacteriol 156:828–836
Lamed R, Setter E, Kenig R, Bayer EA (1983) The cellulosome—a discrete cell surface organelle of Clostridium thermocellum which exhibits separate antigenic, cellulose-binding and various cellulolytic activities. Biotechnol Bioeng Symp 13:163–181
Bayer EA, Belaich J-P, Shoham Y, Lamed R (2004) The cellulosomes: multi-enzyme machines for degradation of plant cell wall polysaccharides. Annu Rev Microbiol 58:521–554
Dror TW, Rolider A, Bayer EA, Lamed R, Shoham Y (2005) Regulation of major cellulosomal endoglucanases of Clostridium thermocellum differs from that of a prominent cellulosomal xylanase. J Bacteriol 187:2261–2266
Gold ND, Martin VJ (2007) Global view of the Clostridium thermocellum cellulosome revealed by quantitative proteomic analysis. J Bacteriol 189:6787–6795
Raman B, Pan C, Hurst GB, Rodriguez M, McKeown CK, Lankford PK, Samatova NF, Mielenz JR (2009) Impact of pretreated Switchgrass and biomass carbohydrates on Clostridium thermocellum ATCC 27405 cellulosome composition: a quantitative proteomic analysis. PLoS ONE 4:e5271
Kahel-Raifer H, Jindou S, Bahari L, Nataf Y, Shoham Y, Bayer EA, Borovok I, Lamed R (2010) The unique set of putative membrane-associated anti-σ factors in Clostridium thermocellum suggests a novel extracellular carbohydrate-sensing mechanism involved in gene regulation. FEMS Microbiol Lett 308:84–93
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The Carbohydrate-Active Enzymes database (CAZy): an expert resource for glycogenomics. Nucl Acids Res 37:D233–D238
Lamed R, Kenig R, Setter E, Bayer EA (1985) Major characteristics of the cellulolytic system of Clostridium thermocellum coincide with those of the purified cellulosome. Enzyme Microb Technol 7:37–41
Demishtein A, Karpol A, Barak Y, Lamed R, Bayer EA (2010) Characterization of a dockerin-based affinity tag: application for purification of a broad variety of target proteins. J Mol Recogit (in press)
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Biochem 31:426–428
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56:658–666
Moraïs S, Barak Y, Caspi J, Hadar Y, Lamed R, Shoham Y, Wilson DB, Bayer EA (2010) Contribution of a xylan-binding module to the degradation of a complex cellulosic substrate by designer cellulosomes. Appl Environ Microbiol 76:3787–3796
Finn RD, Tate J, Mistry J, Coggill PC, Sammut SJ, Hotz H-R, Ceric G, Forslund K, Eddy SR, Sonnhammer ELL, Bateman A (2008) The Pfam protein families database. Nucl Acids Res 36:D281–D288
Hunter S, Apweiler R, Attwood TK, Bairoch A, Bateman A, Binns D, Bork P, Das U, Daugherty L, Duquenne L, Finn RD, Gough J, Haft D, Hulo N, Kahn D, Kelly E, Laugraud A, Letunic I, Lonsdale D, Lopez R, Madera M, Maslen J, McAnulla C, McDowall J, Mistry J, Mitchell A, Mulder N, Natale D, Orengo C, Quinn AF, Selengut JD, Sigrist CJA, Thimma M, Thomas PD, Valentin F, Wilson D, Wu CH, Yeats C (2009) InterPro: the integrative protein signature database. Nucl Acids Res 37:D211–D215
Higgins DG, Thompson JD, Gibson TJ (1996) Using CLUSTAL for multiple sequence alignments. Methods Enzymol 266:383–402
Coutinho PM, Henrissat B (1999) Carbohydrate-active enzymes: an integrated database approach. In: Gilbert HJ, Davies GJ, Henrissat B, Svensson B (eds) Recent advances in carbohydrate bioengineering. R Soc Chem, Cambridge, pp 3–12
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859
Henrissat B, Callebaut I, Fabrega S, Lehn P, Mornon J-P, Davies G (1995) Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases. Proc Natl Acad Sci USA 92:7090–7094
Henrissat B, Davies G (1997) Structural and sequence-based classification of glycoside hydrolases. Curr Opin Struct Biol 7:637–644
MacLeod AM, Lindhorst T, Withers SG, Warren RA (1994) The acid/base catalyst in the exoglucanase/xylanase from Cellulomonas fimi is glutamic acid 127: evidence from detailed kinetic studies of mutants. Biochemistry 33:6371–6376
Tull D, Withers SG, Gilkes NR, Kilburn DG, Warren RA, Aebersold R (1991) Glutamic acid 274 is the nucleophile in the active site of a “retaining” exoglucanase from Cellulomonas fimi. J Biol Chem 266:15621–15625
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Cottrell MT, Yu L, Kirchman DL (2005) Sequence and expression analyses of Cytophaga-like hydrolases in a western Arctic metagenomic library and the Sargasso Sea. Appl Environ Microbiol 71:8506–8513
Ducros V, Czjzek M, Belaich A, Gaudin C, Fierobe H-P, Belaich J-P, Davies GJ, Haser R (1995) Crystal structure of the catalytic domain of a bacterial cellulase belonging to family 5. Structure 3:939–949
Boraston AB, Bolam DN, Gilbert HJ, Davies GJ (2004) Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J 382:769–781
Guillén D, Sánchez S, Rodríguez-Sanoja R (2010) Carbohydrate-binding domains: multiplicity of biological roles. Appl Microbiol Biotechnol 85:1241–1249
Bonnin E, Daviet S, Sorensen JF, Sibbesen O, Goldson A, Juge N, Saulnier L (2006) Behaviour of family 10 and 11 xylanases towards arabinoxylans with varying structure. J Sci Food and Agr 86:1618–1622
Beaugrand J, Chambat G, Wong VW, Goubet F, Rémond C, Paës G, Benamrouche S, Debeire P, O’Donohue M, Chabbert B (2004) Impact and efficiency of GH10 and GH11 thermostable endoxylanases on wheat bran and alkali-extractable arabinoxylans. Carbohydr Res 339:2529–2540
Dias FM, Goyal A, Gilbert HJ, Jose AMP, Ferreira LM, Fontes CM (2004) The N-terminal family 22 carbohydrate-binding module of xylanase 10B of Clostridium themocellum is not a thermostabilizing domain. FEMS Microbiol Lett 238:71–78
Fernandes AC, Fontes CM, Gilbert HJ, Hazlewood GP, Fernandes TH, Ferreira LMA (1999) Homologous xylanases from Clostridium thermocellum: evidence for bi-functional activity, synergism between xylanase catalytic modules and the presence of xylan-binding domains in enzyme complexes. Biochem J 342:105–110
Khasin A, Alchanati I, Shoham Y (1993) Purification and characterization of a thermostable xylanase from Bacillus stearothermophilus T-6. Appl Environ Microbiol 59:1725–1730
Acknowledgments
This research was supported by grants from the United States-Israel Binational Science Foundation (BSF, 2005–186), Jerusalem, Israel, by the Israel Science Foundation (grant numbers 966/09 and 159/07), and by the Mizutani Foundation for Glycoscience. Y.S. holds the Erwin and Rosl Pollak Chair in Biotechnology at the Technion. E.A.B. is the incumbent of The Maynard I. and Elaine Wishner Chair of Bio-organic Chemistry at the Weizmann Institute of Science.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bahari, L., Gilad, Y., Borovok, I. et al. Glycoside hydrolases as components of putative carbohydrate biosensor proteins in Clostridium thermocellum . J Ind Microbiol Biotechnol 38, 825–832 (2011). https://doi.org/10.1007/s10295-010-0848-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-010-0848-9