Abstract
In 2004, Leuconostoc mesenteroides DRC was first used as a starter culture for achieving higher organoleptic effects in Korean kimchi manufacture. For a better understanding of starter growth in a mixed culture system, and for predicting starter predominance in kimchi, a monitoring system for the starter was established. The chloramphenicol resistance marker gene (cat) was randomly integrated into chromosomal DNA of L. mesenteroides DRC using a viral transposon and transposase. The DRC mutant, tDRC2, had a similar growth pattern to the host strain, with no major alteration in phenotypic characteristics. The mutant strain was inoculated into real kimchi, and monitoring of the starter population was successfully achieved. The overall predominance of Leuconostoc in kimchi inoculated with DRC followed the general growth pattern of this genus during kimchi fermentation. Our results also demonstrate the competitive ability of the DRC starter against Leuconostoc from natural flora, maintaining its predominance above 88% during the whole fermentation period. Based on this experiment, the random gene integration method using a transposon was shown to be of utility in transferring any commercial starter into a selectable and monitorable strain for simulation purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leuconostoc spp. are hetero-fermentative lactic acid bacteria that produce lactic acid, acetic acid, alcohol, CO2, and aromatic compounds such as diacetyl, acetoin, and C4 compounds [5]. Leuconostoc is also the predominant bacterial genus in fermented vegetables, including kimchi, sauerkraut, and pickles, and plays an important role in maintaining the quality of those products [9, 16, 20, 29]. In the manufacture of these fermented products, spontaneous fermentation usually results in variations in the sensory quality of the products; the use of a starter culture has been found to help standardize fermentation by controlling the microbial flora [30, 31].
When L. mesenteroides is inoculated as a starter for the manufacture of lactic acid-fermented foods, it is always mixed with natural flora including other Leuconostoc spp. in raw materials; therefore, traditional methods of bacterial enumeration, identification, and typing are insufficient to monitor the starter in such complex, mixed-strain microbial consortia in foods [12]. For clarifying starter growth in lactic acid-fermented foods and for predicting starter predominance during fermentation, starter monitoring in natural or inoculated flora is a primary requirement.
Specific molecular techniques have been developed to identify and quantify populations of microorganisms in complex environments, such as for gram-negative filamentous bacteria in sludge [32], Lactococci in milk [2], or sulfate-reducing bacteria in mixed cultures [1]. Microorganisms can be identified to strain level using DNA fingerprinting techniques such as RAPD-PCR, RFLP, ribotyping, and PCR-DGGE [11, 12, 24]; however, since these techniques are moderately specific and sensitive, they do not allow the detection of individual strains. Furthermore, they are limited by nucleic acid extraction from complex environments and factors limiting PCR efficiency.
Stable marker systems with an easily detectable phenotype provide an alternative strategy to detect microorganisms in complex environments. A few marker genes, such as antibiotic resistance genes, lacZ and lux [7, 8, 18, 26] have been used to detect and monitor microorganisms, including some lactic acid bacteria [10, 13, 28]. However, such specific markers have not yet been determined in L. mesenteroides except for the recent plasmid expression of gfp [22]. Transposable elements are accessible tools for molecular genetic studies when integrated randomly into chromosomes [15], and insertional mutagenesis has been successfully undertaken in gram-positive bacteria [34]. As a result, high levels of segregational instability in many engineered lactic acid bacteria cloning vectors have been circumvented by stably integrating heterologous genes into the chromosome [19].
In this study, we created a genetically stable L. mesenteroides strain by randomly integrating the chloramphenicol acetyltransferase gene (cat) into the chromosomal DNA. The predominance of the starter strain was monitored during kimchi fermentation using this transformant.
Materials and methods
Bacterial strains, plasmids, and culture conditions
Bacterial strains and plasmids used in this study are listed in Table 1. pEK104 [25] was constructed with pUC19 and the cat gene from species of Staphylococcus. Escherichia coli was grown in LB broth at 37 °C with shaking, and ampicillin was used when needed at a final concentration of 50 µg mL−1. L. mesenteroides DRC (KCCM 11318), a starter strain used in the manufacturing of kimchi, was used in this study for cat gene integration. DRC and its transformant were grown in MRS broth at 30 °C. A simulated liquid medium of kimchi (J medium) was prepared by autoclaving and filtering commercial kimchi (Chonggajip, Yongin, Korea).
Reagents and enzymes
Restriction enzymes were obtained from Promega (Madison, WI, USA) and TaKaRa (Kyoto, Japan). T4 DNA ligase, CALP (calf intestinal alkaline phosphatase), and Taq polymerase were purchased from TaKaRa, and pMOD-2<MCS> and EZ::TN transposase were from Epicentre (Madison, WI, USA). The vector contains multiple cloning sites (MCSs) between the hyperactive 19-bp mosaic ends (ME) that are specifically recognized by the transposase. The molecular weight standard DNA was purchased from Bioneer (Daejeon, Korea), and ampicillin and chloramphenicol were obtained from Sigma (St Louis, MO, USA). Lactobacilli MRS broth, LB broth, and phenyl ethyl ethanol agar were from Difco (Detroit, MI, USA). Other chemicals were of reagent grade. Oligonucleotide primers were synthesized with an automated DNA synthesizer by Bioneer. The sequences of oligonucleotides used in this study are listed in Table 1.
DNA techniques
Procedures for DNA manipulations were essentially as described in Sambrook et al. [27]. The chromosomal DNA of L. mesenteroides DRC was prepared according to the methods of the AccuPrep® Genomic DNA Extraction Kit (Bioneer) with minor modifications. All enzymes for DNA modification were used according to the manufacturer’s specifications. For amplification of the partial cat gene, genomic DNA of L. mesenteroides DRC and primers (Table 1) were used. The PCR amplification was carried out with a Mini Cycler™ (MJ Research, Waltham, MA, USA) under standard conditions (94 °C for 5 min, 30 cycles of 30 s at 94 °C, 30 s at 55 °C, and 1 min at 72 °C, with a final elongation at 72 °C for 5 min).
Electrotransformation procedure
Electrocompetent cells of L. mesenteroides DRC were prepared according to the method of Wyckoff and Sandine [33] with minor modifications. For preparation of stable transposomes, 2 µL of transposon DNA (0.2 µg) was incubated with 4 µL (4 units) of transposase in TE Buffer (10 mM Tris–HCl, pH 7.5, with 1 mM EDTA) and 2 µL glycerol in the absence of Mg2+ at room temperature for 30 min. Aliquots (1 µL) of transposomes were transformed by electroporation into competent cells of L. mesenteroides DRC using a Gene-Pulser unit combined with a Pulse Controller (Bio-Rad, Richmond, CA, USA). The electrocompetent cells (50 µL) were mixed with transforming DNA in a microfuge tube, transferred to cold electroporation cuvettes, and placed on ice for 5 min. The mixture was then given a single discharge of 8 kV cm−1, 25 uF, and 400 Ω. Cells were immediately resuspended in 1 mL of MRS broth and incubated for 1 h at 30 °C. The transformed cells (tDRC1 and tDRC2) were selected on MRS agar containing 10 µg mL−1 chloramphenicol after 2 days of incubation at 30 °C.
Preparation and analysis of kimchi
Kimchi was prepared by mixing salted Chinese cabbage with ingredients that included red pepper powder, garlic, ginger, and sugar [3]. Homogenized kimchi samples and broth were serially diluted tenfold with 0.85% sterilized saline. Each diluted solution was spread onto phenyl ethyl alcohol agar containing 2% sucrose (PES medium) for L. mesenteroides DRC, and on PES containing 10 µg mL−1 chloramphenicol for L. mesenteroides tDRC2 at 30 °C. The total LAB count was obtained on MRS agar at 37 °C. The pH of the test solution was determined with a pH meter (IQ240; IQ Scientific Instruments, San Diego, CA, USA). The titratable acidity was determined by titrating with 0.1 N NaOH solution until the pH of the test solution reached 8.3.
Results
Preparation of transposon fragment for integration of the cat gene
To construct a transposon vector (pM-cat), the cat gene in pEK104 was isolated by digestion with Pst I and Xba I, and ligated into MCSs of the pMOD-2 vector (Fig. 1a). The plasmid vector was transformed into E. coli MC1061 and clones were selected on LB agar containing 50 µg mL−1 ampicillin. From the clone, a 1,481-bp fragment of the transposon including CAT and ME (the mosaic end sequences) was obtained by Pvu II digestion (Fig. 1b). The 19-bp ME sites (the boxed sequences) were included to form a transposome (a synaptic complex) with higher binding ability to the EZ-Tn5 transposase.
Chromosomal integration of the transposon in L. mesenteroides
Formation of the transposome was achieved by mixing the transposon DNA obtained above with transposase and glycerol at room temperature. EZ-Tn5 transposase catalyzes a multistep “cut and paste” transposition reaction [14]. The transposase randomly attacks and cleaves the phosphodiester backbone of the target DNA and covalently links the 3′-OH ends of the transposon to the exposed 5′-phosphorylated ends of the target DNA, creating a nine-bp sequence that immediately duplicates the flanking transposon insertion site. For this, aliquots of transposomes were transformed by electroporation into competent cells of L. mesenteroides DRC under optimized conditions obtained in the previous experiment (described in the “Materials and methods”). After electroporation of the transposome, two colonies were chosen on agar plates containing chloramphenicol and were designated tDRC1 and tDRC2. To confirm integration of the cat gene into genomic DNA of L. mesenteroides DRC, PCR analysis was carried out using chromosomal DNAs of the two transformants and primers designed for the cat gene (CAT-For and CAT-Re, see sequences in Table 1). Figure 2 gives the PCR results from amplification of a part (667 bp) of the cat gene in both cell lines and proves the insertion of the cat gene in genomic DNA of L. mesenteroides DRC.
Confirmation of integration of the chloramphenicol acetyltransferase (cat) gene into genomic DNA of L. mesenteroides DRC by PCR. lane M, 1 kb ladder; lane C (+), positive control, PCR product of cat gene from pM-cat using primers CAT-For and CAT-Re, lane C (−), negative control, PCR product from genomic DNA of L. mesenteroides DRC using primers CAT-For and CAT-Re; lanes tDRC1 and tDRC2, PCR product of the cat gene from genomic DNA of L. mesenteroides tDRC 1 and 2 using primers CAT-For and CAT-Re, respectively
Growth of the CAT transposon-inserted DRC
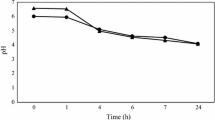
When the two transformants, tDRC1 and tDRC2, were cultivated in MRS medium at 30 °C, they showed growth rates (µ = 0.45 min−1) similar to their parental strain, DRC, but the viable cell counts of tDRC1 rapidly decreased at the late exponential phase, revealing its susceptibility in acidic medium conditions; hence, tDRC2 was used for subsequent experiments (data not shown). To elucidate the growth characteristics of tDRC2 during kimchi fermentation, it was inoculated into sterilized kimchi medium (J medium) and changes in microbial counts and pH were monitored at 10 °C (Fig. 3). Mean log counts of DRC and tDRC2 increased gradually after inoculation and reached a high level of 1 × 109 CFU mL−1 after 6 days; during the same period, the pH decreased to about pH 4. Overall, the two strains showed similar growth profiles for 10 days of fermentation, except for the slightly slower growth rate of the transformant in the exponential phase. Some microorganisms harboring plasmids to express heterologous proteins often resulted in a retarded growth than their host strains [6]. We can therefore assume that after random cat gene integration into the chromosomal DNA of the starter strain, the general physiological characteristics of the transformant were not greatly altered, at least in terms of growth rate and pH-related phenotype.
Growth profiles of Leuconostoc strains DRC and tDRC2 in J medium at 10 °C. Changes in Leuconostoc counts (filled square) and pH (filled circle) were monitored in J medium after addition of the DRC starter. The unfilled symbols refer to Leuconostoc counts and pH after adding the tDRC2 mutant. Symbols refer to the mean of data obtained from six duplicate samples
Monitoring of CAT+ L. mesenteroides (tDRC2) in kimchi
The transformant tDRC2 was next applied to monitor the growth of L. mesenteroides DRC starter during the fermentation of commercial kimchi (Fig. 4a). During kimchi fermentation at 10 °C for 20 days, bacterial counts were measured at intervals by plating diluted aliquots on MRS for total LAB. After addition of DRC or tDRC2 starters in two kimchi preparations, the viable counts of LAB in each sample were 3 × 105 and 4 × 105 CFUg−1 kimchi, respectively. In the control kimchi inoculated with DRC starter, the total LAB count was highest, 8 × 108 CFUg−1 kimchi, after 9 days and then decreased slowly to 2 × 108 CFUg−1 kimchi after 20 days. During this period, the pH changed from 6.3 to 4.3. In the sample kimchi inoculated with tDRC2, the overall total LAB count and pH change were generally same as those of the control. Although the changes in total count and pH were slightly slower with tDRC2, particularly in the exponential phase, after 9 days of incubation, the two cultures reached similar stationary levels, and the plateau values were stable for 20 days.
Monitoring of L. mesenteroides tDRC2 starter after inoculation into commercial kimchi. a Changes in total LAB counts (filled square) and pH (filled circle) were monitored in the control kimchi sample after adding the DRC starter at 10 °C. The unfilled symbols refer to LAB counts and pH in sample kimchi after adding the tDRC2 mutant. b Predominance of the starter was measured using tDRC2 counts among LAB and Leuconostoc as detailed below. Symbols refer to the mean of data obtained from six duplicate samples. Predominance among LAB (filled circle) = (CFU for tDRC2/CFU for total LAB) × 100 (%). Predominance among Leuconostoc (filled square) = (CFU for tDRC2/CFU for total Leuconostoc) × 100 (%)
Then, the competitive ability of the starter strain was investigated by measuring the predominance of tDRC2 in LAB as well as in Leuconostoc (Fig. 4b). During kimchi fermentation, bacterial counts were measured by plating diluted aliquots on MRS for total LAB and on PES for Leuconostoc. First, the predominance of L. mesenteroides tDRC2 among total LAB was measured by dividing CAT+ bacterial counts on chloramphenicol–PES plates by the total LAB counts. At day 0, 62% of the colonies were resistant to chloramphenicol in samples inoculated with strain tDRC2. Meanwhile, in the kimchi sample inoculated with host starter, DRC, none of the bacterial colonies grew on chloramphenicol–PES plates. At day 2, 82% of the colonies were resistant, but the levels were suddenly lowered to 32% at day 5 and they were maintained at 32, 17, and 23%, at days 9, 12, and 19, respectively. The nonresistant colonies were supposed to be species belonging to Lactobacillus or Leuconostoc in the natural flora. Many previous studies reported that the population number of Leuconostoc was highest from the initial to the middle stage of fermentation but it gradually decreased when the pH of kimchi was lowered to 4.0, after which a homo-fermentative type of lactic acid bacteria, Lactobacillus sp., and yeast having strong pH tolerance to high organic acid concentrations, continuously increased to the last stage [17, 23]. Therefore, our results obtained from the experiment using tDRC2 as a monitoring strain show that the overall predominance of DRC starter followed the general growth pattern of this genus during kimchi fermentation. Next, we analyzed the predominance of L. mesenteroides tDRC2 among Leuconostoc genera. When kimchi broth inoculated with tDRC2 was spread on the chloramphenicol–PES plates, up to 100% of colonies were resistant at 0 and 2 days. The apparent absence of nonresistant Leuconostoc colonies might reflect low contamination by indigenous flora of the raw material used in kimchi preparation. At day 5, 88% of colonies were resistant and the colony levels were maintained at 95, 90, and 93%, at days 9, 12, and 19, respectively. The nonresistant colonies, corresponding to Leuconostoc in the natural kimchi flora, were less than 10% during the kimchi distribution period and this result demonstrates the competitive ability of the DRC starter against the same species from natural flora.
Discussion
Kimchi is a general term given to a group of fermented vegetables made in Korea and traditionally served as side dishes with meals. Kimchi is mainly made of Chinese cabbage or radish, and differs in terms of raw ingredients, processing methods, season, and locality [21]. The consumption of factory-made kimchi is increasing and exports are also gradually rising [4]. However, spontaneous fermentation usually results in variations in the sensory quality of the products. Thus, in 2004, Chonggajip Kimchi Inc., the biggest kimchi producer in Korea, began to use a starter strain, L. mesenteroides DRC, in the manufactured kimchi after isolating a strain with superior organoleptic effects. In this study, we developed a laboratory-scale simulation system using a mutant strain inserted with cat gene into chromosomal DNA and it was proven that this mutant was useful to monitor the growth of the starter strain in kimchi product.
Plasmid transformation was frequently used for monitoring of starter strain in various fermented foods. In an experiment of starter monitoring on Enterococcus faecalis [10], a gfp fusion plasmid system was employed, but both plasmid loss and gradual elimination of the GFP+ strain by competitive flora were observed. The same result was also found in the experiment of fermented sausage with Lactobacillus sakei RV1040 [13]. In contrast, our results show that an integration of selection marker gene into the chromosome of Leuconostoc starter can circumvent the segregational instability of a plasmid system. To our knowledge, this is the first report of the successful expression of heterologous genes in Leuconostoc sp. using a random integration method. By using this mutant strain, population monitoring of the starter culture can be achieved in other fermented vegetables, including sauerkraut or pickles.
References
Amann RI, Binder BJ, Olson RJ, Chisholm SW, Devereux R, Stahl DA (1990) Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analysing mixed microbial populations. Appl Environ Microbiol 56:1919–1925
Beimfohr C, Krause A, Amann RI, Ludwig W, Schleifer KH (1993) In situ identification of Lactococci, Enterococci and Streptococci. Syst Appl Microbiol 16:450–456
Cho EJ, Park KY, Rhee SH (1997) Standardization of ingredient ratios of Chinese cabbage kimchi. Korean J Food Sci Technol 29:1228–1235
Choi IK, Jung SH, Kim BJ, Park SY, Kim J, Han HU (2003) Novel Leuconostoc citreum starter culture system for the fermentation of kimchi, a fermented cabbage product. Antonie Van Leeuwenhoek 84:247–253. doi:10.1023/A:1026050410724
Cogan TM, Jordan KN (1994) Metabolism of Leuconostoc bacteria. J Dairy Sci 77:2704–2717
Cooper NS, Brown ME, Caulcott CA (1987) A mathematical method for analysing plasmid stability in micro-organisms. J Gen Microbiol 133:1871–1880
Corthier G, Delorme C, Ehrlich SD, Renault P (1998) Use luciferase genes as biosensors to study bacterial physiology in the digestive tract. Appl Environ Microbiol 64:2721–2722
Duncan S, Glover LA, Killham K, Prosser JI (1994) Luminescence-based detection of activity of starved and viable but nonculturable bacteria. Appl Environ Microbiol 60:1308–1316
Eom HJ, Seo DM, Han NS (2007) Selection of psychrotrophic Leuconostoc spp. producing highly active dextransucrase from lactate fermented vegetables. Int J Food Microbiol 10:61–67. doi:10.1016/j.ijfoodmicro.2007.02.027
Geoffroy MC, Guyard C, Quatannens B, Pavan S, Lange M, Mercenier A (2000) Use of green fluorescent protein to tag lactic acid bacterium strains under development as life vaccine vectors. Appl Environ Microbiol 66:383–391
Giraffa G, Neviani E (2000) Molecular identification and characterisation of food-associated Lactobacilli. Ital J Food Sci 12:403–423
Giraffa G, Rossetti L (2004) Monitoring of the bacterial composition of dairy starter cultures by RAPD-PCR. FEMS Microbiol Lett 237:133–138. doi:10.1111/j.1574-6968.2004.tb09688.x
Gory L, Montel MC, Zagorec M (2001) Use of green fuorescent protein to monitor Lactobacillus sakei in fermented meat products. FEMS Microbiol Lett 194:127–133. doi:10.1111/j.1574-6968.2001.tb09457.x
Goryshin IY, Reznikoff WS (1998) Tn5 in vitro transposition. J Biol Chem 273:7367–7374. doi:10.1074/jbc.273.13.7367
Hamer L, DeZwaan TM, Montenegro-Chamorro MV, Frank SA, Hamer JE (2001) Recent advances in large-scale transposon mutagenesis. Curr Opin Chem Biol 5:67–73. doi:10.1016/S1367-5931(00)00162-9
Han NS, Jung YS, Eom HJ, Koh YH, Robyt JF, Seo JH (2002) Simultaneous biocatalytic synthesis of panose during lactate fermentation in kimchi. J Microbiol Biotechnol 12:46–52. doi:10.1159/000070151
Han HU, Lim CR, Park HK (1990) Determination of microbial community as an indicator of kimchi fermentation. Korean J Food Sci Technol 22:26–32
Hattemer-Frey HA, Brandt EJ, Travis CC (1990) Small-scale field test of the genetically engineered lacZY marker. Regul Toxicol Pharmacol 11:253–261. doi:10.1016/0273-2300(90)90025-7
Hols P, Ferain T, Garmyn D, Bernard N, Delcour J (1994) Use of homologous expression-secretion signals and vector-fress stable chromosomal integration in engineering of Lactobacillus plantarum for α-amylase and levanase expression. Appl Environ Microbiol 60:1401–1413
Johanningsmeier SD, Fleming HP, Breidt F Jr (2004) Malolactic activity of lactic acid bacteria during sauerkraut fermentation. J Food Sci 69:222–227
Koo YJ, Choi SY (1991) Science and Technology of Kimchi, 2nd edn. Korea Food Research Institute, Seoul
Lee KH, Park WJ, Kim JY, Kim HG, Lee JM, Kim JH, Park JW, Lee JH, Chung SK, Chung DK (2007) Development of a monitoring vector for Leuconostoc mesenteroides using the green fluorescent protein. J Microbiol Biotechnol 17:1213–1216
McDonald LC, Fleming HP, Hassan HM (1990) Acid tolerance of Leuconostoc mesenteroides and Lactobacillus plantarum. Appl Environ Microbiol 56:2120–2124
Olive DM, Bean P (1999) Principles and applications of methods for DNA-based typing of microbial organisms. J Clin Microbiol 37:1661–1669
Park MS, Shin DW, Lee KH, Ji GE (1999) Sequence analysis of plasmid pKJ50 from Bifidobacterium longum. Microbiol 145:585–592
Prosser JI (1994) Molecular marker systems for detection of genetically engineered micro-organisms in the environment. Microbiol 140:5–17
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Scott KP, Mercer DK, Richardson AJ, Melville CM, Glover LA, Flint HJ (2000) Chromosomal integration of the green fluorescent protein gene in lactic acid bacteria and the survival of marked strains in human gut simulations. FEMS Microbiol Lett 182:23–27. doi:10.1111/j.1574-6968.2000.tb08867.x
Tamminen M, Joutsjoki T, Sjöblom M, Joutsen M, Palva A, Ryhänen EL, Joutsjoki V (2004) Screening of lactic acid bacteria from fermented vegetables by carbohydrate profiling and PCR–ELISA. Lett Appl Microbiol 39:439–444. doi:10.1111/j.1472-765X.2004.01607.x
de Valdez GF, de Giore GS, Garro M, Mozzi F, Oliver G (1990) Lactic acid bacteria from naturally fermented vegetables. Microbiol Aliment Nutr 8:175–179
Vogel RF, Ehrmann MA, Gänzle MG (2002) Development and potential of starter Lactobacilli resulting from exploration of the sourdough ecosystem. Antonie Van Leeuwenhoek 81:631–638. doi:10.1023/A:1020530227192
Wagner M, Amann R, Kämpfer P, Assmus B, Hartmann A, Hutzler P, Springer N, Schleifer KH (1994) Identification and in situ detection of gram-negative filamentous bacteria in activated sludge. Syst Appl Microbiol 17:405–417
Wyckoff HA, Sandine WE (1991) Transformation of dairy Leuconostoc using plasmid vectors from Bacillis, Escherichia, and Lactococcus hosts. J Dairy Sci 74:1454–1460
Youngman P (1993) Transposons and their applications. In: Sonenshein AL, Hoch JA, Losick R (eds) Bacillus subtilis and other gram-positive bacteria, American Society for Microbiology, Washington, DC, pp 585–596
Acknowledgments
This work was supported by the fund of Research Center for Bioresource and Health (RCBH) at Chungbuk National University and ITEP & MOCIE of Korea.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Eom, HJ., Park, J.M., Seo, M.J. et al. Monitoring of Leuconostoc mesenteroides DRC starter in fermented vegetable by random integration of chloramphenicol acetyltransferase gene. J Ind Microbiol Biotechnol 35, 953–959 (2008). https://doi.org/10.1007/s10295-008-0369-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-008-0369-y