Abstract
Characterization of hepatocellular carcinomas (HCCs) and metastatic carcinomas (METs) from B-mode ultrasound presents a daunting challenge for radiologists due to their highly overlapping appearances. The differential diagnosis between HCCs and METs is often carried out by observing the texture of regions inside the lesion and the texture of background liver on which the lesion has evolved. The present study investigates the contribution made by texture patterns of regions inside and outside of the lesions for binary classification between HCC and MET lesions. The study is performed on 51 real ultrasound liver images with 54 malignant lesions, i.e., 27 images with 27 solitary HCCs (13 small HCCs and 14 large HCCs) and 24 images with 27 MET lesions (12 typical cases and 15 atypical cases). A total of 120 within-lesion regions of interest and 54 surrounding lesion regions of interest are cropped from 54 lesions. Subsequently, 112 texture features (56 texture features and 56 texture ratio features) are computed by statistical, spectral, and spatial filtering based texture features extraction methods. A two-step methodology is used for feature set optimization, i.e., feature pruning by removal of nondiscriminatory features followed by feature selection by genetic algorithm–support vector machine (SVM) approach. The SVM classifier is designed based on optimum features. The proposed computer-aided diagnostic system achieved the overall classification accuracy of 91.6 % with sensitivity of 90 % and 93.3 % for HCCs and METs, respectively. The promising results obtained by the proposed system indicate its usefulness to assist radiologists in diagnosing liver malignancies.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The real-time imaging capabilities offered by widely available ultrasound (US) imaging modality along with its inexpensive, nonradioactive, and noninvasive nature makes it a first-line examination for screening of focal liver lesions (FLLs) [1, 2]. However, there are certain disadvantages associated with the use of conventional gray-scale US for characterization of FLLs. (1) There is limited sensitivity for detection of small FLLs (<2 cm) developed on cirrhotic liver which is already nodular and coarse-textured [3–5]. (2) Sonographic appearance of hepatocellular carcinoma (HCC), primary malignant solid FLL and metastatic carcinoma (MET), secondary malignant solid FLL are highly overlapping [1, 3–6].
The sensitivity of contrast-enhanced US, contrast-enhanced spiral computed tomography, and magnetic resonance imaging modalities for detection and characterization of FLLs is higher than conventional gray-scale US, but these modalities are not widely available, expensive, and pose greater operational inconvenience [1, 3–6]. Therefore, a computer-aided diagnostic (CAD) system for accurate characterization of primary and secondary malignant FLLs based on conventional gray-scale US is highly desired to facilitate radiologists in clinical environment.
Among malignant FLLs, the present study is focused on characterization of HCC (most common primary malignant FLL) and MET (most common secondary malignant FLL). For the present work, benign FLLs such as hemangioma (HEM) and cyst are not considered because experienced participating radiologists (co-authors of this paper) having 13 and 23 years of experience in US imaging opined that HEM and cysts can be easily diagnosed from B-mode US with their classic diagnostic features. Typically, HEM appears as a well-circumscribed and uniformly hyperechoic lesion and cyst appears as a well-defined, rounded, anechoic lesion with thin imperceptible walls and posterior acoustic enhancement [1, 4, 5]. These typical sonographic appearances of benign FLLs can easily be differentiated from malignant FLLs, but differentiation between HCC and MET malignant lesions presents a daunting challenge even for experienced radiologists.
Early and accurate characterization of malignant FLLs is necessary because treatment options like curative surgical resection or successful percutaneous ablation are only possible if these malignancies are detected early [3, 5]. However, the practical problem faced by radiologists during routine practice is highly overlapping sonographic appearances of HCC (small and large HCCs on top of cirrhosis) and MET lesions (atypical METs) [1, 4, 5].
In 85 % of cases, HCC occurs in patients with cirrhosis. In fact in radiology practice, the condition of cirrhosis is seen as a precursor to the development of HCC [1, 3–5]. The only feature which favors the possibility of HCC in differential diagnosis between HCC and other FLLs is that HCC is most commonly associated with cirrhosis [1]. The sonographic appearances of small HCCs (<2 cm) vary from hypoechoic to hyperechoic. Large HCCs appear frequently with mixed echogenicity [4, 5]. A lesion can be labeled as typical in appearance when its subjective diagnosis can be made with a good confidence level by looking at the US examination. Experienced participating radiologists opined that the HCC case series should not be isolated as having typical or atypical sonographic appearance because of wide variability of sonographic appearances even within small and large HCCs; therefore, no sonographic appearance is typical for HCC. A representative dataset for designing the classifier should contain both small and large HCCs.
The occurrence rate of MET is 20 times more than that of HCC eventually because liver is the most common site for metastatic disease [5, 6]. Metastatic deposits may appear as single solitary mass or multiple masses of varying sizes. The sonographic appearance of MET lesions is extremely variable ranging from anechoic, hypoechoic, isoechoic, hyperechoic, and even with mixed echogenicity [1, 3–6]. However, the typical sonographic appearance of MET lesion is the “target” or “bull’s-eye” appearance (i.e., hypoechoic center surrounded by a hyperechoic rim) [1, 7, 8]. Diagnosis of these typical MET lesions can be made easily by an experienced radiologist from B-mode US, but differentiating atypical metastasis from HCCs lesions is considerably difficult. The sample images of small HCC, large HCC, typical MET, and atypical MET lesions from the acquired database are shown in Fig. 1.
The sonographic characterization of HCC and MET lesions is often carried out not only by observing the textural characteristics of regions inside the lesion but also by the texture of the background liver on which the lesion has evolved [1, 9]. The experienced participating radiologists opined that the textural characteristics of the neighboring liver parenchyma surrounding the lesion should contribute for differentiating the HCC and MET lesions from B-mode US. The present work investigates the contribution of texture of surrounding liver parenchyma in characterization of HCC and MET malignant liver lesions.
The related researches in literature for characterization of FLLs are few. The brief details of these studies [8, 10–12] are depicted in Table 1.
The study in [10] reported classification of benign, malignant, and normal liver with statistical texture analysis methods by using linear discriminant analysis and neural network classifier. The study in [12] reported classification of cyst, HEM, strike out this and malignant and normal liver with manually selected optimal statistical and spectral texture features by using a neural network classifier. A CAD system for classification in five classes, namely, HEM, cyst, HCC, MET, and normal liver, is proposed in [8]. However, their proposed CAD system is developed using a large feature vector consisting of 208 features extracted with statistical, spectral, and spatial filtering based methods and neural network classifiers. In studies [10, 12], malignant lesions are considered as single class; however, the characterization of malignant lesions as HCC or MET lesions is clinically significant for effective treatment and management of liver malignancies [3, 5]. The study in [8] used the region of interest (ROI) size of 25 × 25 pixels for computing texture features; however, in [10, 11] the use of ROI size of 10 × 10 pixels is reported. The use of 10 × 10 pixels and even 25 × 25 pixels as ROI size yields a smaller number of pixels in comparison to minimum 800 pixels required to estimate reliable statistics [13–15].
The related research reported in [12] used wavelet packet texture descriptors with neural network classifier for binary classification tasks, i.e., HEM vs. HCC, HEM vs. MET, and HCC vs. MET. Among these, the lowest characterization performance for HCC vs. MET is reported on their data. Their study reports the use of 64 × 64 pixels as ROI size, possibly because they used high-resolution scanned images instead of real US images. It is otherwise difficult to select such a large ROI size keeping in view the size of small lesions and resolution of images obtained from US machines.
According to the best of the authors’ knowledge, all the researches in literature for characterization of FLLs have considered only the texture patterns of regions inside the lesions, and a CAD system for characterization of HCC and MET lesions has not been experimented as yet. The present study investigates the contribution made by texture patterns of regions inside and outside of the lesions for binary classification of HCC and MET lesions.
In the present work, support vector machine (SVM) has been chosen for the classification task because classifier designs which use regularization like SVM are less prone to overfitting and obtain good generalization performance to a certain extent even without feature space dimensionality reduction [16–18]. Extensive literature surveys on texture classification reveal that SVM has shown remarkable performance for classification of medical images [19–27].
Materials and Methods
Data Collection and Description
Data Collection
For the present work, 51 images were collected from 51 different patients; out of these 27 images are HCC images with 27 solitary HCC lesions and 24 images are MET images with 27 MET lesions, i.e., 21 MET images with solitary MET lesion and 3 MET images with 2 MET lesions each. These images were collected from different patients visiting the Department of Radiodiagnosis and Imaging, Post Graduate Institute of Medical Education and Research (PGIMER), Chandigarh, India over the time period from March 2010 to December 2011. Informed consent of patients for using these images for research was taken prior to recording. The medical ethics board of PGIMER, Chandigarh, granted the ethical clearance to carry out this research work. The direct digital images recorded by using Philips ATL HDI 5000 US machine equipped with multifrequency transducer of 2–5 MHz range were used. The size of the images is 800 × 564 pixels with gray scale consisting of 256 tones, and horizontal as well as vertical resolution is 96 dpi. The following protocols were followed for data collection:
(1) The judgment regarding the diagnostic quality (free from artifacts) and representativeness of each image class (HCC and MET) was made by two domain experts (co-authors of this paper) with 13 and 23 years of experience in US imaging. (2) The acquired dataset contained 27 HCC images with 27 solitary HCC lesions comprising of 13 small HCC lesions and14 large HCC lesions. While recording these images, both transverse and longitudinal views of each image were observed to determine the size of each lesion. (The HCC lesion was considered as small if its size was less than or equal to 2 cm.) (3) Only HCCs evolved on cirrhotic liver are considered. (4) The acquired dataset contained 24 MET images with 27 metastatic lesions comprising of 12 lesions with typical bull’s-eye or target appearance and 15 MET lesions with variable sonographic appearances. (5) The labeling of HCC lesions as small HCC and large HCC lesions and labeling of MET lesions as one with typical target bull’s-eye appearance and other atypical sonographic appearances was done during data collection solely for the purpose of having representative data in training set for designing the classifier.
Selection of Regions of Interest (ROIs)
The following protocols were followed for cropping the ROIs from the image database (cropping here refers to extraction of ROIs not to be confused with similar terminology used in photography for removal of unwanted details/objects in the images):
(1) The ROIs were cropped by an experienced participating radiologist by using a specially designed ROI manager software developed in Biomedical Instrumentation Laboratory, Indian Institute of Technology, Roorkee. This ROI manager software provided the radiologist the flexibility to load the image, choose the ROI size and shape, move the ROI to any desired location over the image, freeze the ROI at any location, and crop the ROIs together after the position of all the ROIs for a particular image is frozen. (2) Two types of ROIs are used in this study, within-lesion ROIs (WLROIs) and surrounding lesion ROIs (SLROIs). (3) Maximum nonoverlapping WLROIs were cropped from well within the boundary of each lesion. (4) The areas of necrosis were avoided while cropping WLROIs. (5) For each lesion, a single SLROI was cropped approximately at the same depth as that of the center of the lesion. (6) SLROI were cropped by avoiding the inhomogeneous structures like hepatic ducts and blood vessels, etc.
In the present work, two types of features are considered for analysis, i.e., texture features computed from WLROIs and texture ratio features computed by taking the ratio of texture feature computed from WLROI and texture feature computed from corresponding SLROI.
It can be noted that HCC lesion in Fig. 2a contains five WLROIs and a corresponding SLROI. Thus, five instances of a single texture feature can be obtained with these five WLROIs and five instances of texture ratio feature can be obtained by dividing the texture feature value obtained for each WLROI with the texture feature value obtained for the corresponding SLROI.
Selection of ROI Size
The ROI size should be chosen so as to provide good statistical population for computing texture features. In literature, different ROI sizes ranging from 10 × 10 pixels [9, 11], 25 × 25 pixels [8], and 64 × 64 pixels [12] have been chosen for classification of FLLs. After interaction with the participating radiologists, ROI size of 32 × 32 pixels was considered appropriate for the present study considering the facts mentioned below:
-
1.
There is sufficient evidence in the literature that ROI size must be at least 800 pixels to provide good sampling distribution for estimating reliable statistics [13–15]; as ROI size of 32 × 32 gives 1,024 pixels, it can be believed that the computed texture parameters are reliable estimates.
-
2.
During initial discussions with the participating radiologists, an attempt was made to mark larger ROI sizes, but few practical difficulties were faced. Certain lesions had necrotic area, radiologists opined that the necrotic area inside lesions must be avoided while extracting WLROIs, and it was not possible to consider large ROI size for these lesions. Also, participating radiologists were of the view that SLROI for each lesion must be selected by avoiding the inhomogeneous structures like hepatic ducts and blood vessels, etc., which was practically difficult by considering larger ROI size.
-
3.
For real-time implementation, small ROI size is always favorable as time taken for feature extraction and classification is obviously less in comparison to large ROI size. Also, with small ROI size, more number of samples are available for classifier design.
Interpretation by Radiologists
One of the experienced participating radiologist having more than 13 years of experience confirmed the presence of HCC and MET lesions using liver image assessment criteria including (1) visualization of sonographic appearances, imaging features of FLLs based on their knowledge and expertise, (2) follow-up of clinical history of the patient and other associated findings, and (3) imaging appearance on dynamic helical computed tomography (CT)/ magnetic resonance imaging (MRI)/pathological examinations and biopsy, which is an invasive procedure.
Data Set Description
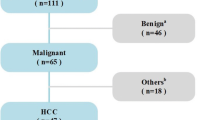
The distribution of clinically acquired database of 51 B-mode liver US images among HCC and MET image categories and the bifurcation of ROIs in training and test data set is described in Fig. 3 below.
Proposed Computer-Aided Diagnostic System
The block diagram of the proposed CAD system is depicted in Fig. 4.
For implementation of the proposed CAD system, the database of 120 nonoverlapping WLROIs and 54 SLROIs was created from 51 clinically acquired US images. The CAD system consisted of feature extraction, feature selection, and classification modules. In feature extraction module, texture features are computed from WLROIs as well as SLROIs by gray-level co-occurrence matrix (GLCM) [28], gray-level run length matrix (GLRLM) [29–31], Fourier power spectrum (FPS) [32], and Laws’ texture feature [33] extraction methods. In feature selection module, initially feature pruning is carried out by removal of nondiscriminatory feature vectors followed by feature selection by genetic algorithm–support vector machine (GA–SVM) approach. The GA–SVM procedure results in optimal reduced set of features. In classification module, a support vector machine (SVM) classifier is designed with the selected optimal features. The SVM classifier is implemented using LibSVM library [34].
Feature Extraction
The general idea of feature extraction is to convert both visually extractable and visually nonextractable sonographic features into mathematical descriptors. These mathematical descriptors are either morphological (based on shape or contour of the lesion) or textural features (based on intensity distribution) [35]. Both these morphological as well as textural features are significant for developing CAD systems for breast lesions from B-mode US [36–39]. Experienced participating radiologists opined that morphological sonographic features of FLLs do not give any significant information for their characterization; as also evident from other related researches, the proposed CAD systems for characterization of FLLs from B-mode US have relied on textural features only [8, 10–12].
Initially, a wide variety of visual and nonvisual echotexture features are extracted by using statistical, spectral, and spatial filtering based feature extraction methods. These features are then applied in the present classification system with a tedious task of combining the most relevant and effective features while discarding the nonperforming features.
Statistical texture features are defined by the spatial distribution of gray-level intensity values in the image. Local features are computed at each point in the image and a set of statistics are derived from the distribution of these local features. Statistical methods are classified as first-order statistics, second-order statistics, or higher-order statistics depending upon the number of pixels used in defining a local feature [8, 40].
Spectral features computed by FPS method such as radial sum and angular sum of the discrete Fourier transform are used to describe texture [32].
Spatial filtering based texture descriptors, i.e., Laws’ texture features, determine texture properties by performing local averaging, edge detection, spot detection, wave detection, and ripple detection in texture [33]. Laws’ texture features are computed by using special 1-D filters of length 3, 5, 7, and 9. Different filter lengths correspond to different resolutions for extraction of texture features from a ROI. In the present work, 1-D filters of length 7, i.e., L7 = [1, 1, 6, 6, 15, 15, 20], E7 = [−1, −4, −5, 0, 5, 4, 1], and S7 = [−1, −2, 1, 4, 1, −2, −1], are used. Special 2-D filters called Laws’ masks are derived by outer vector product of these 1-D kernels with themselves or with each other as shown in Fig. 5.
The texture images are obtained by convolving the ROI of size M × N with these 2D Laws’ masks, for example
The output TIs are processed by texture energy measurement (TEM) filters. The TEM filter performs moving average nonlinear filtering operation as depicted by:
Here, 15 × 15 descriptor windows are used to obtain nine texture energy images (TEIs). Texture energy images obtained by a pair of identical filters, for example, TEIE7L7 and TEIL7E7, are combined to obtain a rotational invariant image (90° rotational invariance) (TR) [41].
Statistics derived from these TR images provide significant texture information of ROI. Five statistics, i.e., mean, standard deviation, skewness, kurtosis, and energy are extracted from each TR image [41, 42]. Thus, 30 Laws’ texture features (6 TR images × 5 statistical parameters) are computed for each ROI.
In the present work, statistical methods, i.e., GLCM and GLRLM methods, spectral method, i.e., FPS method, and spatial filtering based method, i.e., Laws’ texture feature extraction method, are selected for the classification task. The selection of these methods for the classification task is based on other related researches with US images [13, 14, 41, 43] and few other studies for diagnosis of FLLs with US images [8, 10–12].
For extraction of efficient diagnostic features for characterization of liver malignancies, initially 112 features (56 features computed from WLROIs and 56 texture ratio features) are computed using GLCM, GLRLM, FPS, and Laws’ texture feature extraction methods as tabulated in Table 2.
For computation of Laws’ texture features, different 1-D filters of length 5, 7, and 9 were experimented as shown in Table 3.
Statistics derived from TRs are used as feature vectors. In the present work, five features, i.e., mean, standard deviation, skewness, kurtosis, and energy, are computed from TRs. Thus, feature vector of lengths 70, 30, and 70 are obtained with 1-D filter of length 5, 7, and 9, respectively. It was observed that the classification accuracy obtained by SVM classifier by feature vector of length 30 obtained for filter length 7 is higher in comparison with feature vectors of length 70 obtained for filter lengths 5 and 9. Thus, 30 Laws’ texture features computed for filter length 7 are considered for further analysis.
Feature Selection
Feature selection is used to eliminate the interference of irrelevant features which often increases the time taken to perform classification task and also reduces the classification accuracy. In the present work, two-step methodology is followed for feature selection. In the first step, initial feature pruning is carried out by removal of nondiscriminatory individual texture feature vectors (TFVs). The discrimination ability of a TFV is measured by the classification accuracy obtained by SVM classifier. Feature pruning yields a pruned TFV consisting of best-performing individual TFVs.
In the second step, GA–SVM feature selection is applied on pruned TFV; here binary genetic algorithm (GA) is used to evolve subsets of pruned TFV and the classification accuracy obtained by the SVM classifier is used as a fitness function. The GA–SVM feature selection procedure removes irrelevant features from pruned TFV to yield an optimal subset of discriminatory features. The main steps for implementation of binary GA [39] are:
-
Step 1
Define fitness function and select GA run parameters like population size, crossover type, crossover rate, and mutation rate.
-
Step 2
Create initial population (a set of binary coded chromosomes or genotypes).
-
Step 3
Decode chromosomes (binary chromosomes are converted into candidate solutions or phenotypes).
-
Step 4
Fitness function (fitness evaluation of each candidate solution or phenotype).
-
Step 5
Selection (the selection of parents to enter the mating pool based on fitness evaluation).
-
Step 6
Applying crossover and mutation to generate offsprings
-
Step 7
Create next generation (by evaluating offsprings using fitness function)
The GA terminates when there is no improvement in the fitness value or after a fixed number of successive iterations. In the present work, single-point crossover is used and the other run parameters are set as crossover rate equal to 0.7, mutation rate equal to 0.05, and population size equal to 20 after a series of trials.
Classification
SVM Classifier
The SVM classifier attempts to construct an optimum hyperplane in the higher dimensional feature space to separate the training data with minimum expected risk. Kernel functions are used for nonlinear mapping of the training data from input space to higher dimensional feature space. In the present work, the performance of Gaussian Radial Basis Function kernel is investigated. For a detailed description of SVM approach, additional information can be found in [16, 17].
A crucial step for obtaining good generalization performance is correct choice of the regularization parameter C and kernel parameter γ. The regularization parameter C attempts to maximize the margin while keeping low value for training error. In the present work, extensive search is carried out in the parameter space for the values of C ∈ {2−4, 2−3… 215}, γ ∈ {2−12, 2−11… 24} using 10-fold cross-validation on training data. To avoid the bias caused by unbalanced feature values, the extracted features were normalized in the range [0, 1] by using min–max normalization procedure. For the present work, SVM classifier is implemented using LibSVM library [34].
Results
Rigorous experimentations were carried out to identify potential TFVs of texture features and texture ratio features for characterization of HCC and MET FLLs. In all the experiments, the discrimination ability of texture feature vectors (TFVs) has been evaluated by using a SVM classifier.
In experiment 1, the discrimination ability of total 112 texture features, i.e., total eight TFVs (four TFVs corresponding to texture features and four TFVs corresponding to texture ratio features) obtained by GLCM, GLRLM, FPS, and Laws’ feature extraction methods is investigated. The results of experiment 1 are used to obtain a pruned TFV by removal of individual nonperforming TFVs.
Experiments 2 and 3 investigate the discrimination ability of combined TFVs, i.e., combined TFV consisting of all texture features and combined TFV consisting of all texture ratio features. It is observed that combined TFV consisting of texture ratio features has more discrimination ability.
In experiment 4, the discrimination ability of pruned TFV obtained as a result of experiment 1 is examined. Experiment 5 investigates the discrimination ability of optimal reduced TFV obtained by passing pruned TFV to GA–SVM method.
-
Experiment 1
This experiment compares the performance of SVM classifiers by use of various individual TFVs. The results obtained are reported in Table 4. It can be seen from Table 4 that GLRLM texture ratio features provide highest classification accuracy of 71.6 %. Both Laws’ WLROI features and GLCM ratio features provide the second highest accuracy of 70 %. It can also be observed that GLCM and GLRLM ratio features show better characterization performance than corresponding WLROI features. Further, it can be noted that FPS and Laws’ WLROI features show better characterization performance in comparison to corresponding ratio features.
Table 4 Comparison of performance of SVM classifiers for various individual TFVs -
Experiment 2
This experiment evaluates the performance of SVM classifier by use of combined TFV of all 56 WLROI texture features. The results obtained are reported in Table 5. It can be seen from Table 5 that combined TFV of all 56 WLROI texture features provide a classification accuracy of 61.6 % for characterization of HCC and MET FLLs.
Table 5 Performance of SVM classifier for combined TFV of all 56 WLROI texture features -
Experiment 3
This experiment evaluates the performance of SVM classifier by use of combined TFV of all 56 texture ratio features. The results obtained are reported in Table 6. It can be seen from Table 6 that combined TFV of all texture ratio features provide classification accuracy of 78.3 %. For further experimentations, feature pruning is carried out on the basis of classification accuracy obtained by SVM classifier for eight individual TFVs shown in Table 4. The nonperforming individual TFVs are removed and the best-performing individual TFVs (highlighted in Table 4) are combined to form a pruned TFV for adequate discrimination of HCC and MET FLLs.
Table 6 Performance of SVM classifier for combined TFV of all 56 texture ratio features -
Experiment 4
This experiment evaluates the performance of SVM classifier by use of pruned TFV of length 56 consisting of best-performing individual TFVs. The results obtained are reported in Table 7. From Table 7, it can be seen that pruned TFV yields a classification accuracy of 80 %. For further experimentation, this pruned TFV is subjected to GA–SVM feature selection procedure which iteratively removes the irrelevant and interfering features from the pruned TFV and returns an optimal reduced TFV of length 9. Nine texture features, i.e., four GLCM ratio features (angular second moment, sum average, difference entropy, and inverse difference moment), three GLRLM ratio features (long run emphasis, gray level non uniformity, and long run high gray level emphasis), one FPS WLROI feature (radial sum), and one Laws’ WLROI feature (LLmean), are selected by GA–SVM procedure.
Table 7 Performance of SVM classifier for pruned TFV -
Experiment 5
This experiment evaluates the performance of SVM classifier by use of optimal reduced TFV of length 9 consisting of features selected by GA–SVM procedure. The results obtained are reported in Table 8. From Table 8, it can be observed that using optimal reduced TFV consisting of nine features selected by GA–SVM procedure with SVM classifier yields the classification accuracy of 91.6 % and sensitivity of 90 % and 93.3 % for HCC and MET lesions, respectively. It is observed that for the studied population consisting of small HCCs, large HCCs, typical METs, and atypical METs, the proposed CAD system yields high sensitivity values for differentiation between malignant lesions. The generalization ability of the proposed CAD system can be tested by analyzing larger datasets.
Table 8 Performance of SVM classifier for optimal reduced TFV
Discussion
Misclassification Analysis
Analysis of five misclassified cases out of 60 cases in the test data set is reported in Table 9. It can be observed from Table 9 that sensitivity of proposed CAD system for small HCC cases and large HCC cases is 88.8 % and 90.4 %, respectively. In case of MET cases, the sensitivity obtained is 100 % and 91.6 % for typical and atypical MET cases, respectively. However, it can be observed from Table 8 that the classification accuracy of the proposed CAD system is 91.6 % with sensitivity of 90 % for HCC cases and 93.3 % for MET cases. Given the fact that the sonographic appearances of HCC and MET overlap sufficiently, and the sensitivity of conventional B-mode US is limited, the results obtained by the proposed CAD system for the population studied are quite promising specifically in the presence of a comprehensive and representative dataset consisting of SHHCs, LHCCs, and typical as well as atypical MET cases. However, the generalization ability of the proposed CAD system remains to be tested by analyzing larger datasets for which data collection may take a long span of time. At the same time, it is worth mentioning that the selected features significantly account for the textural variations exhibited by primary and secondary liver malignancies as the proposed system performs well on unseen test data.
Conclusions
The ratio features are more discriminatory than WLROI features for characterization of HCC and MET FLLs. Only nine texture features (seven ratio features and two WLROI features) are significant to account for textural variations exhibited by HCC and MET lesions. It can be concluded that the texture of the background liver on which the lesion has evolved do contribute towards characterization of primary and secondary malignant FLLs from B-mode US. The proposed CAD system yields the classification accuracy of 91.6 % with sensitivity of 90 % and 93.3 % for HCC and MET cases, respectively. The results obtained by the proposed CAD are up to the satisfaction of experienced participating radiologists. The promising results of the study indicate that the proposed CAD system can be routinely used in a clinical environment to assist radiologists in diagnosing liver malignancies and thereby facilitate in providing better disease management.
References
Bates J: Abdominal Ultrasound How Why and When, 2nd edition. Churchill Livingstone, Oxford, 2004, pp 80–107
Virmani J, Kumar V, Kalra N, Khandelwal N: A rapid approach for prediction of liver cirrhosis based on first order statistics. In: Proceedings of IEEE International Conference on Multimedia, Signal Processing and Communication Technologies, IMPACT-2011, 212–215, 2011
Soye JA, Mullan CP, Porter S, Beattie H, Barltrop AH, Nelson WM: The use of contrast-enhanced ultrasound in the characterization of focal liver lesions. Ulster Med J 76(1):22–25, 2007
Colombo M, Ronchi G: Focal Liver Lesions—Detection, Characterization, Ablation. Springer, Berlin, 2005, pp 167–177
Harding J, Callaway M: Ultrasound of focal liver lesions. Rad Magazine 36(424):33–34, 2010
Jeffery RB, Ralls PW: Sonography of Abdomen. Raven, New York, 1995
Scheible W, Gossink BB, Leopold G: Gray scale echo graphic patterns of hepatic metastatic disease. Am J Roentgenol 129:983–987, 1977
Mittal D, Kumar V, Saxena SC, Khandelwal N, Kalra N: Neural network based focal liver lesion diagnosis using ultrasound images. Int J Comput Med Imaging Graph 35(4):315–323, 2011
Lee WL, Hsieh KS, Chen YC: A study of ultrasonic liver images classification with artificial neural networks based on fractal geometry and multiresolution analysis. Biomed Eng Appl Basis Commun 16(2):59–67, 2004
Sujana S, Swarnamani S, Suresh S: Application of artificial neural networks for the classification of liver lesions by image texture parameters. Ultrasound Med Biol 22(9):1177–1181, 1996
Poonguzhali S, Deepalakshmi, Ravindran G: Optimal feature selection and automatic classification of abnormal masses in ultrasound liver images. In: Proceedings of IEEE International Conference on Signal Processing, Communications and Networking, ICSCN’07, 503–506, 2007
Yoshida H, Casalino DD, Keserci B, Coskun A, Ozturk O, Savranlar A: Wavelet packet based texture analysis for differentiation between benign and malignant liver tumors in ultrasound images. Phys Med Biol 48:3735–3753, 2003
Kadah YM, Farag AA, Zurada JM, Badawi AM, Youssef AM: Classification algorithms for quantitative tissue characterization of diffuse liver disease from ultrasound images. IEEE Trans Med Imaging 15(4):466–478, 1996
Badawi AM, Derbala AS, Youssef ABM: Fuzzy logic algorithm for quantitative tissue characterization of diffuse liver diseases from ultrasound images. Int J Med Inf 55:135–147, 1999
Fukunaga K: Introduction to Statistical Pattern Recognition. Academic, New York, 1990
Burges CJC: A tutorial on support vector machines for pattern recognition. Data Min Knowl Disc 2(2):1–43, 1998
Guyon I, Weston J, Barnhill S, Vapnik V: Gene selection for cancer classification using support vector machines. J Machine Learn 46(1–3):1–39, 2002
Virmani J, Kumar V, Kalra N, Khandelwal N: SVM-based characterization of liver ultrasound images using wavelet packet texture descriptors. J Digit Imaging, 2012. doi:10.1007/s10278-012-9537-8
Wan J, Zhou S: Features extraction based on wavelet packet transform for b-mode ultrasound images. In: Proceedings of IEEE International Congress on Image and Signal Processing, CISP-2010, 949–955, 2010
Lee C, Chen S H: Gabor wavelets and SVM classifier for liver diseases classification from CT images. In: Proceedings of IEEE International Conference on Systems, Man, and Cybernetics, 548–552, 2006
Nawaz S, Dar A H: Hepatic lesions classification by ensemble of SVMs using statistical features based on co-occurrence matrix. In: Proceedings of 4th IEEE International Conference on Emerging Technologies, ICET-2008, 21–26, 2008
Huang YL, Chen DR, Jiang YR, Kuo J, Wu HK, Moon WK: Computer-aided diagnosis using morphological features for classifying breast lesions on ultrasound. Ultrasound Obstet Gynecol 32:565–572, 2008
Moayedi F, Azimifar Z, Boostani R, Katebi S: Contourlet based mammography mass classification. In: Proceedings of ICIAR 2007, LNCS 4633, 923–934, 2007
Huang YL, Wang KL, Chen DR: Diagnosis of breast tumors with ultrasonic texture analysis using support vector machines. Neural Comput & Applic 15:164–169, 2006
Reddy TK, Kumaravel N: A comparison of wavelet, curvelet and contourlet based texture classification algorithms for characterization of bone quality in dental CT. In: Proceedings of International Conference on Environmental, Biomedical and Biotechnology, IPCBEE 16:60–65, 2011
Tsiaparas N, Golemati S, Andreadis I, Stoitsis J: Multiscale geometric texture analysis of ultrasound images of carotid atherosclerosis. In: Proceedings of 10th IEEE International Conference on Information Technology and Applications in Biomedicine, ITAB-2010, 1–4, 2010
Minhas F, Sabih D, Hussain M: Automated classification of liver disorders using ultrasound images. J Med Syst, 2011. doi:10.1007/s10916-011-9803-1
Haralick R, Shanmugam K, Dinstein I: Textural features for image classification. IEEE Trans Syst Man Cybern SMC-3(6):610–121, 1973
Galloway RMM: Texture analysis using gray level run lengths. Comput Graphics Image Processing 4:172–179, 1975
Chu A, Sehgal CM, Greenleaf JF: Use of gray value distribution of run lengths for texture analysis. Pattern Recognition Lett 11:415–420, 1990
Dasarathy BV, Holder EB: Image characterizations based on joint gray level-run length distributions. Pattern Recognition Lett 12:497–502, 1991
Weszka JS, Dyer CR, Rosenfeld A: A comparative study of texture measures for terrain classification. IEEE Trans Syst Man Cybern SMC-6(4):269–285, 1976
Laws KI: Rapid texture identification. In: SPIE Proceedings of the Seminar on Image Processing for Missile Guidance. 238:376–380, 1980
Chang CC, Lin CJ: LIBSVM, a library of support vector machines. Software available at http://www.csie.ntu.edu.tw/∼cjlin/libsvm. Accessed 15 June 2012
Kim SH, Lee JM, Kim KG, Kim JH, Lee JY, Han JK, Choi BI: Computer-aided image analysis of focal hepatic lesions in ultrasonography: preliminary results. Abdom Imaging 34(2):183–91, 2009
Huang YL, Chen DR, Jiang YR, Kuo SJ, Wu HK, Moon WK: Computer aided diagnosis using morphological features for classifying breast lesions on ultrasound. Ultrasound Obstet Gynecol 32:565–572, 2008
Moayedi F, Azimifar Z, Boostani R, Katebi S: Contourlet-based mammography mass classification. In: Proceedings of 4th International Conference on Image Analysis and Recognition, ICIAR-2007, LNCS series (4633):923–934, 2007
Diao XF, Zhang XY, Wang TF, Chen SP, Yang Y, Zhong L: Highly sensitive computer aided diagnosis system for breast tumor based on color Doppler flow images. J Med Syst 35(5):801–809, 2011
Nandi RJ, Nandi AK, Rangayyan RM, Scutt D: Classification of breast masses in mammograms using genetic programming and feature selection. Med Biol Eng Comput 44(8):683–94, 2006
Srinivasan GN, Shobha G: Statistical texture analysis. Proc World Acad Sci Eng Technol 36:264–1269, 2008
Virmani J, Kumar V, Kalra N, Khandelwal N: Prediction of cirrhosis from liver ultrasound B-mode images based on Laws’ masks analysis. In: Proceedings of IEEE International Conference on Image Information Processing, ICIIP-2011, 1–5, 2011
Rachidi M, Marchadier A, Gadois C, Lespessailles E, Chappard C, Benhamou CL: Laws' masks descriptors applied to bone texture analysis: an innovative and discriminant tool in osteoporosis. Skeletal Radiol 37(6):541–548, 2008
Virmani J, Kumar V, Kalra N, Khandelwal N: Prediction of cirrhosis based on singular value decomposition of gray level co-occurrence matrix and a neural network classifier. In: Proceedings of IEEE International Conference on Developments in E-systems Engineering, DeSe-2011, 146–151, 2011
Acknowledgments
The author Jitendra Virmani would like to acknowledge Ministry of Human Resource Development (MHRD), India for financial support. The authors wish to acknowledge the Department of Electrical Engineering, Indian Institute of Technology, Roorkee, India and Department of Radiodiagnosis and Imaging, Postgraduate Institute of Medical Education and Research, Chandigarh, India for their constant patronage and support in carrying out this research work. The authors would like to thank the anonymous reviewers for their substantive and informed review, which led to significant improvements in the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Virmani, J., Kumar, V., Kalra, N. et al. Characterization of Primary and Secondary Malignant Liver Lesions from B-Mode Ultrasound. J Digit Imaging 26, 1058–1070 (2013). https://doi.org/10.1007/s10278-013-9578-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-013-9578-7
Keywords
- Texture analysis
- B-Mode liver ultrasound
- Hepatocellular carcinoma
- Metastasis
- Primary malignant liver lesion
- Secondary malignant liver lesion
- Genetic algorithm
- Support vector machine classifier
- Small hepatocellular carcinoma
- Large hepatocellular carcinoma
- Typical metastasis
- Atypical metastasis
- Focal liver lesions