Abstract
Glucuronoarabinoxylan is the major hemicellulose in grass cell walls, yet the mechanism of xylan synthesis in monocot plants is still unclear. Unraveling the genes involved in the biosynthesis of xylan in rice will be very important for the utilization of rice straw as a source of bioenergy in the future. In this report, we investigated the functional role of a rice gene homologous to Arabidopsis IRREGULAR XYLEM10 (IRX10), belonging to the glycosyl transferase (GT) gene family 47 (GT47), in the biosynthesis of xylan. The protein sequence of OsGT47A from rice exhibits a 93.49 % similarity to IRX10, which is involved in the biosynthesis of glucuronoxylan in Arabidopsis. Phylogenetic analysis of the GT47 glycosyl transferase family in the rice genome revealed that OsGT47A is a closely related homolog of IRX10 and IRX10L. Expression pattern analysis showed that the OsGT47A gene is highly expressed in the rice stem. Overexpression of OsGT47A in the irx10 irx10L double mutant rescued the plant growth phenotype and restored secondary wall thickness. Analysis of monosaccharides indicated that the rescued plants had levels of xylose identical to those of the wild type plants, and the fluorescence signals were restored in the complementation plants by xylan immunolocalization. The OsGT47A complementation under the native promoter of Arabidopsis IRX10L (ProIRX10L) partially rescued the double mutant, indicating that OsGT47A is functionally equivalent to IRX10L. Together, these results suggest that the IRX10 homolog OsGT47A exhibits functional conservation and is most likely involved in xylan synthesis in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Secondary cell walls are the major form of biomass in plants and provide an important source of renewable and sustainable energy. The major components of the secondary cell wall are cellulose, hemicellulose and lignin, which constitute the most abundant biopolymers produced by plants. The mechanisms of cellulose and lignin synthesis have been well studied for many years (Boerjan et al. 2003; Lerouxel et al. 2006; Scheible and Pauly 2004; Somerville 2006). In contrast, xylan biosynthesis has been elucidated only recently (Brown et al. 2007; Pena et al. 2007; Zhong et al. 2005). The secondary cell wall stores a large source of energy in the form of polysaccharides, yet the presence of lignin and xylan in the secondary cell wall increases the difficulty of degrading cellulose and hence negatively affects the exploitation of plant energy biomass. To date, one of the most effective strategies of increasing access to these energy stores is to alter cell wall structure or composition through the modulation of hemicellulose biosynthesis (Carroll and Somerville 2009; Pauly et al. 2013; Scheller and Ulvskov 2010).
Xylans are the main component of hemicellulose and play a critical structural role in plant cell walls through interactions with cellulose microfibrils (Scheller and Ulvskov 2010). Xylans are the second most abundant polysaccharides in both dicot wood and monocot grass, and the structure of xylans might be quite diverse between different species. The xylans in the green alga Caulerpa have a β-1,3-d-xylan backbone (Ebringerová and Heinze 2005). Dicot xylans, called glucuronoxylan (GX), are composed of a linear backbone of β-(1,4)-linked xylose (Xyl) residues with α-linked side branches of α-d-glucuronic acid (GlcUA) and/or 4-O-methyl-α-d-glucuronic acid (Me-GlcUA) (Pena et al. 2007), which occur on average once every eight Xyl residues (Brown et al. 2007). In vegetative tissues of monocots, the side chain of xylan can also be substituted with arabinosyl or acetyl groups (Ebringerová and Heinze 2000), and the main component is called glucuronoarabinoxylan (GAX). In addition, dicot plants contain a unique tetrasaccharide sequence (called sequence 1), 4-β-d-Xylp-(1,4)-β-d-Xylp-(1,3)-α-l-Rhap-(1,2)-α-d-GalpA-(1,4)-d-Xylp at the reducing end of GX (Andersson et al. 1983; Johansson and Samuelson 1977; Pena et al. 2007). Several genes associated with xylan synthesis have been identified using gene expression profiling approaches in Arabidopsis (Brown et al. 2005; Persson et al. 2005). These genes are named as IRX due to irregular xylem (irx) mutants with secondary cell wall deficiencies (Turner and Somerville 1997), and include members of GT47 (FRA8[IRX7]/F8H[IRX7L] and IRX10/IRX10L), GT43 (IRX9/IRX9L and IRX14/IRX14L), and GT8 (IRX8 and PARVUS) families (Bauer et al. 2006; Brown et al. 2007, 2009; Lee et al. 2007; Pena et al. 2007; Persson et al. 2007; Wu et al. 2009, 2010; Zhong et al. 2005). IRX9/IRX9L, IRX14/IRX14L and IRX10/IRX10L are thought to be responsible for the elongation of the xylan backbone (Brown et al. 2007, 2009; Lee et al. 2007, 2010; Pena et al. 2007; Persson et al. 2007; Wu et al. 2009, 2010) and IRX7(FRA8)/IRX7L(F8H), IRX8 and PARVUS for the synthesis of the tetrasaccharide at the reducing end (Brown et al. 2007; Liepman et al. 2010; Pena et al. 2007; Zhong et al. 2005). Comparative transcript analyses in grasses discovered members of GT43, GT46 and GT61 as candidates for xylan synthesis (Mitchell et al. 2007), and the loss of the GT61 family member protein significantly decreased β-(1,3)-linked arabinosyl substitution of the xylan (Chiniquy et al. 2012). A more recent biochemical study directly associated specific members of wheat GT43, GT47 and GT75 families with GAX biosynthesis (Zeng et al. 2010). Moreover, heterologous expression of wheat and rice GT61s in Arabidopsis can produce arabinoxylan (Anders et al. 2012). Whether GT43 and GT47 members in grass will be functional in xylan backbone elongation still requires further investigation.
To date, evidence has indicated conservation of the GX biosynthetic machinery between herbaceous and woody plants in dicots. For example, it has been shown that PoGT43B, the poplar homolog of Arabidopsis IRX9, can genetically complement the Arabidopsis irx9 mutant (Zhou et al. 2007). PoGT47, the homolog of Arabidopsis FRA8, can complement Arabidopsis fra8 mutant (Zhou et al. 2006), and two poplar homologs of the PARVUS gene are able to complement the Arabidopsis parvus mutant (Kong et al. 2009). The loss of function of OsIRX10 (Os01g0926700/Os01g70200), the rice homologous gene to Arabidopsis IRX10, resulted in short culms, reduced plant size and decreased xylan content (Chen et al. 2013). These observations support that these rice genes have similar functions in xylan biosynthesis to their Arabidopsis homologs.
In this study, another rice GT gene encoding a protein, OsGT47A, with a high sequence similarity to Arabidopsis IRX10, was cloned. To investigate whether rice OsGT47A is a functional homolog of Arabidopsis IRX10, we conducted experiments to complement the Arabidopsis irx10 (−/−) irx10L (−/−) double mutant and demonstrated that the OsGT47A gene can rescue or partially rescue the phenotypes. This gene also leads to a restoration of the wild-type Xyl content in stems. Our results indicate that OsGT47A has the same genetic function as Arabidopsis IRX10 and participates in the xylan biosynthesis in monocots.
Materials and methods
Plant growth conditions
Arabidopsis (Arabidopsis thaliana L.) seed sterilization, growth conditions, medium and resistance screening have been described previously (Wu et al. 2009). In brief, seeds were surface sterilized (Forsthoefel et al. 1992) and sprayed onto Murashige and Skoog medium containing 1 % sucrose and 0.8 % plant agar, buffered to pH 5.8 with 0.5 M KOH. Then the seeds were incubated on plates at 4 °C for 48 h prior to germination under a 16-h light/8-h dark cycle. Next, soil-grown plants were maintained in growth rooms with a controlled light intensity of 120 µmol m−2 s−1 at 23 °C under a 16-h light period. The T-DNA mutants of the IRX10L gene (At5g61840, GABI_179G11) and the IRX10 gene (At1g27440, SALK046368) were identified by polymerase chain reaction (PCR) as previously reported (Wu et al. 2009).
Gene expression analysis
The total RNA was extracted from different tissues using the Aurum Total RNA mini kit (Bio-Rad, Hercules, CA) and digested with DNase I. Next, the iScript cDNA synthesis kit (Bio-Rad) was used for cDNA synthesis. The synthesized cDNA was used as a template for quantitative real time PCR (qRT-PCR) amplification using the SYBR Green PCR Kit (Takara, Japan). The primers for OsGT47A were OsGT47A forward (5′-AGATACTCAACGGGCTGGGAC-3′) and OsGT47A reverse (5′-GAGGACAGCAGCAACTATGTATGG-3′) and the internal standard primers of rice Alpha-tubulin (Os07g38730) were: tub forward (5′-TGTTGATTATGGAAAGAAGTCCAA-3′) and tub reverse (5′-GAGGACACTGTTGTATGGTTCTACA-3′). The PCR program contained an initial denaturation step of 1 min at 95 °C followed by denaturation for 15 s at 95 °C, annealing for 20 s at 60 °C, and extension for 20 s at 72 °C for 40 cycles. All qRT-PCR expression assays were independently performed and analyzed three times under identical conditions. The relative expression level of OsGT47A gene in the root was set to 1.
Complementation
The OsGT47A cDNA sequence from the ATG start codon to the stop codon was amplified by PCR using gene-specific gateway-compatible primers.
The products were cloned into the pDONR207 vector and sequenced. The gene sequences were transferred into the pEarleyGate 100 destination vector (Earley et al. 2006) to produce the 35S CaMV promoter fusion constructs that were transformed into the Arabidopsis irx10L (−/−) irx10 (+/−) mutant plants, because irx10L (−/−) irx10 (−/−) double mutant is sterile, using the floral-dip procedure (Clough and Bent 1998). Transgenic plants were selected by spraying 120 mg l−1 BASTA solution onto one-week-old seedlings in soil.
It should be pointed out that the irx10L (−/−) irx10 (+/−) mutant plants segregate into three genotypes in the next generation (double mutant irx10L (−/−) irx10 (−/−), irx10L (−/−) irx10 (+/−) and irx10L (−/−) in 1:2:1 ratios), only the double mutant irx10L (−/−) irx10 (−/−) have an obviously severe phenotype, and the other two genotypes grow as normally as wild type (Wu et al. 2009). Transgenic BASTA-positive plants will contain different genotype backgrounds, some plants with irx10L (−/−) irx10 (−/−) background, and some plants with irx10L (−/−) irx10 (+/−) or irx10L (−/−) background. Accordingly, to identify background of plants, three-week-old transformed plants were genotyped using the primers IRX10 forward (5′-CCACTCGGAGGACTTGGA-3′) and IRX10 reverse (5′-GGAAAAAGCCATTGAAAGAGG-3′) and the T-DNA-specific primer LBa1 (5′-TGGTTCACGTAGTGGGCCATCG-3′) [due to irx10(−)]. Only double-mutant background plants were picked up and plant growth was examined further.
OsGT47A transcripts in transgenic plants
Total RNA was extracted from 4-week-old plant leaves, then Reverse Transcription-PCR was using to examine expression of the OsGT47A gene in the Arabidopsis irx10 irx10L plants. The internal control transcripts used were Arabidopsis EF1a (28 cycles) employing the primers EF1a forward (5′-TCCAGCTAAGGGTGCC-3′) and EF1a reverse (5′-GGTGGGTACTCGGAGA-3′).
Constructs of proIRX10L:OsGT47A
The construct of proIRX10L:OsGT47A was made as follows: The primers ProIRX10L (5′-GCAAGCTTTGTAAAATGACCACTCGAGC-3′) and proIRX10L rev (5′-GCTCTAGATTTTCTCTCTCAGAAATTTTGGTTCC-3′) that contained HindIII and XbaI sites, respectively, were synthesized and used for PCR; amplified products were then digested and subcloned into the HindIII and XbaI sites of the gateway vector pGWB1; and the ProIRX10L:OsGT47A construct was obtained by gateway LR reaction.
Sectioning of stems
Stem tissues were cut from six-week-old soil-grown plants and were immediately transferred into fixative buffer containing 1.6 % (v/v) paraformaldehyde and 0.2 % (w/v) glutaraldehyde in 25 mM sodium phosphate, pH 7.2. Stem sections (50–80 μm) were cut using a Leica VT1000S vibratome (Leica Microsystems, http://www.leica-microsystems.com) with 3 % agarose as support, stained for 1–2 min in 0.02 % Toluidine blue O (Sigma-Aldrich, http://www.sigmaaldrich.com), rinsed and mounted in 50 % glycerol.
Analysis of the sugar composition of the cell wall
Stem samples were collected from six-week-old plants except for the double mutants, where plants were grown for 9 weeks, and placed into 80 % ethanol before freeze drying. The plant material was treated and fractionated and the sugar composition of alcohol-insoluble residues was analyzed using alditol acetate derivatives as described previously (Englyst and Cummings 1984a, b) with modifications (Wu et al. 2009).
Xylan immunolocalization
The basal stems were collected from six-week-old plants, and 50-μm-thick sections were cut using a vibratome for immunolocalizations (Freshour et al. 1996, 2003; Wu et al. 2009). Sections were incubated for 2 h together with the LM10 monoclonal antibodies (McCartney et al. 2005), and were then washed three times with phosphate-buffered saline (0.1 M sodium phosphate, pH 7.2, 0.5 M NaCl), followed by incubation for 2 h with fluorescein isothiocyanate (FITC)-conjugated antibodies. Following three further washes, the immunofluorescence was observed using a Zeiss LSM710 confocal microscope.
Results
The OsGT47A gene sequence is similar to AtIRX10
To examine whether there are similar mechanisms of xylan synthesis between monocots and dicots, one rice GT47 candidate gene was cloned. According to the rice genome sequence and BLAST results, the open reading frame (ORF) of OsGT47A cDNA, which is located at Os01g0926600/Os01g70190, was amplified from rice (Oryza sativa L. subsp. japonica) by PCR. OsGT47A encodes a protein of 415 amino acid residues with a predicted molecular mass of 46,901 Da and a predicted pI of 6.53 (http://www.scripps.edu/~cdputnam/protcalc.html). Next, we used BLAST searching to compare amino acid similarity in the NCBI database and found the sequence to have a high degree of conservation with Arabidopsis IRX10 and IRX10L. The protein sequence of OsGT47A showed a 93.49 % similarity to Arabidopsis IRX10 and IRX10-L (Fig. 1). The conserved exostosin motif in the GT47 family is extremely similar, including the proline that is altered by a missense mutation in fra8 (Zhong et al. 2005). Similar to IRX10 and IRX10-L, OsGT47A has a predicted signal peptide cleavage site at the N terminus between amino acids 27 and 28 (Fig. S1; http://www.cbs.dtu.dk/services/SignalP).
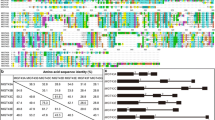
Amino acid sequence alignment of rice OsGT47A and Arabidopsis IRX10 and IRX10L proteins. The numbers at the right of each sequence are the positions of the amino acid residues in the corresponding proteins. Gaps (marked with dots) were introduced to maximize the sequence alignment. Identical and similar amino acid residues are shaded with black and gray colors, respectively. The conserved exostosin motif in pfam03016 is underlined. The proline amino acid residue (P), which is altered in the fra8 missense mutant, is indicated by a star
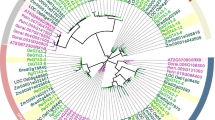
To investigate whether OsGT47A is a close homolog of IRX10 in the rice genome, we further analyzed the phylogeny of rice GT47 genes with several Arabidopsis GT47 genes, such as IRX10, FRA8, F8H and MUR3 (Madson et al. 2003; Wu et al. 2009, 2010; Zhong and Ye 2003). The Rice (Oryza sativa L. subsp. japonica) GT47 family contains 35 members, and their sequences were obtained from the website of the Carbohydrate-Active enZYmes Database (http://www.cazy.org/GT47.html). The overall subgrouping of the phylogenetic tree was very similar to that of Arabidopsis (Zhong and Ye 2003) and OsGT47A was clustered together with Arabidopsis IRX10 in branch I, and not with MUR3, FRA8 or F8H (Fig. 2). The high degree of sequence similarity between OsGT47A and IRX10 suggests that they may exhibit functional conservation, thus OsGT47A possibly providing clues to the xylan biosynthesis in monocot plants.
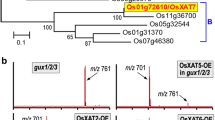
Phylogenetic tree of rice and Arabidopsis GT47 genes. The phylogenetic tree was constructed using MEGA software 5.05 by the CLUSTALW alignment and Neighbor-joining method and 1000 bootstrap replications. The scale bar indicates a branch length of 0.2. Os, Oryza sativa Japonica Group; At, Arabidopsis thaliana
The OsGT47A gene is highly expressed in the rice stem
The Arabidopsis IRX10 and IRX10L genes are highly expressed in vascular tissues and stems (Wu et al. 2009), which mainly contain xylan. To analyze the difference between dicots and monocots, the expression pattern of OsGT47A was detected using real time RT-PCR. The results showed that the expression level of the OsGT47A gene was highest in the stem, with lower expression detected in the leaf and embryo (Fig. 3). Additionally, we found that the OsGT47A gene was also highly expressed in the anther and five-day seeds with unknown reasons. These results demonstrated that the expression of OsGT47A is closely associated with the growth of rice stems.
qRT-PCR analysis of OsGT47A transcripts. OsGT47A has the highest expression level in the stem and a relatively high level in the anther and root, but a low level in the leaf and embryo. RNA isolated from rice tissues was used for gene expression analysis by qRT-PCR. Error bars denote the SE of three biological replicates
The OsGT47A gene can rescue or partially rescue the irx10 irx10L double mutant phenotypes
Overexpression of either Arabidopsis IRX10 or IRX10L gene can complement irx10 irx10L double mutant. To investigate whether OsGT47A is a functional homolog of Arabidopsis IRX10 and IRX10L, a fusion of the cauliflower mosaic virus (CaMV) 35S promoter with the full-length OsGT47A cDNA cassette was constructed and transformed into the Arabidopsis irx10L (−/−) irx10 (+/−) mutant plants. Complementation was examined by testing the genotypic background of transgenic plants, which segregate into three genotypic combinations in the next generation: the double mutant irx10L (−/−) irx10 (−/−), irx10L (−/−) irx10 (+/−), and irx10L (−/−). Finally, we got 36 positive transgenic plants with three different genotype backgrounds, and found only eight plants with the double mutant background from all 36 plants. Three of these eight plants exhibited a rescue of the irx10 irx10L mutant phenotypes, and the remaining five plants could partly rescue the double mutant phenotypes. This may be due to the transcription difference between those lines. Four transgenic lines were analyzed using RT-PCR, and the presence of the OsGT47A transcripts in the homozygous irx10 irx10L double mutant background was confirmed (Fig. 4a).
Complementation of the irx10 irx10L double mutant with OsGT47A. Restoration of plant size and stem growth in the Arabidopsis irx10 irx10L double mutant plants was caused by overexpression of the rice OsGT47A gene. a PCR data from three representative transgenic Arabidopsis lines. The OsGT47A transgene sample (upper) and the EF1a gene were used as an internal control (lower panel). WT represents wild type. b The irx10 irx10L double mutant (middle) has a small dwarf size, and overexpression of OsGT47A in the double mutant restored or partially restore the rosette size to that of the wild type (right), the numbers represent individual lines of transgenic plants
The irx10 irx10L double mutant exhibited a severe dwarf phenotype (Wu et al. 2009). Complementation of the double mutant with the OsGT47A gene can generate plants of the same normal size as the wild type (Fig. 4b). These plants had normal leaf number, size and stem height and a relatively high percentage of sterile siliques (Table 1). These results indicate that OsGT47A has a similar function to the Arabidopsis IRX10 gene.
Overexpressing OsGT47A gene can complement secondary cell wall formation in the irx10 irx10L double mutant
The irx10 irx10L double mutant was found to lose secondary cell wall growth (Wu et al. 2009, 2010), and only very thin cell walls in the interfascicular fibers and xylem vessels were detected in the double mutant (Fig. 5b, d). The thickness of the interfascicular fibers and xylem vessels in the OsGT47A-complemented double mutant plants had a shape that was close to wild type (Fig. 5a–f). These results revealed that the OsGT47A-complemented irx10 irx10L plants restore secondary wall thickness in fibers and vessels.
Secondary cell wall formation in stem tissues is restored in the irx10 irx10L mutant plants by overexpression of rice OsGT47A genes. Transverse sections of stem tissues. a, d Wild type. b, e irx10 irx10L double mutants. c, f OsGT47A plant. Arrows in b point to xylem vessels that have collapsed in irx10 irx10L stems, whereas arrows point to xylem vessels of normal appearance in wild-type (a) and OsGT47A-overexpressing plants (c). co cortex, if interfascicular fiber, ph phloem, ve vessel. Scale bars 20 μm
The OsGT47A gene can restore xylose composition and xylan in the irx10 irx10L mutant
The irx10 irx10L double mutation is mainly defective in xylan backbone synthesis, and the mutation affects the sugar composition of the mutant compared with the wild type (Brown et al. 2009; Wu et al. 2009, 2010). The irx10 irx10L double mutation reduced the xylose content to nearly 8 % of the wild type. The mutant, however, exhibited increased content with regard to all other monosaccharides, such as arabinose, fucose, galactose and glucose. In the complementation plants, the content of xylose was close to that of wild type, suggesting that OsGT47A expression in irx10 irx10L mutants completely complements the level of cell wall monosaccharides to the wild-type level (Table 2).
Immunofluorescence detection of xylem using LM10 antibody shows restoration of xylans in complemented plants
To examine whether overexpression of OsGT47A can rescue the xylan deficiency in irx10 irx10L mutants, the LM10 monoclonal antibody, which recognizes 4-O-methylglucuronoxylan (McCartney et al. 2005), was used to detect xylan localization in stem sections. It has been shown that cross-sections of wild-type stems have strong immunostaining signals in the xylem and interfascicular fibers cell walls. No signal was detected in irx10 irx10L mutant and the xylan fluorescence signals were restored in the complemented plants (Fig. 6).
OsGT47A may be a functional homolog of AtIRX10L
OsGT47A under the control of CaMV 35S promoter can rescue or partially rescue the irx10 irx10L double mutant. To test whether OsGT47A and IRX10/IRX10L proteins are functionally equivalent, the native promoter ProIRX10 was used for complementation experiments. In Arabidopsis, when the proIRX10L:IRX10L transgene was transformed, it partially complemented the irx10 irx10L double mutant, giving an intermediate phenotype similar to irx10 irx10L (+/−) plants. In contrast, proIRX10-L:IRX10 transgenic plants can fully recover from the mutant phenotypes (Wu et al. 2009). When the proIRX10-L:OsGT47A transgene was present in irx10 irx10L mutants, it partially rescued the irx10 irx10L double mutant with an intermediate phenotype as irx10 irx10L (+/−) plants (Fig. 7). The result indicates that OsGT47A might be functionally equivalent to AtIRX10L.
Discussion
The GT47 family shares the β-glucuronyl transferase domain with animal exostosins, and many GT47 family members have been identified in the dicot Arabidopsis and the monocot rice (Cao et al. 2008; Zhong and Ye 2003). On the basis of microarray analyses, we identified 35 genes in the GT47 family after excluding gene fragments (Fig. 2). Phylogenetic analysis suggested that the Arabidopsis GT47 family can be divided into four major groups, and the AtIRX10 and AtIRX10L genes are located in group A (Zhong and Ye 2003). It is interesting that there have been six IRX10-related genes identified in rice, but only two genes (AtIRX10 and AtIRX10L) in Arabidopsis. However, in the branch containing AtFRA8 and AtF8H, there is only one gene in rice Os03g0107900 versus two genes in Arabidopsis (AtFRA8 and AtF8H) (Fig. 2). IRX10 and IRX10L are related to xylan backbone elongation, similar to GT43 family members, such as IRX9 and IRX14 (Brown et al. 2009; Wu et al. 2009) and FRA8 functions on the xylan reducing end (Pena et al. 2007), though the xylan reducing end has not been detected in monocots. Thus, the evolution of more genes for the xylan backbone and fewer genes for the xylan reducing end makes sense. Further, we also found that there are twice as many members of the GT43 family in rice compared with Arabidopsis (data not shown).
In the monocot rice, xylan mainly exists as GAX (Faik 2010). Investigation of the biosynthetic mechanism of xylan at the biochemical and molecular levels in monocots is more challenging and complicated because grasses have more branch structures and unknown backgrounds. Recently, Chen et al. (2013) identified one rice dissociation (DS) transposon insertion mutant in the gene cluster containing GT47 subfamily and named the gene as OsIRX10 (Os01g70200/Os01g0926600). Further mutant analysis showed decreased xylan content, especially in rice culm. Here we analyzed another homologous gene and showed that it can complement the Arabidopsis double mutant and rescue secondary cell wall growth and relative components. Our results support those of Chen et al. (2013) and suggest that the genes involved in xylan backbone biosynthesis might be conserved in the dicot Arabidopsis and the monocot rice. Because GAX is a rich energy resource in rice-producing regions, the understanding of GX biosynthesis would provide novel strategies for the improvement of rice straw utilization.
Recently, Lee et al. (2012) reported that overexpression of either IRX9 or IRX14 alone in the tobacco BY2 protoplast cells can not give rise to XylT activity, but overexpression of IRX9 and IRX14 proteins simultaneously can give rise to XylT activity. Although the IRX10 mutant affected xylan backbone elongations, it is still not evident whether IRX10 exhibits XylT activity (Brown et al. 2009; Wu et al. 2009). We also heterologously expressed Arabidopsis and rice IRX10 genes in BY2 protoplast cells, but no XylT activities were detected (data not shown). This is because the complex formation including IRX10 is require for XylT activity.
In conclusion, we have reported that the monocot rice OsGT47A gene is highly expressed in the rice stem. Overexpression of OsGT47A in Arabidopsis irx10 irx10L double mutant can complement the mutant phenotype and restore the secondary cell wall thickness and monosaccharide content. These results reveal that OsGT47A most likely has an identical biochemical function to that of IRX10. Because IRX10 is required for xylan backbone biosynthesis in Arabidopsis, OsGT47A in rice is most likely involved in xylan biosynthesis during secondary cell wall formation.
Abbreviations
- IRX :
-
IRREGULAR XYLEM
- CaMV:
-
Cauliflower mosaic virus
- GAX:
-
Glucuronoarabinoxylan
- GlcA:
-
Glucuronic acid
- GT:
-
Glycosyltransferase
- qRT-PCR:
-
Quantitative real time-PCR
- GX:
-
Glucronoxylan
- PCR:
-
Polymerase chain reaction
- XylT:
-
Xylosyltransferase
References
Anders N, Wilkinson MD, Lovegrove A, Freeman J, Theodora T, Pellny TK, Weimar T, Mortimer JC, Stott K, Baker JM, Defoin-Platel M, Shewry PR, Dupree P, Mitchell RAC (2012) Glycosyl transferases in family 61 mediate arabinofuranosyl transfer on to xylan in grasses. Proc Natl Acad Sci USA 109:989–993
Andersson SI, Samuelson O, Ishihara M, Shimizu K (1983) Structure of the reducing end-groups in spruce xylan. Carbohydr Res 111:283–288
Bauer S, Vasu P, Persson S, Mort AJ, Somerville CR (2006) Development and application of a suite of polysaccharides degrading enzymes for analyzing plant cell walls. Proc Natl Acad Sci USA 103:11417–11422
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Brown DM, Zeef LAH, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17:2281–2295
Brown DM, Goubet F, Vicky WWA, Goodacre R, Stephens E, Dupree P, Turner SR (2007) Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of xylan synthesis. Plant J 52:1154–1168
Brown DM, Zhang Z, Stephens E, Dupree P, Turner SR (2009) Characterization of IRX10 and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis. Plant J 57:732–746
Cao PJ, Bartley LE, Jung KH, Ronald PC (2008) Construction of a rice glycosyltransferase phylogenomic database and identification of rice-diverged glycosyltransferases. Mol Plant 1:858–877
Carroll A, Somerville C (2009) Cellulosic biofuels. Annu Rev Plant Biol 60:165–182
Chen X, Vega-Sánchez ME, Verhertbruggen Y, Chiniquy D, Canlas PE, Fagerström A, Prak L, Christensen U, Oikawa A, Chern M, Zuo S, Lin F, Auer M, Willats WGT, Bartley L, Harholt J, Scheller HV, Ronald PC (2013) Inactivation of OsIRX10 leads to decreased xylan content in rice culm cell walls and improved biomass saccharification. Mol Plant 6:570–573
Chiniquy D, Sharma V, Schultink A, Baidoo EE, Rautengarten C, Cheng K, Carroll A, Ulvskov P, Harholt J, Keasling JD, Pauly M, Scheller HV, Ronald PC (2012) XAX1 from glycosyltransferase family 61 mediates xylosyltransfer to rice xylan. Proc Natl Acad Sci USA 109:17117–17122
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Earley KW, Haag JR, Pontes O, Opper K, Juehne T, Song K, Pikaard CS (2006) Gateway-compatible vectors for plant functional genomics and proteomics. Plant J 45:616–629
Ebringerová A, Heinze T (2000) Xylan and xylan derivatives—biopolymers with valuable properties. 1. Naturally occurring xylans structures, isolation procedures and properties. Macromol Rapid Commun 21:542–556
Ebringerová A, Heinze T (2005) Hemicellulose. Adv Polym Sci 186:1–67
Englyst HN, Cummings JH (1984a) Simplified method for the measurement of total non-starch polysaccharides by gas-liquid chromatography of constituent sugars as alditol acetates. Analyst (Lond) 109:937–942
Englyst HN, Cummings JH (1984b) Simplified method for the measurement of total non-starch polysaccharides by gas-liquid chromatography of constituent sugars as alditol acetates. Analyst (Lond) 109:937–942
Faik A (2010) Xylan biosynthesis: news from the grass. Plant Physiol 153:396–402
Forsthoefel NR, Wu Y, Schultz B, Bennett MJ, Feldmann KA (1992) T-DNA insertion mutagenesis in Arabidopsis: prospects and perspectives. Aust J Plant Physiol 19:353–366
Freshour G, Clay RP, Fuller MS, Albersheim P, Darvill AG, Hahn MG (1996) Developmental and tissue-specific structural alterations of the cell-wall polysaccharides of Arabidopsis thaliana roots. Plant Physiol 110:1413–1429
Freshour G, Bonin CP, Reiter WD, Albersheim P, Darvill AG, Hahn MG (2003) Distribution of fucose-containing xyloglucans in cell walls of the mur1 mutant of Arabidopsis. Plant Physiol 131:1602–1612
Johansson MH, Samuelson O (1977) Reducing end groups in Birch xylan and their alkaline degradation. Wood Sci Technol 11:251–263
Kong Y, Zhou G, Avic U, Gu X, Jones C, Yin Y, Xu Y, Hahn MG (2009) Two poplar glycosyltransferase genes, PdGATL1.1 and PdGATL1.2 are functional orthologs to PARVUS/AtGATL1 in Arabidopsis. Mol Plant 2:1040–1050
Lee C, O’Neill MA, Tsumuraya Y, Darvill AG, Ye Z-H (2007) The irregular xylem 9 mutant is deficient in xylan xylosyltransferase activity. Plant Cell Physiol 48:1624–1634
Lee C, Teng Q, Huang W, Zhong R, Ye Z-H (2010) The Arabidopsis family GT43 glycosyltransferases from two functionally nonredudant groups essential for elongation of glucuronoxylan backbone. Plant Physiol 153:526–541
Lee C, Zhong R, Ye Z-H (2012) Arabidopsis family GT43 members are xylan xylosyltransferases required for the elongation of the xylan backbone. Plant Cell Physiol 53:135–143
Lerouxel O, Cavalier DM, Liepman AH, Keegstra K (2006) Biosynthesis of plant cell wall polysaccharides—a complex process. Curr Opin Plant Biol 9:621–630
Liepman AH, Wightman R, Geshi N, Turner SR, Scheller HV (2010) Arabidopsis—a powerful model system for plant cell wall research. Plant J 61:1107–1121
Madson M, Christophe D, Li X, Verma R, Vanzin GF, Caplan J, Shoue DA, Carpita NC, Reiter WD (2003) The MUR3 gene of Arabidopsis encodes a xyloglucan galactosyltransferase that is evolutionarily related to animal exostosins. Plant Cell 15:1662–1670
McCartney L, Marcus SE, Knox JP (2005) Monoclonal antibodies to plant cell wall xylans and arabinoxylans. J Histochem Cytochem 53:543–546
Mitchell RA, Dupree P, Shewry PR (2007) A novel bioinformatics approach identifies candidate genes for the synthesis and feruloylation of arabinoxylan. Plant Physiol 144:43–53
Pauly M, Gille S, Liu L, Mansoori N, Souza AD, Schultink A (2013) Hemicellulose biosynthesis. Planta 238:627–642
Pena MJ, Zhong R, Zhou GK, Richardson EA, O’neill MA, Darvill AG, York WS, Ye Z-H (2007) Arabidopsis irregular xylem8 and irregular xylem9: implications for the complexity of glucuronoxylan biosynthesis. Plant Cell 19:549–563
Persson S, Hosmer Caffall K, Freshour G, Hilley MT, Bauer S, Poindexter P, Hahn MG, Mohnen D, Somerville C (2007) The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19:237–255
Scheible WR, Pauly M (2004) Glycosyltransferases and cell wall biosynthesis: novel players and insights. Curr Opin Plant Biol 7:285–295
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289
Somerville CR (2006) Cellulose synthesis in higher plants. Annu Rev Cell Dev Biol 22:53–78
Turner S, Somerville C (1997) Collapsed xylem phenotype of Arabidopsis identifies mutants deficient in cellulose deposition in the secondary cell wall. Plant cell 9:689–701
Wu AM, Rihouey C, Seveno M, Hörnblad E, Singh SK, Matsunaga T, Ishii T, Lerouge P, Marchant A (2009) The Arabidopsis IRX10 and IRX10-LIKE glycosyltransferases are critical for glucuronoxylan biosynthesis during secondary cell wall formation. Plant J 57:718–731
Wu AM, Hornblad E, Voxeur A, Gerber L, Rihouey C, Lerouge P, Marchant A (2010) Analysis of the Arabidopsis IRX9/IRX9-L and IRX14/IRX14-L pairs of glycosyltransferase genes reveals critical contributions to biosynthesis of the hemicellulose glucuronoxylan. Plant Physiol 153:542–554
Zeng W, Jiang N, Nadella R, Killen TL, Nadella V, Faik A (2010) A glucurono(arabino)xylan synthase complex from wheat contains members of the GT43, GT47, and GT75 families and functions cooperatively. Plant Physiol 154:78–97
Zhong R, Ye Z-H (2003) Unraveling the functions of glycosyltransferase family 47 in plants. Trends Plant Sci 8:565–568
Zhong R, Pena MJ, Zhou G-K, Naim CJ, Wood-Jones A, Richardson EA, Morrison WH, Darvill AG, York WS, Ye Z-H (2005) The FRA8 gene which encodes a putative glucuronyltransferase is essential for normal secondary wall synthesis. Plant Cell 17:3390–3408
Zhou GK, Zhong R, Richardson EA, Morrison WH, Nairn CJ, Wood-Jones A, Ye Z-H (2006) The poplar glycosyltransferase GT47C is functionally conserved with Arabidopsis Fragile Fiber8. Plant Cell Physiol 47:1229–1240
Zhou GK, Zhong R, Himmelsbach DS, McPhail BT, Ye Z-H (2007) Molecular characterization of PoGT8D and PoGT43B, two secondary wall-associated glycosyltransferases in poplar. Plant Cell Physiol 48:689–699
Acknowledgments
We thank two anonymous reviewers for their constructive comments and suggestions. This work was supported by the National Natural Science Foundation of China (Grant Number 31170165) and the Natural Science Foundation supported by Guangdong Province (Grant Number S2011010001138).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, B., Zhao, T., Yu, W. et al. Functional conservation of the glycosyltransferase gene GT47A in the monocot rice. J Plant Res 127, 423–432 (2014). https://doi.org/10.1007/s10265-014-0631-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-014-0631-5