Abstract
Green-colored plastids in the dinoflagellates Lepidodinium chlorophorum and L. viride have been widely believed as the remnant of an endosymbiotic prasinophyte. This hypothesis for the origin of the Lepidodinium plastids is solely based on an unpublished result quoted in Elbrächter and Schnepf (Phycologia 35:381–393, 1996) hinting at the presence of a characteristic carotenoid in prasinophytes, prasinoxanthin, in the L. chlorophorum cells. On the other hand, a recent work failed to detect prasinoxanthin in a culture of L. chlorophorum. Unfortunately, we cannot conduct any additional experiments to examine whether the two strains considered in the previous studies are truly of L. chlorophorum, as neither of the two strains is publicly available. We here investigated the pigment composition of L. chlorophorum strain NIES-1868 maintained as a mono-algal culture under laboratory conditions, and detected no sign of prasinoxanthin. The pigment composition of strain NIES-1868 is consistent with previous phylogenetic analyses based on plastid-encoded genes of the same strain, which successfully excluded prasinoxanthin-containing algae from the origin of the L. chlorophorum plastid. We also determined nucleus-encoded 18S ribosomal RNA (rRNA) genes from four Lepidodinium strains (including strain NIES-1868). Analyses of 18S rRNA sequences showed an extremely close relationship among strain NIES-1868 and other Lepidodinium cells/strains originating from different geological locations, suggesting that the cells/strains corresponding to these rRNA sequences lack prasinoxanthin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The majority of photosynthetic dinoflagellates possess plastids containing chlorophylls (Chl) a+c and a unique carotenoid, peridinin (Taylor 1980). The peridinin-containing plastids are most likely derived from a red algal endosymbiont. However, some dinoflagellate lineages are known to bear non-canonical plastids lacking peridinin which are classified into seven types according to Moestrup and Daugbjerg (2007). Traditionally, pigment compositions have been used to infer the origin of the non-canonical plastids; for example, the non-canonical plastids containing 19′-hexanoyloxyfucoxanthin, fucoxanthin, and alloxanthin advocated the evolutionary connections to the plastids in haptophytes, diatoms, and cryptophytes, respectively (Withers and Haxo 1975; Withers et al. 1977; Suzuki and Ishimaru 1992; Meyer-Harms and Pollehne 1998; De Salas et al. 2003; Bergholtz et al. 2005; Tamura et al. 2005). The origins of these non-canonical plastids inferred from pigment compositions were confirmed by phylogenetic analyses of plastid-encoded genes (e.g., Takishita et al. 1999, 2002; Tengs et al. 2000; Horiguchi and Takano 2006). Importantly, the remote relationship amongst the dinoflagellate (host) lineages bearing non-canonical plastids indicates that the plastid replacement events took place in separate branches in dinoflagellate phylogeny (Saldarriaga et al. 2001; Shalchian-Tabrizi et al. 2006).
The genus Lepidodinium bears non-canonical plastids containing Chl-a+b (Watanabe et al. 1987, 1990), but their pigment composition has not been fully settled. Lepidodinium plastids have been stated to contain prasinoxanthin, a carotenoid exclusively found in a few members of the green algal class Prasinophyceae, implying that this dinoflagellate lineage acquired its current non-canonical plastids through the endosymbiosis of a prasinophyte (Elbrächter and Schnepf 1996). Nevertheless, the presence of prasinoxanthin in L. chlorophorum was quoted as a personal communication in Elbrächter and Schnepf (1996), and therefore, needs to be re-examined. To elucidate the precise origin of the Lepidodinium plastids, it would be ideal if the pigment composition, cell morphology, and gene sequences are available for the same strain maintained in the laboratory. Recently, Laza-Martinez et al. (2007) called the popular hypothesis for the origin of the Lepidodinium plastids into question, as no sign of prasinoxanthin was detected in the L. chlorophorum culture. Laza-Martinez et al. (2007) admitted that the L. chlorophorum culture examined was unfortunately not mono-algal, and they did not provide morphological or sequence data for the particular strain. Furthermore, to our knowledge, the prasinoxanthin-lacking ‘L. chlorophorum’ strain has not been deposited in any culture collection.
Some of us have studied the evolutionary characteristics of the L. chlorophorum plastid by sequencing 11 plastid-encoded genes of L. chlorophorum strains (Takishita et al. 2008; Matsumoto et al. 2011a, b). The results showed that the plastids of the L. chlorophorum strains originated from one of “core chlorophytes,” which did not contain prasinoxanthin. These strains have been maintained as mono-algal cultures in the laboratory, and their morphologies have been well characterized. Thus, these strains can be used to address the pigment composition of L. chlorophorum. In this study, we investigated the pigment composition of the L. chlorophorum strain NIES-1868, and revealed that this strain apparently lacked prasinoxanthin. In addition, we newly determined the nucleus-encoded 18S ribosomal RNA (rRNA) genes of strains NIES-1867 and NIES-1868, as well as those of two another Lepidodinium strains. The rRNA sequences from the four Lepidodinium strains determined in this study and four of those deposited under the name of L. chlorophorum or L. viride in GenBank database shared nucleotide identity of ≥99 %, and formed a tight clade in our phylogenetic analyses. Thus, the Lepidodinium species, of which 18S rRNA sequences bear high identities to that of strain NIES-1868, most likely lack prasinoxanthin.
Materials and methods
Algal cells
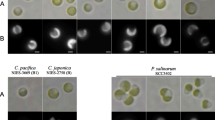
Two strains of L. chlorophorum (NIES-1867 and NIES-1868) were purchased from the Microbial Culture Collection at the National Institute for Environmental Studies (NIES; 16-2, Onogawa, Tsukuba, Ibaraki 305-8506, Japan). We considered the two strains to belong to identical species (or extremely closely related species) because (1) they originated from the exact same water sample collected in the East China Sea (see Takishita et al. 2008), (2) their cell morphologies were identical under the light microscope (data not shown), and (3) most gene sequences available for both strains were identical (see below). Additionally we examined two original clones of Lepidodinium strains: strain ‘MH 360,’ which was confirmed as Lepidodinium sp. by light microscopy, and strain ‘T03-1,’ which was identified as L. viride based on the presence of the body scales on the cell surface. Strain MH 360 was isolated from sea water collected offshore of the Sanriku region, Japan (Pacific Ocean), on September 9, 2009, while strain T03-1 was isolated from sea water collected offshore southeast of Phuket Island, Thailand (Andaman Sea), on September 18, 2003. The four strains were cultured in MNK medium (Noël et al. 2004) at 20 °C under a 14-h light/10-h dark cycle.
The prasinophycean green alga Micromonas pusilla (NIES-1411) was purchased from the Microbial Culture Collection at NIES and cultured in f/2 medium (Guillard and Ryther 1962) at 20 °C under a 14/10 h LD cycle.
Pigment analysis
Lepidodinium chlorophorum strain NIES-1868 and M. pusilla cells were harvested by centrifugation. After removal of a transparent viscosity layer on the cell pellet, pigments were extracted with 100 μl of 100 % methanol, and the pigment extract was collected into a 1.5 ml centrifuge tube after centrifugation. We repeated the extraction until the extract was no longer colored. The extracted pigments were subjected to reverse-phase high-performance liquid chromatography (HPLC) with a Waters Symmetry C8 column (150 mm × 4.6 mm; particle size 3.5 μm; pore size 100 Å). The HPLC was performed as described in Zapata et al. (2000) without any modifications. The eluted pigments were detected by the absorbance at 440 nm and identified by their elution patterns compared to those reported in Zapata et al. (2000).
Preparation of nucleic acids, PCR, cloning, and sequencing
From the Lepidodinium cells, total DNA samples were extracted using the DNA Isolation Reagent (TaKaRa), and then, served to amplify nucleus-encoded 18S rRNA genes. We amplified 18S rRNA genes from total DNA by a pair of primers: 5′-GAAACTGCGAATGGCTCT-3′ and 5′-CCGCAGGTTCACCTAC-3′. The amplification conditions consisted of 30 cycles of 96 °C for 20 s, 55 °C for 20 s, and 72 °C for 3 min. Amplicons were cloned into pGEM T-Easy vector (Promega), and both strands were completely sequenced. The rRNA gene sequences determined in this study were deposited in DDBJ/EMBL/GenBank under the accession numbers AB686253–AB686256.
Phylogenetic analysis
We retrieved nucleus-encoded 18S rRNA sequences of phylogenetically diverged dinoflagellates, including five sequences deposited as Lepidodinium, as well as closely related alveolates, from NCBI GenBank database. These retrieved sequences were then aligned with the four Lepidodinium sequences determined in this study. The details of Lepidodinium 18S rRNA sequences used in this study are listed in Table 1. The initial alignment was prepared by CLUSTALW ver. 1.83 (Thompson et al. 1994), and then manually refined. After exclusion of all ambiguously aligned positions, the final 18S rRNA alignment included 105 taxa with 1,496 nucleotide positions.
The rRNA data set was subjected to the maximum-likelihood (ML) analysis under the GTR model incorporating among-site rate variation approximated by a discrete Γ distribution with four equally probable categories (GTR + Γ model) by using RAxML ver. 7.2.1 (Stamatakis 2006). The ML tree was heuristically searched from 10 distinct parsimony trees. A ML bootstrap analysis (100 replicates) was conducted under the GTR + Γ model with heuristic tree search from a single parsimony tree per replicate.
The same alignment was also subjected to Bayesian analysis with the GTR + Γ model by using MrBayes v.3.1.2 (Ronquist and Huelsenbeck 2003). Two parallel Markov chain Monte Carlo runs, each consisting of four incrementally heated chains, were run for 106 generations, sampling log-likelihoods and trees at 100-generation intervals. The first 20,000 generations were discarded as “burn-in,” and the majority consensus tree and Bayesian posterior probabilities were calculated from the remaining trees.
Results
Pigment composition
Micromonas pusilla is one of the standard algae containing prasinoxanthin (Zapata et al. 2000). As anticipated, our HPLC analysis of M. pusilla successfully identified prasinoxanthin, as well as Mg-3,8-divinyl-pheoporphyrin a5 monomethyl ester (MgDVP), uriolide, 9′-cis-neoxanthin, violaxanthin, micromonal, chlorophyll b (Chl b), chlorophyll a (Chl a), β-carotene, and an unknown carotenoid (Fig. 1, top). In contrast, in the same analysis of L. chlorophorum strain NIES-1868, 9′-cis-neoxanthin, violaxanthin, antheraxanthin, zeaxanthin, lutein, Chl b, Chl a, and β-carotene were detected, while prasinoxanthin was not found (Fig. 1, bottom). The overall pigment composition in strain NIES-1868 is of typical green algae, providing no clue for the origin of the L. chlorophorum plastid.
18S rRNA analysis
We newly determined 18S rRNA sequences from four Lepidodinium strains NIES-1867, NIES-1868, MH 360, and T03-1. The rRNA sequences from our strains determined here and the four sequences deposited in GenBank database—L. viride (AF022199), L. viride (DQ499645), L. chlorophorum (AY331681), and L. chlorophorum (AM184122)—appeared to be identical or almost identical (Table 2). In the 18S rRNA phylogeny, the eight Lepidodinium sequences tightly grouped together, and then, this Lepidodinium clade was placed within the dinoflagellate clade (Fig. 2). Interestingly, the rRNA sequence deposited as ‘L. chlorophorum’ (JF791033) shared lower nucleotide identities with those of other Lepidodinium species (~96 %; Table 2), and it was excluded from the Lepidodinium clade (Fig. 2). Of note, Gymnodinium catenatum, instead of ‘L. chlorophorum’ (JF791033), branched at the base of the Lepidodinium clade, although the relative position of JF791033 to other Lepidodinium sequences was inconclusive (Fig. 2).
Maximum-likelihood phylogeny based on nucleus-encoded 18S rRNA sequences. Only bootstrap values ≥50 % are presented. The bipartitions supported by ≥0.95 Bayesian posterior probabilities are highlighted by thick lines. The Lepidodinium sequences are shaded. The rRNA sequences determined in this study are highlighted by arrowheads
Discussion
We here investigated the pigment composition of L. chlorophorum strain NIES-1868, which is mono-algal culture maintained in the laboratory. Our results presented here indicate that L. chlorophorum strain NIES-1868 lacks prasinoxanthin, and the current Chl-a+b-containing plastid in this strain might be derived from a prasinoxanthin-lacking green alga. These data are consistent with the data presented in Laza-Martinez et al. (2007), suggesting no prasinoxanthin in their L. chlorophorum culture. Importantly, the data on the pigment from strain NIES-1868 can be directly combined with the plastid-encoded gene sequences from the same strain determined in Matsumoto et al. (2011b). Indeed, the absence of prasinoxanthin is apparently consistent with a multigene phylogenetic analysis, which successfully excluded the evolutionary affinity between the plastid in strain NIES-1868 and any of those containing prasinoxanthin (Matsumoto et al. 2011b). Thus, we firmly conclude that (1) the L. chlorophorum strain NIES-1868 lacks prasinoxanthin and (2) the current Chl-a+b-containing plastid in this strain is derived from a prasinoxanthin-lacking green alga, as suggested in Matsumoto et al. (2011b).
The analyses of nucleus-encoded 18S rRNA sequences clearly indicated that the sequence of strain NIES-1868 was very closely related to those of the Lepidodinium cells sampled at various geographical locations. Thus, both pigment composition and phylogenetic data based on plastid-encoded genes from strain NIES-1868 discussed in the previous paragraph are most likely applicable to the cells corresponding to the rRNA sequences, which robustly grouped with the NIES-1868 sequence in the ML analysis. The rRNA sequence deposited as JF791033 in GenBank database, which was excluded from the Lepidodinium clade, may not have originated from a member of the genus Lepidodinium.
References
Bergholtz T, Daugbjerg N, Moestrup Ø, Fernandez-Tejedor M (2005) On the identity of Karlodinium veneficum and description of Karlodinium armiger sp. nov. (Dinophyceae), based on light and electron microscopy, nuclear-encoded LSU rDNA, and pigment composition. J Phycol 42:170–193
De Salas MF, Bolch CJS, Botes L, Nash G, Wright SW, Hallegraeff GM (2003) Takayama gen. nov. (Gymnodiniales, Dinophyceae), a new genus of unarmored dinoflagellates with sigmoid apical grooves, including the description of two new species. J Phycol 39:1233–1246
Elbrächter M, Schnepf E (1996) Gymnodinium chlorophorum, a new, green, bloom-forming dinoflagellate (Gymnodiniales, Dinophyceae) with a vestigial prasinophyte endosymbiont. Phycologia 35:381–393
Guillard RR, Ryther JH (1962) Studies of marine planktonic diatoms. I. Cyclotella nana Hustedt, and Detonula confervacea (Cleve) Gran. Can J Microbiol 8:229–239
Horiguchi T, Takano Y (2006) Serial replacement of a diatom endosymbiont in the marine dinoflagellate Peridinium quinquecorne (Peridiniales, Dinophyceae). Phycol Res 54:193–200
Laza-Martinez A, Seoane S, Zapata M, Orive E (2007) Phytoplankton pigment patterns in a temperate estuary: from unialgal cultures to natural assemblages. J Plankton Res 29:913–929
Matsumoto T, Ishikawa SA, Hashimoto T, Inagaki Y (2011a) A deviant genetic code in the green alga-derived plastid in the dinoflagellate Lepidodinium chlorophorum. Mol Phylogenet Evol 60:68–72
Matsumoto T, Shinozaki F, Chikuni T, Yabuki A, Takishita K, Kawachi M, Nakayama T, Inouye I, Hashimoto T, Inagaki Y (2011b) Green-colored plastids in the dinoflagellate genus Lepidodinium are of core chlorophyte origin. Protist 162:268–276
Meyer-Harms B, Pollehne F (1998) Alloxanthin in Dinophysis norvegica (Dinophysiales, Dinophyceae) from the Baltic Sea. J Phycol 34:280–285
Moestrup Ø, Daugbjerg N (2007) On dinoflagellate phylogeny and classification. In: Brodie J, Lewis J (eds) Unravelling the algae: the past, present, and future of algal systematics. CRC Press, Boca Raton, pp 215–230
Noël MH, Kawachi M, Inouye I (2004) Induced dimorphic life cycle of a coccolithophorid, Calyptrosphaera sphaeroidea (Prymnesiophyceae, Haptophyta). J Phycol 40:112–129
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Saldarriaga JF, Taylor FJR, Keeling PJ, Cavalier-Smith T (2001) Dinoflagellate nuclear SSU rRNA phylogeny suggests multiple plastid losses and replacements. J Mol Evol 53:204–213
Shalchian-Tabrizi K, Minge MA, Cavalier-Smith T, Nedreklepp JM, Klaveness D, Jakobsen KS (2006) Combined heat shock protein 90 and ribosomal RNA sequence phylogeny supports multiple replacements of dinoflagellate plastids. J Eukaryot Microbiol 53:217–224
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Suzuki R, Ishimaru T (1992) Characteristics of photosynthetic pigment composition of Gymnodinium mikimotoi Miyake et Kominami ex ODA. J Oceanogr 48:367–375
Takishita K, Nakano K, Uchida A (1999) Preliminary phylogenetic analysis of plastid-encoded genes from an anomalously pigmented dinoflagellate Gymnodinium mikimotoi (Gymnodiniales, Dinophyta). Phycol Res 47:257–262
Takishita K, Koike K, Maruyama T, Ogata T (2002) Molecular evidence for plastid robbery (kleptoplastidy) in Dinophysis, a dinoflagellate causing diarrhetic shellfish poisoning. Protist 153:293–302
Takishita K, Kawachi M, Noël MH, Matsumoto T, Kakizoe N, Watanabe MM, Inouye I, Ishida K, Hashimoto T, Inagaki Y (2008) Origins of plastids and glyceraldehyde-3-phosphate dehydrogenase genes in the green-colored dinoflagellate Lepidodinium chlorophorum. Gene 410:26–36
Tamura M, Shimada S, Horiguchi T (2005) Galeidinium rugatum gen. et sp. nov. (Dinophyceae), a new coccoid dinoflagellate with as diatom endosymbiont. J Phycol 41:658–671
Taylor FJR (1980) On dinoflagellate evolution. Biosystems 13:65–108
Tengs T, Dahlberg OJ, Shalchian-Tabrizi K, Klaveness D, Rudi K, Delwiche CF, Jakobsen KS (2000) Phylogenetic analyses indicate that the 19′ hexanoyloxy-fucoxanthin-containing dinoflagellates have tertiary plastids of haptophyte origin. Mol Biol Evol 17:718–729
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Watanabe MM, Takeda Y, Sasa T, Inouye I, Suda S, Sawaguchi T, Chihara M (1987) A green dinoflagellate with chlorophylls a and b: morphology, fine structure of the chloroplast and chlorophyll composition. J Phycol 23:382–389
Watanabe MM, Suda S, Inouye I, Sawaguchi T, Chihara M (1990) Lepidodinium viride gen. et sp. nov. (Gymnodiniales, Dinophyta), a green dinoflagellate with a chlorophyll a- and b-containing endosymbiont. J Phycol 26:741–751
Withers N, Haxo FT (1975) Chlorophyll c 1 and c 2 and extraplastidic carotenoids in the dinoflagellate, Peridinium foliaceum Stein. Plant Sci Lett 5:7–15
Withers NW, Cox ER, Tomas R, Haxo FT (1977) Pigments of the dinoflagellate Peridinium balticum and its photosynthetic endosymbiont. J Phycol 13:354–358
Zapata M, Garrido JL, Rodríguez F (2000) Separation of chlorophylls and carotenoids from marine phytoplankton: a new HPLC method using a reversed phase C8 column and pyridine-containing mobile phases. Mar Ecol Prog Ser 195:29–45
Acknowledgments
We thank Drs. R. Kamikawa (University of Tsukuba, Japan), T. Hashimoto (University of Tsukuba, Japan), and K. Takishita (JAMSTEC, Japan) for their critical comments on this manuscript. We also thank Dr. M. H. Noël (NIES, Japan) for providing Lepidodinium strains NIES-1867, 1868, and MH360. TM is a research fellow supported by the Japan Society for Promotion of Sciences (JSPS) for Young Scientists (No. 21508). This work was supported by grants from JSPS and the Ministry of Education, Culture, Sports, Science and Technology of Japan (No. 21370031 and 23117006) awarded to YI.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsumoto, T., Kawachi, M., Miyashita, H. et al. Prasinoxanthin is absent in the green-colored dinoflagellate Lepidodinium chlorophorum strain NIES-1868: pigment composition and 18S rRNA phylogeny. J Plant Res 125, 705–711 (2012). https://doi.org/10.1007/s10265-012-0486-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-012-0486-6