Abstract
In nature, plants can recognize potential pathogens, thus activating intricate networks of defense signals and reactions. Inducible defense is often mediated by the detection of microbe or pathogen associated molecular pattern elicitors, such as flagellin and chitin. Chitosan, the deacetylated form of chitin, plays a role in inducing protection against pathogens in many plant species. We evaluated the ability of chitosan to confer resistance to Botrytis cinerea in Arabidopsis leaves. We subsequently treated Arabidopsis seedlings with chitosan and carried out a transcript profiling analysis using both ATH1 GeneChip microarrays and quantitative RT-PCR. The results showed that defense response genes, including camalexin biosynthesis genes, were up-regulated by chitosan, both in wild-type and in the chitin-insensitive cerk1 mutant, indicating that chitosan is perceived through a CERK1-independent pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants can activate defence responses upon the perception and recognition of numerous pathogen-derived molecules (Dangl and Jones 2001; Nürnberger and Scheel 2001; Parker 2003; Zipfel 2009). The detection of pathogens leads to metabolic reprogramming and the production of an array of antimicrobial compounds (Denoux et al. 2008).
Inducible defense against various pathogenic microorganisms is mediated by the detection of microbe or pathogen associated molecular patterns (MAMPs/PAMPs), which are produced by infectious agents but not by host cells (Asai et al. 2002; Nürnberger et al. 2004; Mackey and McFall 2006; Schwessinger and Zipfel 2008). These pathogen-derived molecules trigger a suite of immune responses that limit pathogen growth and damage to the host (Aziz et al. 2004; Ferrari et al. 2007; Denoux et al. 2008). MAMPs perception triggers an early induction of regulatory factors (e.g. transcription factors) and genes that encode enzymes for the synthesis of antimicrobial compounds, together with genes that encode proteins involved in signal perception and transduction, including kinases and phosphatases (Moscatiello et al. 2006; Zipfel et al. 2006).
Chitin, a major component of fungal cell walls, serves as a MAMP (Gust et al. 2007; Libault et al. 2007; Miya et al. 2007; Denoux et al. 2008). Application of purified chitin oligomers to plants or plant cell cultures elicits various defence reactions (Khan et al. 2003; Tanabe et al. 2006). Chitin pre-treatment of plants also reduces the susceptibility to subsequent fungal pathogen challenge (Tanabe et al. 2006). A large number of Arabidopsis thaliana genes are induced upon chitin elicitation (Ramonell et al. 2002; Ramonell et al. 2005; Libault et al. 2007; Miya et al. 2007; Wan et al. 2008).
Chitosan, a β-1,4-linked glucosamine, is a deacetylated derivative of chitin that induces phytoalexin synthesis in pea pods (Hadwiger and Beckman 1980), in suspension-cultured soybean cells (Köhle et al. 1984), and parsley cells (Conrath et al. 1989). It also induces the lignification response in wounded wheat leaves (Pearce and Ride 1982), and triggers callose formation in suspension-cultured parsley (Conrath et al. 1989), tomato (Grosskopf et al. 1991) and Catharanthus roseus cells (Keen 1975). Moreover, chitosan activates reactive oxygen species (ROS) production (Lee et al. 1999), inhibits fungal growth and elicits a variety of defence reactions in higher plants, therefore enhancing plant resistance against pathogens (Amborabé et al. 2008; Benhamou and Thériault 1992; El Ghaouth et al. 1994).
These abilities highlight the potential importance of chitosan as an elicitor and as a component of host–fungal interactions (Young et al. 1982). For instance, several studies have demonstrated the role of chitosan in protecting different species against Botrytis cinerea (Ben-Shalom et al. 2003; Barka et al. 2004: Trotel-Aziz et al. 2006). Among pathogenic fungi, B. cinerea is the causal agent of grey mould, which is responsible for worldwide yield losses (Aziz et al. 2004). Fungicides can reduce the damage to plants caused by this necrotrophic pathogen, but repeated fungicide applications are harmful to the environment (Siedlecka and Krupa 1996). Furthermore, fungicides have only been partially successful due to the continual appearance and establishment of resistant Botrytis spp. strains to these chemicals (Barka et al. 2004). Consequently, the possibility of stimulating internal plant defences has become an interesting alternative for enhancing natural resistance against B. cinerea and other pathogens. Among the elicitors known to date, chitosan has one of the greatest potentials as a biocontrol agent (El Ghaouth et al. 1994).
Although the role of chitosan in physiological and metabolic processes has been described, transcriptional reprogramming due to chitosan is still largely unknown. Little is known about the mechanisms by which plants can sense chitosan and it is unclear whether the chitin receptor CERK1 could be involved in chitosan perception. Chitosan was recently shown to bind weakly to the chitin receptor CERK1, but this is likely due to the presence of acetylated glucosamine residues (Petutschnig et al. 2010).
In this study, we carried out a transcript profiling analysis in order to evaluate the ability of chitosan to elicit transcriptional defense responses in Arabidopsis seedlings. Microarray results demonstrated the inductive role of chitosan on several genes involved in defense responses. Analysis of the expression of chitosan induced genes using the cerk1 mutant revealed that chitosan perception is uncoupled from CERK1.
Materials and methods
Plant materials and growth condition
Arabidopsis thaliana, ecotype Columbia (Col-0) and cerk1-2 (N409189) seeds were purchased from the European Arabidopsis Stock Centre. Seeds were sterilized for 7 min in 1.7% (v/v) bleach solution, and incubated over night in 4% PPM (Plant Preservative Mixture, Plant Cell Technology, Washington, DC, USA) in a full strength sterilized Murashige-Skoog (MS) salt solution (0.5% sucrose) in Hepes–KOH 50 mM (pH 6.8) with gentle shaking. The seeds were then rinsed in abundant sterile water and transferred into 2.5 ml liquid growing media (MS half strength solution in Hepes–KOH 50 mM pH 6.8 containing 0.5% sucrose) with 0.05% PPM in 6-well plates. The plates were incubated in the dark at 4°C for 2 days and finally transferred to continuous light (90 μmol photons/m−2) with gentle swirling for 4 days in a plant growth chamber at 22°C. We used adult plants for chitosan pre-treatment and B. cinerea inoculation. Plants were grown for 4 weeks in soil, in a growth chamber with a 12-h photoperiod (120 μmol photons/m−2).
Chitosan preparation
Chitin was recovered by successive extraction of the concomitant substances from the chitin-containing raw material (exoskeletons of crab, provided by Algea A.S., Kristiansund, Norway). Exoskeletons of crab were washed, dried and pulverized with pestle and mortar into fine powder. Chitin was extracted from the shell by demineralising and deproteinising the powder. Demineralization was achieved using HCl 1.3 M for 2 h. Deproteinization was carried out with NaOH 4 M, at 70°C for 3 h. Depigmentation was achieved using H2O2 (1.4 M), for 1.5 h at 70°C. After each phase, the sample was filtered using a G1 gooch filter, to recover the supernatant, which was washed with water until the pH returned to neutrality. Deacetylation was carried out in NaOH (8 M) at 120°C for 3 h (raw material:alkali ratio = 1:5). The molecular weight of the chitosan was determined viscometrically in 0.3 M aqueous acetic acid containing 0.2 M sodium acetate at 30°C. The measurements were performed on an Ubbelohde viscometer with a capillary diameter of 0.3 mm at 30°C. The degree of deacetylation in chitosan samples was measured using conductometric titration. The molecular weight of chitosan obtained was 5-10 kDa and the deacetylation was 80%.
Plant treatments
Chitosan was used at a concentration of 150 μg/ml, whereas oligogalacturonides (OGs) were used at a concentration of 50 μg/ml. In some experiments we used the solvent used to solubilise chitosan (final concentration 0.02% acetic acid) as an additional control. Chitosan pre-treatment of Arabidopsis adult leaves was performed by spraying a chitosan solution 6 h before the inoculation of B. cinerea spores. The samples used for the microarray experiment were 4 days old seedlings treated for 3 h with a chitosan solution (150 μg/ml). Chitosan was solubilised using acetic acid. Consequently, we used an acetic acid solution to treat (3 h) the control samples.
Expression analysis
The total RNA, extracted using an RNAqueous kit (Ambion, Austin, TX, USA) according to the manufacturer’s instructions, was subjected to DNase treatment using a TURBO DNA free kit (Ambion, Austin, TX, USA). Two micrograms of each sample were reverse transcribed into cDNA with a “High capacity cDNA archive kit” (Applied Biosystems, Foster City, CA, USA). Quantitative RT-PCR amplification (qPCR) was carried out using an ABI Prism® 7000 Sequence Detection System (Applied Biosystems, Foster City, USA), with primers described in Table S1. Ubiquitin10 (UBQ10) was used as an endogenous control. qPCR reactions were carried out using a SYBR Green PCR Master Mix (Applied Biosystems, Foster City, USA), 50 ng cDNA template, and gene-specific primers in a final reaction volume of 15 μl. The relative quantitation of each single gene expression was performed using the comparative C T method as described in the ABI PRISM 7700 Sequence Detection System User Bulletin #2 (Applied Biosystems).
RNA isolation, cRNA synthesis, and hybridization to Affymetrix GeneChips
Two independent, replicated experiments were performed for each experimental condition. Each independent experiment consisted of four replicated seedling cultures pooled after RNA extraction. RNA was extracted using an Ambion RNAqueous kit (Ambion, Inc., Austin, TX, USA). RNA quality was assessed by agarose gel electrophoresis and spectrophotometry. RNA was processed for use on Affymetrix GeneChip® Arabidopsis ATH1 Genome Arrays, as previously described (Loreti et al. 2005). Hybridization, washing, staining, and scanning procedures were performed by Genopolis (University of Milano-Bicocca), as described in the Affymetrix technical manual. A microarray analysis was performed using an R/Bioconductor (Gentleman et al. 2004). Expression measures were obtained using a GeneChip Robust Multi-Array (Wu and Irizarry 2005), a multi-array analysis method estimating probe set signals, taking into account the physical affinities between probes and targets. Normalization was done using a quantile method (Bolstad et al. 2003). To reduce the number of non-informative genes we applied an interquantile filter (interquantile range, IQR = 0.14). 10,840 probesets passed the filtering. To identify a statistically reliable number of differentially expressed genes among the two conditions, a linear model was performed (Wettenhall and Smyth 2004). To assess a differential expression, an empirical Bayesian method (Smyth and Gordon 2004) was used to moderate the SE of the estimated log-fold changes. Limma, the Bioconductor package that has been used to identify the differentially expressed genes, produces false discovery rate (FDR)-adjusted “p values” that are called “q values”. The technique used for multiple testing correction was the Benjamini and Hochberg (Reiner et al. 2003). What is reported in this paper is the Limma FDR “q value” that for clearness where defined as “adjusted p value”. The differentially expressed genes (DEG, adjusted P value <0.001) were further filtered by selecting genes showing a fold change (≥2 or ≤−2) in a hybridization signal (namely the ratio between the intensity in the treated spot vs the control). Microarray datasets were deposited in a public repository with open access (accession no. GSE17193; http://www.ncbi.nlm.nih.gov/projects/geo).
In order to build gene expression maps the MapMan software (Thimm et al. 2004) from http://gabi.rzpd.de/projects/MapMan/ has been used. Gene categories were used as defined by MapManx BINs.
The chitosan dataset GSE17193 was compared with the publicly available chitin dataset described by Wan et al. (2008) (GSM206278, GSM206277, GSM206279: controls; GSM206274, GSM206275, GSM206276: chitooctaose treatment) using the microarray analysis pipeline described above.
Fungal growth and plant inoculation
B. cinerea (as described by Ferrari et al. 2003) was grown on potato dextrose agar plates for 7–15 days at 22°C with a 12-h photoperiod before collecting the spores as described in Ferrari et al. (2007). Rosette leaves from 4-week-old soil grown Arabidopsis plants were placed in Petri dishes containing 0.8% agar, with the petiole embedded in the medium. Inoculation was performed by placing 5 μl of a suspension of 5 × 105 conidiospores ml−1 in 24 g l−1 potato dextrose broth (PDF; Difco, Detroit, USA) on each side of the middle vein. The plates were incubated at 22°C with a 12-h photoperiod for 5 days. High humidity was maintained by covering the plates with a clear plastic dome. Under these experimental conditions, most inoculations resulted in rapidly expanding water-soaked lesions of a comparable diameter. Lesion size was determined by measuring their area.
Peroxidases activity
Arabidopsis leaves were collected after being treated with chitosan, frozen in liquid nitrogen, and stored at −70°C. Samples were extracted by grinding in 2 volumes of ice-cold sodium phosphate buffer (50 mM, pH 6.5). Homogenates were centrifuged in microfuge tubes at 4°C for 10 min at 14,000g. The supernatant was used to measure enzyme activity. Protein concentration was determined as described by Bradford (1976) with bovine serum albumin (BSA) as the standard. Peroxidase activity was estimated using guaiacol as the substrate. The oxidation of the substrate was measured spectrophotometrically at 450 nm as previously described (Summermatter et al. 1995). The assay solution contained sodium phosphate (10 mM, pH 6), 0.25% (v/v) guaiacol, 100 mM H2O2, and leaf extract.
Results
Chitosan protects Arabidopsis from Botrytis cinerea
Chitosan has been proposed as a strong elicitor of plant defences (Amborabé et al. 2008). We evaluated the ability of this compound to confer resistance to B. cinerea in Arabidopsis (Col-0) adult leaves. Four-week-old leaves were inoculated with a B. cinerea spore suspension. Lesions induced by B. cinerea were measured 5 days after inoculation. Infection of chitosan pre-treated leaves resulted in lesions six times smaller than those on leaves that were not treated with chitosan (Fig. 1a, b). This indicates that chitosan rapidly stimulates defense reactions in Arabidopsis leaves and inhibits the development of B. cinerea.
Effect of chitosan on B. cinerea infection of Arabidopsis leaves. a Wild-type (Col-0) Arabidopsis adult leaves were sprayed with a chitosan solution (150 μg/ml) or with the solvent used to solubilise chitosan as a control. After 6 h, leaves were detached and inoculated with B. cinerea spores. Photos of the lesions induced by B. cinerea were taken 5 days after the inoculation of B. cinerea spores. Enlarged pictures were taken using a Nikon TMS-F microscope (type 104) connected with the NIS-elements F2.20 imaging software (Laboratory Imaging, Nikon, Tokyo, Japan). b Lesion areas were measured 5 days after the inoculation of B. cinerea spores. Data are mean of 10 measurements ± SD. c Expression analysis of Arabidopsis seedlings treated with active oligogalacturonides (OGs). Four-day old Arabidopsis seedlings were treated with 50 μg/ml OGs for 3 h. WRKY40 mRNA level was analyzed by qPCR. Relative expression levels are shown as Fold Change values (1 = control). Data are means of three replicates ± SD. d Effect of chitosan on the expression of WRKY40 and Cyp81f2. mRNA levels were measured in 4-day Arabidopsis seedlings treated with water (control), a solution containing the solvent used to solubilise chitosan, and chitosan (150 μg/ml). Relative expression level, measured by qPCR, is shown. Relative expression levels are shown as Fold Change values (1 = control, time 0). Data are means of three replicates ± SD
Chitosan elicits rapid expression of defence-related genes
To test the effect of chitosan at a molecular level, we selected WRKY40 (At1g80840), a transcription factor (TF) involved in the early response to MAMPs (Xu et al. 2006; Denoux et al. 2008). WRKY40 is responsive to oligogalacturonides (OGs), which can increase its mRNA level (Denoux et al. 2008). OGs were found to be very effective in the experimental set-up used in this study (Fig. 1c). We analysed the pattern of WRKY40 expression in a time-course experiment using chitosan (Fig. 1d). In addition to WRKY40, the expression pattern of CYP81f2 (At5g57220), another gene involved in the early response to MAMPs (Clay et al. 2009) was also tested (Fig. 1d). Both WRKY40 and CYP81f2 were induced by chitosan after a 2 h treatment, although their expression peaked after 3 h (Fig. 1d) and this time-point was chosen to carry out a transcript profiling analysis.
Chitosan elicits a change in the transcript profile
Our microarray results indicated that a treatment with chitosan exerted a dramatic effect on the transcriptome, with 617 probe sets showing an increased expression (≥2-fold, FDR p value <0.001) and 388 probe sets indicating a decreased expression (≤−2-fold, FDR p value <0.001) after data filtering and the selection of differentially expressed genes (Table S2). The list of genes up- regulated >100-fold is reported in Table 1. MapMan (Thimm et al. 2004) highlighted that many chitosan-modulated genes in our experiment are involved in biotic stress responses (Fig. 2a), including defense-related genes and transcription factors related to hormone signalling (Fig. 2b).
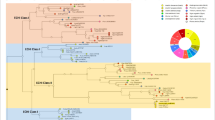
MapMan screenshots showing the effects of chitosan on the expression of chitosan-modulated genes. a, b The MapMan software was queried with the list of differentially regulated genes following chitosan treatment (Table S2). Red shades indicate induction; blue shades indicate repression of gene expression
TFs are the upstream triggers in signalling pathways. Chitosan induces TF genes belonging to different families, including ERF, MYB and WRKY domain transcription factor families (Fig. 2b).
In Arabidopsis over 70% of the entire WRKY gene family of 74 members responds to pathogens or pathogen-mimicking stimuli (Lippok et al. 2007). Eight of them responded to chitosan (Fig. 2b). For example, WRKY40 (Fig. 1c, d) is a pathogen-induced WRKY-TF which is also one of the modulators of jasmonate pathways, functioning as an activator of jasmonate-dependent defence (Xu et al. 2006). Among the differentially expressed WRKY-TF factors, we also identified At5g13080 (WRKY75), At3g01970 (WRKY45), and At2g46400 (WRKY46) and At4g31800 (WRKY18) (Table S3). These WRKY transcription factors are involved in pathogen responses (Dong et al. 2003).
The expression of some MYB TFs was also affected by chitosan (Fig. 2b). MYB31 (At1g74650), one of the MYB factors known to be responsive to chitin (Libault et al. 2007) and MYB15 (At3g23250), a jasmonate responsive gene, were induced by chitosan (Table S3).
At5g47220 (ERF2), a member of the AP2/ERF family of transcription factors, was chitosan-induced (Table S3), in line with previous studies demonstrating the involvement of At5g47220 in early chitin responses and jasmonic acid treatment (Libault et al. 2007).
A large number of PR-proteins were induced by chitosan (Fig. 2b). These include PLP2 (At2g26560, induced 338 fold), an integral component of the plant cell death machinery (La Camera et al. 2009), GH3.12 (At5g13320, induced 42-fold), a gene involved in the regulation of salicylic acid (SA) metabolism (Nobuta et al. 2007), AIG1 (At1g33960, induced 171-fold) induced by avirulence genes avrRpt2 and RPS2 after infection with Pseudomonas syringae (Wang and Li 2009), and the plant defensin PDF1.2B (At2g26020, induced 95-fold).
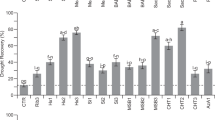
Among the enzymes involved in biotic stress responses, we found a large group of peroxidases (POXs; Figs. 2b, 3a), usually considered enzymes involved in detoxification of ROS (Kawano 2003). The differential expression of genes encoding for peroxidases was mirrored by a corresponding increase in enzyme activity (Fig. 3b).
Chitosan induces genes involved in camalexin biosynthesis
We examined the transcriptional reprogramming induced by chitosan and compared it with the effects of B. cinerea. Arabidopsis eFP browser software (Winter et al. 2007) allowed us to identify a set of genes inducible after B. cinerea inoculation (Ferrari et al. 2007), which are also induced by chitosan treatment. The results of this comparison are summarized in Table S4.
Interestingly, this list includes genes involved in the biosynthesis of camalexin, the indole phytoalexin of Arabidopsis thaliana (Fig. 4a). Camalexin biosynthesis is essential for resistance to B. cinerea in Arabidopsis (Ferrari et al. 2007). Almost all the genes codifying for enzymes which are involved in the biosynthesis of tryptophan, the precursor of camalexin, such as At5g05730 (ASA1), At5g17990 (PAT1), At2g04400 (IGPS), and At3g54640 (TSA1) were induced (Fig. 4a). Camalexin is synthesized from tryptophan via indole-3-acetaldoxime (IAOx) in a reaction catalyzed by CYP79B2 and CYP79B3 (Glawischnig et al. 2004). At4g39950/CYP79B2 was induced fivefold by chitosan (Fig. 4a). In the camalexin pathway, IAOx represents the direct precursor of this compound (Glawischnig et al. 2004). The transcripts encoding At2g30770 (Cyp71A13) and At3g26830 (Cyp71B15, PAD3) showed a very high induction after chitosan treatment (Fig. 4a). Cyp71A13 catalyzes the first committed step in camalexin biosynthesis by dehydrating IAOx to indole-3-acetonitrile (IAN; Nafisi et al. 2007). Cyp71B15/PAD3 on the other hand, encodes an essential enzyme in the camalexin biosynthetic pathway (Zhou et al. 1999), which catalyzes the oxidative decarboxylation of dihydrocamalexic acid to camalexin (Schuhegger et al. 2006).
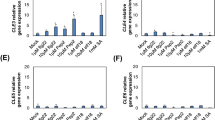
Effects of chitosan on the camalexin biosynthetic pathway. a Chitosan-dependent differentially expressed genes encoding enzymes involved in the camalexin pathway are shown. The fold change measured in the microarray experiment is shown. b Venn diagram showing the overlap between genes differentially regulated by chitin (Wan et al. 2008) and chitosan (this study). c mRNA levels for Cyp71A13 and Cyp71B15 were measured in 4-day Arabidopsis wild-type (Col-0) or cerk1 mutant seedlings treated with water (control), a solution containing the solvent used to solubilise chitosan, and chitosan (150 μg/ml). Relative expression level, measured by qPCR, is shown. Relative expression levels are shown as Fold Change values (1 = control, time 0). Data are means of three replicates ± SD
We also observed a fourfold induction of At3g52430 (PAD4) (Table S2), which is required for the bacterial, but not fungal, induction of camalexin (Jirage et al. 1999; Ferrari et al. 2003).
Chitosan perception is independent of CERK1
A comparison of the transcription profiles of Arabidopsis plants treated with chitin (Wan et al. 2008) or with chitosan (this study), revealed that 33.7% of the genes modulated by chitosan are also regulated by chitin (Fig. 4b; Table S5). This was suggestive of limited overlap with the targets of chitin.
To evaluate whether the effects of chitosan were indeed largely independent from the chitin signalling pathway, we analysed the expression of CYP71A13 and Cyp71B15 in both wild type Arabidopsis and in the chitin-insensitive cerk1 mutant (Miya et al. 2007). Chitosan treatment strongly induced these two genes in both the wild-type and cerk1 (Fig. 4c).
Chitin triggers the regulation of 118 transcription factors (Libault et al. 2007). We identified 49 TF genes up-regulated (more than twofold) after chitosan treatment (Table S3). Only 22 of the 118 TFs regulated by chitin (Libault et al. 2007) were found to be responsive to chitosan (Table S3). These results further suggest that distinct pathways operate following chitin or chitosan treatments.
Some transcription factors are responsive to both chitin (Libault et al. 2007; Wan et al. 2008) and chitosan (Table S3) (e.g. WRKY18, WRKY40, WRKY46 and ERF2) while others are induced by chitosan (Table S3) but not by chitin, e.g. WRKY75 and WRKY45 (Libault et al. 2007; Wan et al. 2008). We tested the induction of these TFs in the cerk1 mutant. The results indicated that (i) chitin-insensitive TFs are induced normally in the cerk1 mutant and that (ii) also chitin-sensitive TFs are responsive to chitosan in cerk1 (Fig. 5). The first point confirms that chitosan activates a CERK1-independent signalling pathway, while the second point indicates that a convergence downstream of CERK1 is possible.
Effects of chitosan on the expression of transcription factors. mRNA levels were measured in 4-day Arabidopsis wild-type (Col-0) or cerk1 mutant seedlings treated with water (control), a solution containing the solvent used to solubilise chitosan, and chitosan (150 μg/ml). Relative expression level, measured by qPCR, is shown. Relative expression levels are shown as Fold Change values (1 = control, time 0). Data are means of three replicates ± SD
Discussion
Chitosan induces defense responses in several plant species. Our findings demonstrated that chitosan protected Arabidopsis leaves against the necrotrophic fungus B. cinerea (Fig. 1). This is because chitosan was perceived by the plant as a powerful elicitor, acting as a pathogen-mimicking stimulus, as demonstrated by the activation of a large number of biotic-stress related genes (Fig. 2). The application of a chitosan solution may sensitize Arabidopsis leaves to respond more rapidly to pathogen attack. For example, camalexin biosynthetic genes were strongly transcriptionally up-regulated after a chitosan treatment (Fig. 4b, c). Some peroxidases were also up-regulated. Extracellular, secreted plant POXs can catalyze the generation of ROS coupled to defense-related compounds, such as salicylic acid (SA), aromatic monoamines (AMAs) and chitooligosaccharide (COSs) elicitors (Kawano 2003). We found that the effect of chitosan at the transcriptional level was mirrored by a corresponding increase in peroxidase activity, indicating that the effects at the transcriptional level are accompanied by an increase in the level of the corresponding proteins.
Whether chitosan is perceived by CERK1, the chitin receptor is under debate. CERK1 was found to bind weakly chitosan (Petutschnig et al. 2010), and this evidence corroborated the idea that CERK1 is involved in chitosan-dependent plant defense activation (Petutschnig et al. 2010). The cerk1-2 mutant is insensitive to partially de-acetylated chitosan in terms of ROS-burst and MAP kinase assays, suggesting that either CERK1 is able to recognize chitosan or that acetylated residues present in commercially available chitosan are recognized by CERK1 (Petutschnig et al. 2010). Iizasa et al. (2010) found very weak binding of chitosan to CERK1, a result suggestive of the existence of a CERK1-independent pathway for chitosan perception. Fully de-acetylated chitooligomers are indeed not effective and this is highly suggestive of the importance of GlcNAc residues for CERK1 binding (Petutschnig et al. 2010).
In this context, a comparison of the transcription profiles of Arabidopsis plants treated with either chitin (Wan et al. 2008) or with chitosan (this study), revealed that the overlap in modulated genes is relatively limited, with only 33.7% of the genes modulated by chitosan also regulated by chitin (Fig. 4b; Table S5).
More direct evidence for a CERK1-independent chitosan signalling arose from the use of the cerk1 mutant. Transcription factors and the key-genes for camalexin biosynthesis (CYP71A13 and Cyp71B15) are responsive to chitosan in the cerk1 mutant, demonstrating that chitosan perception is independent of CERK1 and thus from the chitin signaling pathway.
The signaling pathways for chitin and chitosan thus appear to be independent, but may converge downstream of CERK1 (Fig. 6). Chitin perception by CERK1 activates transcription factors that induce the expression of genes directly involved in plant defence to pathogens, and which contribute to a resistant phenotype. Chitosan is perceived by a still unknown receptor, activating a downstream signaling pathway that largely converges with that induced by chitin. Some transcription factors are known to be induced by both chitin and chitosan (e.g. WRKY18, WRKY40, WRKY46) and demonstrate convergence of the two pathways. The identification of the chitosan receptor is of considerable interest and certainly deserves further studies.
Schematic representation of the chitosan signalling pathway and its relation with chitin signalling. Chitin is perceived by CERK1, while chitosan appears to trigger defense-related genes through a CERK1-independent pathway. The two pathways may converge at the level of TFs such as WRKY18, WRKY40 and WRKY46. Both pathways result in the induction of resistance to pathogens
References
Amborabé BE, Bonmort J, Fleurat-Lessard P, Roblin G (2008) Early events induced by chitosan on plant cells. J Exp Bot 59:2317–2324
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL, Gomez-Gomez L, Boller T, Ausubel FM, Sheen J (2002) MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415:977–983
Aziz A, Heyraud A, Lambert B (2004) Oligogalacturonide signal transduction, induction of defense-related responses and protection of grapevine against Botrytis cinerea. Planta 218:767–774
Barka EA, Eullaffroy P, Clement C, Vernet G (2004) Chitosan improves development, and protects Vitis vinifera L. against Botrytis cinerea. Plant Cell Rep 22:608–614
Benhamou N, Thériault G (1992) Treatment with chitosan enhances resistance of tomato plants to the crown and root rot pathogen Fusarium oxysporum f. sp. radicis-lycopersici. Physiol Mol Plant Pathol 41:33–52
Ben-Shalom N, Ardi R, Pinto R, Aki C, Fallik E (2003) Controlling gray mould caused by Botrytis cinerea in cucumber plants by means of chitosan. Crop Prot 22:285–290
Bolstad BM, Irizarry RA, Astrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on bias and variance. Bioinformatics 19:185–193
Bradford MM (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Clay NK, Adio AM, Denoux C, Jander J, Ausubel FM (2009) Glucosinolate metabolites required for an arabidopsis innate immune response. Science 323:95–101
Conrath U, Domard A, Kauss H (1989) Chitosan-elicited synthesis of callose and of coumarin derivatives in parsley cell suspension cultures. Plant Cell Rep 8:152–155
Dangl JL, Jones JDG (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
Denoux C, Galletti R, Mammarella N, Gopalan S, Werck D, De Lorenzo G, Ferrari S, Ausubel FM, Dewdney J (2008) Activation of defense response pathways by OGs and Flg22 elicitors in Arabidopsis seedlings. Mol Plant 1:423–445
Dong J, Chen C, Chen Z (2003) Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol Biol 51:21–37
El Ghaouth A, Arul J, Grenier J, Benhamou N, Asselin A, Belanger R (1994) Effect of chitosan on cucumber plants: suppression of Pythium aphanidermatum and induction of defence reactions. Phytopathology 84:313–320
Ferrari S, Plotnikova JM, De Lorenzo G, Ausubel FM (2003) Arabidopsis local resistance to Botrytis cinerea involves salicylic acid and camalexin and requires EDS4 and PAD2, but not SID2, EDS5 or PAD4. Plant J 35:193–205
Ferrari S, Galletti R, Denoux C, De Lorenzo G, Ausubel FM, Dewdney J (2007) Resistance to Botrytis cinerea induced in Arabidopsis by elicitors is independent of salicylic acid, ethylene, or jasmonate signaling but requires PHYTOALEXIN DEFICIENT3. Plant Physiol 144:367–379
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Horthorn T, Huber W, Iacus S, Irizarry R, Leisch F, Maechler M, Roosini A, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JYH, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Glawischnig E, Hansen BG, Olsen CE, Halkier BA (2004) Camalexin is synthesized from indole-3-acetaidoxime, a key branching point between primary and secondary metabolism in Arabidopsis. Proc Natl Acad Sci USA 101:8245–8250
Grosskopf DG, Felix G, Boller T (1991) A yeast-derived glycopeptides elicitor and chitosan or digitonin differentially induce ethylene biosynthesis, phenylalanine ammonia-lyase and callose formation in suspension-cultured tomato cells. J Plant Physiol 138:741–746
Gust AA, Biswas R, Lenz HD, Rauhut T, Ranf S, Kemmerling B, Gotz F, Glawischnig E, Lee J, Felix G, Nürnberger T (2007) Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J Biol Chem 282:32338–32348
Hadwiger LA, Beckman JM (1980) Chitosan as a component of pea–Fusarium solani interactions. Plant Physiol 66:205–211
Iizasa E, Mitsutomi M, Nagano Y (2010) Direct binding of a plant LysM receptor-like kinase, LysM RLK1/CERK1, to chitin in vitro. J Biol Chem 285:2996–3004
Jirage D, Tootle TL, Reuber TL, Frost LN, Feys BJ, Parker JE, Ausubel FM, Glazebrook J (1999) Arabidopsis thaliana PAD4 encodes a lipase-like gene that is important for salicylic acid signaling. Proc Natl Acad Sci USA 96:13583–13588
Kawano T (2003) Roles of the reactive oxygen species-generating peroxidase reactions in plant defense and growth induction. Plant Cell Rep 21:829–837
Keen NT (1975) Specific elicitors of plant phytoalexin production-determinants of race specificity in pathogens. Science 187:74–75
Khan W, Prithiviraj B, Smith DL (2003) Chitosan and chitin oligomers increase phenylalanine ammonia-lyase and tyrosine ammonia-lyase activities in soybean leaves. J Plant Physiol 160:859–863
Köhle H, Young DH, Kauss H (1984) Physiological changes in suspension-cultured soybean cells elicited by treatment with chitosan. Plant Sci Lett 33:221–230
La Camera S, Balagué C, Göbel C, Geoffroy P, Legrand M, Feussner I, Roby D, Heitz T (2009) The Arabidopsis patatin-like protein 2 (PLP2) plays an essential role in cell death execution and differentially affects biosynthesis of oxylipins and resistance to pathogens. Mol Plant Microbe Interact 22:469–481
Lee D, Noh HJ, Lee KM (1999) Purification and characterization of a chitinase from Cytophaga sp. HJ isolated from sea sand. J Microbiol Biotechnol 9:839–846
Libault M, Wan JR, Czechowski T, Udvardi M, Stacey G (2007) Identification of 118 Arabidopsis transcription factor and 30 ubiquitin-ligase genes responding to chitin, a plant-defense elicitor. Mol Plant Microbe Interact 20:900–911
Lippok B, Birkenbihl RP, Rivory G, Brummer J, Schmelzer E, Logemann E, Somissich IE (2007) Expression of AtWRKY33 encoding a pathogen- or PAMP-responsive WRKY transcription factor is regulated by a composite DNA motif containing W box elements. Mol Plant Microbe Interact 20:420–429
Loreti E, Poggi A, Novi G, Alpi A, Perata P (2005) A genome-wide analysis of the effects of sucrose on gene expression in Arabidopsis seedlings under anoxia. Plant Physiol 137:1130–1138
Mackey D, McFall AJ (2006) MAMPs and MIMPs: proposed classifications for inducers of innate immunity. Mol Microbiol 61:1365–1371
Miya A, Albert P, Shinya T, Desaki Y, Ichimura K, Shirasu K, Narusaka Y, Kawakami N, Kaku H, Shibuya N (2007) CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc Natl Acad Sci USA 104:19613–19618
Moscatiello R, Mariani P, Sanders D, Maathuis FJM (2006) Transcriptional analysis of calcium-dependent and calcium-independent signalling pathways induced by oligogalacturonides. J Exp Bot 57:2847–2865
Nafisi M, Goregaoker S, Botanga CJ, Glawischnig E, Olsen CE, Halkier BA, Glazebrook J (2007) Arabidopsis cytochrome P450 monooxygenase 71A13 catalyzes the conversion of indole-3-acetaldoxime in camalexin synthesis. Plant Cell 19:2039–2052
Nobuta K, Okrent RA, Stoutemyer M, Rodibaugh N, Kempema L, Wildermuth MC, Innes RW (2007) The GH3 acyl adenylase family member PBS3 regulates salicylic acid-dependent defense responses in Arabidopsis. Plant Physiol 144:1144–1156
Nürnberger T, Scheel D (2001) Signal transmission in the plant immune response. Trends Plant Sci 6:372–379
Nürnberger T, Brunner F, Kemmerling B, Piater L (2004) Innate immunity in plants and animals: striking similarities and obvious differences. Immunol Rev 198:249–266
Parker JE (2003) Plant recognition of microbial patterns. Trends Plant Sci 8:245–247
Pearce RB, Ride JP (1982) Chitin and related compounds as elicitors of lignification response in wounded wheat leaves. Physiol Plant Pathol 20:119–123
Petutschnig EK, Jones AME, Serazetdinova L, Lipka U, Volker L (2010) The lysin motif receptor-like kinase (LysM-RLK) CERK1 is a major chitin-binding protein in Arabidopsis thaliana and subject to chitin-induced phosphorylation. J Biol Chem 285:28902–28911
Ramonell KM, Zhang B, Ewing RM, Chen Y, Xu D, Stacey G, Somerville S (2002) Microarray analysis of chitin elicitation in Arabidopsis thaliana. Mol Plant Pathol 3:301–311
Ramonell K, Berrocal-Lobo M, Koh S, Wan JR, Edwards H, Stacey G, Somerville S (2005) Loss-of-function mutations in chitin responsive genes show increased susceptibility to the powdery mildew pathogen Erysiphe cichoracearum. Plant Physiol 138:1027–1036
Reiner A, Yekutieli D, Benjamini Y (2003) Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics 19:368–375
Schuhegger R, Nafisi M, Mansourova M, Petersen BL, Olsen CE, Svatos A, Halkier BA, Glawischnig E (2006) CYP71B15 (PAD3) catalyzes the final step in camalexin biosynthesis. Plant Physiol 141:1248–1254
Schwessinger B, Zipfel C (2008) News from the frontline: recent insights into PAMP-triggered immunity in plants. News from the frontline: recent insights into PAMP-triggered immunity in plants. Curr Opin Plant Biol 11:389–395
Siedlecka A, Krupa Z (1996) Interaction between cadmium and iron and its effects on photosynthetic capacity of primary leaves of Phaseolus vulgaris. Plant Physiol Biochem 34:833–841
Smyth GK, Gordon K (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3(1) (Article 3)
Summermatter K, Sticher L, Métraux JP (1995) Systemic responses in Arabidopsis thaliana infected and challenged with Pseudomonas syringae pv syringae. Plant Physiol 108:1379–1385
Tanabe S, Okada M, Jikumaru Y, Yamane H, Kaku H, Shibuya N, Minami E (2006) Induction of resistance against rice blast fungus in rice plants treated with a potent elicitor, N-acetylchitooligosaccharide. Biosci Biotechnol Biochem 70:1599–1605
Thimm O, Blaesing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Trotel-Aziz P, Couderchet M, Vernet G, Aziz A (2006) Chitosan stimulates defense reactions in grapevine leaves and inhibits development of Botrytis cinerea. Eur J Plant Pathol 114:405–413
Wan J, Zhang XC, Neece D, Ramonell KM, Clough S, Kim SY, Stacey MG, Stacey G (2008) A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20:471–481
Wang Z, Li X (2009) IAN/GIMAPs are conserved and novel regulators in vertebrates and angiosperm plants. Plant Signal Behav 4:165–167
Wettenhall JM, Smyth GK (2004) limmaGUI: A graphical user interface for linear modeling of microarray data. Bioinformatics 20:3705–3706
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV, Provart NJ (2007) An ‘electronic fluorescent pictograph’ browser for exploring and analyzing large-scale biological data sets. PLoS One 2:e718
Wu Z, Irizarry RA (2005) Stochastic models inspired by hybridization theory for short oligonucleotide arrays. J Comput Biol 12:882–893
Xu XP, Chen CH, Fan BF, Chen ZX (2006) Physical and functional interactions between pathogen-induced Arabidopsis WRKY18, WRKY40, and WRKY60 transcription factors. Plant Cell 18:1310–1326
Young DH, Köhle H, Kauss H (1982) Effect of chitosan on membrane permeability of suspension-cultured Glycine max and Phaseolus vulgaris cells. Plant Physiol 70:1449–1454
Zhou N, Tootle TL, Glazebrook J (1999) Arabidopsis PAD3, a gene required for camalexin biosynthesis, encodes a putative cytochrome P450 monooxygenase. Plant Cell 11:2419–2428
Zipfel C (2009) Early molecular events in PAMP-triggered immunity. Cur Opin Plant Biol 12:414–420
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JDG, Boller T, Felix G (2006) Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125:749–760
Acknowledgments
We thank Dr. Simone Ferrari and Dr. Roberta Galletti (Università di Roma La Sapienza, Italy) for helpful suggestions and for kindly providing us with active oligogalacturonides and Botrytis cinerea inoculum, and Dr. Larisa Angela Swirsky Whitney for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1.
List of primers used in this work (DOC 34 kb)
Table S2.
List of genes up and down-regulated by chitosan in the microarray experiment. The differentially expressed genes (DEG) were filtered by selecting genes showing a fold change (≥ 2 or ≤-2) (see materials and methods) (XLS 452 kb)
Table S3.
List of transcription factors up-regulated by chitosan. Genes induced by chitin (Libault et al. 2007) are also shown. The differentially expressed genes (DEG) were filtered by selecting genes showing a fold change ≥ 2 (DOC 137 kb)
Table S4.
Comparison of the effects of chitosan and B. cinerea infection at the gene level. Expression data relative to B. cinerea 48h treatment were obtained using the eFP Browser software (Winter et al., 2007; http://www.bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi). Expression data of genes induced by chitosan (3h treatment) are averaged transcript level from two biological replicates (this work). The 100 genes most induced by chitosan were selected (XLS 72 kb)
Table S5.
Comparison between the chitosan dataset GSE17193 and the publicly available chitin dataset described by Wan et al. (2008) (GSM206278, GSM206277, GSM206279: controls; GSM206274, GSM206275, GSM206276: chitooctaose treatment) (XLS 1712 kb)
Rights and permissions
About this article
Cite this article
Povero, G., Loreti, E., Pucciariello, C. et al. Transcript profiling of chitosan-treated Arabidopsis seedlings. J Plant Res 124, 619–629 (2011). https://doi.org/10.1007/s10265-010-0399-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-010-0399-1