Abstract
Group II introns are widespread in plant cell organelles. In vivo, most if not all plant group II introns do not self-splice but require the assistance of proteinaceous splicing factors. In some cases, a splicing factor (also referred to as maturase) is encoded within the intronic sequence and produced by translation of the (excised) intron RNA. However, most present-day group II introns in plant organellar genomes do not contain open reading frames (ORFs) for splicing factors, and their excision may depend on proteins encoded by other organellar introns or splicing factors encoded in the nuclear genome. Whether or not the ancestors of all of these noncoding organellar introns originally contained ORFs for maturases is currently unknown. Here we show that a noncoding intron in the mitochondrial cox2 gene of seed plants is likely to be derived from an ancestral reverse transcriptase/maturase-encoding form. We detected remnants of maturase and reverse transcriptase sequences in the 2.7 kb cox2 intron of Ginkgo biloba, the only living species of an ancient gymnosperm lineage, suggesting that the intron originally harbored a splicing factor. This finding supports the earlier proposed hypothesis that the ancient group II introns that invaded organellar genomes were autonomous genetic entities in that they encoded the factor(s) required for their own excision and mobility.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Expression of a number of genes in plant cell organelles is dependent upon removal of intervening sequences (introns) that interrupt reading frames and have to be removed by splicing prior to translation of the mRNA. Most introns in higher plant organellar genomes belong to the group II introns (Michel et al. 1989; Sugita and Sugiura 1996; Bonen and Vogel 2001; Michel and Ferat 1995) although a few group I introns have also been described (Vaughn et al. 1995; Cho et al. 1998; Sugiura 1992; Kuhsel et al. 1990). Both group I and group II introns are catalytically active RNA molecules (Pyle and Cech 1991; Belfort 1991; Dürrenberger and Rochaix 1991; Chowira et al. 1994; Michel and Ferat 1995), and group II introns are thought to be the evolutionary ancestors of spliceosomal introns present in the nuclear genomes of eukaryotes (Copertino and Hallick 1993; Hetzer et al. 1997). Group I and group II introns are often referred to as self-splicing introns and, in fact, at least some group I (Pyle and Cech 1991; Dürrenberger and Rochaix 1991) and group II (Winkler and Kück 1991; Padgett et al. 1994) introns can undergo self-splicing in vitro in the absence of any protein factors. However, in vivo, most if not all introns in plant cell organelles seem to require the assistance of proteinaceous splicing factors to be excised (Deshpande et al. 1995; Li et al. 2002; Hess et al. 1994; Jenkins et al. 1997; Vogel et al. 1999). While many splicing factors for plant organellar group II introns appear to be nuclear-encoded (Jenkins et al. 1997; Vogel et al. 1999; Perron et al. 1999; Jenkins and Barkan 2001; Rivier et al. 2001; Till et al. 2001; Ostheimer et al. 2003; Perron et al. 2004), some are encoded by the plastid or mitochondrial genomes (Neuhaus and Link 1987; Hess et al. 1994; Liere and Link 1995; Schuster and Brennicke 1987; Wissinger et al. 1991; Thomson et al. 1994). These organelle genome-encoded splicing factors are referred to as maturases, and homologous reading frames have also been found in mitochondrial genomes of fungi (Anziano and Butow 1991; Szczepanek and Lazowska 1996; Mohr et al. 1993). Often, these maturase open reading frames (ORFs) are encoded within intronic sequences, suggesting that the splicing factor is produced by translation of the excised group II intron. Maturase proteins encoded by organellar introns are comprised of several distinct domains, some of which can be attributed to defined catalytic activities: a reverse transcriptase domain with seven highly conserved motifs is usually followed by the maturase domain (also referred to as domain X) and sometimes by a DNA-binding motif and an endonuclease domain at the C-terminus (Mohr et al. 1993; Mohr and Lambowitz 2003).
For many maturases, intron substrates are not known. In higher plant chloroplasts, the presence of only a single maturase gene (matK) on the plastid genome and the availability of plastid translation-deficient mutants has facilitated a prediction about which introns require the MatK protein for excision: introns in which splicing is defective in the mutants are thought to depend on MatK protein action whereas introns in which splicing is retained in the mutants are believed to be independent of the MatK maturase and, instead, may require nucleus-encoded splicing factors (Hess et al. 1994; Vogel et al. 1997, 1999).
Reading frames homologous to intron-encoded reverse transcriptase/maturase genes have been identified in the fully sequenced genomes of Arabidopsis and rice (Mohr and Lambowitz 2003), indicating that some formerly organellar reverse transcriptase/maturase genes have been transferred to the nuclear genome in higher plants. Whether or not these genes are actively expressed and their gene products retargeted to plastids or mitochondria is currently unknown.
The evolutionary history of present-day introns in plant organellar genomes is surrounded by many open questions. While some group I introns could be evolutionarily old in that they stem from homologous introns in homologous bacterial genes (Kuhsel et al. 1990), many group II introns in higher plants appear to have been acquired relatively recently. For example, two introns in the mitochondrial cox2 gene and an intron in the nad1 gene first appear in mosses (Qiu et al. 1998), possibly indicating their acquisition with or shortly after the plants’ conquest of terrestrial habitats (for review, see e.g., Knoop 2004). Horizontal gene transfer has been suggested as one mechanism leading to organellar gene invasion by intronic sequences (Vaughn et al. 1995; Cho et al. 1998; Sheveleva and Hallick 2004), and fungal introns have been discussed as one possible source of recently acquired introns in plant mitochondria (Vaughn et al. 1995). Origin and evolution of splicing factors for group II introns represent yet another evolutionary puzzle. One possible scenario is that all introns that invaded plant organellar genomes contained coding sequences for the splicing factor(s) required for their excision. However, whether or not the ancestors of noncoding organellar introns originally contained ORFs for maturases and/or reverse transcriptases is currently unknown. We have searched for traces of reverse transcriptase/maturase genes in ancient groups of seed plants, and we report here on the identification of remnants of a reverse transcriptase/maturase reading frame in a large intron within the cox2 locus of the gymnosperm Ginkgo biloba, suggesting that this intron originally harbored a reverse transcriptase/maturase gene that was lost during seed plant evolution.
Materials and methods
Plant material
Fresh leaf material of Gingko biloba was obtained from a female plant in the Botanical Garden of the University of Freiburg, Germany.
List of oligonucleotides
The following synthetic oligonucleotides were used in this study for polymerase chain reaction (PCR), cDNA synthesis, and DNA sequencing:
- P1401:
-
5′GCAGCGGAACCATGGCAATTAG 3′
- P1402:
-
5′GAGGTACATCAGCGGGTGTTAC 3′
- P1503f::
-
5′ GGATAATCCCTCATCCTCG 3′
- P1502r::
-
5′ CTACGGTCCCTCGAGTCT 3′
- P1501r::
-
5′ GATTCGATCCTATAGTCGTC 3′
- P1500f::
-
5′ CTTTGCACTTATCGAACATCAG 3′
- P1504f::
-
5′ AGTAGTTGCGGAACTACCG 3′
- P1505f::
-
5′ GGAAAACGGCATTCTTTGAG 3′
- P1506r::
-
5′ AGGGGCGGCAGCGTACTTC 3′
- P1507r::
-
5′ TCGTCGAAGTGACCCTTGA 3′
- P1508r::
-
5′ CATATCGATATATATATCCATCC 3′
- P1509r::
-
5′ CTGAAATATTCACATCTCCCAA 3′
- P1510f::
-
5′ GAGAAGAACGAGGACAACC 3′
Isolation of nucleic acids and DNA gel blot hybridization
Total plant nucleic acids were isolated from fresh leaf tissue by a cetyltrimethylammonium bromide (CTAB)-based method (Doyle and Doyle 1990). Total cellular RNA was extracted using the TriFast reagent (Peqlab GmbH, Erlangen, Germany). RNA samples for cDNA synthesis were purified by treatment with RNase-free DNase I (Roche, Mannheim, Germany). For detection of cox2 sequences by Southern blot hybridization, DNA samples were digested with restriction enzymes, separated by gel electrophoresis on 0.8% agarose gels, and transferred onto Hybond nylon membranes (Amersham, Buckinghamshire, UK) by capillary blotting. Hybridizations were performed at 65°C in Rapid-Hyb buffer (Amersham) following the manufacturer’s protocol and using radiolabeled restriction fragments derived from the amplified cox2 gene from Ginkgo biloba as hybridization probes.
cDNA synthesis and polymerase chain reactions (PCR)
Reverse transcription of DNA-free RNA samples was primed with a random hexanucleotide mixture. Elongation reactions were performed with SuperScriptTMII RNase H-free reverse transcriptase (Invitrogen) according to the manufacturer’s instructions. Total cellular DNA or first-strand cDNAs were amplified by 30–35 cycles of 45 s at 94°C, 1.5 min at 55°C, and 2 min at 72°C with a 1-min extension of the first cycle at 94°C and a 5-min final extension at 72°C. Primer pair 1401 (binding to conserved cox2 sequences within exon I) and 1402 (binding to conserved sequences within exon II) were used to amplify the 2.66 kb intronic sequence.
Cloning and DNA sequencing
Amplified cox2 cDNAs or genomic fragments were purified by agarose gel electrophoresis, and subsequent recovery of the PCR products form excised gel slices using the QIAEX II kit (QIAGEN, Hilden, Germany) or the GFX kit (Amersham). Direct sequencing of amplification products was performed by cycle sequencing followed by automated analysis in a MegaBACE capillary sequencer (Amersham). Regions difficult to sequence directly were subcloned into pBluescript vectors and sequenced from plasmid clones using the universal M13 and reverse primers. To exclude mutations introduced by PCR amplification, four to five independent clones were sequenced.
Bioinformatic analyses
cox2 DNA and cDNA sequences were compared with sequences available in the databases using the NCBI BLAST program. Sequences homologous to reverse transcriptase/maturase genes were identified by translating the intron sequence in all three reading frames followed by removal of the stop codons and running BLAST searches for short, nearly exact, matches. Amino acid sequence alignments of reverse transcriptase/maturase proteins were produced using the ClustalW software (http://www.ebi.ac.uk/clustalw/).
Results and discussion
A large intron in the cox2 locus of Ginkgo biloba
The plant mitochondrial cox2 gene encoding cytochrome oxidase subunit II has been intensively used for phylogenetic and evolutionary analyses (e.g., Nugent and Palmer 1991; Covello and Gray 1992; Kudla et al. 2002). In particular, the highly variable intron content of the gene has made the cox2 locus a preferred object of studies on the dynamics of intron evolution in higher plant mitochondria (Hiesel and Brennicke 1983; Laroche et al. 1997; Qiu et al. 1998; Rabbi and Wilson 1993; Albrizio et al. 1994; Kudla et al. 2002). The picture that has emerged from these studies is that the cox2 locus gained two introns in the moss lineage (Qiu et al. 1998). Subsequently, the two introns have undergone multiple independent loss events during seed-plant evolution (Hiesel and Brennicke 1983; Rabbi and Wilson 1993; Albrizio et al. 1994; Kudla et al. 2002). We have conducted a systematic analysis of intron evolution in mitochondrial cox2 genes of seed plants (Albertazzi et al. 1998; Kudla et al. 2002; and our unpublished results). In order to trace back the origin of reverse transcriptase/maturase-encoding introns versus noncoding organellar introns, we also included various representatives of primitive angiosperm lineages and determined the molecular structure of their cox2 loci (Albertazzi et al. 1998 and unpublished data). In the course of this work, we detected an unusually large intron within the cox2 gene of the maidenhair tree, Ginkgo biloba, the only living species of an ancient gymnosperm lineage, the Ginkgoopsida (Zhou and Zheng 2003).
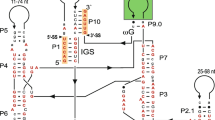
Sequencing of the intron and adjacent exon sequences revealed that the intron inserts in an evolutionary conserved position known from cox2 introns in many angiosperm species, such as Triticum aestivum (Bonen et al. 1984), Petunia hybrida (Pruitt and Hanson 1989, 1991), Beta vulgaris (Mann et al. 1991), and Acorus calamus (Albertazzi et al. 1998). The total size of the intron was determined to be 2,660 bp (EMBL database accession number AJ874265), making it one of the biggest plant mitochondrial introns discovered to date. The secondary structural domains conserved in all group II introns are easily identifiable in the sequence (Fig. 1, and data not shown), and strong homology of these domains with cox2 introns from angiosperm species confirms the singular origin of the intron in seed plants.
Sequence of the cox2 intron from Ginkgo biloba with surrounding exon sequence. Conserved intron domains are shown in color: domain VI in blue, domain V in green and the two complementary sequences forming the stem-loop structure of domain IV in magenta. Scrambled pieces from reverse transcriptase/maturase sequences are shown in red bold letters (see Figs. 2 and 3). Sites of C-to-U mRNA editing are indicated by the lowercase letter c. The affected codon is underlined, the amino acid specified prior to editing is shown in lowercase and followed by the uppercase letter for the amino acid specified by the edited codon
Splicing and RNA editing of the Ginkgo biloba cox2 transcript
A hallmark of mitochondrial gene expression in higher plants is the requirement for an additional RNA processing step, referred to as RNA editing. RNA editing changes the identity of single nucleotides in primary transcripts, predominantly by C-to-U transitions (Gualberto et al. 1989; Covello and Gray 1989; Hiesel et al. 1989; reviewed in Mulligan et al. 1999; Bock 2001). Comparison of exonic sequences of the cox2 gene from Ginkgo biloba with cox2 gene and protein sequences from other plant species suggested the presence of several potential RNA editing sites in Ginkgo biloba (Fig. 1, and data not shown) where conserved amino acid residues could be restored by C-to-U transitions. In order to confirm these potential editing sites experimentally and possibly detect additional sites, we reverse-transcribed DNaseI-treated RNA samples from Ginkgo biloba and amplified cox2 cDNAs. Direct sequencing of the amplified cDNA population confirmed intron excision at the conserved intron-exon borders and revealed numerous sites of C-to-U RNA editing (Fig. 1). The editing sites identified included some that could not be predicted by bioinformatics analyses in that they do not result in changes of the amino acid encoded by the affected triplet. These include a silent CUG-to-UUG transition in exon I (both codons specify leucine) and a CCA-to-UUA transition in exon II that involves two editing events and converts a proline into a leucine codon (Fig. 1). In the latter case, the editing in first codon position seems unnecessary in that a single editing, even in second codon position, would be sufficient to alter the codon identity.
As group II intron splicing and C-to-U RNA editing are absent from the nucleocytosolic compartment in higher plants, intron excision and high-frequency editing of the cox2 transcript confirm the mitochondrial localization of the cox2 gene in Ginkgo biloba.
Mitochondrial genes are occasionally found in more than one copy per genome, usually due to gene duplications that often are confined to a small taxonomic group (e.g., Rothenberg und Hanson 1987; Gutierres et al. 1997). In order to exclude this possibility for the Ginkgo biloba cox2 locus and to confirm that cox2 is a single copy mitochondrial gene, we performed a Southern blot analysis using a cox2 gene-specific probe. A single hybridizing band was detected in all restriction analyses (Fig. 2), ultimately confirming that cox2 is indeed a single copy mitochondrial gene in Ginkgo biloba.
Detection of cox2 sequences in Ginkgo biloba by DNA gel blot analysis. Total DNA was digested with the restriction enzymes indicated, separated by agarose gel electrophoresis, blotted, and hybridized to a cox2 gene-specific probe. Detection of a single hybridizing band in all lanes confirms that cox2 is a single copy mitochondrial gene in Ginkgo biloba
Remnants of an intron-encoded reverse transcriptase/maturase
Presence of an unusually large intron within the cox2 locus of Ginkgo biloba prompted us to search the intron for the presence of potential ORFs. Intron-encoded ORFs potentially encoding reverse transcriptase/maturase proteins most often insert in the loop of domain IV (Schuster and Brennicke 1987; Wissinger et al. 1991) and, as domain IV of the Ginkgo biloba intron contains a large insertion (loop size: 1,967 bp; Fig. 1), it seemed possible that it harbors a maturase-encoding gene. However, conceptual translation of the intron sequence in all three reading frames failed to detect a sufficiently long ORF (Fig. 3, and data not shown). Moreover, bioinformatics analysis of the longest theoretical ORFs revealed no significant homology to any protein in the databases. Also, detailed analysis of the conceptual translation of the 1,967-bp loop of domain IV (Fig. 1) revealed no potential ORF larger than 100 amino acids (Fig. 3, and data not shown). In order to trace back the evolutionary history of this long, presumably noncoding, insertion in intron domain IV, we refined the search criteria by removing stop codons and searching for short, nearly exact, matches at the level of the amino acid sequence. Interestingly, these searches identified altogether three regions with significant homology to intron-encoded reverse transcriptase/maturase proteins in fungal and bryophyte mitochondrial genomes (Figs. 1, 2, and 3). The first two stretches are homologous to conserved regions in the reverse transcriptase domains RT IIa and RT III of intron-encoded reverse transcriptases/maturases (Fig. 4; Mohr and Lambowitz 2003). The third stretch displays homology to the conserved domain X, the presumed intron maturase domain, which is located downstream of the reverse transcriptase domain in most intron-encoded reverse transcriptase/maturase proteins (Fig. 4; Mohr et al. 1993; Mohr and Lambowitz 2003).
Identification of remnants of reverse transcriptase/maturase sequences within the sequenced cox2 intron from Ginkgo biloba. Translation of the relevant parts of the intron in all three reading frames is shown. Nucleotide numbers of the intronic sequence are indicated above the DNA sequence. Amino acid sequence stretches homologous to reverse transcriptase/maturase genes in mitochondrial introns from other organisms are underlined and in bold (cp. Fig. 4), the corresponding DNA sequences are marked in red and bold. Note that all three frames contain numerous stop codons. This, together with the reverse transcriptase/maturase-homologous sequence stretches being in different reading frames, strongly suggests that there is no functional reverse transcriptase/maturase protein encoded in the intron. Instead, these sequences may represent pseudogenic remnants of an intact reverse transcriptase/maturase gene that is likely to have been present in the ancestral intron
Amino acid sequence alignment of reverse transcriptases/maturases with partial homology to the translated intron of the cox2 gene from Ginkgo biloba. N- and C-termini of the proteins display no significant conservation and no homology to the cox2 intron from Ginkgo biloba and, therefore, were omitted from this alignment. Asterisks mark residues that are identical in all sequences in the reverse transcriptase/maturase alignment, colon indicates conserved substitutions, and dot denotes semiconserved substitutions (according to ClustalW; http://www.ebi.ac.uk/clustalw/). Regions exhibiting homology with short sequence stretches in the translated cox2 intron from Ginkgo biloba are aligned with the respective Ginkgo biloba sequences (bold, underlined). Species and accession numbers are as follows: P.a. Podospora anserina (CAA38781 and CAA38778), M.p. Marchantia polymorpha (P38478), A.m. Allomyces macrogynus (S63652), S.o. Schizosaccharomyces octosporus (AF275271.2)
For several reasons, it seems highly unlikely that a functional reverse transcriptase/maturase protein is encoded in the Ginkgo biloba cox2 intron: (1) The sequences surrounding the reverse transcriptase/maturase-related stretches are full of stop codons (Fig. 3, and data not shown); (2) the short sequence stretches homologous to reverse transcriptase/maturase ORFs are in different reading frames (Fig. 3); (3) no regions of homology with the other highly conserved reverse transcriptase/maturase domains could be detected (Fig. 3, and data not shown). Instead, these reverse transcriptase/maturase-related sequences are likely to represent pseudogenic remnants of a functional reverse transcriptase/maturase gene formerly present in this intron.
Evolutionary implications
Removal of group II introns relies on an RNA-catalyzed splicing mechanism but requires protein factors in vivo, presumably to help fold the intronic RNA into the catalytically active secondary and tertiary structures (Michel et al. 1989; Copertino and Hallick 1993; Scott and Klug 1996; Michel and Ferat 1995). In addition to their splicing activity, group II introns can behave as mobile genetic elements, for example, by inserting themselves into intronless alleles, a process known as intron homing (Belfort and Perlman 1995; Curcio and Belfort 1996; Bonen and Vogel 2001). In several cases, bifunctional proteins referred to as reverse transcriptases/maturases mediate both splicing and homing of the intron: while the reverse transcriptase activity plays a role in intron mobility, the maturase domain functions in intron removal (Mohr et al. 1993; Mohr and Lambowitz 2003).
The detection of remnants of a reverse transcriptases/maturase gene in the cox2 intron of Ginkgo biloba has interesting implications. A systematic survey of land plants for the presence of this intron led to the conclusion that the intron was acquired once during early land-plant evolution in a common ancestor of all land plants, exclusive of liverworts (Qiu et al. 1998). Our data suggests that the intron, when it was acquired, possessed a reverse transcriptases/maturase gene and hence was a mobile group II intron. Subsequently, degeneration of the reverse transcriptase/maturase gene has occurred, with the intron in Ginkgo biloba reflecting an intermediate on the way to complete loss of the reverse transcriptase/maturase sequences. Consequently, the intron in present-day angiosperms no longer shows any evidence of the mobile past of the intron.
What could have been the source of the intron when it was acquired in early land-plant evolution? Our bioinformatics analyses show that the closest relatives of the reverse transcriptase/maturase sequences in Ginkgo biloba come from fungal homologues and a reverse transcriptase/maturase gene within an intron in the mitochondrial 18S rRNA gene from the liverwort Marchantia polymorpha (Fig. 4). Hence, two scenarios seem feasible: (1) transposition of the intron from a mitochondrial gene with a mobile intron (e.g., the 18S rRNA) into the cox2 locus; (2) acquisition from a fungus via horizontal gene transfer. Recent studies have provided evidence for horizontal gene transfer of mitochondrial genes and introns both from fungi to higher plants and between higher plant species (Vaughn et al. 1995; Marienfeld et al. 1997; Cho et al. 1998; Bergthorsson et al. 2003; Won and Renner 2003; Davis and Wurdack 2004; Mower et al. 2004). At present, we cannot distinguish between the two above possibilities of cox2 intron acquisition, but molecular phylogenetic analysis of the homologous introns in mosses, hornworts, and ferns may help to resolve this issue.
A recent analysis of mitochondrial introns has provided evidence for ORF remnants in several introns of the liverwort M. polymorpha and a few other bryophytes but failed to detect such sequences in the higher plant Arabidopsis thaliana (Toor et al. 2001). Our finding that remnants of a reverse transcriptase/maturase gene are present in the cox2 intron of Ginkgo biloba represents the first case of traces of reverse transcriptase/maturase sequences in an intron of a higher plant lineage. Due to their high degree of evolutionary degeneration (Figs. 1, 3, and 4), ORF remnants were detectable only by refined bioinformatics searches, and the presence of rather short sequence traces in the gymnosperm lineage (as represented by Ginkgo biloba) provides an explanation as to why they are undetectable in the angiosperm lineage (Toor et al. 2001). It thus seems conceivable that the last recognizable traces of the reverse transcriptase/maturase ORF in the cox2 intron were lost during seed-plant evolution. This underscores the importance of including early branching seed-plant lineages in reconstruction of the evolutionary history of plant group II introns.
Another interesting question is how the maturase function lost by degeneration of the reverse transcriptase/maturase gene in the cox2 intron was compensated for. The two most plausible possibilities may be: (1) that the gene was transferred to the nucleus; (2) that the splicing function was taken over by another intron-encoded maturase in the mitochondrial genome. The single chloroplast-genome-encoded maturase gene matK provides precedence for the latter scenario in that it appears to be involved not only in the splicing of the group II intron it resides in (trnK) but also in the splicing of several other plastid introns (Hess et al. 1994; Jenkins et al. 1997; Vogel et al. 1999).
References
Albertazzi FJ, Kudla J, Bock R (1998) The cox2 locus of the primitive angiosperm plant Acorus calamus: molecular structure, transcript processing and RNA editing. Mol Gen Genet 259:591–600
Albrizio M, de Gara L, De Benedetto C, Arrigoni O, Gallerani R (1994) Investigations of the coxII intron structure in the mitochondrial genomes of angiosperms. Plant Sci 100:179–186
Anziano PQ, Butow RA (1991) Splicing-defective mutants of the yeast mitochondrial coxI gene can be corrected by transformation with a hybrid maturase gene. Proc Natl Acad Sci USA 88:5592–5596
Belfort M (1991) Self-splicing introns in prokaryotes: migrant fossils? Cell 64:9–11
Belfort M, Perlman PS (1995) Mechanisms of intron mobility. J Biol Chem 270:30237–30240
Bergthorsson U, Adams KL, Thomason B, Palmer JD (2003) Widespread horizontal transfer of mitochondrial genes in flowering plants. Nature 424:197–201
Bock R (2001) RNA editing in plant mitochondria and chloroplasts. In: Bass B (ed) Frontiers in molecular biology: RNA editing, Oxford University Press, NewYork, pp 38–60
Bonen L, Vogel J (2001) The ins and outs of group II introns. Trends Genet 17:322–323
Bonen L, Boer PH, Gray MW (1984) The wheat cytochrome oxidase subunit II gene has an intron and three radical amino acid changes relative to maize. EMBO J 3:2531–2536
Cho Y, Qiu Y-L, Kuhlman P, Palmer JD (1998) Explosive invasion of plant mitochondria by a group I intron. Proc Natl Acad Sci USA 95:14244–14249
Chowira BM, Berzal-Herranz A, Burke JM (1994) Novel RNA polymerization reaction catalyzed by a group I ribozyme. EMBO J 12:3599–3605
Copertino DW, Hallick RB (1993) Group II and group III introns of twintrons: potential relationship with nuclear pre-mRNA introns. Trends Biochem Sci 18:467–471
Covello PS, Gray MW (1989) RNA editing in plant mitochondria. Nature 341:662–666
Covello PS, Gray MW (1992) Silent mitochondrial and active nuclear genes for subunit 2 of cytochrome c oxidase (cox2) in soybean: evidence for RNA-mediated gene transfer. EMBO J 11:3815–3820
Curcio MJ, Belfort M (1996) Retrohoming: cDNA-mediated mobility of group II introns requires a catalytic RNA. Cell 84:9–12
Davis CC, Wurdack KJ (2004) Host-to-parasite gene transfer in flowering plants: phylogenetic evidence from Malpighiales. Science 305:676–678
Deshpande NN, Hollingsworth M, Herrin DL (1995) The atpF group-II intron-containing gene from spinach chloroplasts is not spliced in transgenic Chlamydomonas chloroplasts. Curr Genet 28:122–127
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Dürrenberger F, Rochaix J-D (1991) Chloroplast ribosomal intron of Chlamydomonas reinhardtii: in vitro self-splicing, DNA endonuclease activity and in vivo mobility. EMBO J 10:3495–3501
Gualberto JM, Lamattina L, Bonnard G, Weil J-H, Grienenberger J-M (1989) RNA editing in wheat mitochondria results in the conservation of protein sequences. Nature 341:660–662
Gutierres S, Lelandais C, De Paepe R, Vedel F, Chetrit P (1997) A mitochondrial sub-stoichiometric orf87–nad3–nad1 exonA co-transcription unit present in Solanaceae was amplified in the genus Nicotiana. Curr Genet 31:55–62
Hess WR, Hoch B, Zeltz P, Hübschmann T, Kössel H, Börner T (1994) Inefficient rpl2 splicing in barley mutants with ribosome-deficient plastids. Plant Cell 6:1455–1465
Hetzer M, Wurzer G, Schweyen RJ, Mueller MW (1997) Trans-activation of group II intron splicing by nuclear U5 snRNA. Nature 386:417–420
Hiesel R, Brennicke A (1983) Cytochrome oxidase subunit II gene in mitochondria of Oenothera has no intron. EMBO J 2:2173–2178
Hiesel R, Wissinger B, Schuster W, Brennicke A (1989) RNA editing in plant mitochondria. Science 246:1632–1634
Jenkins BD, Barkan A (2001) Recruitment of a peptidyl-tRNA hydrolase as a facilitator of group II intron splicing in chloroplasts. EMBO J 20:872–879
Jenkins BD, Kulhanek DJ, Barkan A (1997) Nuclear mutations that block group II RNA splicing in maize chloroplasts reveal several intron classes with distinct requirements for splicing factors. Plant Cell 9:283–296
Knoop V (2004) The mitochondrial DNA of land plants: peculiarities in phylogenetic perspective. Curr Genet 46:123–139
Kudla J, Albertazzi FJ, Blazevic Hermann M, Bock R (2002) Loss of the mitochondrial cox2 intron 1 in a family of monocotyledonous plants and utilization of mitochondrial intron sequences for the construction of a nuclear intron. Mol Genet Genomics 267:223–240
Kuhsel MG, Stickland R, Palmer JD (1990) An ancient group I intron shared by eubacteria and chloroplasts. Science 250:1570–1573
Laroche J, Li P, Maggia L, Bousquet J (1997) Molecular evolution of angiosperm mitochondrial introns and exons. Proc Natl Acad Sci USA 94:5722–5727
Li F, Holloway SP, Lee J, Herrin DL (2002) Nuclear genes that promote splicing of group I introns in the chloroplast 23S rRNA and psbA genes in Chlamydomonas reinhardtii. Plant J 32:467–480
Liere K, Link G (1995) RNA-binding activity of the matK protein encoded by the chloroplast trnK intron from mustard (Sinapis alba L.). Nucleic Acids Res 23:917–921
Mann V, Ekstein I, Nissen H, Hiser C, McIntosh L, Hirschberg J (1991) The cytochrome oxidase II gene in mitochondria of the sugar-beet Beta vulgaris L.. Plant Mol Biol 17:559–566
Marienfeld JR, Unseld M, Brandt P, Brennicke A (1997) Viral nucleic acid sequence transfer between fungi and plants. Trends Genet 13:260–261
Michel F, Ferat J-L (1995) Structure and activities of group II introns. Annu Rev Biochem 64:435–461
Michel F, Umesono K, Ozeki H (1989) Comparative and functional anatomy of group II catalytic introns—a review. Gene 82:5–30
Mohr G, Lambowitz AM (2003) Putative proteins related to group II intron reverse transcriptase/maturases are encoded by nuclear genes in higher plants. Nucleic Acids Res 31:647–652
Mohr G, Perlman PS, Lambowitz AM (1993) Evolutionary relationships among group II intron-encoded proteins and identification of a conserved domain that may be related to maturase function. Nucleic Acids Res 21:4991–4997
Mower JP, Stefanovic S, Young GJ, Palmer JD (2004) Gene transfer from parasitic to host plants. Nature 432:165–166
Mulligan RM, Williams MA, Shanahan MT (1999) RNA editing site recognition in higher plant mitochondria. J Hered 90:338–344
Neuhaus H, Link G (1987) The chloroplast tRNALys(UUU) gene from mustard (Sinapis alba) contains a class II intron potentially coding for a maturase-related polypeptide. Curr Genet 11:251–257
Nugent JM, Palmer JD (1991) RNA mediated transfer of the gene coxII from the mitochondrion to the nucleus during flowering plant evolution. Cell 66:473–481
Ostheimer GJ, Williams-Carrier R, Belcher S, Osborne E, Gierke J, Barkan A (2003) Group II intron splicing factors derived by diversification of an ancient RNA-binding domain. EMBO J 22:3919–3929
Padgett RA, Podar M, Boulanger SC, Perlman PS (1994) The stereochemical course of group II intron self-splicing. Science 266:1685–1688
Perron K, Goldschmidt-Clermont M, Rochaix J-D (1999) A factor related to pseudouridine synthases is required for chloroplast group II intron trans-splicing in Chlamydomonas reinhardtii. EMBO J 18:6481–6490
Perron K, Goldschmidt-Clermont M, Rochaix J-D (2004) A multiprotein complex involved in chloroplast group II intron splicing. RNA 10:704–711
Pruitt KD, Hanson MR (1989) Cytochrome oxidase subunit II sequences in petunia mitochondria: two intron containing genes and an intron-less pseudo gene associated with cytoplasmic male sterility. Curr Genet 16:281–291
Pruitt KD, Hanson MR (1991) Splicing of the Petunia cytochrome oxidase subunit II intron. Curr Genet 19:191–197
Pyle AM, Cech TR (1991) Ribozyme recognition of RNA by tertiary interactions with specific ribose 2′-OH groups. Nature 350:628
Qiu Y-L, Cho Y, Cox JC, Palmer JD (1998) The gain of three mitochondrial introns identifies liverworts as the earliest land plants. Nature 394:671–674
Rabbi MF, Wilson KG (1993) The mitochondrial coxII intron has been lost in two different lineages of dicots and altered in others. Am J Bot 80:1216–1223
Rivier C, Goldschmidt-Clermont M, Rochaix J-D (2001) Identification of an RNA-protein complex involved in chloroplast group II intron trans-splicing in Chlamydomonas reinhardtii. EMBO J 20:1765–1773
Rothenberg M, Hanson MR (1987) Different transcript abundance of two divergent ATP synthase subunit 9 genes in the mitochondrial genome of Petunia hybrida. Mol Gen Genet 209:21–27
Schuster W, Brennicke A (1987) Plastid, nuclear and reverse transcriptase sequences in the mitochondrial genome of Oenothera: is genetic information transferred between organelles via RNA? EMBO J 6:2857–2863
Scott WG, Klug A (1996) Ribozymes: structure and mechanism in RNA catalysis. Trends Biochem Sci 21:220–223
Sheveleva EV, Hallick RB (2004) Recent horizontal intron transfer to a chloroplast genome. Nucleic Acids Res 32:803–810
Sugita M, Sugiura M (1996) Regulation of gene expression in chloroplasts of higher plants. Plant Mol Biol 32:315–326
Sugiura M (1992) The chloroplast genome. Plant Mol Biol 19:149–168
Szczepanek T, Lazowska J (1996) Replacement of two non-adjacent amino acids in the S. cerevisiae bi2 intron-encoded RNA maturase is sufficient to gain a homing-endonuclease activity. EMBO J 15:3758–3767
Thomson MC, Macfarlane JL, Beagley CT, Wolstenholme DR (1994) RNA editing of mat-r transcripts in maize and soybean increases similarity of the encoded protein to fungal and bryophyte group II intron maturases: evidence that mat-r encodes a functional protein. Nucleic Acids Res 22:5745–5752
Till B, Schmitz-Linneweber C, Williams-Carrier R, Barkan A (2001) Crs1 is a novel group II intron splicing factor that was derived from a domain of ancient origin. RNA 7:1227–1238
Toor N, Hausner G, Zimmerly S (2001) Coevolution of group II intron RNA structures with their intron-encoded reverse transcriptases. RNA 7:1142–1152
Vaughn JC, Mason MT, Sper-Whitis GL, Kuhlman P, Palmer JD (1995) Fungal origin by horizontal gene transfer of a plant mitochondrial group I intron in the chimeric coxI gene of Peperomia. J Mol Evol 41:563–572
Vogel J, Hübschmann T, Börner T, Hess WR (1997) Splicing and intron-internal RNA editing of trnK-matK transcripts in barley plastids: support for matK as an essential splicing factor. J Mol Biol 270:179–187
Vogel J, Börner T, Hess WR (1999) Comparative analysis of splicing of the complete set of chloroplast group II introns in three higher plant mutants. Nucleic Acids Res 27:3866–3874
Winkler M, Kück U (1991) The group IIB intron from the green alga Scenedesmus obliquus mitochondrion: molecular characterization of the in vitro splicing products. Curr Genet 20:495–502
Wissinger B, Schuster W, Brennicke A (1991) Trans splicing in Oenothera mitochondria: nad1 mRNAs are edited in exon and trans-splicing group II intron sequence. Cell 65:473–482
Won H, Renner SS (2003) Horizontal gene transfer from flowering plants to Gnetum. Proc Natl Acad Sci USA 100:10824–10829
Zhou Z, Zheng S (2003) The missing link in Ginkgo evolution. Nature 423:821–822
Acknowledgments
We thank Dragica Blazević for excellent technical assistance during the early phase of this work. This research was supported by grants from the Deutsche Forschungsgemeinschaft (DFG) to RB and JK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ahlert, D., Piepenburg, K., Kudla, J. et al. Evolutionary origin of a plant mitochondrial group II intron from a reverse transcriptase/maturase-encoding ancestor. J Plant Res 119, 363–371 (2006). https://doi.org/10.1007/s10265-006-0284-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-006-0284-0