Abstract
Rearrangement of cellulose microfibrils within cell-wall matrices is considered one of the most critical steps in the regulation of both the orientation and extent of cell expansion in plants. Xyloglucan endotransglucosylase/hydrolases (XTHs) are a family of enzymes that mediate the construction and restructuring of load-bearing cross links among cellulose microfibrils. The Arabidopsis thaliana XTH genes AtXTH17, 18, 19, and 20 are phylogenetically closely related to one another and are preferentially expressed in the roots. However, they exhibit different expression profiles within the root and respond to hormonal signals differently. To investigate their functions in root growth, we examined phenotypes of loss-of-function mutants for these genes using T-DNA insertion lines and RNAi plants. These functional analyses disclosed a principal role for the AtXTH18 gene in primary root elongation. Of the four XTH genes, AtXTH18 exhibits the highest level of mRNA expression. We also determined auxin-signaling pathways for these genes using a mutant with a defect in the AXR2/IAA7 gene and found that the expression of AtXTH19 in the elongation/maturation region of the root is under the control of the AXR2/IAA7 signaling pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Understanding cell expansion control is a key step in elucidating the mechanism of plant growth and development. Alternation in cell-wall properties is considered one of the most critical steps in the regulation of cell expansion processes (Cosgrove 1997a, b; Fry 2004). Although several hormonal signals, particularly those of auxin and gibberellin, are considered to play principal roles in the regulation of root growth (see references cited in Fu and Harberd 2003; Goda et al. 2004), the molecular events linking hormonal signaling to cell-wall modification raise questions that are still unanswered.
The plant cell wall is a supramolecular architecture composed of crystalline cellulose microfibrils and various types of macromolecules (Carpita and Gibeaut 1993), and many genes encoding enzymes and structural proteins are required for its construction and remodeling processes (reviewed by Yokoyama and Nishitani 2004). In this complex architecture, xyloglucans function as load-bearing cross links among microfibrils, where they play a central role in modulating the mechanical properties of cell walls (reviewed by Nishitani 1997). Whereas auxin signaling has been implicated as the principal regulatory pathway for xyloglucan metabolism during cell-wall loosening (e.g., Nishitani and Masuda 1981) and xyloglucan endotransglucosylase/hydrolases (XTHs) have been implicated in auxin-mediated xyloglucan metabolism (e.g., Catala et al. 2001; Nakamura et al. 2003), the function of XTH in the cell expansion process has not been demonstrated.
XTHs are a family of enzymes that catalyze molecular grafting and/or the hydrolysis of xyloglucans (Fry et al. 1992; Nishitani and Tominaga 1992; Okazawa et al. 1993; Rose et al. 2002). Therefore, they play a principal role in the construction and restructuring of load-bearing cross links among cellulose microfibrils and hence the framework of the cell wall. Thirty-three members of the XTH gene family occur in the genome of Arabidopsis thaliana (Yokoyama and Nishitani 2001; Yokoyama et al. 2004). Extensive expression analyses of these XTH genes revealed that most family members exhibit temporally and spatially distinct expression patterns and respond differently to various hormonal signals, including auxin (Akamatsu et al. 1999; Goda et al. 2004; Hyodo et al. 2003; Nakamura et al. 2003; Xu et al. 1995, 1996; Yokoyama and Nishitani 2000, 2001; Yokoyama et al. 2004). Recently, we showed that AtXTH27, a member of the A. thaliana XTH family of proteins, is involved in the cell-wall modification of tracheary elements at a specific stage of rosette leaf development and is essential for tertiary vein development (Matsui et al. 2005). Data thus far obtained support our hypothesis that each member of the XTH family of genes is regulated by specific cues and is committed to cell-wall dynamics specific to certain tissues or cell types (Nishitani 2002). However, despite extensive studies on many XTH genes from various plant species, no XTH member responsible for the cell elongation process has been identified.
The Arabidopsis XTH genes AtXTH17, 18, 19, and 20 are phylogenetically closely related to one another and are preferentially expressed in the roots. Among the four genes, cis-regulatory sequences implicated in the root-preferential expression profile are conserved (Vissenberg et al. 2005a; Yokoyama and Nishitani 2001). Detailed expression analyses of the four genes showed that AtXTH17 and AtXTH18 were expressed in all cell types in the elongating and differentiating regions of the root. AtXTH19 was expressed in the apical dividing and elongating regions as well as in the differentiation zone and was upregulated by auxin. In contrast, AtXTH20 was specifically expressed in vascular tissues in the mature regions of the root (Vissenberg et al. 2005a). These results strongly suggest that the four XTH genes, which were generated by gene duplication, have diversified their expression profiles within the root in such a way as to take responsibility for particular physiological roles in the roots.
To investigate the roles of these XTH genes in A. thaliana root growth, we analyzed a loss-of-function mutant phenotype for each of the four genes using T-DNA insertion lines or RNAi plants. This functional analysis disclosed a principal role for the AtXTH18 gene in primary root elongation. We also focused on the AtXTH18 and AtXTH19 genes and determined their gene expressions using a mutant defective in auxin signaling. This disclosed that expression of AtXTH19 is controlled by auxin signaling.
Materials and methods
Plant materials and hormone treatment
Seeds of A. thaliana (L.) Heynh. ecotype Columbia and its transgenic plants were sown and grown on either rock wool moistened with MGRL medium (Tsukaya et al. 1991) or sterilized, germinated, and grown on solid Murashige and Skoog (MS) medium (Murashige and Skoog 1962) supplemented with 3% sucrose at 22°C under continuous illumination at 70 μmol m–2 s–1. For β-glucuronidase (GUS) enzymatic activity analyses, transgenic seeds were sown on vertically oriented MS plates under continuous light and temperature conditions for 7 days unless otherwise specified. To examine the action of indole-3-acetic acid (IAA) on the expression of XTH promoter::GUS fusion genes, seedlings were grown on vertically oriented MS plates under continuous light and temperature conditions for 5 days. They were then transferred to a new solid MS plate with or without 0.1 nM IAA (Wako Junyaku Co., Osaka, Japan) and grown for an additional 2 days under the same conditions.
GUS activity assay
The GUS activities of 7-day-old light-grown seedlings were assayed using 4-methylumbelliferyl β-glucuronide as a substrate according to the method of Jefferson et al. (1987) with some modification (Vissenberg et al. 2005a). Protein concentrations were estimated using the Bradford method (1976) (Bio-Rad Laboratories, Hercules, CA, USA). The specific activity of the GUS enzyme in extracts was calculated as nanomoles of 4-methylumbelliferone produced per minute per milligram of total protein.
For the histochemical analysis of GUS expression, seedlings were incubated in GUS staining solution (0.1 M Na2HPO4–NaH2PO4 buffer pH 7.0, 1 mM X-Gluc) at 37°C for 2–20 h, followed by washing in 70% ethanol. The procedure was undertaken according to Jefferson et al. (1987). Stained seedlings were observed under a Leica MZ APO binocular microscope (Leica, Cherbourg, Switzerland), and images were recorded using a Nikon DXM1200 digital camera (Tokyo, Japan).
Crossing
Seeds of axr2-1 (Wilson et al. 1990) and ein2-1 (Guzman and Ecker 1990) mutants were supplied by the Arabidopsis Biological Resource Center. The homozygous double mutants axr2-1/pAtXTH18::GUS, axr2-1/pAtXTH19::GUS, ein2-1/pAtXTH18::GUS, and ein2-1/pAtXTH19::GUS were generated by crossing pollen of the homozygous axr2-1 mutant or ein2-1 mutant to homozygous transgenic plants expressing pAtXTH18::GUS or pAtXTH19::GUS.
Isolation and characterization of T-DNA insertion lines
T-DNA insertion lines for AtXTH17 (SALK_015077 and SALK_008429), AtXTH19 (SALK_034274), and AtXTH20 (SAIL_575_H09) genes were supplied by the Salk Institute Genome Analysis Laboratory. Insertions of T-DNA into individual XTH genes were confirmed by polymerase chain reaction (PCR) and sequencing using primers specific to T-DNA left-border sequences (SALK left border (LBb1) 5′-gcgtggaccgcttgctgcaact-3′ and SAIL left border (LB3s) 5′-gaatttcataaccaatctcgatacac-3′) and primers specific to the XTH genes SALK_015077 (5′-atgggctaatggaaaatcatcttgtt-3′), SALK_008429 (5′-tactttgcacacctttcattcttgtc-3′), SALK_034274 (5′-actggatccttgtatatattttggaggtgtattga-3′), and SAIL_575_H09 (5′-ccgtggatccgaatgttgttactgttccagctg-3′). Isolated homozygous lines for SALK_015077, SALK_008429, SALK_034274, and SAIL_575_H09 were designated xth17-1, xth17-2, xth19, and xth20, respectively. The expression level of mRNA in each of the four homozygous lines was measured by quantitative real-time reverse-transcriptase PCR (RT-PCR) according to the procedure described by Yokoyama and Nishitani (2001).
XTH18 RNAi construct
We designed an AtXTH18 RNAi construct in which the transcription of a hairpin-shaped AtXTH18 mRNA fragment was driven by the 1.5 kb AtXTH18 promoter. A 1,839-bp fragment of the XTH gene containing a 5′ untranscribed region (1,460 bp), the first exon (282 bp), and the first intron (97 bp) was amplified by PCR using the following primers containing restriction sites as indicated in boldface: the forward primer 5′-gcctgcatgcgaggttggcatcttattttgtatg-3′ (SphI), and the reverse primer 5′-ccgtggatcctaagctacacaaagtcatcataaacac-3′(BamHI). A 309-bp fragment containing the first exon was amplified by PCR using the following primers containing restriction sites as indicated in boldface: forward primer 5′-cctcgagctctatcaatacacctccaaaatatatacaaaa-3′ (SacI), and reverse primer 5′-ccgtggatccgaatgttgttactgttccagctg-3′ (BamHI). The amplified 1,839-bp fragment was TA-cloned into plasmid pBluescript SK II, and the 309-bp PCR fragment was cloned into the plasmid at the downstream end of the 1,839-bp fragment using the BamHI/SacI restriction sites in antisense orientation. AtXTH18 RNAi constructs thus cloned in pBluescript SK II were excised as a 2.2-kbp SphI/SacI fragment and cloned into a binary vector pBI101 (Clontech, Mountain View, CA, USA). After the binary vector was transformed into Agrobacterium tumefaciens strain C58, A. thaliana Col 0 plants were transformed with the RNAi construct according to the floral-dip method of Clough and Bent (1998).
RNA extraction and real-time RT-PCR
Seven-day-old seedlings of A. thaliana Columbia lines (Col 0, xth17-1, xth17-2, xth19, xth20, and AtXTH18 RNAi lines) were frozen in liquid nitrogen and subjected to RNA extraction according to a conventional SDS–phenol protocol. Oligo DNA primer sets for the AtXTH17, 18, 19, and 20 genes were prepared and used, as previously described (Yokoyama and Nishitani 2001). Quantitative one-step real-time RT-PCR was performed using the SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) in an ABI Prism TM 5700 Sequence Detection System (Applied Biosystems) according to the protocol provided by the supplier.
Measurements of root dimensions
Primary root lengths of Arabidopsis seedlings grown on vertically placed MS plates were measured under a binocular microscope equipped with an ocular micrometer (MZ APO, Leica). For micrometer measurements of root epidermal cell length, fresh root sections ranging 10–20 mm from the root tip of 7-day-old seedlings were prepared and stained with a solution of propidium iodide (10 μm m–1) for 30 s, followed by observations of length of cells with root hair under a laser scanning confocal microscope (Fluoview; Olympus, Tokyo, Japan) equipped with a filter set (extraction filter 360/40 nm, barrier filter; 420 nm). Root radii at 15 mm from the root tip of 7-day-old seedlings were measured under a binocular microscope (MZ APO, Leica) equipped with a digital camera (CAMEDIA C-5050 ZOOM, Olympus).
For measurements of tissue transverse sections, 5-day-old seedlings were fixed in 2% glutaraldehyde for 2 h, followed by successive dehydration in an ethanol series. The samples were then embedded in Technovit 7100 resin (Kulzer), sectioned using a rotary microtome (Microm HM 325) at 10 μm, and observed under a differential interference contrast microscope (DMRPX, Leica) equipped with a digital camera (CAMEDIA C-5050 ZOOM, Olympus).
Results
Phenotypes of transgenic A. thaliana plants with modified XTH gene expression levels
T-DNA insertion lines
To gain insight into the role of each of the four XTH genes AtXTH17, 18, 19, and 20, we undertook phenotypic characterizations of their loss-of-function transgenic plants. We identified T-DNA insertion lines for AtXTH17, 19, and 20 in the Salk Institute Genome Analysis Laboratory and isolated their homozygous lines. No T-DNA insertion line was identified for AtXTH18. Figure 1 shows a schematic drawing for T-DNA insertion sites and mRNA expression levels in individual T-DNA insertion lines. In 7-day-old seedlings of xth17-1 and xth17-2, the AtXTH17 mRNA level, as studied by real-time RT-PCR, was reduced to less than 10% of wild-type expression. In the xth19 line, the AtXTH19 mRNA level was reduced to 66% of wild-type expression. On the other hand, in the xth20 line, the AtXTH20 mRNA expression level was not reduced: rather, it was elevated four times. Given the presence of enhancer sequence in the CaM35S promoter for the T-DNA transgene inserted at the 5′ upsteam of the AtXTH20 gene, it is possible that transcription of the AtXTH20 gene is stimulated by action of the CaM35S enhancer.
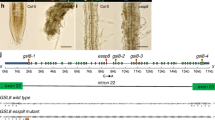
Schematic presentation of T-DNA insertion sites on the three xyloglucan endotransglucosylase/hydrolase (XTH) genes and mRNA expression levels in individual T-DNA insertion lines. a AtXTH17 mRNA expression in wild-type Columbia, xth17-1, and xth17-2 mutants. b AtXTH19 mRNA expression in wild-type Columbia and xth19 mutants. c AtXTH20 mRNA expression in wild-type Columbia and xth20 mutants. Horizontal arrows show directions of individual genes. Black boxes indicate exons and solid lines indicate noncoding regions of individual genes. T-DNA insertion sites are shown as vertical lines with arrows. Abundance of individual mRNA species in 7-day-old seedlings was measured by quantitative real-time reverse-transcriptase polymerase chain reaction (RT-PCR). Mean copy numbers for individual mRNAs per total RNA are shown with standard deviations as vertical lines (n=3)
Figure 2 shows primary root lengths of the 7-day-old seedlings of the wild-type and four T-DNA insertion lines. No significant differences in root growth were observed either in the three insertion lines xth17-1, xth17-2 and xth19, in which XTH gene expression is significantly reduced, or in a gain-of-function insertion line xth20, in which the XTH gene expression was enhanced by four times.
Primary root lengths of 7-day-old seedlings of four T-DNA insertion lines and wild-type (wt) Columbia. Mean values with standard errors are shown (n=9 for wt, n=10 for mutants). According to t tests, no statistically significant difference was found between the wild-type and each of the four mutants (P<0.05)
RNAi plants
Because no T-DNA insertion line for AtXTH18 was obtained, we prepared RNAi plants for the AtXTH18 gene. We isolated four homozygous transgenic lines for the RNAi construct and examined their phenotypes and AtXTH18 mRNA levels in 7-day-old seedlings. We also examined levels of AtXTH17, 19, and 20 mRNA in individual plant lines because sequence similarities of 93%, 79%, and 84% were observed between the first exon of AtXTH18 and that of AtXTH17, 19, and 20, respectively. Figure 3 shows the mRNA levels in a typical AtXTH18 RNAi plant, in which the AtXTH18 mRNA level was reduced to 86% of that in wild-type plants. The extent of mRNA reduction was small but significant. Similar reduction levels of mRNA were observed in other RNAi lines (data not shown). On the other hand, no significant reduction in the mRNA levels of the other three genes was seen in AtXTH18 RNAi plants, indicating that the abundance of AtXTH18 mRNA was specifically interfered with in this transgenic line.
Expression levels of the four xyloglucan endotransglucosylase/hydrolase (XTH) mRNAs in the XTH18 RNAi plant line. Abundance of AtXTH17, 18, 19, and 20 gene transcripts in 7-day-old seedlings of wild-type Columbia (black bars) and the XTH18 RNAi line (open bars) was measured using the quantitative real-time reverse-transcriptase polymerase chain reaction (RT-PCR) procedure. Mean copy numbers for individual mRNAs per total RNA are shown with standard deviations as vertical lines (n=3). Asterisk indicates statistical significance (P<0.05). The AtXTH18 mRNA level in the RNAi plant was reduced to 86% of wild-type Columbia
Since the AtXTH18 gene is expressed preferentially in roots (Vissenberg et al. 2005a; Yokoyama and Nishitani 2001), we focused on root growth in the RNAi lines. Figure 4a shows that the growth rate of primary roots was significantly reduced in the RNAi line compared with that in wild-type plants. The development of lateral roots was not significantly affected in the RNAi plants (data not shown). Measurements of epidermal cell length at 10–20 mm from the root tip of 7-day-old seedling showed that the average cell length at the mature zone was reduced to 85% of that of wild-type plants (Fig. 4b). Thus, one can attribute the reduction in primary root length in the RNAi plant to that in the final cell length as disclosed in the epidermal cells. In addition to reduced cell length, aberrant trichomes were more often observed in RNAi plants compared with wild-type plants (cf. Fig. 5c, d).
Root-growth parameters in XTH18 RNAi plants. a Time course of primary root elongation of wild-type Columbia (solid square) and the RNAi plant (open circle) grown on vertical solid MS plates. Mean root lengths are shown with standard deviations as vertical lines (n=10). Asterisk indicates that the lengths are statistically significant between the wild-type and RNAi line (P<0.05). b Epidermal cell lengths at the mature zone of the primary root of the wild-type and XTH18 RNAi plants. Cell lengths of stained primary roots were measured under a laser scanning confocal microscope. Mean values are shown with standard deviations as vertical lines (n=45 for wild-type, n=59 for RNAi). Asterisk indicates that the cell lengths are statistically significant between the wild-type and RNAi line (P<0.05). c Root width 2 mm from the root tip of 7-day-old seedlings of the wild-type and RNAi plants as measured under a binocular microscope. Mean values are shown with standard deviations as vertical lines (n=13 + for wild-type, n=14 for RNAi)
Morphological phenotypes of XTH18 RNAi plants. a–d Primary roots of 7-day-old seedlings of the wild-type (a, c) and XTH18 RNAi plant line (b, d). In the RNAi plants, aberrant morphologies of epidermal cells, particularly in the root hairs, are observed in addition to reduced epidermal cell length. e , f Differential interference contrast images of transverse sections of primary roots of 5-day-old wild-type plants (e) and RNAi plants (f). No difference in the cell file pattern was found between the wild-type and RNAi plants. Scale bars 1 mm (a, b), 250 μm (c, d), 50 μm (e, f)
However, other root phenotypic traits such as root width (Figs. 4c, 5a, b) and cell files within roots (Fig. 5e, f), as disclosed in transverse sections, were not drastically affected. Similar phenotypic traits were observed in the other RNAi lines (data not shown). These results indicate that only a slight reduction in the AtXTH18 mRNA level caused a significant reduction in the final cell length of the primary root. This indicates an essential role for AtXTH18 in the root elongation process.
Auxin signaling
Auxin signaling is considered to play a principal role in the regulation of root growth. Our previous study showed that application of IAA to intact seedlings of A. thaliana causes significantly elevated expression of both AtXTH19 mRNA (Yokoyama and Nishitani 2001) and the AtXTH19 promoter::GUS fusion gene (Vissenberg et al. 2005a), but causes little or no effect on expressions of the AtXTH18 gene. To determine auxin action on XTH gene expression in the root, we analyzed the expression of promoter::GUS fusion genes for AtXTH18 and AtXTH19 in the axr2-1 mutant, an auxin-insensitive mutant in which the expression of early auxin genes are suppressed (Abel and Theologis 1996; Abel et al. 1995; Gil et al. 1994). Since the axr2-1 is a pleitropic mutant capable of conferring resistance to other hormonal signals, including ethylene (Wilson et al. 1990), as a control experiment, we also examined the expression of the two XTH genes in the ein2-1 mutant, an ethylene-insensitive mutant (Guzman and Ecker 1990; Roman et al. 1995) in which the ethylene-signaling pathway is defective (Alonso et al. 1999).
In the wild-type background, the AtXTH18 promoter::GUS fusion gene (pAtXTH18::GUS) and a 5′-truncated AtXTH18 promoter::GUS fusion gene (-329pAtXTH18) was expressed in the elongating and differentiating regions of the primary root (Vissenberg et al. 2005a, Fig. 6a, d, f). Although the expression level of the pAtXTH18::GUS gene was slightly elevated by 50% in the axr2-1 mutant compared with that in wild-type plants (Fig. 7a), spatial expression profiles of the two gene constructs in the root were not prominently affected either in axr2-1 (Fig. 6a, b, d, e, f, h) or in ein2-1 mutant plants (Fig. 6a, c, f, h).
Expression profiles of AtXTH18 promoter::GUS and AtXTH19 promoter::GUS fusion genes in 7-day-old seedlings of wild-type, axr2-1, and ein2-1 mutant backgrounds. a–c pAtXTH18::GUS fusion gene expression in the primary roots of wild-type (a), axr2-1 (b), and ein2-1 (c) backgrounds. d, e Expression of the −339pAtXTH18::GUS fusion gene in which a 5′ upstream region is truncated in the primary root of wild-type (d) and axr2-1 (e) plants. f–h pAtXTH19::GUS fusion gene expression in lateral root primordia in wild-type (f), axr2-1 (g), and ein2-1 (h) backgrounds. i–k pAtXTH19::GUS fusion gene expression in primary roots of wild-type (i), axr2-1 (j), and ein2-1 (k) backgrounds. l and m Expression of a 5′ truncated promoter gene construct,−330pAtXTH19::GUS, in the primary root of wild-type (l) and axr2-1 (m) plants. n–p pAtXTH19::GUS fusion gene expression in lateral root primordia in wild-type (n), axr2-1 (o), and ein2-1 (p) backgrounds. pAtXTH18::GUS is expressed in elongation and differentiation regions in the wild-type background (a), while pAtXTH19::GUS is expressed in both the dividing and basal regions (i). In the axr2-1 background, pAtXTH19::GUS expression in the basal regions was reduced (j, l, m, n, o). Expression of the pAtXTH18::GUS and pAtXTH19::GUS fusion genes was not affected in the ein2-1 background (c, k). Scale bar 1 mm
Comparison of GUS activity in the pAtXTH18::GUS (a) and pAtXTH19::GUS fusion genes (b, c) in 7-day-old seedlings of wild-type, axr2-1, and ein2-1 backgrounds. a and b Wild-type (WT), axr2-1(axr2), and ein2-1 (ein2) seedlings were grown on vertically oriented solid MS medium for 7 days and subjected to GUS activity measurements. c WT and axr2-1 seedlings were grown on vertically oriented solid MS medium for 5 days, and then transferred to new MS plates containing either 0.1 nM IAA (+IAA) or no IAA (−IAA). Following this, the seedlings were allowed to grow for an additional 2 days and their GUS activity measured. GUS activities relative to wild-type (a, b) and wild-type without IAA treatment (WT–IAA) (c) were calculated and shown, with standard deviations as vertical lines (n=3)
On the other hand, pAtXTH19::GUS was expressed throughout both the main and lateral roots, with intensive expression being seen at the apical region (Fig. 6i, n). An auxin responsive element, TGTCTC, is found in the −956 to −951-bp region upstream of the AtXTH19 gene. Deletion of the upstream region to −330 bp of the promoter caused loss of GUS activity in the apical dividing region (Fig. 6l). This is consistent with our previous results (Vissenberg et al. 2005a).
In the axr2-1 mutant plants, expression of the pAtXTH19::GUS gene was only observed in the apical region of the main and lateral roots (Fig. 6j, o). Its expression was drastically reduced in the differentiating regions. GUS activity in the axr2-1 mutant was also reduced to 70% of that in wild-type plants (Fig. 7b). In the ein2-1 mutant background, neither expression level nor profile of the pAtXTH19::GUS was affected (Figs. 6k, p, 7b). Interestingly, the expression of −330pAtXTH19::GUS, which lacks the TGTCTC sequence, was also completely suppressed in the elongation and differentiation zones of the root. This suggests that transcriptional activity of the −330pAtXTH19::GUS gene in wild type is still under regulation by auxin signaling via the AXR2/IAA7. It is worthy of note that TGTCAC sequence is found in the −82 to −67-bp region upstream of the AtXTH19 gene, a sequence that is supposed to be another potential auxin-responsive element (Okushima et al. 2005).
Finally, we examined the effects of exogenous auxin on pAtXTH19::GUS expression (Fig. 7c). Whereas its GUS expression was significantly elevated in wild-type plants, little or no enhancement of GUS activity was observed in the axr2-1 plants.
Discussion
An essential role for AtXTH18 in root elongation
Despite many studies aimed at elucidating physiological roles of the XTH family of proteins in cell expansion, no particular gene or protein has been identified as being responsible for a specific cell expansion process in plants. Very recently, we showed that loss-of-function mutants for an A. thaliana XTH gene, AtXTH27, which belongs to the XTH class III subfamily and is predicted to encode an enzyme capable of mediating the hydrolysis of xyloglucan molecules in the cell wall, exhibited aberrant tracheary elements in rosette leaves. This provides the first evidence showing an essential role of a specific member of the XTH family in cell differentiation in a specific cell type (Matsui et al. 2005).
Unlike AtXTH27, AtXTH18 belongs to the XTH class II subfamily, each member of this subfamily being considered to encode a transferase that exclusively mediates molecular grafting between xyloglucan molecules (Nishitani and Tominaga 1992; Rose et al. 2002; Yokoyama and Nishitani 2001). In order to gain insight into role of this member of XTH family of genes, we employed native promoter for the XTH18 RNAi construct because the XTH family of proteins consist of many members with similar primary structures and because we aimed at elucidating the role of a particular member of this family but not its ectopic effects in plants. The results indicate that a slight decrease in AtXTH18 mRNA abundance by RNAi resulted in a small but significant reduction in the epidermal cell length of the primary root, providing the first evidence demonstrating that a particular member of the XTH class II subfamily is responsible for cell elongation. These results are also consistent with our hypothesis that individual XTH genes have specific physiological roles for cell-wall dynamics in a specific site of plants (Nishitani 2002; Vissenberg et al. 2005a).
Of the four XTH genes, AtXTH17, 18, 19, and 20, examined in the present study, AtXTH18 exhibited prominent expression levels in terms of mRNA abundance as estimated by real-time RT-PCR (Fig. 3). In particular, this was in the elongation and differentiation regions of the primary root (Fig. 6). Data also indicate that reductions in the AtXTH18 mRNA level did not alter the expression levels of other XTHs, particularly AtXTH17, which is coexpressed in the same tissue in the roots and exhibits similar enzymatic activity (cf. Vissenberg et al. 2005a). Therefore, it is unlikely that the transcriptions of closely related XTH genes are regulated interactively. Rather, their transcription processes are individually regulated.
Furthermore, in all the independent homozygous RNAi lines characterized in the present study, the reduction rate of the AtXTH18 mRNA level was 14% compared with wild-type plants. No RNAi line with a higher reduction rate of the AtXTH18 mRNA was isolated in the present study (data not shown). This result implies that unrestrained expression of AtXTH18 mRNA is essential for this plant.
Inversely, two T-DNA insertion lines for AtXTH17, in which reduction rates of the AtXTH17 mRNA level were more than 80%, did not show any prominent phenotypic traits in the root. This consideration leads to the conclusion that a high level of AtXTH18 expression is essential and cannot be compensated for by the presence of coexpressing lower levels of isozymes, such as AtXTH17 and AtXTH19, in the same tissues. Thus, AtXTH18 protein plays a critical role in mediating the construction and/or remodeling of the xyloglucan/cellulose framework required for cell elongation and the maturation of root cells (Vissenberg et al. 2005a). These results also suggest the possibility that the extent of cell elongation can be precisely regulated via the transcription levels of the AtXTH18 gene. This conclusion is consistent with the findings of Vissenberg et al. (2000, 2001, 2003) that XTH exhibits the most prominent xyloglucan endotransglycosylase (XET) action in epidermal cells in the elongation region and in the trichoblasts in the differentiation zone of the primary root.
Whereas the mechanism by which XTH causes cell-wall loosening has not been fully demonstrated, several studies using fluorescent xyloglucan probes have revealed that XTH actually modifies xyloglucans anchored to cellulose microfibrils (Ito and Nishitani 1999; Vissenberg et al. 2005b). The present study provides further evidence for the scheme that molecular grafting of xyloglucans, which is considered to be mediated by classes I and II XTH family members, is essential for cell-wall loosening and hence cell extension (Nishitani 1997).
Auxin regulation of XTH expression
To gain insight into role of auxin signaling in the XTH gene expression in root, we examined expressions of AtXTH18 and AtXTH19 promoter::GUS constructs in axr2-1 mutant, a gain-of-function mutant of A. thaliana. This mutation is caused by a single amino acid change in domain II of the AXR2/IAA7 protein (Nagpal et al. 2000). Domain II of this protein is shown to be necessary and sufficient for ubiquitin ligase SCFTIR1-mediated degradation (Gray et al. 2001). Although specific auxin-responsive factor (ARF) protein(s) inactivated by interaction with the AXR2/IAA has not been identified, auxin signal perceived by the TIR1 protein (Dharmasiri et al. 2005; Kepinski and Leyser 2005) is considered to affect transcription activity via auxin-responsive element (AuxRE) (reviewed by Dharmasiri and Estelle 2004).
A potential AuxRE, TGTCTC (Ulmasov et al. 1997), is found −956 to −951 bp upstream of the AtXTH19 gene. Whereas the expression of pAtXTH19::GUS was observed in both the apical dividing region and basal elongating/maturing regions in the wild type, under axr2-1 background, the expression was maintained only in the apical region, indicating that regulation of AtXTH19 expression in elongating/maturing regions of the root is mediated by auxin signaling via AXR2/IAA7. It is reported that the AuxRE element is also found in the 5′ upstream region of TCH4 (AtXTH22) (Iliev et al. 2002), another member of the class II XTH gene family, which was shown to be up-regulated by both auxin and brassinolide application in A. thaliana (Goda et al. 2004; Xu et al. 1995, 1996).
Both −329pAtXTH18::GUS and −330pAtXTH19::GUS were expressed in the elongating and maturing regions of the primary root in wild-type plants. Under the axr2-1 mutant background, although −329pAtXTH18::GUS was still expressed as in the wild-type background (Fig. 6d, e), expression of -330pAtXTH19::GUS was no longer observed (Fig. 6l, m). This result indicates that expression of −330pAtXTH19::GUS is also under regulation by auxin signaling via AXR2/IAA7 in the elongating and maturating regions of wild-type plant roots. In this context, it is worthy of note that TGTCAC sequence is found in −82 to −67 bp upstream of the 330pAtXTH19::GUS gene. In addition to the TGTCTC sequence, to which the ARF1 transcription factor is shown to bind (Ulmasov et al. 1997), TGTCnC and GnGACA sequences are also supposed to be potential AuxREs (Okushima et al. 2005). Thus, it may be possible that the TGTCAC sequence in the -330pAtXTH19::GUS construct might be involved in the auxin signaling pathway.
The axr2-1 is one of the pleiotropic auxin-resistant mutants that confers resistance to not only auxin but to abscisic acid and ethylene (Wilson et al. 1990). Thus, altered expression of pAtXTH19:: GUS genes in the axr2-1 mutant plants might be due to indirect action derived from other hormonal signals. The present result that pAtXTH19::GUS is not affected in the ein2-1 mutant plant indicates that, at least, EIN2-mediated ethylene signaling is not involved in the regulation of the AtXTH19 gene in axr2-1 mutant plants.
Given an essential role for auxin in the regulation of root-cell expansion, and a principal role for AtXTH18 protein in modifying cell-wall architecture during root cell elongation, a question arises as to whether AtXTH18 gene expression is regulated via auxin signaling pathways or not. Our previous study showed that auxin application caused a slight elevation in AtXTH18 mRNA level as quantified by real-time RT-PCR (Vissenberg et al. 2005a). Although no TGTCTC element is found in the 5′ region of the AtXTH18 gene, GTGACA and TGTCGC sequences are found in −1,501 to −1,496-bp and −70 to −65-bp regions, respectively, of the gene. Thus, it is possible that these putative AuxREs might be involved in the responsiveness of the AtXTH18 gene to exogenous auxin. The pAtXTH18::pro GUS construct used in the present study, however, does not contain the −1,501−1,496 GTGACA sequence, and at present, it is not known whether this element is actually responsive to auxin or not.
Alternatively, our previous result showed that gibberellic acid enhanced the AtXTH18 mRNA level (Yokoyama and Nishitani 2001). Fu and Harberd (2003) elegantly showed that auxin controls the growth of roots by modulating the gibberellin-mediated degradation of DELLA proteins. Therefore, it is probable that the expression of AtXTH18 in root tissue is regulated directly or indirectly via the gibberellin signaling system, which is regulated by auxin. Considering the essential role of AtXTH18 in root-cell elongation, molecular dissection of its transcriptional regulation might offer opportunities for exploring the still unknown signaling pathway downstream of the DELLA proteins and the enzyme actions responsible for cell-wall modifications.
In either event, AtXTH18 and 19 genes are functionally different in terms of responses to hormonal signaling and spatial expression profile. This is despite the fact that they share a highly conserved sequence located between −250 and −150 bp in individual XTH genes (Vissenberg et al. 2005a). Molecular dissection of this potential cis region needs further investigation for full characterization of the auxin-mediated regulation of the two XTH genes in root-cell elongation.
Conclusions
To our knowledge, the present study provides the first evidence demonstrating that a specific member of the XTH protein family plays an indispensable role in the cell elongation process. This study also provides an important clue as to the regulatory mechanism by which auxin and gibberellin signaling regulates downstream molecules in charge of cell-wall modification, which is critical in controlling cell expansion and hence plant growth and morphogenesis. Putative cis-acting regions disclosed in the present study will offer opportunities for exploring components of signaling pathways or networks through which environmental as well as developmental cues will be integrated and transmitted to downstream molecules responsible for cell-wall modifications.
References
Abel S, Theologis A (1996) Early genes and auxin action. Plant Physiol 111:9–17
Abel S, Nguyen MD, Chow W, Theologis A (1995) ACS4, a primary indoleacetic acid-responsive gene encoding 1-aminocyclopropane-1-carboxylate synthase in Arabidopsis thaliana. J Biol Chem 270:19093–19099. DOI 10.1074/JBC.270.32.19093
Akamatsu T, Hanzawa Y, Ohtake Y, Takahashi T, Nishitani K, Komeda Y (1999) Expression of endoxyloglucan transferase genes in acaulis mutants of Arabidopsis. Plant Physiol 121:715–722
Alonso JM, Hirayama T, Roman G, Nourizadeh S, Ecker JR (1999) EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 284:2148–2152. DOI 10.1126/science.284.5423.2148
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. DOI 10.1016/0003-2697(76)90527-3
Carpita NC, Gibeaut DM (1993) Structural models of primary cell walls in flowering plants: consistency of molecular structure with the physical properties of the walls during growth. Plant J 3:1–30. DOI 10.1046/j.1365-313X.1993.00999.x
Catala C, Rose JKC, York WS, Albersheim P, Darvill AG, Bennett AB (2001) Characterization of a tomato xyloglucan endotransglycosylase gene that is downregulated by auxin in etiolated hypocotyls. Plant Physiol 127:1180–1192
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. DOI 10.1046/j.1365-313x.1998.00343.x
Cosgrove DJ (1997a) Assembly and enlargement of the primary cell wall in plants. Ann Rev Cell Devel Biol 13:171–201
Cosgrove DJ (1997b) Relaxation in a high-stress environment—the molecular basis of extensible cell walls and cell enlargement. Plant Cell 9:1031–1041
Dharmasiri N, Dharmasiri S, Estelle M (2005) The F-box protein TIR1 is an auxin receptor. Nature 435:441–445. DOI 10.1038/nature03543
Dharmasiri N, Estelle M (2004) Auxin signaling and regulated protein degradation (2004). Trend Plant Sci 9:302–308
Kepinski S, Leyser O (2005) The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435:446–451. DOI 10.1038/nature03542
Fry SC (2004) Primary cell wall metabolism: tracking the careers of wall polymers in living plant cells. New Phytol 161:641–675. DOI 10.1111/j.1469-8137.2004.00980.x
Fry SC, Smith RC, Renwick KF, Martin DJ, Hodge SK, Matthews KJ (1992) Xyloglucan endotransglycosylase, a new wall-loosening enzyme activity from plants. Biochem J 282:821–828
Fu X, Harberd NP (2003) Auxin promotes Arabidopsis root growth by modulating gibberellin response. Nature 421:740–743
Gil P, Liu Y, Orbovic V, Verkamp E, Poff KL, Green PJ (1994) Characterization of the auxin-inducible SAUR-AC1 gene for use as a molecular genetic tool in Arabidopsis. Plant Physiol 104:777–784
Goda H, Sawa S, Asami T, Fujioka S, Shimada Y, Yoshida S (2004) Comprehensive comparison of auxin-regulated and brassinosteroid-regulated genes in Arabidopsis. Plant Physiol 134:1555–1573. DOI 10.1104/pp.103.034736
Gray WM, Kepinski S, Rouse D, Leyser O, Estelle M (2001) Auxin regulates SCFTIR1-dependent degradation of AUX/IAA proteins. Nature 414:271–276. DOI 10.1038/35104500
Guzman P, Ecker JR (1990) Exploiting the triple response of Arabidopsis to identify ethylene-related mutants. Plant Cell 2:513–523
Hyodo H, Yamakawa S, Takeda Y, Tsuduki M, Yokota A, Nishitani K, Kohchi T (2003) Active gene expression of a xyloglucan endotransglucosylase/hydrolase gene, XTH9, in inflorescence apices is related to cell elongation in Arabidopsis thaliana. Plant Mol Biol 52:473–482. DOI 10.1023/A:1023904217641
Iliev EA, Xu W, Polisensky DH, Oh M-H, Torisky RS, Clouse SD, Braam J (2002) Transcriptional and posttranscriptional regulation of Arabidopsis TCH4 expression by diverse stimuli. Roles of cis regions and brassinosteroids. Plant Physiol 130:770–783. DOI 10.1104/pp.008680
Imoto K, Yokoyama R, Nishitani K (2005) Comprehensive approach to genes involved in cell wall modifications in Arabidopsis thaliana. Plant Mol Biol 58:177–192
Ito H, Nishitani K (1999) Visualization of EXGT-mediated molecular grafting activity by means of a fluorescent-labeled xyloglucan oligomer. Plant Physiol 40:1172–1176
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Matsui A, Yokoyama R, Seki M, Ito T, Shinozaki K, Takahashi T, Komeda Y, Nishitani K (2005) AtXTH27 plays an essential role in cell wall modification during the development of tracheary elements. Plant J 42:525–534. DOI 10.1111/j.1365-313X.2005.02395.x
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–496
Nagpal P, Walker LM, Young JC, Sonawala A, Timpte C, Estelle M, Reed JW (2000) AXR2 encode a member of the Aux/IAA protein family. Plant Physiol 123:563–573
Nakamura T, Yokoyama R, Tomita E, Nishitani K (2003) Two azuki bean XTH genes, VaXTH1 and VaXTH2, with similar tissue-specific expression profiles, are differently regulated by auxin. Plant Cell Physiol 44:16–24
Nishitani K (1997) The role of endoxyloglucan transferase in the organization of plant cell walls. Int Rev Cytol 173:157–206
Nishitani K (2002) A genome-based approach to study the mechanisms by which cell-wall type is defined and constructed by the collaborative actions of cell-wall-related enzymes. J Plant Res 115:303–307. DOI 10.1007/s10265-002-00032-z
Nishitani K, Masuda Y (1981) Auxin-induced changes in the cell wall structure: changes in the sugar compositions, intrinsic viscosity and molecular weight distribution of matrix polysaccharides of the epicotyl cell wall of Vigna angularis. Physiol Plant 52:482–494
Nishitani K, Tominaga R (1992) Endo-xyloglucan transferase, a novel class of glycosyltransferase that catalyzes transfer of a segment of xyloglucan molecule to another xyloglucan molecule. J Biol Chem 267:21058–21064
Okazawa K, Sato Y, Nakagawa T, Asada K, Kato I, Tomita E, Nishitani K (1993) Molecular cloning and cDNA sequencing of endoxyloglucan transferase, a novel class of glycosyltransferase that mediates molecular grafting between matrix polysaccharides in plant cell walls. J Biol Chem 268:25364–25368
Okushima Y, Overvoorde PJ, Arima K, Alonso JM, Chan A, Chang C, Ecker JR, Hughes B, Lui A, Nguyen D, Onodera C, Quach H, Smith A, Yu G, Theologis T (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17:444–463
Roman G, Lubarsky B, Kieber JJ, Rothenberg M, Ecker JR (1995) Genetic analysis of ethylene signal transduction in Arabidopsis thaliana: five novel mutant loci integrated into a stress response pathway. Genetics 139:1393–1409
Rose JKC, Braam J, Fry SC, Nishitani K (2002) The XTH family of enzymes involved in xyloglucan endotransglucosylation and endohydrolysis: current perspectives and a new unifying nomenclature. Plant Cell Physiol 43:1421–1435
Tsukaya H, Ohshima T, Naito S, Chino M, Komeda Y (1991) Sugar-dependent expression of the CHS-A gene for chalcone synthase from Petunia in transgenic Arabidopsis. Plant Physiol 97:1414–1421
Ulmasov T, Hagen G, Guilfoyle TJ (1997) ARF1, a transcription factor that binds to auxin response elements. Science 276:1865–1868
Vissenberg K, Martinez-Vilchez IM, Verbelen J-P, Miller JG, Fry SC (2000) In vivo co-localization of xyloglucan endotransglycosylase activity and its donor substrate in the elongation zone of Arabidopsis roots. Plant Cell 12:1229–1237
Vissenberg K, Fry SC, Verbelen JP (2001) Root hair initiation is coupled to a highly localized increase of xyloglucan endotransglycosylase action in Arabidopsis roots. Plant Physiol 127:1125–1135
Vissenberg K, Van Sandt V, Fry SC, Verbelen J-P (2003) Xyloglucan endotransglucosylase action is high in the root elongation zone and in the trichoblasts of all vascular plants from Selaginella to Zea mays. J Exp Bot 54:334–344. DOI 10.1093/jxb/erg024
Vissenberg K, Oyama M, Osato Y, Yokoyama R, Verbelen J-P, Nishitani K (2005a) Differential expression of AtXTH17, AtXTH18, AtXTH19 and AtXTH20 genes in Arabidopsis roots. Physiological roles in specification in cell wall construction. Plant Cell Physiol 46:192–200. DOI 10.1093/pcp/pci013
Vissenberg K, Fry SC, Pauly M, Höfte H, Verbelen JP (2005b) XTH acts at the microfibril-matrix interface during cell elongation. J Exp Bot 56:673–683. DOI 10.1093/jxb/eri048
Wilson AK, Pickett FB, Turner JC, Estelle M (1990) A dominant mutation in Arabidopsis confers resistance to auxin, ethylene and abscisic acid. Mol Gen Genet 222:377–383
Xu W, Purugganan MM, Polisensky DH, Antosiewicz DM, Fry SC, Braam J (1995) Arabidopsis TCH4, regulated by hormones and the environment, encodes a xyloglucan endotransglycosylase. Plant Cell 7:1555–1567
Xu W, Campbell P, Vargheese AK, Braam J (1996) The Arabidopsis XET-related gene family: environmental and hormonal regulation of expression. Plant J 9:879–889. DOI 10.1046/j.1365-313X.1996.9060879.x
Yokoyama R, Nishitani K (2000) Functional diversity of xyloglucan-related proteins and its implications in the cell wall dynamics in plants. Plant Biol 2:598–604
Yokoyama R, Nishitani K (2001) A comprehensive expression analysis of all members of a gene family encoding cell-wall enzymes allowed us to predict cis-regulatory regions involved in cell-wall construction in specific organs of Arabidopsis. Plant Cell Physiol 42:1025–1033
Yokoyama R, Nishitani K (2004) Genomic basis for cell-wall diversity in plants. A comparative approach to gene families in rice and Arabidopsis. Plant Cell Physiol 45:1111–1121. DOI 10.1093/pcp/pch151
Yokoyama R, Rose JKC, Nishitani K (2004) A surprising diversity and abundance of xyloglucan endotransglucosylase/hydrolases in rice. Classification and expression analysis. Plant Physiol 134:1088–1099. DOI 10.1104/pp.103.035261
Acknowledgements
This work was supported by a Grant-in-Aid for Scientific Research on Priority Areas (17027001) and Scientific Research (B) (17370012) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, and by the Program for “Development of Fundamental Technologies for Controlling the Process of Material Production of Plants” from the New Energy and Industrial Technology Development Organization, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Osato, Y., Yokoyama, R. & Nishitani, K. A principal role for AtXTH18 in Arabidopsis thaliana root growth: a functional analysis using RNAi plants. J Plant Res 119, 153–162 (2006). https://doi.org/10.1007/s10265-006-0262-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-006-0262-6