Abstract
The activity of indole-3-acetamide (IAM) hydrolase from rice cells was enriched ca. 628-fold by gel filtration and anion exchange column chromatography. The molecular masses of the IAM hydrolase estimated by gel filtration and sodium dodecyl sulfate polyacrylamide gel electrophoresis were approximately 50.5 kD and 50.0 kD, respectively. The enzyme exhibited maximum activity at pH 6.0–6.5. The enzyme was stable against heat treatments between 4 and 50°C and works optimally at 52°C. The activity remained constant at 4°C for at least 143 days. The purified enzyme fraction hydrolyzed indoleacetic acid ethyl ester (Et-IAA) in addition to IAM and its homologue, 1-naphthalene-acetamide, but not indole-3-acetonitrile. Km values of the enzyme were 0.96 mM and 0.55 mM for IAM and Et-IAA, respectively. Although the molecular mass of the enzyme was very similar to that of IAM hydrolase of Agrobacterium tumefaciens involved in tumor formation, the biochemical properties of the enzyme including its high Km value were considerably different from those of the A. tumefaciens enzyme. Based on these enzyme properties, we will discuss whether the amidohydrolase is involved in auxin biosynthesis in rice cells.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The auxin biosynthetic pathway via indole-3-acetamide (IAM) from l-tryptophan (IAM pathway) was first identified by Magie et al. (1963) in the plant-pathogenic bacterium, Pseudomonas savastanoi, which causes the production of tumorous outgrowths on olive and oleander plants. The pathway was unequivocally established by the cloning of the genes responsible for the biosynthesis of indole-3-acetic acid (IAA) via the IAM pathway (Comai and Kosuge 1980, 1982). The IAM pathway consists of biochemical reactions catalyzed by two enzymes, a tryptophan monooxygenase and an IAM hydrolase. The monooxygenase converts l-tryptophan to IAM, while the second enzyme, amidohydrolase, hydrolyzes IAM to produce IAA.
The same pathway is also known to be required for tumorous outgrowths induced by the infection of Agrobacterium tumefaciens (Follin et al. 1985; Inzé et al. 1984; Klee et al. 1984; Schröder et al. 1984; Thomashow et al. 1984, 1986; Van Onckelen et al. 1985, 1986) and A. rhizogenes (White et al. 1985; Offringa et al. 1986; Camilleri and Jouanin 1991).
Although Agrobacterium is prokaryotic, the genes, tms1 and tms2, that encode the two enzymes reside on T-DNA and have eukaryotic structures consisting of a CAAT box, TATAA box and a polyA signal; they are actually expressed in eukaryotic cells after the transfer of T-DNA from Agrobacterium to plant cells (Akiyoshi et al. 1983; Surico et al. 1985; Cardarelli et al. 1987).
The origin of such genes with eukaryotic structures in prokaryotes is of great interest. In general, these genes are considered to be of prokaryotic origin, since they include nucleotide sequences homologous to the Pribnow box and closely resemble the genes in P. savastanoi (Weiler and Schröder 1987; Zambryski et al. 1989). Alternatively, it can be suggested that Agrobacterium might have captured the genes from a plant genome during its evolution. Although the IAM pathway is not generally known in higher plants, there are reports demonstrating the presence of an endogenous intermediate (Isogai et al. 1967; Takahashi et al. 1975; Rausch et al. 1985; Saotome et al. 1993; Rajagopal et al. 1994) and the enzymatic activities that seem to be involved in the IAM pathway (Kawaguchi et al. 1991, 1993; Rajagopal et al. 1994). Therefore, there is a clear need to define the nature of the plant enzymes and to compare them with those of Agrobacterium when it comes to considering the latter possibility.
Through the survey of the enzymatic activities associated with the IAM pathway among the cultured cell lines of various plants, we found that rice calli have stable activity that converts IAM to IAA (Kawaguchi et al. 1991). Since it is possible to obtain a large amount of rice calli as an experimental material, we attempted to purify the enzyme. In this paper, we partially purified amidohydrolase from rice cells and characterized the properties of the enzyme to compare them with those of Agrobacterium. Based on the enzymatic properties and today’s genomic information, we will discuss the question of whether or not the rice enzyme operates in IAA biosynthesis.
Materials and methods
Plant materials
Rice (Oryza sativa L. cv. Nipponbare) seeds, pretreated at high temperature (48°C) for 8 days, were sterilized with 100% ethanol and 20% sodium hypochlorite and germinated aseptically on Murashige and Skoog (MS) basal medium. The sterilized seedlings were grown under continuous light for 10 days. The root segments were then transferred to MS medium containing 1 mg/l of 2,4-dichlorophenoxyacetic acid (2,4-D) and 0.1 mg/l of kinetin to induce callus tissues (Kawaguchi et al. 1991). The induced callus tissues were successively subcultured at 26°C under light on medium of the same composition as that use for the callus induction.
For the preparation of enzyme, the callus tissues cultured for 43–46 days were harvested, immediately frozen with liquid nitrogen and stored at −80°C until use.
Enzyme preparation
The frozen tissues of rice callus and an equal weight of sand were suspended in an equal volume of 0.1 M Tris-HCl buffer, pH 7.6, containing 10% sucrose, 10 mM 2-mercaptoethanol, 5 mM EDTA and 10 mM MgCl2 and homogenized with a mortar and a pestle after leaving the tissues to thaw. The homogenates were filtered through two layers of nylon cloth and centrifuged at 15,000 g for 15 min at 4°C. The supernatant was used as crude enzyme preparation. The crude enzyme was used for further purification by ammonium sulfate fractionation, gel filtration and ion exchange column chromatography.
Enzyme assays
During enzyme purification, amidohydrolase activity was assayed by adding IAM to enzyme solution to a final concentration of 5 mM. For the analysis of the biochemical properties of enzyme, amidohydrolase activity was assayed in 0.l ml of 50 mM Tris-HCl buffer (pH 7.6) with 5 mM IAM, 5 mM MgCl2, 5% (v/v) glycerol, 5 mM 2-mercaptoethanol, 20 mM KCl and a fixed amount of the enzyme in an Eppendorf tube. Citrate buffer (25 mM), 50 mM phosphate buffer or 25 mM Tris-HCl buffer was also used to analyze the effects of pH on the enzyme activity. After incubation for 0.5 h for enzyme purification or 1 h for other enzyme characterization at 37°C, the reaction was stopped by the addition of 0.1 ml of 0.1 M citric acid to make the pH 2–3. Then 0.4 ml ethyl acetate was added to the tube to extract the reaction product (IAA). The tube was vibrated by Vortex mixer at the maximum vibration rate for 10 min twice and centrifuged at 15,000 rpm for 1 min. The layer of ethyl acetate was transferred to a new Eppendorf tube and evaporated to dryness in vacuo. The residue was analyzed by high performance liquid chromatography (HPLC).
We define 1 U of enzyme activity as the activity that produces 1 nmole IAA in 0.5 h. Specific activity is units of activity per mg of protein.
Protein concentrations were determined according to the method of Lowry et al. (1951) except in the case of DEAE-Sepharose CL-6B fractionation where BCA protein assay reagent (PIEPCE) was used and of MiniQ PC3.2/3 fractionation where absorbance at 280 nm was used.
The HPLC system consisted of an HPLC model-576 pump unit (Gas Chromatography Industry) coupled to the UV detector (Spectro detector 502U; Gas Chromatography Industry) and the Chromato Pak CR-1B (Shimadzu, Kyoto).
The HPLC analysis was done under the following conditions: the column was HPLC-packed Nucleosil 100-5C18 (150×4.6 mm internal diameter) (Gas Chromato Industry); the mobile phase was 30% acetonitrile in 0.5% CH3COOH; and the flow rate was 1.5 ml/min. IAA was quantified by absorption at 280 nm.
Liquid chromatography-mass spectrometry
IAA was identified by liquid chromatography-mass spectrometry (LC-MS) using LC/APCI-MS M-1,000 (Hitachi). The conditions of analysis were as follows: multi 1.8 kV; focus voltage 120 V; filter 5; resolution 55; scan range 5–500/2 s; nebulization temperature 280°C; desorption temperature 339°C.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis
Mini-protean II (Bio-Rad) was used for a polyacrylamide gel electrophoresis (PAGE) system. Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) was carried out according to the method of Laemmli (1970). Proteins were visualized by silver stain kit (Wako Pure Chemical Industries). The calibration protein combithek (Boehringer Mannheim) was used as the standard marker of molecular weight.
Results
Identification of reaction product
Crude enzyme fraction was incubated with 20 mM IAM for 30 min at 37°C and the reaction product was separated by HPLC as described previously. One major peak was detected at the same retention time (4.6 min) as that of authentic IAA (data not shown).
LC-MS analysis revealed that the mass spectrum of the reaction product showed fragments at mass-to-charge ratios 130, 146 and 176 (M++1, base peak), characteristic of authentic IAA (data not shown).
IAM hydrolase activity during tissue culture
The time course of IAM hydrolase activity during tissue culture was surveyed to specify the culture period appropriate for harvest of callus tissues for use in purification of the enzyme and also to examine the physiological function of the enzyme in relation to the growth of calluses (Fig. 1).
The time courses of callus growth of Oryza sativa L. cv. Nipponbare and indole-3-acetamide (IAM) hydrolase activity. The rice calli were cultured on Murashige and Skoog agar medium supplemented with 1 mg/l 2,4-dichlorophenoxyacetic acid (2,4-D) and 0.1 mg/l kinetin at 25±1°C in the dark. Relative growth was calculated as (W t−W 0)/W 0, where W 0 is the initial weight and W t is the weight recorded on the day of assay. Open squares (□) relative growth, (W t−W 0)/W 0; solid circles (●) specific activity (U/ml); open circles (○) total protein (mg/g fresh weight of callus)
Fresh weight of callus tissues increased over a period of 30 days after transfer onto the new medium and thereafter decreased slightly. Protein content increased rapidly over 5–10 days and reached a maximum at 15 days after transfer. Thereafter the content gradually decreased up to a rapid decrease after 30 days of culture. The protein content related to the growth of callus fairly well.
Specific activity of IAM hydrolase was relatively constant during the culture period of 46 days, except at 30 days after the transfer where the specific activity decreased temporarily (Fig. 1). We did not obtain any reproducibility of this transient decrease in enzymatic activity, but in order to prepare a large collection of material, we used callus tissues cultivated for 43–46 days for enzyme purification.
Partial purification of IAM hydrolase
Ammonium sulfate fractionation
The crude enzyme preparation was saturated with solid ammonium sulfate to give a saturation level of 20% and left under agitation by magnetic stirrer for 20 min. After centrifugation (10,000 g) for 15 min at 4°C to remove the precipitates, the resultant supernatant was again saturated with ammonium sulfate to a level of 60%. After agitation for 20 min, the precipitated proteins were collected by centrifugation as described above. The activity was exclusively detected at this saturation level (20–60%). The precipitates were suspended in 10 mM Tris-HCl buffer (pH 7.6) containing 10% (v/v) glycerol and 1 mM dithiothreitol (DTT) (buffer A) and passed through a Sephadex G-25 column (1.5×50 cm). The active fraction desalted with Sephadex G-25 was named the ammonium sulfate fraction. Through this procedure, proteins in the crude enzyme fraction were concentrated to a small volume, but no increase of specific activity was observed.
Gel filtration
The ammonium sulfate fraction was further purified using a Sephacryl S-200HR gel filtration column (3×89 cm) with buffer A as the eluent. Each 4.5 ml fraction was collected and assayed for IAM hydrolase activity. We found that fraction numbers 74–77 yielded 81.4% of the total activity of the ammonium sulfate fraction (Fig. 2a).
Partial purification of amidohydrolase. a Gel filtration of the Sephadex G-25 fraction with Sephacryl S-200HR (3×89 cm). The column was equilibrated and eluted with 10 mM Tris-HCl buffer (pH 7.6). Fraction nos. 74–77 were collected. Open circles (○) absorbance at 280 nm; solid circles (●) IAM hydrolase activity (U/ml). b Elution profile of IAM hydrolase from DEAE-Sepharose CL-6B column chromatography. The active fraction from Sephacryl S-200HR gel filtration was put on a column (3×15 cm) of DEAE-Sepharose CL-6B equilibrated with 20 mM Tris-HCl buffer (pH 7.6). Elution was by a linear gradient of 0–500 mM NaCl in the equilibration buffer. Fraction nos. 77–79 were collected. Open circles (○) absorbance at 280 nm; solid circles (●) IAM hydrolase activity (U/ml); line NaCl content. c Anion exchange chromatography of the active fractions from the DEAE-Sepharose CL-6B column on a SMART-HPLC system on a MiniQ PC3.2/3 column (3.2×30 mm), eluted with a linear NaCl gradient in 10 mM Tris-HCl buffer (pH 7.5). Fraction no. 27 was collected. Open circles (○) absorbance at 280 nm; solid circles (●) IAM hydrolase activity (U/ml); line concentration of NaCl; arrows start or stop points of fraction collection
DEAE-Sepharose CL-6B chromatography
The Sephacryl S-200 HR fraction was dialyzed overnight against 20 mM Tris-HCl buffer (pH 7.0) containing 10% glycerin and 1 mM DTT (buffer B). The dialyzed fraction was loaded onto a DEAE-Sepharose CL-6B anion exchange column and the column was washed first with 100 ml buffer B. The bound proteins were then eluted at a flow rate 25.7 ml/h with a linear NaCl gradient (0–500 mM) in 500 ml of the same buffer. Each 6 ml fraction was collected and assayed for IAM hydrolase activity. In fractions 77–79, 87.7% of the total activity of the Sephacryl S-200 HR fraction was obtained (Fig. 2b). These fractions were combined and named the DEAE-Sepharose CL-6B fraction. A part of this fraction was used for the characterization of IAA hydrolase.
MiniQ PC3.2/3 anion exchange chromatography
The DEAE-Sepharose CL-6B fraction was dialyzed against 10 mM Tris-HCl buffer (pH 7.5) containing 5% (v/v) glycerol and 1 mM DTT (buffer C) overnight. The dialyzed sample was applied to a MiniQ PC3.2/3 column and eluted at a flow rate of 80 μl/min with buffer C (from 0 to 8.2 min) and followed with a linear NaCl gradient (0–304 mM between 8.2 and 13.6 min, 304–386 mM between 13.6 and 20.0 min and 386–500 mM between 20.2 and 23.2 min) in the same buffer. In fraction 27 at 12 min, 67.3% of total activity of the DEAE-Sepharose CL-6B fraction was obtained (Fig. 2c). This fraction was named the MiniQ PC3.2/3 fraction.
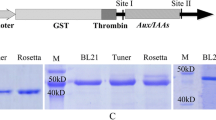
SDS-PAGE after each purification step is shown in Fig. 3. The one major band was detected by silver stain between 39.2-kD and 55.6-kD molecular markers. Through this four-step-protocol, the amidohydrolase activity was enriched ca. 628-fold compared to the specific activity of the crude extract (Table 1). Final yield through the four-step purification is 1.6% of the crude extract. Due to the limited size of the MiniQ PC3.2/3 fraction, the DEAE-sepharose CL-6B fraction was used for the characterization of IAM hydrolase.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) of amidohydrolase after each purification step, showing silver-stained gel. Lane 2 crude enzyme column; lane 3 (NH4)2SO4 fraction; lane 4 after Sephacryl S-200HR column; lane 5 after DEAE-Sepharose CL-6B column; lane 6 after MiniQ PC 3.2/3 column; lane 7 standard protein
Molecular mass estimation
For the estimation of M r for the IAM hydrolase from rice cells, a Sephacryl S-200 gel filtration column [2×83 cm (Pharmacia LKB Biotechnology)] was equilibrated with buffer A supplemented with 0.1 M NaCl and the DEAE-Sepharose CL-6B fraction was applied to the column. The column was eluted with the same buffer described above at a flow rate of 15 ml/h. The M r of the enzyme was estimated to be 50.5 kD by measuring the active fraction(s) (Fig. 4a). The following proteins were used as M r standards: bovine serum albumin (67,000), ovalbumin (43,000), chymotrypsinogen A (25,000) and ribonuclease A (13,700).
Molecular weight estimation of amidohydrolase in rice. a Estimation of molecular mass of IAM hydrolase by gel filtration on a Sephacryl S-200HR column. Elution was with 10 mM Tris-HCl buffer at pH 7.6. The elution volume of the IAM hydrolase was 127.4 ml and corresponds to a molecular mass of 50.5 kD. b Estimation of molecular mass of IAM hydrolase by SDS-PAGE. The R f value of the IAM hydrolase was 0.30, which corresponds to a molecular mass of 50.0 kD
As shown in Fig. 4b, the M r was also estimated to be 50 kD by SDS-PAGE. For calibration proteins, β-fructose-6-phosphate-kinase (85,200), glutamate dehydrogenase (55,600), aldolase (39,200), and triosephosphate isomerase (26,600) were used.
The pH optimum
Under the assay conditions described in “Materials and Methods”, IAM hydrolase has a pH optimum at 6.0-6.6. At pH 4.7, the activity of the enzyme sharply decreased to 8.6% of that at the optimum pH (Fig. 5a).
The pH optimum and temperature effects. a pH-dependence of the activity of IAM hydrolase. Open circles (○) 25 mM citrate, pH 4.7–5.8; open squares (□) 50 mM phosphate, pH 6.0–7.6; open triangles (△) 25 mM Tris-HCl buffer, pH 7.6–8.85. b Effects of temperature on IAM hydrolase activity and stability. Solid circles (●) temperature sensitivity curve; open circles (○) temperature stability curve. Temperature stability of the enzyme was calculated by measuring the remaining activity of IAM hydrolase after incubation for 15 min at various temperatures
Effects of temperature
Effects of temperature on the activity and the stability of IAM hydrolase were investigated at various temperatures. As shown in Fig. 5b, the optimum temperature was 52°C at pH 6.6 and the activity fell off sharply on either side of this temperature. For the assay of stability, enzyme preparations were heated for 15 min at various temperatures. After cooling in ice for 5 min, the remaining activities were measured. Above 50°C, the activity decreased sharply and was completely lost at 65°C. When stored at 4°C, the enzyme was stable for 143 days (data not shown).
Substrate specificity
Besides IAM, this enzyme was capable of converting 1-naphthaleneacetamide to its corresponding acid, but at a reduced velocity (Table 2). The ethyl ester of IAA (Et-IAA) was also a good substrate for this amidohydrolase. The reaction product (IAA) was confirmed by LC-MS (Fig. 6). The amount of IAA formed from the ester was ca. 57% of that from IAM. The preparation that had been further purified (the MiniQ PC3.2/3 fraction) also showed this activity (data not shown).
The enzyme did not convert indole-3-acetonitrile (IAN) or the amino acid conjugates such as indole acetylaspartic acid at all (Table 2).
Km value
The rate of enzyme reaction was measured in the presence of various concentrations of substrates, IAM and Et-IAA. The maximum velocity of enzyme reaction was 339.2 and 372.5 nmole/mg protein h-1 for IAM and Et-IAA, respectively. The rate was reduced slightly by substrate concentrations higher than those that gave the maximum velocity for both substrates (Fig. 7).
From the Lineweaver-Burk plot, the Km values for IAM and Et-IAA were estimated to be 0.96 and 0.55 mM, respectively (Fig. 7).
Discussion
The molecular weight (M r) (ca. 50 kD) of the hydrolase from rice cells estimated by both SDS-PAGE and gel filtration was very similar to that from Agrobacterium tumefaciens (M r 49,000 D) (Schröder et al. 1984). The properties of the enzyme (Table 2) were, however, quite different from those of the enzymes from A. tumefaciens (Kemper et al. 1985). For example the amidohydrolase from rice is stable under low temperature whereas the enzyme purified from crown gall is unstable even at 4°C. The stability under low temperature and optimal pH of rice enzyme rather resemble amidohydrolase purified from etiolated squash seedlings (Rajagopal et al. 1994). However they are different in terms of substrate specificity, namely squash amidohydrolase has an ability to hydrolyze IAN whereas rice enzyme has no such ability (Table 2).
Km value of the rice enzyme (0.96 mM) was considerably higher for IAM than that of crown gall (1.2 μM) (Kemper et al. 1985). In order for the rice amidohydrolase to serve in auxin biosynthesis, high concentrations of substrate must be present endogenously. However no endogenous IAM was detected from various organs such as shoots, roots, calli and young fruits of rice (data not shown). Therefore it is unlikely that the rice enzyme is involved in auxin biosynthesis via the IAM pathway.
It is interesting to note that the fraction purified ca. 628-fold can hydrolyze not only IAM but also Et-IAA. This observation suggests that the one enzyme has dual functions as amidase and esterase. Although there are no data demonstrating the presence of Et-IAA in rice, it has been reported that rice ears contain a large amount of esterified IAA at the stage of anthesis (Kobayashi et al. 1989). Therefore the rice amide hydrolase purified here may serve to control IAA accumulation via hydrolysis of the endogenous esterified IAA. Further purification of the enzyme and the examination of substrate specificity will be required to address this possibility.
The activity of IAM hydrolase is prominent in wild and cultivated rice. The activity is also found in various organs of rice plants (Kawaguchi et al. 1991). Therefore the enzyme may have a basic function specific for the physiology of rice. BLAST search for homologues of IAM hydrolase from A. tumefaciens indicated that there are two genes showing similarity in the rice genome (BAC clones on chromosome 4 and 12). Regions showing similarity with IAM hydrolase of A. tumefaciens are identical to conserved regions found between A. tumefaciens, Pseudomonas savastanoi, and Bradyrhizobium japonicum (Yamada et al. 1985; Sekine et al. 1989). RNA-induced gene silencing strategies are likely to be effective for investigating the function of these genes in rice.
References
Akiyoshi DE, Morris RO, Hinz R, Mischke BS, Kosuge T, Garfinkel DJ, Nester EW (1983) Cytokinin/auxin balance in crown gall tumors is regulated by specific loci in the T-DNA. Proc Natl Acad Sci USA 80:407–411
Camilleri C, Jouanin L (1991) The TR-DNA region carrying the auxin synthesis genes of the Agrobacterium rhizogenes agropine-type plasmid pRiA4: nucleotide sequence analysis and induction into tobacco plants. Mol Plant Microbe Interact 4:155–162
Cardarelli M, Spano L, Mariotti D, Mauro ML, Van Sluys MA, Costantino P (1987) The role of auxin in hairy root induction. Mol Gen Genet 208:457–46
Comai L, Kosuge T (1980) Involvement of plasmid deoxyribonucleic acid in indoleacetic acid synthesis in Pseudomonas savastanoi. J Bacteriol 143:950–957
Comai L, Kosuge T (1982) Cloning and characterization of iaaM, a virulence determinant of Pseudomonas savastanoi. J Bacteriol 149:40–46
Follin A, Inzé D, Budar F, Genetero C, Van Montagu M, Schell J (1985) Genetic evidence that the tryptophan 2-monooxygenase gene of Pseudomonas savastanoi is functionally equivalent to one of the T-DNA genes involved in plant tumor formation by Agrobacterium tumefaciens. Mol Gen Genet 201:178–185
Inzé D, Follin A, Van Lijsebettens M, Simons C, Genetello C, Van Montagu M, Schell J (1984) Genetic analysis of the individual T-DNA genes of Agrobacterium tumefaciens: further evidence that two genes are involved in indole-3-acetic acid synthesis. Mol Gen Genet 194:265–274
Isogai Y, Okamoto T, Koizumi T (1967) Studies on plant growth regulators. I. Isolation of indole-3-acetamide, 2-phenylacetamide, and indole-3-carboxyaldehyde from etiolated seedlings of Phaseolus. Chem Pharm Bull 15:151–158
Kawaguchi M, Kobayashi M, Sakurai A, Syono K (1991) The presence of an enzyme that converts indole-3-acetamide into IAA in wild and cultivated rice. Plant Cell Physiol 32:143–149
Kawaguchi M, Fujioka S, Sakurai A, Yamaki YT, Syono K (1993) Presence of a pathway for the biosynthesis of auxin via indole-3-acetamide in trifoliata orange. Plant Cell Physiol 34:121–128
Kemper E, Waffenschmidt S, Weiler EW, Rausch T, Schröder J (1985) T-DNA-encoded auxin formation in crown gall cells. Planta 163:257–262
Klee H, Montana A, Hododyski F, Lichtenstein CD, Garfinkel D, Fuller S, Flores C, Peshon J, Nester EW, Gordon MP (1984) Nucleotide sequence of the tms genes of the pTiA6NC octopine Ti plasmid: two gene products involved in plant tumorigenesis. Proc Natl Acad Sci USA 81:1728–1732
Kobayashi M, Sakurai A, Saka H, Takahashi N (1989) Fluctuation of the endogenous IAA level in rice during its life cycle. Agric Biol Chem 53:1089–1094
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 277:680–685
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265
Magie AR, Wilson EE, Kosuge T (1963) Indole-acetoamide as an intermediate in the synthesis of indoleacetic acid in Pseudomonas savastanoi. Science 141:1281–1282
Offringa IA, Melchers LS, Regensburg-Tunink AJG, Costantino P, Schilperoort RA, Hooykaas PJJ (1986) Complementation of Agrobacterium tumefaciens tumor-inducing aux mutants by genes from the TR-region of the Ri plasmid of Agrobacterium rhizogenes. Proc Natl Acad Sci USA 83:6935–6939
Rajagopal R, Tsurusaki K, Kannangara G, Kuraishi S, Sakurai N (1994) Natural occurrence of indoleacetamide and amidohydrolase activity in etiolated aseptically-grown squash seedlings. Plant Cell Physiol 35:329–339
Rausch T, Minocha SC, Hilgenberg W, Kahl G (1985) l-Tryptophan metabolism in wound-activated and Agrobacterium tumefaciens transformed potato tuber cells. Physiol Plant 63:335–344
Saotome M, Shirahata K, Nishimura R, Yahaba M, Kawaguchi M, Syono K, Kitsuwa T, Ishii Y, Nakamura T (1993) The identification of indole-3-acetic acid and indole-3-acetamide in the hypocothyls of Japanese cherry. Plant Cell Physiol 34:157–159
Schröder G, Waffenschmidt S, Weiler EW, Schröder J (1984) The T-region of Ti plasmids codes for an enzyme synthesizing indole-3-acetic acid. Eur J Biochem 138:387–391
Sekine M, Watanabe K, Syono K (1989) Nucleotide sequence of a gene for indole-3-acetamide hydrolase from Bradyrhizobium japonicum. Nucleic Acids Res 17:6400.
Surico G, Iacobellis NS, Sisto A (1985) Studies on the role of indole-3-acetic acid and cytokinins in the formation of knot on olive and oleander plants by Pseudomonas syringae pv. savastanoi. Physiol Plant Pathol 26:309–320
Takahashi N, Yamaguchi I, Kono T, Igoshi M, Hirose K, Suzuki K (1975) Characterization of plant growth substances in Citrus unshiu and their change in fruit development. Plant Cell Physiol 16:1101–1111
Thomashow LS, Reeves S, Thomashow MF (1984) Crown gall oncogenesis: evidence that a T-DNA gene from the Agrobacterium Ti plasmid pTiA6 encodes an enzyme that catalyze synthesis of indoleacetic acid. Proc Natl Acad Sci USA 81:5071–5075
Thomashow MF, Hugly S, Buchholz WG, Thomashow LS (1986) Molecular basis for the auxin-dependent phenotype of crown gall tumor tissue. Science 231:616–618
Van Onckelen HA, Rudelsheim P, Inzé D, Follin A, Messens E, Horemans S, Schell J, Van Montagu M, De Greef J (1985) Tobacco plants transformed with the Agrobacterium T-DNA gene 1 contain high amounts of indole-3-acetamide. FEBS Lett 181:373–376
Van Onkelen HA, Prinsen P, Inzé D, Rudelsheim P, Van Lijsebettens M, Schell J, Van Montagu M, De Greef J (1986) Agrobacterium T-DNA gene 1 codes for tryptophan-2-monooxygenase activity in tobacco crown gall cells. FEBS Lett 198:357–360
Weiler EW, Schröder J (1987) Hormone genes and crown gall disease. Trends Biochem Sci 12:271–275
White FF, Taylor BH, Huffman GA, Gordon MP, Nester EW (1985) Molecular and genetic analysis of the transferred DNA regions of the root-inducing plasmid of Agrobacterium rhizogenes. J Bacteriol 164:33–44
Yamada T, Palm CJ, Brooks B, Kosuge T (1985) Nucleotide sequence of the Pseudomonas savastanoi indoleacetic acid genes show homology with Agrobacterium tumefaciens T-DNA. Proc Natl Acad Sci USA 82:6522–6526
Zambryski P, Tempe J, Schell J (1989) Transfer and function of T-DNA genes from Agrobacterium Ti and Ri plasmids in plants. Cell 56:193–201
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Arai, Y., Kawaguchi, M., Syono, K. et al. Partial purification of an enzyme hydrolyzing indole-3-acetamide from rice cells. J Plant Res 117, 191–198 (2004). https://doi.org/10.1007/s10265-004-0146-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-004-0146-6