Abstract
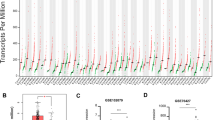
Hepatocellular carcinoma (HCC) remains an incurable malignancy despite the treatment methods being continually updated. Matrix metalloproteinases (MMPs) promote the progression of HCC; however, no consensus exists on which MMP plays the predominant role in HCCs. In the present study, we analyzed differentially expressed genes in HCCs, especially MMPs, compared with adjacent tissues using the Cancer Genome Atlas database. The KEGG enrichment pathway using differentially expressed genes included extracellular matrix–receptor interaction, which was correlated with MMPs. We found that among the MMP family, only MMP1, MMP3, MMP8, MMP9, MMP11, MMP12, MMP14, MMP15, MMP20, MMP21, and MMP24 significantly increased in HCCs compared with adjacent tissues. Crucially, survival and univariate analyses indicated that only MMPs 1, 9, 12, and 14 predict poor overall survival; however, multivariate Cox analysis and a nomogram demonstrated that only MMP1 is a poor prognostic biomarker for HCCs. In addition, we observed significant enrichment of uncharacterized cells but decreased macrophages in HCC tissues. Consistent with decreased macrophages in HCCs, MMP1 was negatively associated with macrophages but positively correlated with uncharacterized cells, indicating that the main producer of MMP1 is uncharacterized cells. Furthermore, MMP1 expression was negatively correlated with immune responses of HCCs. Taken together, our findings indicated that MMP1 is a poor and predominant prognostic biomarker for patients with HCC and that anti-MMP1 may be a novel therapy that is worth studying in depth.









Similar content being viewed by others
Data availability statement
All data generated in the study are included in the present article and supplementary data.
Change history
31 January 2023
A Correction to this paper has been published: https://doi.org/10.1007/s10238-023-01001-8
References
Wang H, Naghavi M, Allen C, Barber RM, Carter A, Casey DC, et al. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. 2016;388:1459–544.
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021;71:7–33.
Bruix J, Chan SL, Galle PR, Rimassa L, Sangro B. Systemic treatment of hepatocellular carcinoma: An EASL position paper. J Hepatol [Internet]. 2021; 75:960–74. Available from: https://www.sciencedirect.com/science/article/pii/S0168827821019036.
Duarte S, Baber J, Fujii T, Coito AJ. Matrix metalloproteinases in liver injury, repair and fibrosis. Matrix Biol. 2015;44–46:147–56.
Cui N, Hu M, Khalil RA. Biochemical and biological attributes of matrix metalloproteinases. Prog Mol Biol Transl Sci. 2017;147:1–73.
Egeblad M, Werb Z. New functions for the matrix metalloproteinases in cancer progression. Nat Rev Cancer. 2002;2:161–74. https://doi.org/10.1038/nrc745.
Coussens LM, Werb Z. Matrix metalloproteinases and the development of cancer. Chem Biol. 1996;3:895–904.
Masson R, Lefebvre O, Noël A, Fahime ME, Chenard MP, Wendling C, et al. In vivo evidence that the stromelysin-3 metalloproteinase contributes in a paracrine manner to epithelial cell malignancy. J Cell Biol. 1998;140:1535–41.
Itoh T, Tanioka M, Yoshida H, Yoshioka T, Nishimoto H, Itohara S. Reduced angiogenesis and tumor progression in gelatinase A-deficient mice. Cancer Res. 1998;58:1048–51.
Itoh T, Tanioka M, Matsuda H, Nishimoto H, Yoshioka T, Suzuki R, et al. Experimental metastasis is suppressed in MMP-9-deficient mice. Clin Exp Metastasis. 1999;17:177–81.
Sternlicht MD, Lochter A, Sympson CJ, Huey B, Rougier JP, Gray JW, et al. The stromal proteinase MMP3/stromelysin-1 promotes mammary carcinogenesis. Cell. 1999;98:137–46.
Coussens LM, Tinkle CL, Hanahan D, Werb Z. MMP-9 supplied by bone marrow-derived cells contributes to skin carcinogenesis. Cell. 2000;103:481–90.
Ha HY, Moon HB, Nam MS, Lee JW, Ryoo ZY, Lee TH, et al. Overexpression of membrane-type matrix metalloproteinase-1 gene induces mammary gland abnormalities and adenocarcinoma in transgenic mice. Cancer Res. 2001;61:984–90.
Wang S-W, Tai H-C, Tang C-H, Lin L-W, Lin T-H, Chang A-C, et al. Melatonin impedes prostate cancer metastasis by suppressing MMP-13 expression. J Cell Physiol. 2021;236:3979–90.
Azevedo Martins JM, Rabelo-Santos SH, do Amaral Westin MC, Zeferino LC. Tumoral and stromal expression of MMP-2, MMP-9, MMP-14, TIMP-1, TIMP-2, and VEGF-A in cervical cancer patient survival: a competing risk analysis. BMC Cancer. 2020;20:660. https://doi.org/10.1186/s12885-020-07150-3.
Määttä M, Soini Y, Liakka A. Autio-Harmainen H. Differential Expression of Matrix Metalloproteinase (MMP)-2, MMP-9, and Membrane Type 1-MMP in Hepatocellular and Pancreatic Adenocarcinoma: Implications for Tumor Progression and Clinical Prognosis. Clin Cancer Res [Internet]. 2000; 6:2726 LP – 2734. Available from: http://clincancerres.aacrjournals.org/content/6/7/2726.abstract.
Yu J, Ma S, Tian S, Zhang M, Ding X, Liu Y, et al. Systematic construction and validation of a prognostic model for hepatocellular carcinoma based on immune-related genes. Front Cell Dev Biol. 2021;9:700553.
Jacob A, Prekeris R. The regulation of MMP targeting to invadopodia during cancer metastasis. Front Cell Dev Biol. 2015;3:4.
Sakamoto Y, Mafune K, Mori M, Shiraishi T, Imamura H, Mori M, et al. Overexpression of MMP-9 correlates with growth of small hepatocellular carcinoma. Int J Oncol. 2000;17:237–43.
Cao W, Fan W, Wang F, Zhang Y, Wu G, Shi X, et al. GM-CSF impairs erythropoiesis by disrupting erythroblastic island formation via macrophages. J Transl Med. 2022;20(1):1–7.
Wang Y, Li W, Schulz VP, Zhao H, Qu X, Qi Q, et al. Impairment of human terminal erythroid differentiation by histone deacetylase 5 deficiency. Blood Internet. 2021;1(138):1615–27. https://doi.org/10.1182/blood.2020007401/476012/Impairment-of-human-terminal-erythroid.
Li W, Wang Y, Zhao H, Zhang H, Xu Y, Wang S, et al. Identification and transcriptome analysis of erythroblastic island macrophages. Blood Am Soc Hematol. 2019;134:480–91.
Yu G, Wang LG, Han Y, He QY. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omi A J Integr Biol. 2012;10801(16):284–7.
Li B, Severson E, Pignon J-C, Zhao H, Li T, Novak J, et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 2016;17:174.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7.
Sturm G, Finotello F, Petitprez F, Zhang JD, Baumbach J, Fridman WH, et al. Comprehensive evaluation of transcriptome-based cell-type quantification methods for immuno-oncology. Bioinformatics. 2019;35:i436–45.
Zhou T, Cai Z, Ma N, Xie W, Gao C, Huang M, et al. A novel ten-gene signature predicting prognosis in hepatocellular carcinoma. Front cell Dev Biol. 2020;8:629.
Lin W, Wu S, Chen X, Ye Y, Weng Y, Pan Y, et al. Characterization of hypoxia signature to evaluate the tumor immune microenvironment and predict prognosis in glioma groups. Front Oncol. 2020;10:796.
Li W, Li T, Sun C, Du Y, Chen L, Du C, et al. Identification and prognostic analysis of biomarkers to predict the progression of pancreatic cancer patients. Mol Med. 2022;28:43. https://doi.org/10.1186/s10020-022-00467-8.
Xiong Y, Yuan L, Xiong J, Xu H, Luo Y, Wang G, et al. An outcome model for human bladder cancer: a comprehensive study based on weighted gene co-expression network analysis. J Cell Mol Med. 2020;24:2342–55.
Jeong S-H, Kim RB, Park SY, Park J, Jung E-J, Ju Y-T, et al. Nomogram for predicting gastric cancer recurrence using biomarker gene expression. Eur J Surg Oncol J Eur Soc Surg Oncol Br Assoc Surg Oncol. 2020;46:195–201.
Liu Z, Mi M, Li X, Zheng X, Wu G, Zhang L. A lncRNA prognostic signature associated with immune infiltration and tumour mutation burden in breast cancer. J Cell Mol Med. 2020;24:12444–56.
Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell. 2014;158:929–44.
Iglesia MD, Parker JS, Hoadley KA, Serody JS, Perou CM, Vincent BG. Genomic analysis of immune cell infiltrates across 11 tumor types. J Natl Cancer Inst. 2016. https://doi.org/10.1093/jnci/djw144.
Malta TM, Sokolov A, Gentles AJ, Burzykowski T, Poisson L, Weinstein JN, et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell. 2018;173:338-354.e15.
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24:1550–8.
Van den Eynden GG, Majeed AW, Illemann M, Vermeulen PB, Bird NC, Høyer-Hansen G, et al. The multifaceted role of the microenvironment in liver metastasis: biology and clinical implications. Cancer Res [Internet]. 2013; 73:2031 LP – 2043. Available from: http://cancerres.aacrjournals.org/content/73/7/2031.abstract.
Hu B, Jarzynka MJ, Guo P, Imanishi Y, Schlaepfer DD, Cheng S-Y. Angiopoietin 2 induces glioma cell invasion by stimulating matrix metalloprotease 2 expression through the αvβ1 integrin and focal adhesion kinase signaling pathway. Cancer Res [Internet]. Am Assoc Cancer Res. 2006; 66:775–83. Available from: https://cancerres.aacrjournals.org/content/66/2/775.
Eke I, Cordes N. Focal adhesion signaling and therapy resistance in cancer. Semin Cancer Biol. 2015;31:65–75.
Shi B, Chu J, Huang T, Wang X, Li Q, Gao Q, et al. The scavenger receptor MARCO expressed by tumor-associated macrophages are highly associated with poor pancreatic cancer prognosis. Front Oncol. 2021;11:771488.
Pinter M, Scheiner B, Peck-Radosavljevic M. Immunotherapy for advanced hepatocellular carcinoma: a focus on special subgroups. Gut. 2021;70:204–14.
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang T-H, et al. The immune landscape of cancer. Immunity. 2018;48:812-830.e14.
Bonneville R, Krook MA, Kautto EA, Miya J, Wing MR, Chen H-Z, et al. Landscape of microsatellite instability across 39 cancer types. JCO Precis Oncol. 2017. https://doi.org/10.1200/PO.17.00073.
Tsuchiya H, Shiota G. Immune evasion by cancer stem cells. Regen Ther [Internet]. 2021; 17: 20–33. Available from: https://www.sciencedirect.com/science/article/pii/S2352320421000134.
Espinoza-Sánchez NA, Chimal-Ramírez GK, Mantilla A, Fuentes-Pananá EM. IL-1β, IL-8, and matrix metalloproteinases-1, -2, and -10 are enriched upon monocyte-breast cancer cell cocultivation in a matrigel-based three-dimensional system. Front Immunol. 2017;8:205. https://doi.org/10.3389/fimmu.2017.00205.
Kessenbrock K, Plaks V, Werb Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell. 2010;141:52–67.
Iredale JP. Tissue inhibitors of metalloproteinases in liver fibrosis. Int J Biochem Cell Biol. 1997;29:43–54.
Consolo M, Amoroso A, Spandidos DA, Mazzarino MC. Matrix metalloproteinases and their inhibitors as markers of inflammation and fibrosis in chronic liver disease (Review). Int J Mol Med. 2009;24:143–52.
Yang JD, Nakamura I, Roberts LR. The tumor microenvironment in hepatocellular carcinoma: Current status and therapeutic targets. Semin Cancer Biol [Internet]. 2011; 21:35–43. Available from: https://www.sciencedirect.com/science/article/pii/S1044579X1000091X.
Ogasawara S, Yano H, Momosaki S, Nishida N, Takemoto Y, Kojiro S, et al. Expression of matrix metalloproteinases (MMPs) in cultured hepatocellular carcinoma (HCC) cells and surgically resected HCC tissues. Oncol Rep. 2005;13:1043–8. https://doi.org/10.3892/or.13.6.1043.
Chen R, Cui J, Xu C, Xue T, Guo K, Gao D, et al. The significance of MMP-9 over MMP-2 in HCC invasiveness and recurrence of hepatocellular carcinoma after curative resection. Ann Surg Oncol. 2012;19(Suppl 3):S375–84.
Jin Y, Liang Z-Y, Zhou W-X, Zhou L. High MMP14 expression is predictive of poor prognosis in resectable hepatocellular carcinoma. Pathology [Internet]. 2020; 52:359–65. Available from: https://www.sciencedirect.com/science/article/pii/S0031302520304621.
Yokoi A, Yoshioka Y, Yamamoto Y, Ishikawa M, Ikeda S, Kato T, et al. Malignant extracellular vesicles carrying MMP1 mRNA facilitate peritoneal dissemination in ovarian cancer. Nat Commun. 2017;8:14470. https://doi.org/10.1038/ncomms14470.
Sunami E, Tsuno N, Osada T, Saito S, Kitayama J, Tomozawa S, et al. MMP-1 is a prognostic marker for hematogenous metastasis of colorectal cancer. Oncologist. 2000;5:108–14.
Yen C-Y, Chen C-H, Chang C-H, Tseng H-F, Liu S-Y, Chuang L-Y, et al. Matrix metalloproteinases (MMP) 1 and MMP10 but not MMP12 are potential oral cancer markers. Biomarkers. 2009;14:244–9. https://doi.org/10.1080/13547500902829375.
Poola I, DeWitty RL, Marshalleck JJ, Bhatnagar R, Abraham J, Leffall LD. Identification of MMP-1 as a putative breast cancer predictive marker by global gene expression analysis. Nat Med. 2005;11:481–3. https://doi.org/10.1038/nm1243.
Liu H, Lan T, Li H, Xu L, Chen X, Liao H, et al. Circular RNA circDLC1 inhibits MMP1-mediated liver cancer progression via interaction with HuR. Theranostics. 2021;11:1396–411.
Kim E, Kim D, Lee J-S, Yoe J, Park J, Kim C-J, et al. Capicua suppresses hepatocellular carcinoma progression by controlling the ETV4-MMP1 axis. Hepatology. 2018;67:2287–301.
Yu C-L, Yu Y-L, Yang S-F, Hsu C-E, Lin C-L, Hsieh Y-H, et al. Praeruptorin A reduces metastasis of human hepatocellular carcinoma cells by targeting ERK/MMP1 signaling pathway. Environ Toxicol. 2021;36:540–9.
Llovet JM, Montal R, Sia D, Finn RS. Molecular therapies and precision medicine for hepatocellular carcinoma. Nat Rev Clin Oncol. 2018;15:599–616. https://doi.org/10.1038/s41571-018-0073-4.
Rizzo A, Ricci AD. PD-L1, TMB, and other potential predictors of response to immunotherapy for hepatocellular carcinoma: how can they assist drug clinical trials? Expert Opin Invest Drugs. 2021. https://doi.org/10.1080/13543784.2021.1972969.
Zitvogel L, Kroemer G. Targeting PD-1/PD-L1 interactions for cancer immunotherapy. Oncoimmunology. 2012;1:1223–5.
Ren D, Hua Y, Yu B, Ye X, He Z, Li C, et al. Predictive biomarkers and mechanisms underlying resistance to PD1/PD-L1 blockade cancer immunotherapy. Mol Cancer. 2020;19:19. https://doi.org/10.1186/s12943-020-1144-6.
Sharpe AH, Pauken KE. The diverse functions of the PD1 inhibitory pathway. Nat Rev Immunol. 2018;18:153–67. https://doi.org/10.1038/nri.2017.108.
Siddhartha R, Garg M. Molecular and clinical insights of matrix metalloproteinases into cancer spread and potential therapeutic interventions. Toxicol Appl Pharmacol. 2021;426:115593.
Acknowledgements
We thank all of the authors listed in this manuscript.
Funding
This study was supported by Key Research Projects of Henan Higher Education Institutions (21A320049) and Henan Province Medical Science and Technology Research Project (SBGJ202102063).
Author information
Authors and Affiliations
Contributions
Wei Li and Linping Xu designed, wrote, and edited the manuscript; analyzed the data; and finished the figures. Hui Yang revised the manuscript. Meimei Yan analyzed the data. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests relating to the publication of this manuscript.
Ethics approval
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10238_2022_897_MOESM1_ESM.tif
Supplementary Figure 1 (A) Upregulated KEGG pathways in HCC tissues. (B) Downregulated KEGG pathways in HCC tissues. (C) Upregulated GO terms in HCC tissues. (D) Downregulated GO terms in HCC tissues (TIF 7872 KB)
10238_2022_897_MOESM2_ESM.tif
Supplementary Figure 2 (A) The dotted line represents the median risk score and divides the patients into low-risk and high-risk groups. The curve of risk score. Survival status of the patients. More dead patients correspond to a higher risk score. Heat map of the expression profiles of the MMP3, -11, -15, -24, and -21 in the low- and high-risk groups. (B) Kaplan–Meier survival analysis of MMP3, -11, -15, -24, and -21 in the low- and high-risk groups. (C) Time-dependent ROC analysis of MMP3, -11, -15, -24, and -21 in the low- and high-risk groups (TIF 16729 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, L., Yang, H., Yan, M. et al. Matrix metalloproteinase 1 is a poor prognostic biomarker for patients with hepatocellular carcinoma. Clin Exp Med 23, 2065–2083 (2023). https://doi.org/10.1007/s10238-022-00897-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-022-00897-y