Abstract
Despite improved survival for the patients with diffuse large B cell lymphoma (DLBCL), the prognosis after relapse is poor. LincRNA-p21 is a long intergenic noncoding RNA, which is located on chromosome 17, approximately 15 kb upstream from the Cdkn1a (p21) gene. However, its clinical importance and biological role in DLBCL prognosis are unknown. In this study, we conducted quantitative reverse-transcription polymerase chain reaction to investigate the lincRNA-p21 expression in DLBCL. We found that lincRNA-p21 levels were markedly decreased in DLBCL tissues compared with normal. Its expression level was significantly correlated with Ann Arbor stages, B symptoms, performance status, IPI score and serum LDH. Moreover, patients with high levels of LincRNA-p21 expression had a favorable overall survival and progression-free survival. Furthermore, ectopic expression of lincRNA-p21 inhibited cell proliferation, arrested cycle progression and modulated cyclin D1, CDK4 and p21 expression in DLBCL cell lines. These results demonstrated lincRNA-p21 can be identified as a potential novel prognostic biomarker for prognosis in DLBCL and regulate cell proliferation and cycle in vitro. Our findings highlight the value of integrated comprehensive analysis to identify prognostic markers and genetic driver events not previously implicated in DLBCL.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse large B cell lymphoma (DLBCL) is the most common lymphoid malignancy, accounting for approximately 25–30 % of all newly diagnosed cases and with rapidly rising incidence [1]. The introduction of rituximab plus cyclophosphamide/doxorubicin/vincristine/prednisone (R-CHOP) chemotherapy is considered as the standard treatment for DLBCL patients, which dramatically improves the treatment outcome and prognosis [2, 3]. Despite important advances in treatment, approximately one-third of patients will develop relapsed/refractory disease shortly after initial remission and will eventually die as a result [4–6]. Because DLBCL has heterogeneous biological features and clinical behaviors, current attempts to evaluate prognosis in DLBCL rely mainly on clinical parameters, as used most notably in the international prognostic index (IPI) score [6–8]. But it does not fully reflect the biological spectrum of DLBCL. Thus, novel prognostic factors are being explored to predict treatment outcome.
Long noncoding RNAs (lncRNAs) are defined as RNA molecules more than 200 nt in length with no open reading frame [9, 10]. Long intergenic noncoding RNAs (lincRNAs) are those whose transcription loci are within genomic interval between two protein coding genes [10, 11]. To date, numbers of large intergenic noncoding RNAs (lincRNAs) play pivotal roles in cancer-related gene regulatory system, and the disorder of their expression is thought to promote cancer cell proliferation, invasion and metastasis [11, 12]. However, the biological functions and prognostic values of lincRNAs in bladder cancer are still largely unexplored.
In the present work, we focus on lincRNA-p21. LincRNA-p21 is a long intergenic noncoding RNA (3100 nt) and located on chromosome 17, approximately 15 kb upstream from the Cdkn1a (p21) gene [13, 14]. It was originally identified as a direct transcriptional target of p53 and to mediate p53-dependent apoptosis but not cell-cycle arrest in doxorubicin-treated mouse embryo fibroblasts (MEFs) [14, 15]. Wang et al. [16] showed that lincRNA-p21 enhances the sensitivity of radiotherapy for human colorectal cancer by targeting the wnt/beta catenin signaling pathway. In addition, Dimitrova et al. [17, 18] showed that deregulation of lincRNA-p21 expression promotes polycomb target gene expression and enforces the G1/S checkpoint. However, the role of lincRNA-p21 in DLBCL remains largely elusive.
The aim of this study is to identify and characterize the expression pattern and clinical significance of lincRNA-p21 in DLBCL and further explore the biological functions and molecular mechanism of lincRNA-p21 in DLBCL. Firstly, we showed that lincRNA-p21 was downregulated in both DLBCL tissues and cell lines and could be used to distinguish DLBCL from normal. Next, we demonstrated the clinical significance of lincRNA-p21 in DLBCL and determined that it could be a novel biomarker for poor prognosis of DLBCL. Moreover, overexpression of lincRNA-p21 could inhibit cell proliferation and cycle progression of DLBCL. Furthermore, we found that alteration of lincRNA-p21 expression can influence the molecular events that occur downstream of lincRNA-p21, indicating that lincRNA-p21 affected DLBCL tumorigenesis. Our results suggest that lincRNA-p21 may represent a novel indicator of poor prognosis and may be a potential therapeutic target for the diagnosis and gene therapy of DLBCL. This study advances our understanding of the role of lincRNAs such as lincRNA-p21 as regulators of DLBCL pathogenesis.
Materials and methods
Patients
All of the experiments were approved by the Ethics Board of Jiangsu Cancer Hospital of Nanjing Medical University. The informed consents were provided by all the patients, and all the clinical investigations were conducted according to the principles expressed in the Declaration of Helsinki. R-CHOP- or R-CHOP-like treatment as a frontline regimen was administrated to 105 patients whose clinical efficacy was evaluable for this study. A total of 105 patients, who were diagnosed with DLBCL at Jiangsu Cancer Hospital or First People’s Hospital of Yunnan Province from 2007 to 2009 and treated with rituximab-based chemotherapy, were enrolled. Tissue samples were washed with sterile phosphate-buffered saline before being snap-frozen in liquid nitrogen for future RNA and protein extraction. The patients’ histologic slides and clinical medical records were reviewed by two experienced pathologists and oncologists, respectively.
Quantitative real-time reverse transcriptase PCR (qRT-PCR)
Total RNA from frozen tissues or cultured cells was isolated with TRIzol reagent (Invitrogen, GrandIsland, USA) according to the manufacturer’s instructions. RNA was reverse-transcribed to cDNA by using a PrimeScript™ First-Strand cDNA Synthesis Kit (Takara, Dalian, China). The SYBR® Premix Ex Taq™ II (Takara, Dalian, China) was used to detect lincrna-p21 expression according to the manufacturer’s protocol. The real-time PCR primers for lincRNA-p21 and GAPDH were as follows: lincRNA-p21 sense, 5′-GCTCGACGCTAGGATCTGAC-3′ and reverse, 5′-GCTTTCCACGACGGTGAC-3′; cyclin D1 sense, 5′-GGCGGATTGGAAATGAACTT-3′ and reverse, 5′-TCCTCTCCAAAATGCCAGAG-3′; CDK4 sense, 5′-GTCGGCTTCAGAGTTTCCAC-3′ and reverse, 5′-TGCAGTCCACATATGCAACA-3′; p21 sense, 5′-CATGGGTTCTGACGGACAT-3′ and reverse, 5′-AGTCAGTTCCTTGTGGAGCC-3′; GAPDH sense, 5′-AATGAAGGGGTCATTGATGG-3′ and reverse, 5′-AAGGTGAAGGTCGGAGTCAA-3′. Real-time PCR and data collection were performed on ABI 7300 (Applied Biosystems, Waters, USA). Results were normalized to the expression of GAPDH [18].
Cell culture
Human DLBCL cell lines (SU-DHL-2, OCI-LY-3, OCI-LY-10, SU-DHL-4 and OCI-LY-7) were purchased from the American Type Culture Collection (Manassas, USA). All the cells were cultured in RPMI1640 (#11875-093, Gibco;) medium supplemented with penicillin–streptomycin (#15140-122, Gibco) and 10 % fetal bovine serum (#10100-147, Gibco) at 37 °C with 5 % CO2 [19].
LincRNA-p21 overexpression
Overexpression of lincRNA-p21 was performed as previously described [14–16]. A plasmid expressing lincRNA-p21 was used to construct plasmid plincRNA-p21 pcDNA3.1 (+) (pc-lincRNA-p21). The plasmid was provided by GenePharma Corporation (Shanghai, China).
Cell proliferation assay
Cell proliferation was counted using Cell Proliferation Reagent Kit (MTT) (Beyotime, Haimen, China). Twenty-four hours after transfection, 1 × 103 cells per well were plated in 96-well plates. MTT (Beyotime) was added to each well, and cells were further cultured for 4 h at 37 °C. The reaction was stopped by 150 μl DMSO, and optical density at 490 nm was detected on a microplate reader [19].
Flow cytometric analysis of cell cycle
Transfected cells were harvested after transfection. Cells for cell-cycle analysis were stained with propidium oxide by Cell Cycle Analysis Kit (Beyotime, Haimen, China) following the protocol and analyzed by FACScan (BD Biosciences, San Jose, USA). The percentage of the cells in G1–G0, S and G2–M phase were counted and compared [18].
Western blot analysis
Western blot analysis was performed according to our previous study [20]. Total proteins were extracted and separated using sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to polyvinylidene fluoride membranes (Millipore, Billerica, USA). The membranes were blocked with 10 % nonfat milk at room temperature for 1 h and then incubated overnight with rabbit anti-cyclin D1 (#2978, Cell Signaling, Boston, USA), anti-CDK4 (#12790, Cell Signaling), anti-p21 (#2947, Cell Signaling) or anti-GAPDH (#2118, Cell Signaling) antibodies at 4 °C. After incubation with horseradish peroxidase-conjugated anti-rabbit IgG (#3423, Cell Signaling), the specific protein band was visualized by enhanced chemiluminescence (Millipore). GAPDH was used as an internal control, and each experiment was repeated at least thrice.
Statistical analysis
All experiments were independently repeated at least thrice, and all data were analyzed using the SPSS software package (version 19.0, SPSS, IBM, Armonk, USA). The significant differences in the expression of lincRNA-p21 in the DLBCL and normal tissues were analyzed by the Wilcoxon rank-sum test. The Pearson Chi-square test was used to examine the correlation between the expression of lincRNA-p21 and clinicopathological characteristics. The Kaplan–Meier method was used to estimate the probability of patient survival. A one-way ANOVA test was adopted to investigate the difference in cell proliferation assays, and the least significant difference t test was used to analyze two groups. The Student’s t test was used to compare data from the densitometry analysis of qRT-PCR and Western blot analysis.
Data were expressed as mean ± SD. The results were considered to be statistically significant at P < 0.05.
Results
Decreased expression and diagnostic value of lincRNA-p21 in DLBCL
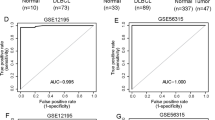
To investigate the role of lincRNA-p21 in DLBCL, we first quantified the expression levels of lincRNA-p21 in tissues by qRT-PCR. As shown in Fig. 1a, tumor tissues expressed evidently lower levels of lincRNA-p21 than normal (P < 0.0001). Levels of lincRNA-p21 in the DLBCL cell lines, SU-DHL-2, OCI-LY-3, OCI-LY-10, SU-DHL-4 and OCI-LY-7 were consistently lower than the average levels of lincRNA-p21 expression in the normal tissues (P < 0.0001, Fig. 1b). To explore whether the levels of lincRNA-p21 expression could be used as a diagnostic biomarker of DLBCL, we produced a ROC curve by comparing the different levels of lincRNA-p21 expression in DLBCL tissues and normal. Area under the ROC curve was up to 0.8718 (P < 0.0001, Fig. 1c).
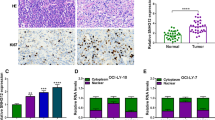
LincRNA-p21 expression in DLBCL. a LincRNA-p21 expression levels assessed by qRT-PCR in cancerous (T) and non-cancerous tissues (N) from 105 DLBCL samples. LincRNA-p21 levels were normalized to GAPDH. LincRNA-p21 levels in T were significantly lower than those in N (P < 0.0001). Horizontal lines, mean value of each sample. b Relative expression of lincRNA-p21 between five DLBCL cell lines (SU-DHL-2, OCI-LY-3, OCI-LY-10, SU-DHL-4 and OCI-LY-7). Each cell line was duplication analyzed three times, and lincRNA-p21 expression level of five DLBCL cell lines was significantly lower than the average levels of normal tissues. c Receiver operating characteristic (ROC) curve analysis was employed to determine whether lincRNA-p21 is really good candidates to discriminate tumor tissues from non-tumorous tissues. d qRT-PCR analysis of lincRNA-p21 in 105 DLBCL tissues and the classification based on lincRNA-p21 level (lincRNA-p21/GAPDH = −1.783). Vertical line, borderline of lincRNA-p21 high (n = 55) or low (n = 50)
Clinicopathological significance of lincRNA-p21
To gain further insights into the clinical role of lincRNA-p21 in DLBCL, we analyzed the correlation between lincRNA-p21 expression and patient clinicopathological feathers. According to the median ratio of relative lincRNA-p21 expression (−1.784-fold) in tumor tissues, 105 DLBCL patients were classified into two groups: high group (n = 55, lincRNA-p21 expression ratio >−1.784-fold) and low group (n = 50, lincRNA-p21 expression ratio <−1.784-fold, Fig. 1d). We found that lincRNA-p21 expression was irrelevant with age (P = 0.828), gender (P = 0.315) and extra nodal status (P = 1.000). However, lincRNA-p21 expression was closely associated with Ann Arbor stages (P = 0.043), B symptoms (P = 0.046), performance status (P = 0.021), IPI score (P = 0.000) and serum LDH (P = 0.002, Table 1).
Prognostic importance of lincRNA-p21
Log-rank tests showed that the 5-year OS rates in the group with low-lincRNA-p21 were significantly poorer than those in the high-lincRNA-p21 (P = 0.0001, Fig. 2a). Similarly, the PFS rates in the low-lincRNA-p21 group were significantly shorter than those in the high-lincRNA-p21 group (P < 0.0001, Fig. 2b). Furthermore, Cox regression analysis suggested that lincRNA-p21 expression was an independent factor that forecasts OS and PFS (Table 2). Collectively, these data implicated that lincRNA-p21 could be used as a prognostic biomarker in DLBCL.
LincRNA-p21 overexpression arrested cell growth and cycle progression of DLBCL cells
Using qRT-PCR, we confirmed that lincRNA-p21 expression levels of OCI-LY-7 cells transfected with pc-lincRNA-p21 were significantly lower than control (Fig. 3a). Twenty-four hours after transfection with pc-lincRNA-p21-2, MTT assays revealed that cell proliferation was repressed in OCI-LY-7 cells compared with control (Fig. 3b). Next, flow cytometric analysis was conducted to further explore the effect of pc-lincRNA-p21 on proliferation of DLBCL cells by regulating cell-cycle progression. The results revealed that the cell-cycle progression of OCI-LY-7 cells transfected with pc-lincRNA-p21-2 was significantly stalled at the G1–G0 phase compared with the control (Fig. 3c). As shown in Fig. 3d, lincRNA-p21 overexpression downregulated the expression levels of cyclin D1 and CDK4 and upregulated p21. Taken together, these results showed that lincRNA-p21 overexpression could obviously suppress tumor growth of DLBCL cells.
Effect of lincRNA-p21 overexpression on DLBCL cell growth in vitro. a The relative expression level of lincRNA-p21 overexpression in OCI-LY-7 cells, transfected with empty vector (pc-control) or lincRNA-p21 plasmids (pc-lincRNA-p21), was tested by qRT-PCR. b At 48 h after transfection, MTT assay was performed to determine the proliferation of OCI-LY-7 cells. c Cell cycle of OCI-LY-7 was analyzed by flow cytometry. The bar chart represents the percentage of cells in G1–G0, S or G2–M phase, as indicated. d Western blot analysis of expression levels of cyclin D1, CDK4 and p21 in DLBCL cells
Association of lincRNA-p21 with cyclin D1, CDK4 and p21 in DLBCL
Then, we sought to determine whether there was any interaction between cyclin D1, CDK4, p21 and lincRNA-p21. To further validate it, the expression of lincRNA-p21, cyclin D1, CDK4 and p21 was analyzed by qRT-PCR in DLBCL tissues. As shown in Fig. 4a, b, the expression levels of lincRNA-p21 were negatively correlated with the mRNA expression levels of cyclin D1 and CDK4. The levels of lincRNA-p21 expression were positively associated with p21 mRNA (Fig. 4c). Taken together, all these results confirmed a functional link between lincRNA-p21 and cyclin D1, CDK4 and p21 in human DLBCL.
Correlation of expression levels of lincRNA-p21 with cyclin D1, CDK4 and p21 in DLBCL tissues. Expression levels of lincRNA-p21 are negatively correlated with cyclin D1 (a) and CDK4 (b) and positively associated with p21 (c) in human DLBCL tissues. Pearson’s correlation analysis of the relative expression levels of lincRNA-p21 (normalized to GAPDH) and the relative expression levels of cyclin D1, CDK4 and p21 mRNA (normalized to GAPDH) determined using qRT-PCR in 105 human DLBCL tissue samples
Discussion
DLBCL is the most common lymphoid malignancy and is biologically and pathologically heterogeneous [1, 21]. Mainstream tumorigenic processes involved in DLBCL are characterized by phenotypic multistep progression cascades and gene expression patterns [7, 22]. It is difficult to predict its prognosis, and there is no definite prognostic marker yet. The reliable identification of DLBCL progression-specific targets has huge implications for its prevention and treatment [4, 23]. However, identification of the molecular markers that correlate with the development and progression of DLBCL still remains a challenge.
Currently, a large number of cancer-related lncRNAs have been discovered, and they are demonstrated to be involved in the development of human diseases, including tumor cell proliferation and cell cycle [12, 18, 19, 24]. Several associations between altered lincRNAs in cancers and clinical significance were observed [10, 11]. Until now, emerging evidences show that some lincRNAs can be used as biomarkers for the prediction of diagnosis and prognosis of or as tumor therapeutic targets in human cancer. However, the DLBCL-related lincRNAs have rarely been reported yet.
In present study, we focused on the lincRNA-p21. We found that lincRNA-p21 was downregulated in DLBCL compared with normal. Consistently, previous studies have showed that lincRNA-p21 was downregulated in CRC [16]. And lincRNA-p21 is correlated with Ann Arbor stages, B symptoms, performance status, IPI score and serum LDH. These observations indicate that lincRNA-p21 may function as an anti-oncogenic factor in human tumor progression. To determine the relationship between lincRNA-p21 expression and prognosis of DLBCL patients, we attempted to evaluate the correlation between lincRNA-p21 expression and clinical outcomes. Kaplan–Meier analysis showed that patients with low levels of lincRNA-p21 expression had remarkably shorter survival time than those with high levels. Multivariate analysis further revealed that lincRNA-p21 expression was a significant independent predictor of poor survival of DLBCL patients. To our best knowledge, this is the first report that showed lincRNA-p21 may be a predictor of survival in DLBCL in a sizable group of DLBCL patients.
Our work showed that cyclin D1, CDK4 and p21 were functional targets of lincRNA-p21 in DLBCL cells. Cyclin D1 is a member of highly conserved cyclin family [25]. Overexpression of cyclin D1, which alters cell-cycle progression, is observed frequently in a variety of tumors, related to the development of many cancers and may contribute to tumorigenesis [25, 26]. Cyclin D1 activates CDK4, which subsequently allows the cell cycle to progress through G1 into S [27]. Furthermore, p21 expression has been shown to be reduced or lost in a variety of cancer types [28]. A possible explanation is that p21 exerts its inhibitory control over the cell cycle primarily through direct binding to cyclins and CDKs, thereby preventing cell proliferation [28, 29]. We showed p21 was a downstream regulator involved in lincRNA-p21-mediated growth arrest in DLBCL cells. Here, these data indicate that lincRNA-p21 may function as a tumor suppressor and its deficiency or decreased expression could contribute to DLBCL development. Taken together, our findings indicate that lincRNA-p21 arrests DLBCL cell growth and cycle progression maybe partly via regulating cyclin D1, CDK4 and p21 expression.
In conclusion, we demonstrate that the decreased lincRNA-p21 expression is a common event underlying DLBCL and can be considered a novel diagnostic and independent prognostic factor in patients with DLBCL. It indicates that lincRNA-p21 may play a key tumor-suppressive as an indicator of favorable survival rate and a positive prognostic factor for DLBCL patients. Further, well understanding of the mechanisms of lincRNA-p21 in the molecular etiology of DLBCL will supply a strategy and facilitate the development of lincRNA-directed diagnostic, prognostic and therapeutic agents against this malignancy.
References
Martelli M, Ferreri AJ, Agostinelli C, Di Rocco A, Pfreundschuh M, Pileri SA. Diffuse large B-cell lymphoma. Crit Rev Oncol Hematol. 2013;87(2):146–71.
Feugier P, Van Hoof A, Sebban C, Solal-Celigny P, Bouabdallah R, Ferme C, et al. Long-term results of the R-CHOP study in the treatment of elderly patients with diffuse large B-cell lymphoma: a study by the Groupe d’Etude des Lymphomes de l’Adulte. J Clin Oncol. 2005;23(18):4117–26.
Lee HJ, Shin DH, Kim KB, Shin N, Park WY, Lee JH, et al. Polycomb protein EZH2 expression in diffuse large B-cell lymphoma is associated with better prognosis in patients treated with rituximab, cyclophosphamide, doxorubicin, vincristine and prednisone. Leuk Lymphoma. 2014;55(9):2056–63. doi:10.3109/10428194.2013.858816.
Vaidya R, Witzig TE. Prognostic factors for diffuse large B-cell lymphoma in the R(X)CHOP era. Ann Oncol. 2014;25(11):2124–33. doi:10.1093/annonc/mdu109.
Hu Y, Ding N, Jin X, Feng L, Ping L, Song Y, et al. Genetic polymorphisms of STAT3 correlated with prognosis in diffuse large B-cell lymphoma patients treated with rituximab. Cancer Cell Int. 2014;14(1):25.
Taskinen M, Louhimo R, Koivula S, Chen P, Rantanen V, Holte H, et al. Deregulation of COMMD1 is associated with poor prognosis in diffuse large B-cell lymphoma. PLoS One. 2014;9(3):e91031. doi:10.1371/journal.pone.0091031.
Roschewski M, Staudt LM, Wilson WH. Diffuse large B-cell lymphoma-treatment approaches in the molecular era. Nat Rev Clin Oncol. 2014;11(1):12–23. doi:10.1038/nrclinonc.2013.197.
Koh YW, Park C-S, Yoon DH, Suh C, Huh J. Should the cut-off values of the lymphocyte to monocyte ratio for prediction of prognosis in diffuse large B-cell lymphoma be changed in elderly patients? Eur J Haematol. 2014;93(4):340–8. doi:10.1111/ejh.12354.
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–41.
Tsai M-C, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010;329(5992):689–93.
Tsai M-C, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71(1):3–7.
Feng S, Yao J, Chen Y, Geng P, Zhang H, Ma X, et al. Expression and functional role of reprogramming-related long noncoding RNA (lincRNA-ROR) in glioma. J Mol Neurosci. 2015;. doi:10.1007/s12031-014-0488-z.
Hall JR, Messenger ZJ, Tam HW, Phillips SL, Recio L, Smart RC. Long noncoding RNA lincRNA-p21 is the major mediator of UVB-induced and p53-dependent apoptosis in keratinocytes. Cell Death Dis. 2015;6:e1700. doi:10.1038/cddis.2015.67.
Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010;142(3):409–19. doi:10.1016/j.cell.2010.06.040.
Yoon JH, Abdelmohsen K, Srikantan S, Yang XL, Martindale JL, De S, et al. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012;47(4):648–55. doi:10.1016/j.molcel.2012.06.027.
Wang GY, Li ZW, Zhao Q, Zhu YY, Zhao C, Li X, et al. LincRNA-p21 enhances the sensitivity of radiotherapy for human colorectal cancer by targeting the Wnt/beta-catenin signaling pathway. Oncol Rep. 2014;31(4):1839–45. doi:10.3892/or.2014.3047.
Dimitrova N, Zamudio JR, Jong RM, Soukup D, Resnick R, Sarma K, et al. LincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol Cell. 2014;54(5):777–90. doi:10.1016/j.molcel.2014.04.025.
Peng W, Gao W, Feng J. Long noncoding RNA HULC is a novel biomarker of poor prognosis in patients with pancreatic cancer. Med Oncol. 2014;31(12):346. doi:10.1007/s12032-014-0346-4.
Peng W, Wu G, Fan H, Wu J, Feng J. Long noncoding RNA SPRY4-IT1 predicts poor patient prognosis and promotes tumorigenesis in gastric cancer. Tumour Biol. 2015;. doi:10.1007/s13277-015-3376-4.
Peng W, Zhang J, Liu J. URG11 predicts poor prognosis of pancreatic cancer by enhancing epithelial-mesenchymal transition-driven invasion. Med Oncol. 2014;31(7):64. doi:10.1007/s12032-014-0064-y.
Lossos IS. Molecular pathogenesis of diffuse large B-cell lymphoma. J Clin Oncol. 2005;23(26):6351–7. doi:10.1200/jco.2005.05.012.
Martelli M, Ferreri AJM, Agostinelli C, Di Rocco A, Pfreundschuh M, Pileri SA. Diffuse large B-cell lymphoma. Crit Rev Oncol Hematol. 2013;87(2):146–71. doi:10.1016/j.critrevonc.2012.12.009.
Lossos IS, Morgensztern D. Prognostic biomarkers in diffuse large B-cell lymphoma. J Clin Oncol. 2006;24(6):995–1007. doi:10.1200/jco.2005.02.4786.
Peng W, Fan H, Wu G, Wu J, Feng J. Upregulation of long noncoding RNA PEG10 associates with poor prognosis in diffuse large B cell lymphoma with facilitating tumorigenicity. Clin Exp Med. 2015. doi:10.1007/s10238-015-0350-9
Barnes DM. Cyclin D1 in mammary carcinoma. J Pathol. 1997;181(3):267–9. doi:10.1002/(sici)1096-9896(199703)181:3<267:aid-path783>3.0.co;2-x.
Nakamura S, Yatabe Y, Seto M. Cyclin D1 overexpression in malignant lymphomas. Pathol Int. 1997;47(7):421–9. doi:10.1111/j.1440-1827.1997.tb04519.x.
Kitagawa M, Higashi H, Jung HK, SuzukiTakahashi I, Ikeda M, Tamai K, et al. The consensus motif for phosphorylation by cyclin D1-Cdk4 is different from that for phosphorylation by cyclin A/E-Cdk2. EMBO J. 1996;15(24):7060–9.
Kumar R, Gururaj AE, Barnes CJ. p21-activated kinases in cancer. Nat Rev Cancer. 2006;6(6):459–71. doi:10.1038/nrc1892.
Abbas T, Dutta A. p21 in cancer: intricate networks and multiple activities. Nat Rev Cancer. 2009;9(6):400–14. doi:10.1038/nrc2657.
Acknowledgments
This work was supported by grants from Agency of Jiangsu Province Science and Technology (No. 2013035) and Research Office of Jiangsu Cancer Hospital (No. ZK201401).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Rights and permissions
About this article
Cite this article
Peng, W., Wu, J. & Feng, J. LincRNA-p21 predicts favorable clinical outcome and impairs tumorigenesis in diffuse large B cell lymphoma patients treated with R-CHOP chemotherapy. Clin Exp Med 17, 1–8 (2017). https://doi.org/10.1007/s10238-015-0396-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-015-0396-8