Abstract
In recent years, biodiversity loss has become one of the most serious environmental issues worldwide, especially in aquatic ecosystems. To avoid diversity loss, it is necessary to monitor biological communities, and environmental DNA (eDNA) metabarcoding has been developed as a rapid, noninvasive, and cost-effective method for aquatic biodiversity monitoring. Although this method has been applied to various environments and taxa, a detailed assessment of the efficient sampling methods for monitoring is still required. In this study, we explored eDNA metabarcoding sampling methods for fish at a single site to maximize the number of detected species using realistic effort in a natural, small river. We considered the following three parameters: sample type (water or sediment), sample position at a site (right and left shore and center of the river), and water volume (10–4000 mL). The results suggested that the number of detected species from sedimentary eDNA was equivalent to that from aqueous eDNA, although the species composition was different. The number of detected species could be saturated by collecting a 1000 mL water sample, regardless of sampling position within a survey site. However, sedimentary eDNA showed a spatially heterogeneous species composition between sampling positions within a survey site despite the short distance (5 m) between positions, without apparent differences in physical properties such as velocity and sediment particle distribution. By completing eDNA biodiversity monitoring of fish with 1000 mL water samples across the whole river, we detected more fish species than in previous traditional surveys conducted at the same sites. Thus, the aqueous eDNA metabarcoding method is as efficient as traditional surveys, while sedimentary eDNA metabarcoding could complement the results of aqueous eDNA metabarcoding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biodiversity loss is a major environmental concern (Butchart et al. 2010), and a particularly critical issue in freshwater environments (Dudgeon et al. 2006; WWF 2018). For biodiversity conservation, rapid and noninvasive underwater biomonitoring methods are required (Dudgeon et al. 2006) because traditional survey methods are costly and their results (e.g., types and numbers of fish species collected) may vary depending on the skill levels of investigators and survey tools used.
Environmental DNA (eDNA) metabarcoding is a rapid and noninvasive method that is also cost-effective (Valentini et al. 2016; Bista et al. 2017; Deiner et al. 2017; Yamamoto et al. 2017). Especially for freshwater fish species, eDNA metabarcoding reveals comparable or more fish species than traditional surveys (Hänfling et al. 2016; Shaw et al. 2016; Nakagawa et al. 2018). However, the maximization of species detection in each ecosystem has not been fully assessed, although some studies have started (Evans et al. 2017; Hayami et al. 2020).
To improve the detection capability of eDNA metabarcoding within a single site, the amount of DNA in a sample is important (Schultz and Lance 2015). In general, the larger the amount of water sampled, the greater the number of species detected (Miya et al. 2016). However, a large volume of water causes difficulties such as a long filtration time in the laboratory, heavy samples to transport, or both. Additionally, large volumes of turbid water are difficult to sift owing to the clogging of the filters. Therefore, it is necessary to explore the appropriate volume of water samples.
Sedimentary eDNA (eDNA included in the sediment), which has a lower decay rate and includes higher concentrations of fish eDNA than aqueous eDNA, is considered a potential alternative medium for eDNA studies (Turner et al. 2015; Sakata et al. 2020). However, only a few studies have been conducted on eDNA metabarcoding for fish species using sediment samples (Shaw et al. 2016; Sakata et al. 2020). Moreover, the heterogeneity of detected species among replications and the small spatial differences within sampling sites have rarely been investigated in both aqueous and sedimentary eDNA studies. Therefore, to increase detection capability, these factors need to be examined within a single site. In addition, when biodiversity monitoring is carried out in a large survey area, economizing the sampling effort by considering the number of sampling sites will improve the effectiveness of eDNA biomonitoring. Therefore, to improve eDNA metabarcoding effectiveness, sampling methods and number of sampling sites need to be examined for each ecosystem.
In this study, we investigated an effective sampling method for eDNA metabarcoding by considering three parameters (sample type, sampling position, and filtered water volume) in the Koide River watershed, and assessed the effectiveness of biomonitoring using eDNA metabarcoding with a determined sampling method. We investigated the effects of these parameters on the number of species detected by eDNA metabarcoding to determine the most effective sampling method for each site. To compare the results of eDNA metabarcoding with those of traditional surveys previously conducted along the whole river, we performed eDNA surveys at the traditional survey performed sites. Finally, we discussed the number of sampling sites required to perform effective biomonitoring using eDNA metabarcoding.
Materials and methods
Field sample collection and filtration
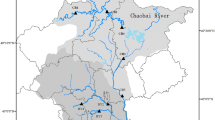
A field survey was conducted in the Koide River watershed, which flows through Kanagawa Prefecture, Japan (Fig. 1). On August 3, 2017, water and sediment samples were collected for eDNA analysis. To determine the sampling method for this river in relation to sample type, sample position, and filtered water volume, we collected water and sediment samples from three transverse positions at site C at approximately 5-m intervals on the left and right shore and the center of the stream (Left, Right, and Center, respectively; Fig. 1). These three positions were selected for two reasons: (1) the difference in the physical environment between the center and shores, and (2) the difference in habitat preference for each fish species. Site C, which is located at the center of the study area, was chosen for our initial investigation because (1) it shows a typical riverscape of the surveyed river in terms of its environmental features such as vegetation and shape of the river; (2) it is far from the estuary and, therefore, should not be affected by marine fish, and (3) sites upstream of site C seem to be unsuitable for method comparisons because of the low fish species number owing to the lack of vegetation. At this site, the river width was 10 m, and shoreside vegetation on both sides consisted of similar emerging plants.
Survey area map. All samples were collected from the Koide River watershed, Kanagawa Prefecture, Japan. Red points show sampling sites and letters indicate site ID. Left, Right, and Center represent sample positions within site C. The traditional survey was performed at all eDNA sampling sites in a previous study
We measured the physical properties (pH, DO, water temperature, velocity, and sediment particle distribution) using a water quality meter (model WQC-24; TOA-DDK, Japan) and an electromagnetic current meter (model AEM-1D; JFE Advantech, Japan) at each position on July 31, 2020. The sediment particle size distribution was measured according to the test method for particle size distribution of soils “JIS A1204” (Japanese Industrial Standards Committee 2009).
At each position, approximately 8 L of surface water was collected using a plastic bottle. Benzalkonium chloride (0.1% final concentration) was added to each sample and mixed to prevent eDNA degradation (Yamanaka et al. 2017). After mixing well, each water sample was then dispensed into six subsamples of 10, 100, 500, 1000, 2000, and 4000 mL for the filtered water volume evaluation. To monitor potential contamination during the filtration and eDNA extraction process, 1000 mL of distilled water was processed in the same way as the samples (negative control).
In addition, to compare sample types, 50 g of sediment was also collected at the same positions as water samples in Site C. Sediment samples were scooped from the surface of the river bottom using a 50-mL tube and subsequently separated into five subsamples (9 g each) from one bulk sample (a total of 15 subsamples). The sediment samples primarily consisted of mud. All sediment samples were stored at − 25 °C until DNA extraction.
To compare the fish monitoring performance between eDNA metabarcoding and traditional surveys, 1000 mL of surface water were collected from 11 sites at various locations along the river (Fig. 1), and previously collected traditional survey data were obtained (Kimura et al. 2015). This traditional survey was performed with constant capture efforts from downstream to upstream in a 50-m section per site. A 50-min survey with four people was conducted at each site using hand, casting, and scoop nets.
After water collection, benzalkonium chloride was added to a final concentration of 0.1%. Within 24 h of sampling, all water samples were filtered using glass-fiber filters with a nominal pore size of 0.7 μm (GF/F; GE Healthcare, Chicago, IL, USA) in a dedicated eDNA laboratory at Kobe University (Kobe, Japan). To avoid cross-contamination, reverse osmosis membrane water (manufactured water by Elix Essential UV; Merck, Japan) was filtered as an equipment blank. Filters were stored at − 25 °C until DNA extraction.
When handling water and sediment samples, the collection bottles, tweezers, filter funnels, and filter holders used were decontaminated with chlorine bleach (0.1% effective chlorine concentration) to prevent cross-contamination among samples (The eDNA Society 2019). Disposable gloves were worn during all procedures to minimize the risk of contamination.
eDNA extraction
The aqueous eDNA on the filters (eDNA from water samples) was extracted using Salivette (Sarstedt, Nümbrecht, Germany) and DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) and stored at − 25 °C according to previously described methods (Minamoto et al. 2019). Briefly, the Salivette tubes were incubated at 56 °C for 30 min, and after incubation, the tubes were centrifuged at 3000×g for 3 min to collect the DNA. To increase DNA yield, 300 μL of Tris–EDTA (TE) buffer was added to the filters and re-centrifuged at 3000×g for 1 min. The collected DNA was purified using the DNeasy Blood & Tissue Kit (Qiagen) according to the manufacturer's protocol. The extracted DNA samples (100 μL) were stored at − 25 °C until the PCR assay.
Extraction of sedimentary eDNA was performed following a previous method with minor modifications (Sakata et al. 2020). Sedimentary eDNA was extracted from 9 g sediment samples by combining alkaline DNA extraction (Kouduka et al. 2012) with ethanol precipitation and a fecal/soil DNA extraction kit (PowerSoil DNA Isolation Kit; Qiagen). A DNA enhancer (G2 DNA/RNA Enhancer; Ampliqon, Odense, Denmark) was added during extraction (Jacobsen et al. 2018). Additionally, to detect cross-contamination, 9 mL ultra-pure water was used as a negative control and treated in the same way as the sediment samples. The final volume of eDNA was 100 μL for both sample types. All tools used were decontaminated with chlorine bleach (0.1% effective chlorine concentration).
Next-generation sequencing library preparation and bioinformatics
To investigate fish species composition, eDNA metabarcoding was performed with MiFish-U primers (forward: 5ʹ-ACACTCTTTCCCTACACGACGCTCTTCCGATCT NNNNNNGTCGGTAAAACTCGTGCCAGC-3ʹ, reverse: 5ʹ-GTGACTGGAGTTCAG ACGTGTGCTCTTCCGATCTNNNNNNCATAGTGGGGTATCTAATCCCAGTTTG-3ʹ), which are universal primers for fish species targeting the 12S rRNA region (Miya et al. 2015). Six random bases were used to enhance cluster separation on the flow cells during initial base call calibrations on the MiSeq platform. The following six-step procedure was carried out according to Sakata et al. (2020): (1) first-round PCR (first PCR) was followed by purification using the SPRIselect Reagent Kit (Beckman Coulter, Brea, CA, USA); (2) quantification of purified DNA was performed using a Qubit dsDNA HS assay kit and a Qubit fluorometer 3.0 (ThermoFisher Scientific, Waltham, MA, USA); (3) the second PCR was run; (4) DNA size selection was carried out by electrophoresis using E-Gel SizeSelect 2% (ThermoFisher Scientific) and the E-Gel Precast Agarose Electrophoresis System (ThermoFisher Scientific); (5) the size distribution of amplicons was confirmed using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA); (6) the library was sequenced using an Illumina MiSeq v2 Reagent kit for 2 × 150 bp paired end (Illumina, San Diego, CA, USA).
MiSeq raw reads were preprocessed and analyzed using USEARCH v. 10.0.240 (Edgar 2010) under the same conditions as those described by Sakata et al. (2020). After data preprocessing and analysis, we performed the following species processing, which included two steps: (1) reads assigned to fish species that were detected in both samples and negative controls were regarded as possible contamination, and the number of species reads detected in the negative controls (i.e., filtration blanks or PCR blanks) was subtracted from the corresponding samples; (2) saltwater fish and migratory salmonid fish DNA was judged as contamination from drainage, and those were excluded because those species did not inhabit the Koide River watershed. For all samples, MiSeq sequencing depth was sufficient to detect fish species because the number of species was saturated (Figs. S1, S2, and S3).
Statistical analysis
All analyses were performed in R v. 3.6.0 (R Core Team 2019) using the “vegan” package v. 2.5-5 and “exactRankTests” package v. 0.8-30. The read data were converted to presence/absence of each species for all analyses. First, to verify different fish species compositions between sampling positions within aqueous or sedimentary eDNA, non-metric multidimensional scaling (NMDS) was performed with “jaccard index” and 10,000 permutations. In addition, permutational multivariate analysis of variance (PERMANOVA) was performed with “jaccard index” and 10,000 permutations using the “adonis” function. Next, to compare the change in number of species owing to the difference in filtered water volume, ANOVA and a post hoc Tukey–Kramer test were performed. To compare the number of species between the 1000 mL water samples and sediment samples, a Wilcoxon test was performed. In this analysis, samples collected from the three positions at site C were regarded as three replicates at this site because the fish species composition detected by aqueous eDNA did not change depending on the water sampling position at this site (Fig. S4). In addition, samples with less than 1000 mL filtered water volume were excluded because those samples should have low confidence (see “Results”).
Wilcoxon signed rank test was performed to compare detected fish species between the whole river aqueous eDNA survey and traditional survey. All graphs were plotted with the "ggplot2" package v. 3.1.1.
Finally, we explored the number of sampling sites that would be sufficient to detect fish fauna in this study river. First, we classified the survey sites based on detected fish communities using nonhierarchical cluster analysis with the k-mean method (“pam” function in package “cluster”). Second, because sampling sites were classified into two clusters (see “Results”; cluster 1 included sites A, G, and H; cluster 2 included all the other sites; Fig. S5), the detected fish community structures were visualized by Venn diagram (“venn.diagram” function in “VennDiagram” package), and fish community structures between the two clusters were compared. Finally, because the fish composition of cluster 1 (minor cluster) was contained in that of cluster 2 (major cluster; Fig. S6), we used species accumulation curves to compare the detected number of species to cumulative the number of sampling sites using cluster 2 only (“specaccum” function in “vegan” package with 1000 permutations).
Results
At site C, the physical properties were similar between sampling positions, except for velocity and sediment particle size distribution. At the center, velocity was faster (Table S1A) and sediment particles were larger than those at both shores (Table S1B; Fig. S7).
To determine the sampling method at a single site, 1,762,797 and 1,918,090 raw reads were obtained for aqueous and sedimentary eDNA, respectively, from explorations at site C. After the bioinformatics processing steps (see “Materials and methods”), 1,276,817 and 1,416,482 reads were retained, respectively, corresponding to 73.17% of the total reads. Using aqueous and sedimentary eDNA, 57 and 88 fish species were detected, respectively, from which 30 and 34 species, respectively, of freshwater and brackish water fish were used for further analysis (Tables 1 and 2).
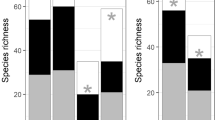
For aqueous eDNA, the number of species from eDNA metabarcoding increased with water volume. However, the number of detected species did not increase significantly at more than 1000 mL [ANOVA: p < 0.05, post hoc Tukey–Kramer test: p > 0.05 (among 1000, 2000, and 4000 mL); Fig. 2]. NMDS showed a difference in composition by water volume (PERMANOVA: p < 0.01; Fig. 3). However, when only 1000, 2000, and 4000 mL samples were analyzed, there was no significant difference in species composition (PERMANOVA: p > 0.05; data not shown). Additionally, results did not show any significant difference related to sampling position (PERMANOVA: p > 0.05; Fig. S4). Meanwhile, the number of species from sedimentary eDNA metabarcoding differed among sampling positions (ANOVA: p < 0.05; Fig. 4), with the highest number of species observed at the right position (Tukey–Kramer test: p < 0.05; Fig. 4), and species composition showing a difference with sampling position in NMDS (PERMANOVA: p < 0.001; Fig. 5). Moreover, species composition significantly differed between aqueous and sedimentary eDNA (PERMANOVA: p < 0.05; Fig. S8; Table 3). However, the detected number of species was equivalent between sedimentary eDNA and aqueous eDNA (Wilcoxon test: p > 0.05; Table 3).
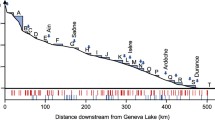
For the whole river, 1,107,750 MiSeq reads were obtained. After the bioinformatics processing steps, 816,807 reads were retained, corresponding to 73.7% of the total reads. Aqueous eDNA detected 68 fish species across the whole river and, after species processing, 33 species were used for analysis (Table 4). Our comparison of these results with those of traditional surveys indicated that eDNA analysis detected more species than the traditional survey (Wilcoxon signed rank test: p < 0.001; Fig. 6; Table 4).
All survey sites were classified into two clusters using nonhierarchical cluster analysis with the k-means method: cluster 1 included sites A, G, and H, while cluster 2 included all other sites (Fig. S5). The Venn diagram showed that the fish community structure of cluster 2 encompassed that of cluster 1 (Fig. S6). Moreover, the species accumulation curve showed that the average cumulative number of detected species reached 95% of all species detected by eDNA metabarcoding after sampling at six sites (Fig. 7).
Discussion
In this study, we showed the effect of filtered water volume, sampling position, and sample type on metabarcoding results within a single site. Based on our results, 1000 mL of water is considered sufficient for fish monitoring in this river, which is inhabited by dozens of fish species. Regarding the number of required sampling sites, the average cumulative number of detected species reached 95% of all species detected by eDNA metabarcoding by taking six samples within the major cluster. In addition, the detected number of species in sedimentary eDNA was equivalent to that of aqueous eDNA. However, sedimentary eDNA showed spatially heterogeneous species composition despite the short distance among sampling positions.
Biodiversity monitoring results from aqueous eDNA metabarcoding were consistent between sampling positions at a single site (site C). Using aqueous eDNA, there was no significant difference in fish species composition between sampling positions at the same site despite the difference in velocity and sediment particle size distribution between the center and shores (Fig. S4; Table S1). Therefore, aqueous eDNA may have a spatially homogeneous distribution; however, additional validation using more number of samples will be required to make this result more robust. In comparison, sedimentary eDNA showed significant differences in fish species composition among positions at the same site despite the short distance (5 m) between sampling positions (Fig. 5). For sedimentary eDNA metabarcoding at site C, the detected number of species was different between the right and the other two positions (Fig. 4); however, the physical characteristics of both shores were similar (Table S1). Therefore, sedimentary eDNA may be spatially heterogeneous in distribution. In this case, several samples are needed to compensate for such spatial variation. Additional information is required regarding the variations in detected species using sedimentary eDNA, potentially on a site-by-site basis.
Physical environment differences may affect the heterogeneity of detected species in sedimentary eDNA more than in aqueous eDNA. In the three positions at site C, the velocity at the center was faster than that at both shores, and larger sediment was present in the center (Table S1). Although it has been reported that aqueous eDNA is influenced by stream velocity and substrate (Jerde et al. 2016; Wilcox et al. 2016; Shogren et al. 2017), the number of species and fish composition detected in the aqueous eDNA showed no difference between positions despite the difference in velocity and substrate size. Contrastingly, more fish species were detected using sedimentary eDNA at the right than at the center and left. However, there were no clear differences in the physical environment between the right and left shore. The difference in detected species between the right and left sampling points might be caused by other environmental parameters not measured in this study, such as the amounts of minerals and organic matter in the sediment, which potentially affect DNA sorption to the sediment (Kanbar et al. 2020). Alternatively, this result may be caused by differences in microbial abundance or chlorophyll concentration, which affect eDNA persistence (Barnes et al. 2014). From these results, aqueous eDNA, which seems to be homogeneously distributed, is less affected by the physical environment, whereas sedimentary eDNA seems to be more heterogeneously distributed, possibly owing to the influence of the physical environment.
Sedimentary eDNA metabarcoding may be a complementary method to aqueous eDNA metabarcoding. Although the number of detected species from sedimentary eDNA was equivalent to that from aqueous eDNA, the species composition differed between them (Fig. S8; Table 3; Siegenthaler et al. 2019). For example, the benthic fish Rhinogobius flumineus and Odontobutis obscura were only detected in sediment samples (Table 3), suggesting that eDNA released by benthic fish may be more detectable from the sediment. However, because DNA extraction methods varied between sample types, eDNA yields and quality, such as the average lengths of collected eDNA, may differ. Therefore, care should be taken when comparing them. Overall, for wide-scale biodiversity monitoring, surveying through water samples is effective and easy because aqueous eDNA would be homogeneously distributed despite differences in fish habitat at a site. In addition, eDNA metabarcoding using sediment samples may detect some species that were not detected by aqueous eDNA alone. Therefore, the most effective eDNA metabarcoding methods to obtain the maximum number of species will include both aqueous and sedimentary eDNA.
In this river, 1000 mL water samples were sufficient to detect fish species through eDNA metabarcoding. The number of species detected using water samples of 500 mL or less was lower than that in samples of 1000 mL or more (Fig. 2). Additionally, the number of species did not vary between water volumes of 1000 mL or more (Fig. 2). In our comparison of fish species composition, composition tended to be similar as water volume increased (Fig. 3), and those obtained from water samples of 1000 mL or more showed no difference among them. These results suggested that a water sample of 1000 mL is sufficient to investigate the number and composition of fish species for biomonitoring in a river that is inhabited by dozens of species. Using 1000 mL water samples for eDNA metabarcoding, we could detect almost all fish species identified by traditional survey methods (Table 4). However, previous studies have shown that detection rates might vary by the number of species in the study area or a combination of target species and the types of environments such as lentic or lotic (Mächler et al. 2016; Bylemans et al. 2018); Therefore, it will be necessary to assess the water volume required for exhaustive detection in each ecosystem.
For biodiversity monitoring, aqueous eDNA metabarcoding was as effective as traditional surveys. In comparing eDNA metabarcoding with 1000 mL water samples and traditional surveys, eDNA metabarcoding provided a higher number of detected species at each site (Fig. 6; Table 4). Fish species captured in the traditional survey included several common species such as Cyprinus carpio, Carassius spp., and Misgurnus anguillicaudatus. Traditional surveys were performed primarily using a hand net and casting net. Thus, fast swimming species such as Tribolodon brandtii maruta (detected only by eDNA metabarcoding in this study) may have been difficult to catch. Furthermore, nocturnal species such as Silurus asotus were detected at more sites with eDNA metabarcoding than with traditional survey methods. In contrast, Lefua echigonia was only detected by traditional survey. Species with localized habitat requirements and low population size may be less likely to be detected by eDNA metabarcoding. Such a small population may be difficult to detect owing to the small concentration of released eDNA and short transport distance (Nukazawa et al. 2018). Therefore, although aqueous eDNA metabarcoding is effective for biodiversity monitoring, combining it with a traditional survey or sediment samples that can better detect benthic fish could be effective in improving monitoring accuracy.
In this study, we presented the usefulness of eDNA metabarcoding compared to that of traditional survey methods. However, following eDNA detection, traditional surveys are important for confirming that the detected species truly inhabit the area (Sakata et al. 2017) because eDNA has a risk of false positives (Ficetola et al. 2016; Guillera-Arroita et al. 2017). Moreover, although previous studies have identified the effects of external water quality factors such as pH and temperature on eDNA (Strickler et al. 2015; Eichmiller et al. 2016; Andruszkiewicz et al. 2017; Kessler et al. 2019), only a few elaborate on how such factors affect eDNA metabarcoding results. Therefore, future studies should focus on clarifying the effects of not only water volume but also environmental factors such as water quality or the presence of PCR inhibitors on metabarcoding results such as detected number of species, composition, and differences in detection.
Although 1000 mL water sampling was considered sufficient for surveying our study river containing dozens of species, a greater water volume is needed to detect hundreds of target species (Cantera et al. 2019; Bessey et al. 2020). Therefore, the required filtered water volume depends partly on the number of species in the study area. In addition, considering the number of sample replicates may be important to reduce sampling effort (Evans et al. 2017; Cantera et al. 2019; Doi et al. 2019). To fully assess the fish fauna in the entire river, it is also important to consider the transportation distance of aqueous eDNA by water flow (Deiner and Altermatt 2014; Jane et al. 2015; Shogren et al. 2017). Therefore, to use eDNA metabarcoding efficiently for monitoring, it will be necessary to consider filtered water volume, sample replicates, and the distance between sampling sites for each survey area. In addition, in environments such as subtropical habitats, where several hundred fish species occur, estimates of species richness based on monitoring results can be used to predict the necessary number of samples (Oka et al. 2020). Such consideration will also be important for eDNA-based biomonitoring in the future.
In this study, the average cumulative number of detected species from water sampling at six sites within the major cluster provided 95% of all species detected by eDNA metabarcoding in all 11 sites. Furthermore, the grouping and visualization of fish communities within each cluster may help determine representative sites in the study area. In addition, the species accumulation curve can suggest a reasonable amount of effort required for survey, as previously suggested (Sato et al. 2017; Sigsgaard et al. 2019; Bessey et al. 2020). However, it is difficult to set a general rule regarding a reasonable amount of effort required because the number of species and species composition is different in each river. Furthermore, the results of such assessments may only be applicable during certain seasons as the sampling season affects eDNA metabarcoding results (Hayami et al. 2020).
In addition, sampling at downstream sites is important when the eDNA survey is performed on a river because released eDNA is transported in stream systems (Deiner and Altermatt 2014; Shogren et al. 2017; Carraro et al. 2018). Our results showed that fish composition at downstream sites (Sites D, E, and F) almost included those of upstream sites (the other site) (see Table 4). This result seemed to be caused by the downstream transportation eDNA, as shown in previous studies (Deiner and Altermatt 2014; Shogren et al. 2017; Carraro et al. 2018). Therefore, it may also be important for sampling design to consider focusing on downstream sites to detect representative fish species of the river. However, to detect a rare species only inhabiting upstream sites, sampling at the downstream site and sampling upstream may be required. Thus, the sampling design should be adapted for the purpose of the monitoring.
Overall, by considering three parameters (sample type, sampling position, and water volume), we could determine the sampling method that would maximize the number of detected species at a single survey site. In addition, considering the number of sampling sites will allow for more cost-effective eDNA biomonitoring. We showed that sedimentary eDNA is spatially heterogeneous in distribution and may complement aqueous eDNA metabarcoding by detecting different fish species. Examination of ecosystem-specific sampling methods at a single site and on a number of sampling sites, as performed in this study, will be important prior to large-scale or long-term surveys, potentially allowing to increase biodiversity monitoring efficiency through eDNA metabarcoding.
References
Andruszkiewicz EA, Sassoubre LM, Boehm AB (2017) Persistence of marine fish environmental DNA and the influence of sunlight. PLoS ONE 12(9):e0185043. https://doi.org/10.1371/journal.pone.0185043
Barnes MA, Turner CR, Jerde CL et al (2014) Environmental conditions influence eDNA persistence in aquatic systems. Environ Sci Technol 48:1819–1827. https://doi.org/10.1021/es404734p
Bessey C, Jarman SN, Berry O et al (2020) Maximizing fish detection with eDNA metabarcoding. Environ DNA. https://doi.org/10.1002/edn3.74
Bista I, Carvalho GR, Walsh K et al (2017) Annual time-series analysis of aqueous eDNA reveals ecologically relevant dynamics of lake ecosystem biodiversity. Nat Commun 8:14087. https://doi.org/10.1038/ncomms14087
Butchart SHM, Walpole M, Collen B et al (2010) Global biodiversity: indicators of recent declines. Science 328(5982):1164–1168. https://doi.org/10.1126/science.1187512
Bylemans J, Gleeson DM, Lintermans M et al (2018) Monitoring riverine fish communities through eDNA metabarcoding: determining optimal sampling strategies along an altitudinal and biodiversity gradient. Metabarcoding Metagenomics 2:1–12. https://doi.org/10.3897/mbmg.2.30457
Cantera I, Cilleros K, Valentini A et al (2019) Optimizing environmental DNA sampling effort for fish inventories in tropical streams and rivers. Sci Rep 9:3085. https://doi.org/10.1038/s41598-019-39399-5
Carraro L, Hartikainen H, Jokela J et al (2018) Estimating species distribution and abundance in river networks using environmental DNA. Proc Natl Acad Sci 115(46):11724–11729. https://doi.org/10.1073/pnas.1813843115
Deiner K, Altermatt F (2014) Transport distance of invertebrate environmental DNA in a natural river. PLoS ONE 9(2):e88786. https://doi.org/10.1371/journal.pone.0088786
Deiner K, Bik HM, Mächler E et al (2017) Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26:5872–5895. https://doi.org/10.1111/mec.14350
Doi H, Fukaya K, Oka S et al (2019) Evaluation of detection probabilities at the water-filtering and initial PCR steps in environmental DNA metabarcoding using a multispecies site occupancy model. Sci Rep 9:3581. https://doi.org/10.1038/s41598-019-40233-1
Dudgeon D, Arthington AH, Gessner MO et al (2006) Freshwater biodiversity: importance, threats, status and conservation challenges. Biol Rev Camb Philos Soc 81:163–182. https://doi.org/10.1017/S1464793105006950
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Eichmiller JJ, Best E, Sorensen PW et al (2016) Effects of temperature and trophic state on degradation of environmental DNA in lake water. Environ Sci Technol 50:1859–1867. https://doi.org/10.1021/acs.est.5b05672
Evans NT, Li Y, Renshaw MA et al (2017) Fish community assessment with eDNA metabarcoding: effects of sampling design and bioinformatic filtering. Can J Fish Aquat Sci 1374:1–13. https://doi.org/10.1139/cjfas-2016-0306
Ficetola GF, Taberlet P, Coissac E (2016) How to limit false positives in environmental DNA and metabarcoding? Mol Ecol Resour 16:604–607. https://doi.org/10.1111/1755-0998.12508
Guillera-Arroita G, Lahoz-Monfort JJ, van Rooyen AR et al (2017) Dealing with false-positive and false-negative errors about species occurrence at multiple levels. Methods Ecol Evol 8:1081–1091. https://doi.org/10.1111/2041-210X.12743
Hänfling B, Lawson Handley L, Read DS et al (2016) Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol Ecol 25(13):3101–3119. https://doi.org/10.1111/mec.13660
Hayami K, Sakata MK, Inagawa T et al (2020) Effects of sampling seasons and locations on fish environmental DNA metabarcoding in dam reservoirs. Ecol Evol 10(12):5354–5367. https://doi.org/10.1002/ece3.6279
Jacobsen CS, Nielsen TK, Vester JK et al (2018) Inter-laboratory testing of the effect of DNA blocking reagent G2 on DNA extraction from low-biomass clay samples. Sci Rep 8:5711. https://doi.org/10.1038/s41598-018-24082-y
Jane SF, Wilcox TM, Mckelvey KS et al (2015) Distance, flow and PCR inhibition: eDNA dynamics in two headwater streams. Mol Ecol Resour 15:216–227. https://doi.org/10.1111/1755-0998.12285
Japanese Industrial Standards Committee (2009) Test method for particle size distribution of soils (JIS A1204)
Jerde CL, Olds BP, Shogren AJ et al (2016) Influence of stream bottom substrate on retention and transport of vertebrate environmental DNA. Environ Sci Technol 50:8770–8779. https://doi.org/10.1021/acs.est.6b01761
Kanbar HJ, Olajos F, Englund G, Holmboe M (2020) Geochemical identification of potential DNA-hotspots and DNA-infrared fingerprints in lake sediments. Appl Geochem. https://doi.org/10.1016/j.scitotenv.2019.135577
Kessler EJ, Davis MA, Ash KT et al (2019) Radiotelemetry reveals effects of upstream biomass and UV exposure on environmental DNA occupancy and detection for a large freshwater turtle. Environ DNA 2(1):13–23. https://doi.org/10.1002/edn3.42
Kimura K, Saitou K, Moriue Y (2015) Third report of freshwater fish fauna in Chigasaki City. Bull Chigasaki City Museum Heritage 24:21–46.
Kouduka M, Suko T, Morono Y et al (2012) A new DNA extraction method by controlled alkaline treatments from consolidated subsurface sediments. FEMS Microbiol Lett 326:47–54. https://doi.org/10.1111/j.1574-6968.2011.02437.x
Mächler E, Deiner K, Spahn F, Altermatt F (2016) Fishing in the water: effect of sampled water volume on environmental DNA-based detection of macroinvertebrates. Environ Sci Technol 50:305–312. https://doi.org/10.1021/acs.est.5b04188
Minamoto T, Hayami K, Sakata MK, Imamura A (2019) Real-time polymerase chain reaction assays for environmental DNA detection of three salmonid fish in Hokkaido, Japan: application to winter surveys. Ecol Res 34:237–242. https://doi.org/10.1111/1440-1703.1018
Miya M, Sato Y, Fukunaga T et al (2015) MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. R Soc Open Sci 2:150088. https://doi.org/10.1098/rsos.150088
Miya M, Minamoto T, Yamanaka H et al (2016) Use of a filter cartridge for filtration of water samples and extraction of environmental DNA. J Vis Exp 117:e54741. https://doi.org/10.3791/54741
Nakagawa H, Yamamoto S, Sato Y et al (2018) Comparing local- and regional-scale estimations of the diversity of stream fish using eDNA metabarcoding and conventional observation methods. Freshw Biol 63:569–580. https://doi.org/10.1111/fwb.13094
Nukazawa K, Hamasuna Y, Suzuki Y (2018) Simulating the advection and degradation of the environmental DNA of common carp along a river. Environ Sci Technol 52:10562–10570. https://doi.org/10.1021/acs.est.8b02293
Oka S, Doi H, Miyamoto K et al (2020) Environmental DNA metabarcoding for biodiversity monitoring of a highly diverse tropical fish community in a coral reef lagoon: Estimation of species richness and detection of habitat segregation. Environ DNA. https://doi.org/10.1002/edn3.132
R Core Team (2019). R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. Retrieved from https://www.R-project.org/
Sakata MK, Maki N, Sugiyama H, Minamoto T (2017) Identifying a breeding habitat of a critically endangered fish, Acheilognathus typus, in a natural river in Japan. Sci Nat 104:100. https://doi.org/10.1007/s00114-017-1521-1
Sakata MK, Yamamoto S, Gotoh RO et al (2020) Sedimentary eDNA provides different information on timescale and fish species composition compared with aqueous eDNA. Environ DNA 2(4):505–518 https://doi.org/10.1002/edn3.75
Sato H, Sogo Y, Doi H, Yamanaka H (2017) Usefulness and limitations of sample pooling for environmental DNA metabarcoding of freshwater fish communities. Sci Rep 7:14860. https://doi.org/10.1038/s41598-017-14978-6
Schultz MT, Lance RF (2015) Modeling the sensitivity of field surveys for detection of environmental DNA (eDNA). PLoS ONE 10(10):e0141503. https://doi.org/10.1371/journal.pone.0141503
Shaw JLA, Clarke LJ, Wedderburn SD et al (2016) Comparison of environmental DNA metabarcoding and conventional fish survey methods in a river system. Biol Conserv 197:131–138. https://doi.org/10.1016/j.biocon.2016.03.010
Shogren AJ, Tank JL, Andruszkiewicz E et al (2017) Controls on eDNA movement in streams: transport, retention, and resuspension /704/158/2464 /704/242 /45/77 article. Sci Rep 7:5065. https://doi.org/10.1038/s41598-017-05223-1
Siegenthaler A, Wangensteen OS, Soto AZ et al (2019) Metabarcoding of shrimp stomach content: harnessing a natural sampler for fish biodiversity monitoring. Mol Ecol Resour 19:206–220. https://doi.org/10.1111/1755-0998.12956
Sigsgaard EE, Torquato F, Frøslev TG et al (2019) Using vertebrate environmental DNA from seawater in biomonitoring of marine habitats. Conserv Biol 34(3):697–710. https://doi.org/10.1111/cobi.13437
Strickler KM, Fremier AK, Goldberg CS (2015) Quantifying effects of UV-B, temperature, and pH on eDNA degradation in aquatic microcosms. Biol Conserv 183:85–92. https://doi.org/10.1016/j.biocon.2014.11.038
The eDNA Society (2019) Environmental DNA sampling and experiment manual. Version 2.1. https://ednasociety.org/eDNA_manual_Eng_v2_1_3b.pdf
Thomsen PF, Kielgast J, Iversen LL et al (2012) Monitoring endangered freshwater biodiversity using environmental DNA. Mol Ecol 21:2565–2573. https://doi.org/10.1111/j.1365-294X.2011.05418.x
Turner CR, Uy KL, Everhart RC (2015) Environmental DNA fish environmental DNA is more concentrated in aquatic sediments than surface water. Biol Conserv 183:93–102. https://doi.org/10.1016/j.biocon.2014.11.017
Valentini A, Taberlet P, Miaud C et al (2016) Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol Ecol 25:929–942. https://doi.org/10.1111/mec.13428
Wilcox TM, McKelvey KS, Young MK et al (2016) Understanding environmental DNA detection probabilities: a case study using a stream-dwelling char Salvelinus fontinalis. Biol Conserv 194:209–216. https://doi.org/10.1016/j.biocon.2015.12.023
WWF (2018) Living planet report—2018: aiming higher. https://www.worldwildlife.org/pages/living-planet-report-2018
Yamamoto S, Masuda R, Sato Y et al (2017) Environmental DNA metabarcoding reveals local fish communities in a species-rich coastal sea. Sci Rep 7:40368. https://doi.org/10.1038/srep40368
Yamanaka H, Minamoto T, Matsuura J et al (2017) A simple method for preserving environmental DNA in water samples at ambient temperature by addition of cationic surfactant. Limnology 18:233–241. https://doi.org/10.1007/s10201-016-0508-5
Acknowledgements
We would like to thank Mr. R. Osawa (Kobe University) for assistance with filtration. This study was partly supported by the Japan Society for the Promotion of Science (JSPS; KAKENHI Grant Numbers 19J11126 and 20H03326) and by a fund from a private company to which five of the authors (T.W., N.M., K.I., T.K., and H.O.) belong.
Author information
Authors and Affiliations
Contributions
MKS, TW, and TM conceived and designed the research. NM, KI, TK, and HO collected samples and performed filtration. MKS, HY, TS, and MM performed the experiments, along with bioinformatic and statistical analyses. MKS wrote the first draft of the manuscript. All authors discussed the results and contributed to the development of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
T.W., N.M., K.I., T.K., and H.O. belong to a private company.
Research involving animal rights
No animal experiments were performed in this study. All experiments were performed according to the current laws of Japan.
Additional information
Handling Editor: Teruhiko Takahara.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sakata, M.K., Watanabe, T., Maki, N. et al. Determining an effective sampling method for eDNA metabarcoding: a case study for fish biodiversity monitoring in a small, natural river. Limnology 22, 221–235 (2021). https://doi.org/10.1007/s10201-020-00645-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10201-020-00645-9