Abstract
Background
Published data indicate that common genetic variants in immune/inflammatory response genes can affect the outcome of diffuse large B-cell lymphomas (DLBCL). This study investigated the association of interleukin (IL)-10 (−3575, −1082), tumor necrosis factor (TNF)-α −308 and transforming growth factor (TGF)-β Leu10Pro gene polymorphisms with clinical characteristics and outcome of DLBCL patients treated with rituximab–CHOP therapy.
Methods
Between January 2004 and December 2007, a total of 84 patients with newly diagnosed DLBCL entered into this study. Genotypes were determined with PCR-based methodology.
Results
Patients presenting with B symptoms had IL-10 −3575 TA/AA genotypes more frequently than TT genotype [odds ratio (OR) 2.89, 95 % confidence interval (CI) 1.11–7.57; p = 0.03]. Carriers of TGF-β Pro10 allele more frequently had an advanced clinical stage III/IV (OR 4.65, 95 % CI 1.33–16.19; p = 0.016) and intermediate-high/high IPI score (OR 5.37, 95 % CI 1.45–20.0; p = 0.012). In rituximab–CHOP-treated patients (n = 64), the TNF-α −308 AG/AA carriers had shorter overall (p = 0.048) and event-free survival (p = 0.07) compared to GG carriers. In multivariate analysis of prognostic factors for survival, the TNF-α AG/AA genotypes were significantly associated with inferior survival of lymphoma patients (OR 0.23, 95 % CI 0.07–0.78; p = 0.018).
Conclusion
Our results indicate the association of IL-10 −3575 and TGF-β Leu10Pro gene variations with clinical characteristics. In patients treated with rituximab–CHOP therapy, the TNF-α −308 AG/AA genotypes showed a significantly less favorable survival than the GG genotype.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse large B-cell lymphomas (DLBCL) represent a heterogeneous group of lymphoproliferative disorders with highly variable clinical course and outcome [1]. The addition of rituximab to CHOP (R-CHOP) or CHOP-like chemotherapy has markedly improved the outcome of all subgroups of patients with DLBCL and has established R-CHOP therapy as the standard of care in DLBCL [2–5]. Despite significant advances in immunochemotherapy, response to treatment is heterogeneous, ranging from cure to treatment failure, relapse, and death. This variability in clinical course and outcome is largely due to biological differences between tumors and patients. The etiology of DLBCL remains unknown although there are data that support a role of genetic and immune-related factors in the pathogenesis of lymphoma [6, 7].
The balance of different cytokine signals plays an important role in the regulation of normal immune function. Some evidence indicates that the disturbance of this balance participates in development or evolution of numerous immune-regulated disorders, including lymphoma [8].
Several studies have suggested that cytokine production is influenced by genetic factors [9, 10]. Recently, many single-nucleotide polymorphisms (SNP) were detected within cytokine genes, particularly within their promoter regions [11]. Some of these polymorphisms may be associated with different levels of cytokine expression and also modulate expression of other cytokines involved in the immune response. Numerous studies have provided evidence for the role of interleukin (IL)-10, tumor necrosis factor (TNF)-α and transforming growth factor (TGF)-β in the pathogenesis of B-cell lymphomas [12–15]. These cytokines exploit different models to exert their effects in immunoregulation and inflammation. They influence, in a paracrine or autocrine manner, growth and survival of normal and malignant cells, including B lymphocytes [16–18]. These cytokines can also contribute to a drug-resistant phenotype in many tumors [19]. Considering their properties, a number of studies investigated the association between IL-10, TNF-α and TGF-β polymorphisms and lymphoma susceptibility or prognosis [20–27].
The aim of the present study was to investigate the association of genetic variation within IL-10 (−3575, −1082), TNF-α −308 and TGF-β Leu10Pro genes with clinical features at presentation and outcome of DLBCL patients treated with R-CHOP therapy.
Materials and methods
Patients
This study included a total of 84 patients with DLBCL who were diagnosed and treated at the Clinic of Hematology, MMA, Belgrade, Serbia between January 2004 and December 2007. Diagnosis was based on histopathology and immunohistochemistry according to the World Health Organization (WHO) classification [1]. The patients had mandatory baseline examinations that included clinical examination, laboratory tests, chest radiograph, computer tomography of chest and abdomen, and a bone marrow biopsy. The extent of the disease was categorized according to the Ann Arbor classification and the risk score was determined by the International Prognostic Index [28].
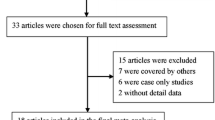
Samples of 94 patients were eligible for the study. After excluding patients with previous history of low-grade lymphoma (7 patients), cancer (2 patients), and HIV-related DLBCL (1 patient), the final study population consisted of 84 patients (aged 22–76 years, median 48).
Among the 84 patients with DLBCL, 20 patients received CHOP (6–8 cycles), and 64 patients received R-CHOP therapy (6–8 cycles). After R-CHOP therapy, radiotherapy was applied in 15 (23.4 %) patients with bulky or residual masses. Response to the therapy was assessed using the International Working Group criteria [29]. Patients were systematically followed up from diagnosis to September 2010. The overall characteristics of the 84 patients eligible for this study and the 64 R-CHOP-treated patients are shown in Table 1.
Informed consent was obtained from all DLBCL patients or patients’ close relatives. The study was approved by the Ethics Committee of MMA.
Methods
Extraction of DNA
Each patient’s DNA was extracted from whole blood by Blood Prep™ Chemistry for ABI PRISM™ 6100 Nucleic Acid PrepStation (Applied Biosystems, USA).
TNF-α −308 genotyping
TNF-α −308 genotypes were determined by restriction fragment length polymorphism (RFLP) of polymerase chain reaction (PCR) products [30]. PCR products were digested with restriction endonuclease NcoI according to the manufacturer’s recommendations (Fermentas, Lithuania). Digested DNA was analyzed on 10 % polyacrylamide gels (PAGs) after electrophoresis and silver nitrate staining [31]. G at position 308 was represented with the presence of two bands (325 and 20 bp), while A at 308 was visualized as a 345-bp band on the gel. All samples were analyzed in duplicate.
IL-10 −1082, IL-10 −819 and TGF-β Leu10Pro genotyping
The genotyping for the single-base-pair polymorphisms at these loci was performed by an amplification refractory mutation system (ARMS) PCR method previously described by Perrey and co-workers [32]. This method involves a specific sense primer complementary to the wild-type allele or a sense primer complementary to the variant allele in combination with the generic antisense primer. For each sample and each locus, two parallel PCR reactions were performed. As an internal control, primers amplifying a human growth hormone sequence were added to each PCR reaction. The presence/absence of PCR products was visualized on 2 % agarose gels after electrophoresis and ethidium bromide staining.
For each of these gene loci we chose ten samples (blind controls) and repeated the analysis.
IL-10 −3575 genotyping
IL-10 −3575 genotypes were determined by TaqMan® SNP genotyping assay rs1800890 (Applied Biosystems, USA) on a 7500 Real Time PCR System (Applied Biosystems, USA). All samples were analyzed in duplicate.
Statistical analysis
The association between SNP genotypes and clinical characteristics at diagnosis and outcome were analyzed using the chi-squared test. The prognostic relevance of different parameters was examined by stepwise logistic regression analysis. Event-free survival (EFS) was defined as the time from first day of treatment to progressive disease under therapy (PD), relapse or death from lymphoma. Overall survival (OS) was defined as the time from first day of treatment to death from any cause or to date of last follow-up. Survival curves were generated using the method of Kaplan and Meier and compared by the log-rank test. The Cox proportional hazards model was used to estimate effect of gene polymorphisms along with DLBCL characteristics for OS. The Cox model included only parameters that showed statistical significance in univariate analyses.
Statistical analyses were performed using SPSS software for Windows, version 15 (SPSS, Inc., Chicago, IL, USA). p < 0.05 was considered to indicate statistical significance.
Results
Genotype analysis and clinical characteristics
The observed genotype frequencies IL-10 (−3575, −1082), TNF-α and TGF-β were in Hardy–Weinberg equilibrium.
Factors examined for prognostic significance included age (≤60years vs. >60 years), sex, Ann Arbor stage (I/II vs. III/IV), B symptoms, bulky disease, and IPI risk score (low/low–intermediate vs. intermediate–high/high). Table 2 summarizes the distribution of genotypes with respect to the clinical characteristics of disease.
IL-10 −1082 A/G and TNF-α −308 G/A polymorphisms were not associated with established prognostic factors (shown in Table 2). We observed an association between IL-10 −3575 and TGF-β Leu10 Pro polymorphism and clinical characteristics of disease. For IL-10 −3575 polymorphism, patients presenting with B symptoms had TA/AA genotypes (70.7 %) more frequently than TT genotype (45.5 %) [odds ratio (OR) 2.89, 95 % confidence interval (CI) 1.11–7.57; p = 0.03]. Analyses of the distribution of TGF-β genotypes with respect to the stage and IPI score have shown significantly more frequent stage III/IV (OR 4.65, 95 % CI 1.33–16.19; p = 0.016) and intermediate–high/high IPI score (OR 5.37, 95 % CI 1.45–20.0; p = 0.012) in patients with the Pro allele variant (LeuPro/ProPro genotypes) than in carriers of the LeuLeu genotype.
Genotype analysis and outcome of disease
We analyzed the association of IL-10, TNF-α and TGF-β gene polymorphisms with the outcome (OS, EFS) in the cohort of 64 DLBCL patients treated with R-CHOP therapy.
In this group of patients, 46 (73 %) achieved complete remission (CR), 11 (17.5 %) experienced disease progression during therapy (PD) and 15 (23.4 %) patients died. With a median follow-up of 36 months (range, 0.5–77), the 3-year OS and EFS for all R-CHOP-treated patients were 71.7 % (95 % CI 63.7–79.7) and 59.0 % (95 % CI 49.5–68.5), respectively.
Analysis of genotypes and outcome did not show any association of IL-10 (− 3575, −1082) and TGF-β Leu10Pro polymorphisms with outcome for patients with DLBCL (data not shown).
However, we observed an association between the TNF-α –308 G/A polymorphism and outcome in the 64 R-CHOP treated patients. The carriers of the GG genotype showed higher sensitivity to R-CHOP therapy. Clinical characteristics of GG and GA/AA carriers and their response to therapy are shown in Table 3. Patients with GA/AA genotypes had significantly decreased OS compared to homozygous G-allele carriers (3-year OS 61.3 % for AG/GG vs. 82.5 % for GG genotype; p = 0.048) (Fig. 1b). With regard to EFS, there was a trend toward a less favorable outcome for GA/AA genotypes (3-year EFS 48.1 %) than GG genotype (3 years 70 %), but statistical significance was not reached (p = 0.07) (Fig. 1a).
In multivariate analysis of prognostic factors for survival, including the clinical characteristics (IPI ≥3, B symptoms, male sex) and TNF-α −308 G/A, GA/AA genotypes (OR 0.23; 95 % CI 0.07–0.78; p = 0.018) and advanced IPI score (OR 0.08; 95 % CI 0.02–0.43; p = 0.003) were significantly associated with the OS (Table 4).
We also analyzed the association between the polymorphism combinations and the survival of R-CHOP-treated patients. The results obtained were not statistically significant, probably due to the small group of patients.
Discussion
The present study examined the association of IL-10 (−3575, −1082), TNF-α −308 and TGF-β Leu10Pro polymorphisms with the clinical characteristics and outcome of DLBCL patients treated with R-CHOP therapy.
Published data indicate that common genetic variants in immune/inflammatory response genes are associated with the risk of developing lymphomas [21–24]. The largest multicentre epidemiological study identified TNF-α −308 G/A and IL-10 −3575 T/A as a risk polymorphism for development of DLBCL [20]. However, only a limited number of studies have analyzed the clinical characteristics and treatment outcome of DLBCL in association with IL-10, TNF-α and TGF-β polymorphisms [25–27, 33].
To date, only a few studies have been published that addressed the association of TNF-308 polymorphism and prognosis of patients with DLBCL [26, 33]. Warzocha and co-workers [26] reported that an extended haplotype in TNF and LT-α was associated with higher TNF production at the time of lymphoma diagnosis, and DLBCL patients with two or more TNF/LT-α high-producing alleles had lower progression-free survival and OS. In our study, we observed a relationship between TNF-α −308 G/A polymorphism and outcome of 64 patients treated with R-CHOP. We found significantly worse survival in patients treated with R-CHOP who had AG/AA genotypes than carriers of the TNF-α −308 GG genotype. The TNF-α −308 A allele is associated with higher constitutional and inducible expression of TNF-α [9]. In-vitro studies have suggested that high levels of TNF-α can reduce cellular sensitivity to apoptosis-inducing chemotherapeutic agents and contribute to the emergence of drug-resistant disease [34]. However, it has been reported that rituximab inhibits the constitutive NF-κB signaling pathway in selected non-Hodgkin lymphoma B-cell lines, leading to increased sensitivity to chemotherapy [35]. Results of studies that have investigated the influence of TNF-α and LT-α gene polymorphisms on infections and predisposition to death from sepsis suggested that patients with high-producing genotypes can manifest significantly more severe inflammatory reactions [36]. However, in our patients R-CHOP therapy was not accompanied by pronounced toxic effects or life-threatening infections or required a significant delay in immunochemotherapy. In the present study, carriers of TNF-α GG showed higher sensitivity to R-CHOP therapy than carriers of the AG/AA genotypes. In addition, patients with AG/AA genotypes showed significantly more resistant/PD during the early course of therapy and died from lymphoma progression within the first 18 months.
Furthermore, we observed a positive association between IL-10 −3575 TA/AA genotypes and the presence of B symptoms in DLBCL patients. Several reports have demonstrated that increased serum levels of IL-10 were associated with poor outcome and adverse prognostic factors in patients with aggressive non-Hodgkin lymphoma [25, 37]. However, previous in-vitro studies have suggested that the IL-10 −3575 A allele is associated with low, and the T allele with high production of IL-10 [38]. There is only one study on DLBCL, published by Kube et al. [27], reporting that the IL-10 −3575 AA carriers have an increased risk of displaying intermediate–high/high IPI score. In addition, we found no difference in OS or EFS in relation to IL-10 −3575 T/A and IL-10 −1082 A/G polymorphism. Similar findings were also reported by other investigators [27, 39, 40].
Several polymorphisms in the TGF-β gene have been reported [41]. Dunning and co-workers [42] found that TGF-β Pro variant of codon 10 was associated with a higher level of serum TGF-β and increased risk of invasive breast cancer. This and other studies have suggested the importance of the TGF-β Leu10Pro polymorphism for the incidence and clinical course of various diseases [43–45]. To date, only a few studies have analyzed the clinical course and outcome in patients with non-Hodgkin lymphoma in association with TGF-β polymorphisms. In the present study, we found that the Pro variant (LeuPro/ProPro genotypes) of TGF-β codon 10 was associated with the unfavorable phenotypic features of DLBCL—advanced clinical stage III/IV and intermediate–high/high IPI score. The relationship between TGF-β Leu10Pro and more aggressive non-Hodgkin lymphoma had been recognized previously by Mazur and co-workers [46]. However, in both studies, the present one and Mazur et al., TGF-β polymorphism was not a prognostic parameter that affected survival of lymphoma patients.
In summary, the present study included DLBCL patients who were diagnosed, uniformly treated, and controlled in one institution. In this ethnically matched group of patients, the frequencies of the analyzed genotypes were similar to those previously reported for other European populations. Our results indicate an association between IL-10 −3575 and TGF-β Leu10Pro gene polymorphisms and clinical characteristics. Furthermore, TNF-α −308 was associated with treatment outcome of DLBCL patients treated with R-CHOP. According to our knowledge, there are no published studies on the association of cytokine gene polymorphisms and survival of DLBCL patients treated with R-CHOP. Herein, we report that the TNF-α −308 A allele is associated with adverse response to R-CHOP therapy. In addition to the TNF-α −308 A allele, unfavorable IPI score (intermediate–high/high) remained an independent risk factor for survival of the patients analyzed. This finding has possible clinical implications regarding treatment. In a selected group of patients, the use of immunomodulatory agents or TNF inhibitors may be an attractive approach [47]. Preclinical and early clinical data demonstrated a synergistic action of immunomodulatory drugs (especially lenalidomide) and rituximab [48–50]. Considering these findings, immunomodulatory therapy, alone or in combination with rituximab, represents a promising strategy for improving prognosis in DLBCL.
References
Gatter KC, Warnke RA (2001) Diffuse large B-cell lymphoma. In: Jaffe ES, Harris NL, Stein H, Vardiman JW (eds) World Health Organization Classification of Tumours. Pathology and genetics of tumours of haematopoietic and lymphoid tissues. IARC Press, Lyon, pp 171–174
Coiffier B, Thieblemont C, Van Den Neste E et al (2010) Long-term outcome of patients in the LNH-98.5 trial, the first randomized study comparing rituximab-CHOP to standard CHOP chemotherapy in DLBCL patients: a study by the Groupe d’Etudes des Lymphomes de I’Adulte. Blood 116:2040–2045
Habermann TM, Weller EA, Morrison VA et al (2006) Rituximab-CHOP versus CHOP alone or with maintenance rituximab in older patients with diffuse large B-cell lymphoma. J Clin Oncol 24:3121–3127
Pfreundschuh M, Trumper L, Osterborg A et al (2006) CHOP-like chemotherapy plus rituximab versus CHOP-like chemotherapy alone in young patients with good-prognosis diffuse large B-cell lymphoma: a randomised controlled trial by the MabThera International Trial (MInT) Group. Lancet Oncol 7:379–391
Pfreundschuh M, Schubert J, Ziepert M et al (2008) Six versus eight cycles of bi-weekly CHOP-14 with or without rituximab in elderly patients with aggressive CD20+ B-cell lymphomas: a randomised controlled trial (RICOVER-60). Lancet Oncol 9:105–116
Chatterjee N, Hartge P, Cerhan JR et al (2004) Risk of non-Hodgkin’s lymphoma and family history of lymphatic, hematologic, and other cancers. Cancer Epidemiol Biomarkers Prev 13:1415–1421
Alexander DD, Mink PJ, Adami HO et al (2007) The non-Hodgkin lymphomas: a review of the epidemiologic literature. Int J Cancer 120(Suppl 12):1–39
Keen LJ (2002) The extent and analysis of cytokine and cytokine receptor gene polymorphism. Transpl Immunol 10:143–146
Wilson AG, Symons JA, McDowell TL et al (1997) Effects of a polymorphism in human tumor necrosis factor alpha promoter on transcriptional activation. Proc Natl Acad Sci USA 94:3195–3199
Escdale J, Gallagher G, Verweij C et al (1998) Interleukin 10 secretion in relation to human IL-10 locus haplotypes. Proc Natl Acad Sci USA 95:9465–9470
Bidwell J, Keen L, Gallagher G et al (2001) Cytokine gene polymorphism in human disease: on-line databases, supplement 1. Genes Immun 2:61–70
Czarneski J, Lin YC, Chong S et al (2004) Studies in IL-10 knockout mice of the requirement of IL-10 for progression of B-cell lymphoma. Leukemia 18:597–606
Korner H, Cretney E, Wilhelm P et al (2000) Tumor necrosis factor sustains the generalized lymphoproliferative disorder (gld) phenotype. J Exp Med 191:89–96
Karin M, Greten FR (2005) NF-kB: linking inflammation and immunity to cancer development and prognosis. Nat Rev Immunol 5:749–759
Douglas RS, Capocasale RJ, Lamb RJ et al (1997) Chronic lymphocytic leukemia B cells are resistant to the apoptotic effects of transforming growth factor-beta. Blood 89:941–947
Moore KW, deWaal Malefyt R, Coffman RL et al (2001) Interleukin-10 and the interleukin-10 receptor. Annu Rev Immunol 19(68):3–765
Pasparakis M, Alexopulou L, Episkopou V et al (1996) Immune and inflammatory responses in TNFα-deficient mice: a critical requirement for TNFα in formation of primary B cell follicles, follicular dendritic networks and germinal centers, and in the maturation of the humoral immune response. J Exp Med 184:1397–1411
Ruscetti FW, Akel S, Bartelmez SH (2005) Autocrine transforming growth factor-β regulation of hematopoiesis: many outcomes that depend on the context. Oncogene 24:5751–5763
Alas S, Emmanouilides Ch, Bonavida B (2001) Inhibition of interleukin-10 by rituximab results in down-regulation of bcl-2 and sensitization of B-cell non-Hodgkin’s lymphoma to apoptosis. Clin Cancer Res 7:709–723
Rothman N, Skibola CF, Wang SS et al (2006) Genetic variation in TNF and IL10 and risk of non-Hodgkin lymphoma: a report from the Inter-Lymph Consortium. Lancet Oncol 7:27–38
Purdue MP, Lan Q, Kricker A et al (2007) Polymorphisms in immune function genes and risk of non-Hodgkin lymphoma: findings from the New South Wales non-Hodgkin lymphoma study. Carcinogenesis 28:704–712
Lan Q, Zheng T, Rothman N et al (2006) Cytokine polymorphisms in the Th1/Th2 pathway and susceptibility to non-Hodgkin lymphoma. Blood 107:4101–4108
Wang S, Cerhan J, Hartge P et al (2006) Common genetic variants in proinflammatory and other immunoregulatory genes and risk for non-Hodgkin lymphoma. Cancer Res 66:9771–9780
Spink CF, Keen LJ, Mensah FK et al (2006) Association between non-Hodgkin lymphoma and haplotypes in the TNF region. BJ Haematol 133:293–300
Lech-Maranda E, Baseggio L, Bienvenu J et al (2004) Interleukin-10 gene promoter polymorphisms influence the clinical outcome of diffuse large B-cell lymphoma. Blood 103:3529–3534
Warzocha K, Ribeiro P, Bienvenu J et al (1998) Genetic polymorphisms in the tumor necrosis factor locus influence non-Hodgkin’s lymphoma outcome. Blood 91:357–381
Kube D, Hua TD, von Bonin F et al (2008) Effect of interleukin-10 gene polymorphisms on clinical outcome of patients with aggressive non-Hodgkin’s lymphoma: an exploratory study. Clin Cancer Res 14:3777–3784
Shipp MA, Harrington DP, Anderson JR et al (1993) A predictive model for aggressive non-Hodgkin’s lymphoma. N Engl J Med 329:987–994
Cheson BD, Horning SJ, Coiffier B et al (1999) Report of an international workshop to standardize response criteria for non-Hodgkin’s lymphoma. NCI Sponsored International Working Group. J Clin Oncol 17(4):1244–1253
Tang GJ, Huang SL, Yien HW et al (2000) Tumor necrosis factor gene polymorphism and septic shock in surgical infection. Crit Care Med 28:2733–2736
Sambrook J, Fritsch EF, Maniatis T (1989) Preparation of organic reagents. In: Nolan C (ed) Molecular cloning, a laboratory manual. Cold Spring Harbor, Cold Spring Harbor Laboratory Press, pp B4–B5
Perrey C, Turner SJ, Pravica V et al (1999) ARMS-PCR methodologies to determine IL-10, TNF-α, TNF-β, and TGF-β1 gene polymorphisms. Transpl Immunol 7:127–128
Habermann TM, Wang SS, Maurer MJ et al (2008) Host immune gene polymorphisms in combination with clinical and demographic factors predicts late survival in diffuse large B-cell lymphoma patients in the pre-rituximab era. Blood 112:2694–2702
Kobayashi D, Watanabe N, Yamauchi N et al (1997) Endogenous tumor necrosis factor as a predictor of doxorubicin sensitivity in leukemic patients. Blood 89:2472–2479
Jazirehi AR, Huerta-Yepez S, Cheng G et al (2005) Rituximab (chimeric anti CD20 monoclonal antibody) inhibits the constitutive nuclear factor-kB signaling pathway in non-Hodgkin’s lymphoma B-cell lines: role in sensitisation to chemotherapeutic drug-induced apoptosis. Cancer Res 65:264–276
Mira J-P, Cariou A, Grall F et al (1999) Association of TNF2, a TNF-alpha promoter polymorphism, with septic shock susceptibility and mortality. JAMA 282:561–568
Blay J-Y, Burdin N, Rousset F et al (1993) Serum interleukin-10 in non-Hodgkin’s lymphoma: a prognostic factor. Blood 82:2169–2174
Gibson AW, Edberg JC, Wu J et al (2001) Novel single nucleotide polymorphisms in the distal IL-10 promoter affect IL-1-production and enhance the risk of systemic lupus erythematosus. J Immunol 166:3915–3922
Kube D, Hua TD, Kloss M et al (2007) The interleukin-10 gene promoter polymorphism −1087AG does not correlate with clinical outcome in non-Hodgkin’s lymphoma. Genes Immun 8:164–167
Berglund M, Thunberg U, Roos G et al (2005) The interleukin-10 gene promoter polymorphism (−1082) does not correlate with clinical outcome in diffuse large B-cell lymphoma. Blood 105:4894–4895 (author reply 5)
Cambien F, Ricard S, Troesch A et al (1996) Polymorphisms of the transforming growth factor-β1 gene in relation to myocardial infarction and blood pressure: the Etude cas-Temoin de L’infarctus du Myocarde (ECTIM) Study. Hypertension 28:881–887
Dunning AM, Ellis PD, McBride S et al (2003) A transforming growth factor β1 signal peptide variant increases secretion in vitro and is associated with increased incidence of invasive breast cancer. Cancer Res 63:2610–2615
Crilly A, Hamilton J, Clark CJ et al (2002) Analysis of transforming growth factor beta1 gene polymorphisms in patients with systemic sclerosis. Ann Rheum Dis 61:678–681
Arkwright PD, Laurie S, Super M et al (2000) TGF-beta(1) genotype and accelerated decline in lung function of patients with cystic fibrosis. Thorax 55:459–462
Yokota M, Ichihara S, Lin TL et al (2000) Association of a T29>C polymorphism of the transforming growth factor-beta1 gene with genetic susceptibility to myocardial infarction in Japanese. Circulation 101:2783–2787
Mazur G, Bogunia-Kubik K, Wrobel T et al (2006) TGF-β1 gene polymorphisms influence the course of the disease in non-Hodgkin’s lymphoma patients. Cytokine 33:145–149
Padyukov L, Lampa J, Heimburger M et al (2003) Genetic markers for the efficacy of tumor necrosis factor blocking therapy in rheumatoid arthritis. Ann Rheum Dis 62:526–529
Reddy N, Hernandez-Ilizaliturri FJ, Deeb G et al (2008) Immunomodulatory drugs stimulate natural killer-cell function, alter cytokine production by dendritic cells, and inhibit angiogenesis enhancing the anti-tumour activity of rituximab in vivo. Br J Haematol 140:36–45
Ivanov V, Tabouret E, Chuto G et al (2010) Rituximab-lenalidomide-dexamethasone induces complete and durable remission in relapsed refractory diffuse large B-cell non-Hodgkin lymphoma. Leuk Lymphoma 51:1758–1760
Nowakowski GS, LaPlant B, Habermann TM et al (2011) Lenalidomide can be safely combined with R-CHOP (R2 CHOP) in the initial chemotherapy for aggressive B-cell lymphomas: phase I study. Leukemia 25:1877–1881
Conflict of interest
The authors declare that they have no conflict of interest. All financial and material support for this research and work was obtained from Military Medical Academy, Belgrade, Serbia.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Tarabar, O., Cikota-Aleksić, B., Tukić, L. et al. Association of interleukin-10, tumor necrosis factor-α and transforming growth factor-β gene polymorphisms with the outcome of diffuse large B-cell lymphomas. Int J Clin Oncol 19, 186–192 (2014). https://doi.org/10.1007/s10147-013-0531-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-013-0531-z