Abstract
The alterations of the intestinal proteome were observed in intrauterine growth restriction (IUGR) piglets during early life by gel-based approaches. Nevertheless, how IUGR affects the intestinal membrane proteome during neonatal development remains unclear. Here, we applied the iTRAQ-based proteomics technology and biochemical analysis to investigate the impact of IUGR on the membrane proteome of the jejunal mucosa in the piglets. Three hundred sixty-one membrane proteins were screened by functional prediction. Among them, eight, five, and one differentially expressed membrane proteins were identified between IUGR and NBW piglets at day 0, day 7, and day 21 after birth, respectively. Differentially expressed membrane proteins (DEMPs) including F1SBL3, F1RRW8, F1S539, F1S2Z2, F1RIR2, F1RUF2 I3LP60, Q2EN79, and F1SIH8 were reduced while the relative abundance of I3L6A2, F1SCJ1, F1RI18, I3LRJ7, and F1RNN0 were increased in IUGR piglets than NBW piglets. From the aspects of function, F1RRW8, F1S539, F1S2Z2, and F1RIR2 are mainly associated with D2 dopamine receptor binding, transmembrane transport of small molecules, signal transduction, and translocation of GLUT4, respectively, and F1SIH8, I3LRJ7, and F1RNN0 are related to autophagy, metabolism of vitamins, and intracellular protein transport. Additionally, IUGR decreased the level of proteins (F1RRW8, Q2EN79, and F1RI18) that are involved in response to oxidative stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intrauterine growth restriction (IUGR), accounting for approximately 5–10% of human neonates worldwide and is the main mediator of neonatal mortality and morbidity (Li et al. 2017a; Sangild et al. 2006; Tomasz et al. 2007). In the livestock species, neonatal piglets exhibit the most severe naturally occurring IUGR (15–25%) (Li et al. 2017a; Wang et al. 2017). IUGR is defined as the birth weight lower than 10th percentile for the corresponding gestational age, which is derived from impaired growth and development of the mammalian embryo/fetus or organs during the pregnancy (D’Inca et al. 2010a; Wang et al. 2016; Wu et al. 2006). As a major organ for nutrient digestion and absorption (Sangild et al. 2006), as well as defensive barrier against endogenous pathogens and dietary contaminants (Hsueh et al. 2003), the poor intestinal structure and function in IUGR piglets (Regev and Reichman 2004) such as decreased intestinal mass, inferior gastrointestinal tract barriers, antioxidant systems, and immune response (D’Inca et al. 2011), as well as abnormal absorption (Wang et al. 2008), has led to feeding intolerance and elevated risk of morbidity (Baserga et al. 2004; Bozzetti et al. 2013).
Results from our previous studies revealed that the expression level of proteins associated with development reprogramming are affected in the gut of IUGR pigs at birth, during the neonatal stage and even during pregnancy (Wang et al. 2014; Wang et al. 2010; Wang et al. 2018). During birth and lactation (days 1, 7, and 21), the results of our proteomics analysis reveal increased levels of proteins related to oxidative stress and immune response, as well as the decreased abundance of proteins required for digestion, absorption, and metabolism of nutrients (Wang et al. 2010). Thus, the alteration of the proteome in the jejunal mucosa in IUGR piglets predisposes the gut to multiple metabolic abnormalities later in life. Furthermore, from days 60 to 110 of gestation (mid to late gestation) (Wang et al. 2014), our results indicated increased abundances of proteins and enzymes associated with oxidative stress, apoptosis, and protein degradation, as well as decreased abundances of proteins that are required for maintenance of cell structure and motility, absorption and transport of nutrients, energy metabolism, and protein synthesis in the fetal gut.
Based on previous studies, the intestinal mucosa plays important roles in the intestinal barrier, gut microbiota colonization, substance transport, oxidative stress, and immune response in health and during disease (D’Inca et al. 2010b; Glover et al. 2016; Haq et al. 2019). Under normal physiological conditions, the membrane proteins are an important component of the intestinal mucosa and serve a variety of functions, including mucus barrier, substance transport, signal transduction, and immune regulation (Turner 2009). However, none of the prior studies included a global assessment of membrane proteins related to membrane signaling pathways of impaired intestine from IUGR piglets. Meanwhile, compared with iTRAQ-based proteomics, gel-based approaches can only detect proteins with relatively high abundance. There is little knowledge about the mechanism of the impaired intestinal development in IUGR piglets after birth. We hypothesized that the changes in membrane protein expression may reveal its mechanism.
Therefore, the current study endeavored to analyze the protein profiles at D0, D7, D21, and D35 after birth between IUGR and normal birth weight (NBW) piglets and figured out the membrane proteins which are responsible for the differences between groups by using the up-to-date iTRAQ.
Methods and materials
Ethics statement
All experiments were carried out with the approval of China Agricultural University Animal Care and Use Committee (CAU20170114–1, Beijing, China).
Experiment design and sample harvesting
In this study, a total of 16 multiparous sows (Yorkshire, 2~4 parities) were selected and raised in a commercial pig breeding farm in Sichuan province, China. In the experimental period, the selected sows were fed with the same commercial gestation diet and had free access to clean water. Porcine neonates (Landrace × Yorkshire) were spontaneously delivered from sows at 113~114 days of gestation, and their body weights were recorded immediately at birth (D0). A piglet was defined as an IUGR when its body weight was less than two standard deviations (SD) below the mean body weight for gestational age (Wang et al. 2010). In our study, one IUGR piglet with a mean body weight of 0.95 (± 0.04) kg and one NBW with 1.50 (± 0.05) kg were marked in each of the 16 litters (12–14 piglets/litter). The selected piglets (n = 32, 16 IUGR vs. 16 NBW) were housed and nursed by their sow along with the rest of their littermates. All the piglets were weaned at day 21 (D21; before weaning) and transferred into the nursery pens with free access to commercial weaning diet and water. From ages 3 to 5 days, piglets also started to receive commercial creep feed and drinking water ad libitum. None of the piglets was administered antibiotics or other drugs throughout this experiment. On day 0 (D0, before suckling), day 7 (D7), day 21 (D21, before weaning), and day 35 (D35, 2 weeks after weaning) after birth, piglets (4 LBW and 4 NBW piglets) from randomly selected 4 litters were killed after anesthesia for sample collection. The piglets were weighed individually and sacrificed for sampling after fasting for 2~4 h. The preparation and fractionation of total jejunal mucosa from the jejunal segment was performed as previously described (Wang et al. 2008; Wang et al. 2014; Wang et al. 2007; Wang et al. 2018), and then the membrane protein isolation, 50 mg of frozen jejunal mucosa (Fig. 1) from each sample was used. The plasma membrane proteins were isolated according to the Minute Plasma Membrane Protein Isolation kit manufacturer’s protocol as previously described (Invent Biotechnologies, Eden Prairie, MN) (Nemoto et al. 2019; Yu et al. 2018). After membrane fractionation, the membrane protein concentrations were determined according to a Bradford assay, and equal amounts of membrane protein for further experiments. Intestinal mucosa samples for iTRAQ assays were snap frozen with liquid nitrogen and then stored at − 80 °C. The flow of the experiment is presented in Fig. 2a.
Comparative membrane proteomic workflow for the analysis of jejunum mucosa from NBW and IUGR neonatal piglets (a) and results of all identified proteins and membrane proteins (b). a Experimental design and workflow of this study. Four jejunal mucosa samples from each group per treatment were labeled with different iTRAQ tags and used for MS analysis in both NBW and IUGR piglets at D0 (a), D7 (B), D21 (c), and D35 (d) after birth. B The total number of all identified proteins and membrane proteins from the NBW and IUGR piglets at D0, D7, D21, and D35 after birth
Histomorphology analysis
The paraformaldehyde-fixed mid-jejunum was dehydrated in graded alcohol and embedded in paraffin wax. Then, hematoxylin and eosin (H&E)-stained paraffin sections were viewed under bright field on a Zeiss Axio Imager microscope as we described (Huang et al. 2019b; Wang et al. 2010). The degree of intestinal tissue damage was scored as described previously (Huang et al. 2019b; Ji et al. 2018; Nishiyama et al. 2012), which the extent of epithelial loss on intestinal villi and inflammatory infiltration were evaluated and included in the histopathological examination.
Protein preparation, digestion, and iTRAQ labeling
Protein concentration was determined using the Pierce™ BCA Protein Assay Kit (Thermo Scientific, USA) and confirmed by SDS-PAGE. The homogenate proteins were subjected to acetone precipitation before being redissolved in lysis buffer. Then, 100 μg protein was reduced by 10 mM DDT at 36 °C for 1 h and alkylated by 35 mM iodoacetamide for 1 h in the dark. Twelve hours of trypsin (Promega Corporation, Madison, WI, USA) digestion was facilitated by diluting urea to 1 M by 50 mM NH4HCO3, and resulting peptides were desalted by Strata-X C18 SPE column (Phenomenex, Torrance, CA, USA).
iTRAQ labeling was performed according to the manufacturer’s instructions (Applied Biosystems, Foster City, CA, USA). Briefly, isobaric tags were solubilized in 70 mL isopropanol. Protein samples from the NBW group were labeled with tags 113, 114, 115, and 116, while those from the IUGR group received tags 117, 118, 119, and 121 (four replicates/group). The labeled peptides were incubated for 2 h at room temperature and then mixed and dried by vacuum centrifugation.
LC−MS/MS analysis of the membrane proteome
Strong cationic-exchange chromatography (SCX) peptide fractions were pooled to obtain 10 final fractions and reduce the sample number and collection time. A 10-μL sample from each fraction was injected 3 times to the Proxeon Easy Nano-LC system. Peptides were separated on a C18 analytical reverse phase column (ID 75 μm × 10 cm, 200 Å, 3 μm particles) at a flow rate of 250 nL/min. A linear LC gradient profile was used to elute peptides from the column. After liquid-phase separation, peptides were subjected to nanoelectrospray ionization followed by tandem mass spectrometry (MS/MS) in a Q Exactive mass spectrometer (Thermo Fisher Scientific, MA, USA) coupled online to the HPLC. Data was acquired using a data-dependent acquisition mode in which, the 10 most abundant multiply-charged peptides with values between 350 and 2000 m/z in each cycle were selected for MS/MS analysis with the 15-s dynamic exclusion setting. The fragment intensity multiplier was set to 20 and the maximum accumulation time was 2 s. Peak areas of the iTRAQ reporter ions reflected the relative abundance of proteins in the samples. All of the mass spectrometry data were collected after being analyzed by related software.
Data analysis
For iTRAQ/Shotgun protein identification, the mass spectrometry data were transformed into MGF files with Proteome Discovery 1.3 (Thermo, Pittsburgh, PA, USA) and analyzed using MASCOT software 2.3.02 (Matrix Science, London, U.K.). Mascot was searched with a fragment ion mass tolerance of 0.05 Da and peptide mass tolerance of 20.0 ppm. The results were averaged using medians and quantified. Proteins with fold change > 1.2 or < 0.83 in comparison and unadjusted significance level P < 0.05 were considered differentially expressed.
At each time point (D0, D7, D21, and D35), numbers of DEMPs between IUGR and NBW piglets were summed to obtain the overall DEMPs number. For protein identification, the filters were set as follows: significance threshold P < 0.05 (with 95% confidence) and an ion score or expected cut-off of less than 0.05 (with 95% confidence). For protein quantitation, the filters were set as follows: “median” was chosen for the protein ratio type (http://www.matrixscience.com/help/quant_config_help.html); the minimum precursor charge was set to 2 t, and the minimum peptide was set to 2; only unique peptides were used to quantify the proteins. The median intensities were set for normalization, and the outliers were removed automatically. The peptide threshold was set as above for identity.
To predict the functions of DEMPs, the proteins were analyzed with regard to three aspects. Proteins were annotated using blast against Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) database to obtain their functions. Significant GO functions and pathways were examined within DEPs with P ≤ 0.05. The category gene enrichment test of all proteins was performed using Blast2GO to determine whether the DEMPs were significantly enriched in any functional subcategories. Lastly, we allocated the DEMPs to biological pathways using the KEGG resource (www. genome.jp/kegg/). Proteins with a fold change greater than 1.2 (P ≤ 0.05) were considered to be differentially expressed.
Quantitative real-time PCR analysis of gene expression
Total RNA from jejunal mucosa was extracted using the TRIzol kit (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. Primers for RT-qPCR were synthesized by Shanghai Generay Biotech Co., Ltd. (Supplementary Table S1). RNA extraction, cDNA synthesis, and RT-qPCR were conducted according to the method as described previously (Huang et al. 2019b). Amplifications were performed in triplicate for each sample. The relative expression of genes was calculated using the 2−ΔΔCt method.
Results
Analysis of Jejunal morphology
Histological sectioning determined the different morphological changes in the jejunum between NBW piglets and IUGR piglets at D0, D7, D21, and D35. As shown in Fig. 1, there was no difference in the jejunum of piglets between the two groups at birth. However, at D7, D21 and D35, IUGR-induced piglets had increased (P < 0.05) infiltration of inflammatory cells in the mucosal layer of jejunal tissue compared with the NBW group.
Analysis of the overall proteins
A total of 5176 proteins were identified, with 980 at D0, 1295 at D7, 2165 at D21, and 736 at D35 (Fig. 2b). All identified proteins were annotated to GO function entries (Fig. 3), and the second-level GO terms were applied to classify proteins regarding their involvement in 3 main categories: molecular function (MF), cellular component (CC), and biological process (BP). Although proteins from each time point were annotated to similar main categories and subcategories, the exact protein number in each term was slightly different. Details are presented in Fig. 3. Next, the proteins were annotated against KEGG to identify the pathways related to different time points. Results showed that all proteins were mainly involved in organismal systems, metabolism, human diseases, genetic information processing, environmental information processing, and cellular processes, which were consistent in different time points (Fig. 4). In addition, the top twenty most abundant pathways identified at each time point were still consistent overall, regardless of the slight differences in certain pathways (Fig. 5). For example, mammalian target of rapamycin (mTOR) pathway was enriched at D0 with 29 proteins assigned, but was not present in other time points. Besides, phosphatidylinositol-3-kinase/protein kinase B (PI3K-Akt) signaling pathway only enriched at D21 and D35, with 33 and 31 proteins clustered.
Analysis of membrane proteins
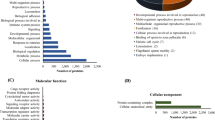
After being annotated against GO and KEGG databases, from D0 to D35, a total of 361 proteins related to the component and function of the membrane were selected and subjected to GO and KEGG assignment (Fig. 6a and Fig. 7). Firstly, to understand the putative functional implications of membrane protein abundances in IUGR and NBW piglets at each time point, we performed a GO term over-representation analysis. Analysis of the main molecular function (MF) GO category revealed the terms “binding,” “catalytic activity,” “transporter activity,” and “signal transducer activity.” Under the cellular component (CC) category GO terms, these membrane proteins are predominantly involved in the terms of cell part, membrane part, organelle part, and organelle. In terms of GO Biological Process (BP), membrane proteins, as expected, belonging to the cellular process, localization, biological regulation, and cellular component organization or Biogen. Besides, on the basis of their biological functions, these proteins are classified into the top 20 pathways. For example, the top 5 pathways identified were phagosome, ABC transporters, endocytosis, bile secretion, and tight junction. Details are presented in Fig. 6b.
Analysis of differentially expressed membrane proteins between NBW and IUGR piglets
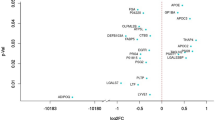
Among the 316 membrane proteins, there were 14 differentially expressed membrane proteins (DEMPs) with fold change > 1.2 (P < 0.05) (Table 1) were identified between IUGR and NBW piglets at different postnatal days (D0, D7, D21, and D35). Among them, 8 were found (F1SBL3, F1RRW8, F1S539, F1S2Z2, F1RIR2, F1RUF2, I3L6A2, and F1SCJ1) at D0, 5 (I3LP60, Q2EN79, F1RI18, F1SIH8, and I3LRJ7) at D7, 1 (F1RNN0) at D21. At D0, the abundance of F1SBL3, F1RRW8, F1S539, F1S2Z2, F1RIR2, and F1RUF2 was lower in the jejunal mucosa of IUGR neonates compared with NBW pigs, but I3L6A2 and F1SCJ1 were increased in the IUGR piglets. At D7, the abundance of I3LP60, Q2EN79, and F1SIH8 was lower in the IUGR group. In contrast, the levels of F1RI18 and I3LRJ7 were increased in the IUGR piglets. Finally, at D21, only one membrane protein was differentially expressed between IUGR and NBW group, for instance, F1RNN0 was increased in the IUGR piglets.
Functional analysis of differentially expressed membrane proteins
All the DEMPs were compared against NCBI and Uniprot databases to identify protein name and biological functions. Details are presented in Table 1. Among them, DEMPs at birth including F1RRW8, F1S539, F1S2Z2, F1R1R2, F1RUF2, I3L6A2, and F1SCJ1 are involved in D2 dopamine receptor binding, transmembrane transport of small molecules, signal transduction, translocation of GLUT4 to the plasma membrane, metal ion binding, and mediated vesicle transport. Meanwhile, the proteins such as the DEMPs at D7 after birth including F1RI18, F1SIH8, and I3LRJ7 are associated with electron transport chain, autophagy, and metabolism of vitamins and cofactors. The DEMPs at D21 after birth, F1RNN0, is involved in intracellular protein transport.
Validation of gene expression by RT-qPCR
Changes in protein expression may be due to changes in the mRNA levels; then, we used RT-qPCR to determine mRNA levels of seven selected differential membrane proteins that were related to signal transduction, immune response, and transmembrane transport at D0, D7, and D21. As shown in Fig. 8, the mRNA expression levels of PTPRK, DNM1, ABC4, and STX4 were significant decreased (P < 0.05) in the jejunal mucosa of IUGR piglets at birth. Compared with the NBW piglets, IUGR piglets had lower (P < 0.05) mRNA expression levels of UQCR10 and VCP at D7. Meanwhile, the gene expression of VPS29 in jejunal mucosa was higher (P < 0.05) in the IUGR piglets than in the NBW piglets. Collectively, the changes in mRNA expression of genes were generally corrected with the corresponding changes in differential membrane proteins expression detected by the iTRAQ approach.
Validation analysis of the mRNA expression levels of seven selected differential membrane proteins between NBW and IUGR groups at D0 (a, b, c, and d), D7 (e and f), and D21 (g) after birth. Data are mean ± SEM,n = 4 for each group at D0, D7, and D21. *P < 0.05, **P < 0.01, ***P < 0.001 versus the NBW group at each time phase
As shown in Fig. 9, the autophagy marker LC3B, the pro-apoptotic gene Bax, and the anti-apoptotic gene Bcl-2 were detected by RT-qPCR in the jejunal mucosa of piglets during early life. The mRNA expression levels of LC3B and Bax were increased (P < 0.05) and Bcl-2 was decreased (P < 0.05) in the IUGR piglets at D0, D7, D21, and D35, respectively, similar with previous studies. Furthermore, the results indicated that the enhanced autophagy activity and apoptosis may have a key role in IUGR-induced intestinal membrane impaired during the postnatal period.
Discussion
To identify the differentially expressed membrane proteins between NBW and IUGR piglets, an iTRAQ temporal proteomic analysis was performed to assess the intestinal development in piglets with IUGR at different postnatal days. Among the 361 identified membrane proteins, 14 membrane proteins involved in material transport, inflammatory response, oxidative stress, autophagy, apoptosis, and nervous regulation were significantly different between NBW and IUGR pigs at D0, D7, and D21.
At D0, there were 4 proteins associated with transport, 2 related to immune response, and 1 for neuromodulation. As a GTPase, dynamin 1 (F1RRW8) participates in the division of organelles, cytokinesis, and microbial pathogen resistance, which is critical for endocytosis and sustained neurotransmission (Boumil et al. 2010; Ferguson et al. 2007). A previous study found that depletion of both dynamin 1 and 3 seriously impaired the formation and development of the large nerve terminal in the auditory brainstem of mice during the first postnatal week (Fan et al. 2016), revealing the importance of dynamin in synaptic development and subsequent maintenance in the mammalian brain. In the current study, the relative abundance of dynamin 1 protein was lower in IUGR piglets than NBW piglets at birth. Consistent with the results of protein expression, the mRNA level of DNM1 was decreased in the jejunal mucosa of IUGR piglets at birth. According to previous studies (Miller et al. 2016), many neurodevelopmental disorders in motor and cognitive function were due to intrauterine growth restriction at birth. Meanwhile, dynamin-1-like protein is a fundamental component of mitochondrial fission, thus dysfunction in the dynamin-related protein (DRP) activity may result in mutated DNA or malfunctioning proteins diffusing throughout the mitochondrial system (Archer 2013; Fujimaki et al. 2011). Finally, these results increase the potential of producing reactive oxygen species, which can disrupt normal biochemical processes inside of cells. According to previous studies (Wang et al. 2008; Wang et al. 2016), IUGR piglets were more susceptible to oxidative damage or imbalance in antioxidant systems at birth, which is in agreement with our results.
Syntaxin 4 (F1RIR2) is one of the plasma membrane proteins associated with the immune system, exocytosis, and transport, which has been identified in mast cells and phagosomal and plasma membranes by immunoblotting (Band and Kuismanen 2005). Previous studies reported that syntaxin 4 promotes myoblast differentiation and plays a critical role in insulin-stimulated glucose uptake by promoting translocation of glucose transporter 4 (GLUT4) (Yoo et al. 2015). Meanwhile, syntaxin 4 is involved in the key signaling of p38MAPK and AKT in myoblast differentiation (Yoo et al. 2015). However, the abundance of syntaxin 4 was decreased in the IUGR piglets at birth, suggesting that abnormal expression of syntaxin 4 may cause growth retardation in IUGR piglets at birth.
According to the Uniprot database, an ATP-binding cassette subfamily A member 4 (F1S539) is responsible for transmembrane transport of small molecules. The protein is encoded by the ABCA4 gene, a member of the ATP-binding cassette transporter gene sub-family A (Tsybovsky et al. 2010). ABCA4 may function as an inward-directed retinoid flippase. Flippase is a transmembrane protein that has the conformation to transport materials across a membrane (Yatsenko et al. 2005). It is also related to autosomal-recessive disease Stargardt macular dystrophy (Maeda et al. 2008). Many previous studies reported that IUGR or low birth weight could impair intestinal development and reduced the efficiency of nutrient absorption during early life (Li et al. 2017a; Wang et al. 2008; Wu et al. 2006). Meanwhile, the current study showed that the small molecule transport protein was lower in the IUGR pigs compared with those in NBW pigs, which suggested the protein may play an important role in the IUGR-induced piglets.
Protein tyrosine phosphatase, receptor type K (F1S2Z2) has been implicated in regulating the phosphorylation state of many important signaling molecules, such as the MAP kinase family (Tonks 2006). The protein encoded by this gene is a member of the protein tyrosine phosphatase (PTP) family. PTPs are known to be signaling molecules that regulate a variety of cellular processes including cell growth, differentiation, mitotic cycle, and oncogenic transformation (Huang et al. 2010). In the previous studies, the PTP kappa protein was observed in neural cells and radial glial cells of the developing mouse brain (Huang et al. 2010). Our study may support that the protein could impact the brain development in the IUGR-induced pigs at birth.
At D7, there are 3 proteins for nervous system regulation including both inflammatory response and transport. Membrane protein Q2EN79 and F1RI18 are both from the ubiquinol-cytochrome c reductase family, which are reported to be associated with Alzheimer’s disease, development of cerebellar Purkinje cell layer, hippocampus, hypothalamus, midbrain, and pyramidal neuron as well as mitochondrial activity according to the Uniprot database. There is also a small core-associated protein and a subunit of ubiquinol-cytochrome c reductase complex III, which is part of the mitochondrial respiratory chain (Barel et al. 2008). The protein might be involved in the production of superoxide and oxygen free radicals. Superoxide anions produced by mitochondria appear to be an important source of oxidative stress and contribute to an overall increase in blood and lipid peroxidation in the intestine (Mittal et al. 2014; Ott et al. 2007; Oxidative Stress and Antioxidant Defense). Although a physiological balance between lipid peroxides and antioxidative processes is maintained in normal birth weight piglets, an imbalance may increase oxidative stress (Wang et al. 2016; Wang et al. 2010). Increased concentrations of these two proteins, along with altering oxidative and antioxidative homeostasis. Many reports have described that the intestinal protein localization and regulatory contribution to intestine integrity, which prevented the oxidant stress in the IUGR piglets (Wang et al. 2016; Wang et al. 2010). Also, decreased levels of membrane protein Q2EN79 and F1RI18 suggest systemic oxidative stress in IUGR neonatal piglets.
Transitional endoplasmic reticulum ATPase (F1SIH8), also recognized as valosin-containing protein (VCP) or p97, is a multi-functional protein related to autophagy, membrane traffic, cell cycle regulation, nuclear envelope reconstruction, and DNA damage repair (Ju et al. 2009; Kittler et al. 2004; Meerang et al. 2011; Woodman 2003; Ye et al. 2001; Zhang et al. 1994; Zhong et al. 2004), as well as various neurodegenerative disorders, such as frontotemporal dementia (Watts et al. 2004) and spinocerebellar ataxia type 3 (Fujita et al. 2013). It is well known that IUGR may be a delay in the development of the antioxidant system in the intestine. Many reports have described that oxygen stress-induced DNA damage may be particularly deleterious because it can produce mutations [26]. In this study, according to the Uniprot database, our results showed that the valosin-containing protein is involved in DNA damage repair. However, the level of VCP was decreased in the IUGR piglets, which suggests that the DNA repair and redox regulatory activities are damaged in the intestine of IUGR-induced piglets. Especially, our findings reveal abnormal expression levels of genes and membrane proteins related to oxidative stress, autophagy, and apoptosis in the jejunal mucosa of IUGR piglets, in agreement with previous studies (D’Inca et al. 2011; D’Inca et al. 2010b; Wang et al. 2016; Wang et al. 2014; Wang et al. 2018). This may be the major mechanism responsible for reduced growth and impaired development of jejunum in IUGR piglets. It is well known that autophagy is a self-digesting mechanism responsible for clearance of damaged organelles and long-lived proteins and plays a key role in regulating cell survival, cell death, and cell growth (Codogno and Meijer 2005; Zhu et al. 2015). Consistent with the previous study, the role of enhanced autophagy had been revealed in the small intestinal mucosa of IUGR fetal piglets (Zhu et al. 2015), and higher abundances of apoptotic proteins have also been reported in the small intestine (D’Inca et al. 2010b; Wang et al. 2014).
There is only one protein that functions as a transporter at D21. Vacuolar protein sorting-associated protein 29 (F1RNN0) is the only differentially expressed membrane protein at D21, which is also an intercellular transporter (McNally et al. 2017). Furthermore, it can be noticed that the number of significantly different membrane proteins tended to decrease along with piglet’s age, indicating the importance of prompt intervention on IUGR neonates to avoid any retardation in nervous system development, nutrient transport, and metabolism as they grow.
More importantly, emerging evidence has demonstrated that the gut microbiota plays a key role in nutrient absorption, intestinal development, metabolic equilibrium, and immune response. Consistent with our previous studies, the results of the dynamic changes in small intestinal structure, microbiota composition, mucosal mutation, and proteome were revealed in both NBW piglets and IUGR piglets during early life (Wang et al. 2014; Wang et al. 2018). In the present study, dynamic variations of intestinal membrane proteins involved in biological pathways including oxidative stress, immune response, autophagy, and apoptosis were consistent with the IUGR-induced gut microbiota disequilibrium. Compared with the NBW piglets, IUGR piglets had significantly different communities and microbial gene functions in the intestine and feces during early life (Huang et al. 2019a; Li et al. 2019). The results implied that IUGR significantly impairs intestinal structure, modifies gut microbiota colonization, and disturbs inflammatory at birth and during postnatal periods (D’Inca et al. 2011; D’Inca et al. 2010b; Huang et al. 2019a; Li et al. 2019). In addition to the gut microbiota, glycosylation is an important posttranslational modification, which regulates several critical biological processes including signal transduction and disease progression (Li et al. 2017b). Probing the glycosylation status on membrane protein enables an in-depth understanding of the role of glycosylation on membrane protein structure and function. Based on the above results, further attention in revealing differential membrane protein function could focus on glycosylation, intestinal mucosa development, and gut microbiota colonization in the intestine of IUGR piglets.
In summary, we conducted, for the first time, a temporal differential membrane proteomic study in porcine jejunal mucosa between NBW and IUGR piglets at postnatal D0, D7, D21, and D35 using iTRAQ technology. These differentially expressed membrane proteins focused on never regulation, nutrient transport, oxidative stress, autophagy, apoptosis, and inflammatory response. The decreased membrane proteins in IUGR piglets are mainly associated with substance transportation, signal transduction, polypeptide synthesis, and intestinal integrity, while the increased proteins are mainly involved in the regulation of oxidative stress and dysfunction. These findings provide the first evidence for jejunal mucosa membrane protein alterations for IUGR neonates and thus render new insights into the mechanisms responsible for intestinal dysfunction and the need for new nutritional strategies targeting IUGR piglets.
References
Archer SL (2013) Mitochondrial dynamics--mitochondrial fission and fusion in human diseases. N Engl J Med 369:2236–2251. https://doi.org/10.1056/NEJMra1215233
Band AM, Kuismanen E (2005) Localization of plasma membrane t-SNAREs syntaxin 2 and 3 in intracellular compartments. BMC Cell Biol 6:26. https://doi.org/10.1186/1471-2121-6-26
Barel O, Shorer Z, Flusser H, Ofir R, Narkis G, Finer G, Shalev H, Nasasra A, Saada A, Birk OS (2008) Mitochondrial complex III deficiency associated with a homozygous mutation in UQCRQ. Am J Hum Genet 82:1211–1216. https://doi.org/10.1016/j.ajhg.2008.03.020
Baserga M, Bertolotto C, Maclennan NK, Hsu JL, Pham T, Laksana GS, Lane RH (2004) Uteroplacental insufficiency decreases small intestine growth and alters apoptotic homeostasis in term intrauterine growth retarded rats. Early Hum Dev 79:93–105. https://doi.org/10.1016/j.earlhumdev.2004.04.015
Boumil RM, Letts VA, Roberts MC, Christine L, Mahaffey CL, Zhong-Wei Z, Tobias M, Frankel WN (2010) A missense mutation in a highly conserved alternate exon of dynamin-1 causes epilepsy in fitful mice. PLoS Genet 6:182–188. https://doi.org/10.1371/journal.pgen.1001046
Bozzetti V, Tagliabue PE, Visser GH, van Bel F, Gazzolo D (2013) Feeding issues in IUGR preterm infants. Early Hum Dev 89(Suppl 2):S21–S23. https://doi.org/10.1016/j.earlhumdev.2013.07.006
Codogno P, Meijer AJ (2005) Autophagy and signaling: their role in cell survival and cell death. Cell Death Differ 12(Suppl 2):1509–1518. https://doi.org/10.1038/sj.cdd.4401751
D’Inca R, Che L, Thymann T, Sangild PT, Huërou-Luron IL (2010a) Intrauterine growth restriction reduces intestinal structure and modifies the response to colostrum in preterm and term piglets. Livest Sci 133:20–22. https://doi.org/10.1016/j.livsci.2010.06.015
D’Inca R, Kloareg M, Gras-Le Guen C, Le Huerou-Luron I (2010b) Intrauterine growth restriction modifies the developmental pattern of intestinal structure, transcriptomic profile, and bacterial colonization in neonatal pigs. J Nutr 140:925–931. https://doi.org/10.3945/jn.109.116822
D’Inca R, Gras-Le Guen C, Che L, Sangild PT, Le Huerou-Luron I (2011) Intrauterine growth restriction delays feeding-induced gut adaptation in term newborn pigs. Neonatology 99:208–216. https://doi.org/10.1159/000314919
Fan F, Funk L, Lou X (2016) Dynamin 1- and 3-mediated endocytosis is essential for the development of a large central synapse in vivo journal of neuroscience the official. J Soc Neurosci 36:6097–6115. https://doi.org/10.1523/jneurosci.3804-15.2016
Ferguson SM, Brasnjo G, Hayashi M, Wolfel M, Collesi C, Giovedi S, Raimondi A, Gong LW, Ariel P, Paradise S, O’Toole E, Flavell R, Cremona O, Miesenbock G, Ryan TA, De Camilli P (2007) A selective activity-dependent requirement for dynamin 1 in synaptic vesicle endocytosis. Science 316:570–574. https://doi.org/10.1126/science.1140621
Fujimaki A, Watanabe K, Mori T, Kimura C, Shinohara K, Wakatsuki A (2011) Placental oxidative DNA damage and its repair in preeclamptic women with fetal growth restriction. Placenta 32:367–372. https://doi.org/10.1016/j.placenta.2011.02.004
Fujita K, Nakamura Y, Oka T, Ito H, Tamura T, Tagawa K, Sasabe T, Katsuta A, Motoki K, Shiwaku H, Sone M, Yoshida C, Katsuno M, Eishi Y, Murata M, Taylor JP, Wanker EE, Kono K, Tashiro S, Sobue G, La Spada AR, Okazawa H (2013) A functional deficiency of TERA/VCP/p97 contributes to impaired DNA repair in multiple polyglutamine diseases. Nat Commun 4:1816. https://doi.org/10.1038/ncomms2828
Glover LE, Lee JS, Colgan SP (2016) Oxygen metabolism and barrier regulation in the intestinal mucosa. J Clin Invest 126:3680–3688. https://doi.org/10.1172/JCI84429
Haq S, Grondin J, Banskota S, Khan WI (2019) Autophagy: roles in intestinal mucosal homeostasis and inflammation. J Biomed Sci 26:19. https://doi.org/10.1186/s12929-019-0512-2
Hsueh W, Caplan MS, Qu XW, Tan XD, De Plaen IG, Gonzalez-Crussi F (2003) Neonatal necrotizing enterocolitis: clinical considerations and pathogenetic concepts. Pediatr Dev Pathol 6:6–23. https://doi.org/10.1007/s10024-002-0602-z
Huang JB, Liu YL, Sun PW, Lv XD, Du M, Fan XM (2010) Molecular mechanisms of congenital heart disease. Cardiovasc Pathol 19:e183–e193. https://doi.org/10.1016/j.carpath.2009.06.008
Huang S, Li N, Liu C, Li T, Wang W, Jiang L, Li Z, Han D, Tao S, Wang J (2019a) Characteristics of the gut microbiota colonization, inflammatory profile, and plasma metabolome in intrauterine growth restricted piglets during the first 12 hours after birth. J Microbiol 57:748–758. https://doi.org/10.1007/s12275-019-8690-x
Huang S, Wu Z, Liu C, Han D, Feng C, Wang S, Wang J (2019b) Milk fat globule membrane supplementation promotes neonatal growth and alleviates inflammation in low-birth-weight mice treated with lipopolysaccharide. Biomed Res Int 2019:4876078–4876010. https://doi.org/10.1155/2019/4876078
Ji Y, Dai Z, Sun S, Ma X, Yang Y, Tso P, Wu G, Wu Z (2018) Hydroxyproline attenuates dextran sulfate sodium-induced colitis in mice: involvment of the NF-κB signaling and oxidative stress. Mol Nutr Food Res e1800494. https://doi.org/10.1002/mnfr.201800494
Ju JS, Fuentealba RA, Miller SE, Jackson E, Piwnica-Worms D, Baloh RH, Weihl CC (2009) Valosin-containing protein (VCP) is required for autophagy and is disrupted in VCP disease. J Cell Biol 187:875–888. https://doi.org/10.1083/jcb.200908115
Kittler R, Putz G, Pelletier L, Poser I, Heninger AK, Drechsel D, Fischer S, Konstantinova I, Habermann B, Grabner H, Yaspo ML, Himmelbauer H, Korn B, Neugebauer K, Pisabarro MT, Buchholz F (2004) An endoribonuclease-prepared siRNA screen in human cells identifies genes essential for cell division. Nature 432:1036–1040. https://doi.org/10.1038/nature03159
Li N, Wang W, Wu G, Wang J (2017a) Nutritional support for low birth weight infants: insights from animal studies. Br J Nutr 117:1390–1402. https://doi.org/10.1017/S000711451700126X
Li X, Jiang X, Xu X, Zhu C, Yi W (2017b) Imaging of protein-specific glycosylation by glycan metabolic tagging and in situ proximity ligation. Carbohydr Res 448:148–154. https://doi.org/10.1016/j.carres.2017.06.015
Li N, Huang S, Jiang L, Dai Z, Li T, Han D, Wang J (2019) Characterization of the early life microbiota development and predominant Lactobacillus species at distinct gut segments of low- and normal-birth-weight piglets. Front Microbiol 10:797. https://doi.org/10.3389/fmicb.2019.00797
Ma J, Chen T, Wu S, Yang C, Bai M, Shu K, Li K, Zhang G, Jin Z, He F, Hermjakob H, Zhu Y (2019) iProX: an integrated proteome resource. Nucleic Acids Res 47:1211–1217. https://doi.org/10.1093/nar/gky869
Maeda A, Maeda T, Golczak M, Palczewski K (2008) Retinopathy in mice induced by disrupted all-trans-retinal clearance. J Biol Chem 283:26684–26693. https://doi.org/10.1074/jbc.M804505200
McNally KE, Faulkner R, Steinberg F, Gallon M, Ghai R, Pim D, Langton P, Pearson N, Danson CM, Nagele H, Morris LL, Singla A, Overlee BL, Heesom KJ, Sessions R, Banks L, Collins BM, Berger I, Billadeau DD, Burstein E, Cullen PJ (2017) Retriever is a multiprotein complex for retromer-independent endosomal cargo recycling. Nat Cell Biol 19:1214–1225. https://doi.org/10.1038/ncb3610
Meerang M, Ritz D, Paliwal S, Garajova Z, Bosshard M, Mailand N, Janscak P, Hubscher U, Meyer H, Ramadan K (2011) The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks. Nat Cell Biol 13:1376–1382. https://doi.org/10.1038/ncb2367
Miller SL, Huppi PS, Mallard C (2016) The consequences of fetal growth restriction on brain structure and neurodevelopmental outcome. J Physiol 594:807–823. https://doi.org/10.1113/JP271402
Mittal M, Siddiqui MR, Tran K, Reddy SP, Malik AB (2014) Reactive oxygen species in inflammation and tissue injury. Antioxid Redox Signal 20:1126–1167. https://doi.org/10.1089/ars.2012.5149
Nemoto Y, Kumagai T, Ishizawa K, Miura Y, Shiraishi T, Morimoto C, Sakai K, Omizo H, Yamazaki O, Tamura Y, Fujigaki Y, Kawachi H, Kuro OM, Uchida S, Shibata S (2019) Phosphate binding by sucroferric oxyhydroxide ameliorates renal injury in the remnant kidney model. Sci Rep 9:1732. https://doi.org/10.1038/s41598-018-38389-3
Nishiyama Y, Kataoka T, Yamato K, Taguchi T, Yamaoka K (2012) Suppression of dextran sulfate sodium-induced colitis in mice by radon inhalation. Mediat Inflamm 2012:239617–239611. https://doi.org/10.1155/2012/239617
Ott M, Gogvadze V, Orrenius S, Zhivotovsky B (2007) Mitochondria, oxidative stress and cell death. Apoptosis 12:913–922. https://doi.org/10.1007/s10495-007-0756-2
Regev RH, Reichman B (2004) Prematurity and intrauterine growth retardation--double jeopardy? Clin Perinatol 31:453–473. https://doi.org/10.1016/j.clp.2004.04.017
Sangild PT, Siggers RH, Schmidt M, Elnif J, Bjornvad CR, Thymann T, Grondahl ML, Hansen AK, Jensen SK, Boye M, Moelbak L, Buddington RK, Westrom BR, Holst JJ, Burrin DG (2006) Diet- and colonization-dependent intestinal dysfunction predisposes to necrotizing enterocolitis in preterm pigs. Gastroenterology 130:1776–1792. https://doi.org/10.1053/j.gastro.2006.02.026
Tomasz D, Daniel G, Rafal R, Agnieszka S, Jaroslaw S, Romuald Z, Ryszard O (2007) Urinary excretion rates of 8-oxoGua and 8-oxodG and antioxidant vitamins level as a measure of oxidative status in healthy, full-term newborns. Free Radic Res 41:997-1004. doi.org/https://doi.org/10.1080/10715760701468757
Tonks NK (2006) Protein tyrosine phosphatases: from genes, to function, to disease. Nat Rev Mol Cell Biol 7:833–846. https://doi.org/10.1038/nrm2039
Tsybovsky Y, Molday RS, Palczewski K (2010) The ATP-binding cassette transporter ABCA4: structural and functional properties and role in retinal disease. Adv Exp Med Biol 703:105–125. https://doi.org/10.1007/978-1-4419-5635-4_8
Turner JR (2009) Intestinal mucosal barrier function in health and disease. Nat Rev Immunol 9:799–809. https://doi.org/10.1038/nri2653
Wang X, Qiao S, Yin Y, Yue L, Wang Z, Wu G (2007) A deficiency or excess of dietary threonine reduces protein synthesis in jejunum and skeletal muscle of young pigs. J Nutr 137:1442–1446. https://doi.org/10.1093/jn/137.6.1442
Wang J, Chen L, Li D, Yin Y, Wang X, Li P, Dangott LJ, Hu W, Wu G (2008) Intrauterine growth restriction affects the proteomes of the small intestine, liver, and skeletal muscle in newborn pigs. J Nutr 138:60–66. https://doi.org/10.1093/jn/138.1.60
Wang X, Wu W, Lin G, Li D, Wu G, Wang J (2010) Temporal proteomic analysis reveals continuous impairment of intestinal development in neonatal piglets with intrauterine growth restriction. J Proteome Res 9:924–935. https://doi.org/10.2527/jas.2015-8928
Wang X, Lin G, Liu C, Feng C, Zhou H, Wang T, Li D, Wu G, Wang J (2014) Temporal proteomic analysis reveals defects in small-intestinal development of porcine fetuses with intrauterine growth restriction. J Nutr Biochem 25:785–795. https://doi.org/10.1016/j.jnutbio.2014.03.008
Wang W, Degroote J, Van Ginneken C, Van Poucke M, Vergauwen H, Dam TM, Vanrompay D, Peelman LJ, De Smet S, Michiels J (2016) Intrauterine growth restriction in neonatal piglets affects small intestinal mucosal permeability and mRNA expression of redox-sensitive genes. FASEB J 30:863–873. https://doi.org/10.1096/fj.15-274779
Wang J, Feng C, Liu T, Shi M, Wu G, Bazer FW (2017) Physiological alterations associated with intrauterine growth restriction in fetal pigs: causes and insights for nutritional optimization. Mol Reprod Dev 84:897–904. https://doi.org/10.1002/mrd.22842
Wang X, Zhu Y, Feng C, Lin G, Wu G, Li D, Wang J (2018) Innate differences and colostrum-induced alterations of jejunal mucosal proteins in piglets with intra-uterine growth restriction. Br J Nutr 119:734–747. https://doi.org/10.1017/S0007114518000375
Watts GD, Wymer J, Kovach MJ, Mehta SG, Mumm S, Darvish D, Pestronk A, Whyte MP, Kimonis VE (2004) Inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia is caused by mutant valosin-containing protein. Nat Genet 36:377–381. https://doi.org/10.1038/ng1332
Woodman PG (2003) p97, a protein coping with multiple identities. J Cell Sci 116:4283–4290. https://doi.org/10.1242/jcs.00817
Wu G, Bazer FW, Wallace JM, Spencer TE (2006) Board-invited review: intrauterine growth retardation: implications for the animal sciences. J Anim Sci 84:2316–2337. https://doi.org/10.2527/jas.2006-156
Yatsenko AN, Wiszniewski W, Zaremba CM, Jamrich M, Lupski JR (2005) Evolution of ABCA4 proteins in vertebrates. J Mol Evol 60:72–80. https://doi.org/10.1007/s00239-004-0118-4
Ye Y, Meyer HH, Rapoport TA (2001) The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414:652–656. https://doi.org/10.1038/414652a
Yoo M, Kim BG, Lee SJ, Jeong HJ, Park JW, Seo DW, Kim YK, Lee HY, Han JW, Kang JS, Bae GU (2015) Syntaxin 4 regulates the surface localization of a promyogenic receptor Cdo thereby promoting myogenic differentiation. Skelet Muscle 5:28. https://doi.org/10.1186/s13395-015-0052-8
Yu Y, Zhao Y, Teng F, Li J, Guan Y, Xu J, Lv X, Guan F, Zhang M, Chen L (2018) Berberine improves cognitive deficiency and muscular dysfunction via activation of the AMPK/SIRT1/PGC-1a pathway in skeletal muscle from naturally aging rats. J Nutr Health Aging 22:710–717. https://doi.org/10.1007/s12603-018-1015-7
Zhang L, Ashendel CL, Becker GW, Morre DJ (1994) Isolation and characterization of the principal ATPase associated with transitional endoplasmic reticulum of rat liver. J Cell Biol 127:1871–1883. https://doi.org/10.1083/jcb.127.6.1871
Zhong X, Shen Y, Ballar P, Apostolou A, Agami R, Fang S (2004) AAA ATPase p97/valosin-containing protein interacts with gp78, a ubiquitin ligase for endoplasmic reticulum-associated degradation. J Biol Chem 279:45676–45684. https://doi.org/10.1074/jbc.M409034200
Zhu YH, Lin G, Dai ZL, Zhou TJ, Yuan TL, Feng CP, Chen F, Wu GY, Wang JJ (2015) Developmental changes in polyamines and autophagic marker levels in normal and growth-restricted fetal pigs. J Anim Sci 93:3503–3511. https://doi.org/10.2527/jas.2014-8743
Acknowledgments
We thank the Mianyang New-hope Livestock Farming Co. Ltd. in Sichuan Province, China, for their assistance in this study. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository (Ma et al. 2019) with the dataset identifier PXD013315.
Funding
This work was supported by the National Natural Science Foundation of China (31630074), the Beijing Municipal Natural Science Foundation (S170001), the National Key Research and Development Program of China (2016YFD0500506, 2018YDF0501002), the 111 Project (B16044), and Jinxinnong Animal Science Developmental Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All experiments were carried out with the approval of China Agricultural University Animal Care and Use Committee (CAU20170114–1, Beijing, China).
Competing interests
The authors declared that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Huang, S., Liu, C., Li, N. et al. Membrane proteomic analysis reveals the intestinal development is deteriorated by intrauterine growth restriction in piglets. Funct Integr Genomics 20, 277–291 (2020). https://doi.org/10.1007/s10142-019-00714-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-019-00714-y