Abstract
Atherosclerosis, a multifactorial and chronic immune inflammatory disorder, is the main cause of multiple cardiovascular diseases. Researchers recently reported that lncRNAs may exert important functions in the progression of atherosclerosis (AS). Some studies found that lncRNAs can act as ceRNAs to communicate with each other by the competition of common miRNA response elements. However, lncRNA-associated ceRNA network in terms of atherosclerosis is limited. In present study, we pioneered to construct and systematically analyze the lncRNA-mRNA network and reveal its potential roles in carotid atherosclerotic rabbit models. Atherosclerosis was induced in rabbits (n = 3) carotid arteries via a high-fat diet and balloon injury, while age-matched rabbits (n = 3) were treated with normal chow as controls. RNA-seq analysis was conducted on rabbits carotid arteries (n = 6) with or without plaque formation. Based on the ceRNA mechanism, a ternary interaction network including lncRNA, mRNA, and miRNA was generated and an AS-related lncRNA-mRNA network (ASLMN) was extracted. Furthermore, we analyzed the properties of ASLMN and discovered that six lncRNAs (MSTRG.10603.16, 5258.4, 12799.3, 5352.1, 12022.1, and 12250.4) were highly related to AS through topological analysis. GO and KEGG enrichment analysis indicated that lncRNA MSTRG.5258.4 may downregulate inducible co-stimulator to perform a downregulated role in AS through T cell receptor signaling pathway and downregulate THBS1 to conduct a upregulated function in AS through ECM-receptor interaction pathway. Finally, our results elucidated the important function of lncRNAs in the origination and progression of AS. We provided an ASLMN of atherosclerosis development in carotid arteries of rabbits and probable targets which may lay the foundation for future research of clinical applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Atherosclerosis (AS), a chronic inflammatory disorder of blood vessel walls characterized by an imbalance between inflammatory response and lipid metabolism (Weber and Noels 2011), is the principal cause of multiple cardiovascular disorders that have become the main cause of mortality and long-term morbidity worldwide (Husain et al. 2015). The pathogenesis of AS is complicated because sundry cell types, growth factors, cytokines, and regulatory molecules are involved in regulating its initiation and progression (Koenen and Weber 2010). Additionally, the disease is accompanied by plethoric intima fibrosis, inflammation cascade reaction, fatty streaks, and plaque formation, further vascular smooth muscle cell proliferation as well as migration of monocytes. Yet, underlying molecular mechanisms of each event in AS remain unclear. Currently, medication for AS primarily reduces the plasma concentration of cholesterol and blood pressure (Parsons et al. 2018). However, mortality and morbidity regarding to atherosclerosis still retain high. After novel molecular pathways responsible for AS development being discovered, abundant prognostic biomarkers and valuable therapeutic target medicines for AS may be provided.
LncRNAs were recently recognized to be broadly transcribed in eukaryotic organisms’ genome ranging from nematodes to humans (Guttman et al. 2009; Ponting et al. 2009). LncRNAs comprise of transcripts > 200 nucleotides in length and may regulate wide-ranging biological mechanisms (Tian et al. 2019), such as genomic imprinting and chromatin modification (Bernstein and Allis 2005; Pandey et al. 2008). Additionally, accumulating evidence indicated that lncRNAs had extensive and complex impact on the development and progression of AS. For example, lncRNA Zfas1, SNHG6, and GAS5 were found distinctly increased in the sufferers with atherosclerotic plagues compared with the normal (Chen et al. 2016). Li et al. summarized that lncRNAs participated to regulate AS-related endothelial cell, macrophage, smooth muscle cell, and lipid metabolism, which was relevant to comprehend the underlying functional mechanisms of lncRNAs in AS progression (Li et al. 2016).
Recently, high-throughput RNA sequencing (RNA-seq) is pervasively available and used in multiple biomedical researches, which can accurately quantify gene expression of low-abundance lncRNAs. As for analyzing transcriptome data, particularly in AS, networks play an important role for people to comprehend the molecular and biological mechanisms of multiple pathophysiological processions and relating different genes (Schramm et al. 2010; Wheelock et al. 2009). For example, Chen et al. constructed a rat atherosclerosis model and characterized a global transcriptomic network of AS development (Chen et al. 2016). Additionally, Salmena et al. proposed competitive endogenous RNA (ceRNA) as a new complex rectified and control mechanism between coding and non-coding RNA (Salmena et al. 2011). Long non-coding RNA competitively interacted with micro-RNA, therefore suppressed mRNA degradation on which the same micro-RNA targeted (Pilyugin and Irminger-Finger 2014). The lncRNA-associated ceRNA network participates in many diseases such as carcinomas and cardiac hypertrophy as a novel form of post-transcriptional control (Salmena et al. 2011; Song et al. 2016). Nevertheless, the ceRNA mechanism related to AS remains largely unknown. Thus, in present research, we used high-throughput RNA-seq analysis method on the basis of carotid atherosclerotic rabbit models and constructed a ternary interaction network based on the theory of ceRNA to describe properties of the transcriptome during AS process containing long non-coding RNA, micro-RNA, and mRNA and analyzed related GO enrichment and KEGG pathways. Results of our study may further improve the comprehension of AS development in molecular level and provide a systematic structure and rationale for clinical studies on human AS diagnosis, therapy, and even prognosis in the future.
Methods and materials
Carotid atherosclerotic animal models
All rabbits (New Zealand white adult male rabbits) of uniform age were recruited from the model animal center of the Second Affiliated Hospital of Harbin Medical University. Experimental protocols were conducted strictly based on guidelines, which are formulated by the National Institutes of Health to make every endeavor to minimize rabbits suffering or discomfort. All performed procedures involved in this study were endorsed by the Medical Ethics Committee on Animal Research of the Second Affiliated Hospital of Harbin Medical University (Ethics No.KY2016-090) and were in compliance with the principles and regulations of laboratory animal care.
We randomly divided all rabbits (2.5–3.5 kg) (n = 6) into two groups. The rabbits divided in atherosclerosis group (n = 3) were fed with a high-fat diet including 10% lard (Shandong Shiyuantianjiaji Factory), 1% cholesterol (Shanghai Lanji technology), and 3% yolk powder (Shandong Shiyuantianjiaji Factory) and accepted an endothelial injury caused by a 2F Fogarty balloon catheter (Boston Scientific, Temecula, California) to induce rabbit carotid atherosclerosis. The three rabbits were intramuscularly anesthetized with ketamine (35 mg/kg), xylazine (5 mg/kg), and acepromazine (0.75 mg/kg). During the procedure, anesthesia was kept with isoflurane inhalation. As a previous research reported (Meng et al. 2008), the right common carotid arteries (CCA) were injured with a balloon catheter which was advanced into the CCA through the external carotid artery gently. The balloon was gently inflated at 2 atm and retracted. This procedure was repeated three times in each rabbit. Then, the balloon catheter was removed, the incision was closed with a suture, and the rabbits were allowed to recover. After that, rabbits were fed with a high-fat diet for approximately 12 weeks. The progress of plaques in these rabbits was monitored by two-dimensional ultrasound weekly. Age-matched rabbits (n = 3) treated with normal chow served as the control group. After 12 weeks, three high-fat diet rabbits were sacrificed by air embolism; the right carotid artery was taken to obtain the carotid atherosclerosis plaque and the intima. These blood vessels obtained were immediately preserved in liquid nitrogen. The normal food-treated age-matched rabbits which considered as controls were served as the same. Obtained arteries were fixed with 4% paraformaldehyde fixative and embedded in paraffin for subsequent HE staining. Serial cross section with a thickness of 3 μm was stained with HE and observed by light microscopy (Olympus, BX41, Tokyo, Japan.).
Total RNA isolation, library construction, and sequencing
TRIzol reagent (Life Technologies, Beijing, China) based on the manufacturer’s recommended protocol was utilized to isolate total RNAs of carotid atherosclerotic animal models. The RNA purity was inspected with a NanoPhotometer® spectrophotometer (IMPLEN, CA), RNA concentration was measured by Qubit® RNA Assay Kit in Qubit® 2.0 Flurometer (Life Technologies, CA, USA), and RNA integrity was measured with the RNA Nano 6000 Assay Kit of the Bioanalyzer 2100 system (Agilent Technologies, CA, USA).
The mRNA or lncRNA samples were prepared using 3 μg RNA per sample as input material. Firstly, ribosomal RNA was removed by Epicentre Ribo-zero™ rRNA Removal Kit (Epicentre, USA), and rRNA-free residue was cleaned up by ethanol precipitation. Next, we used the rRNA-depleted RNA by NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (NEB, USA) to obtain sequencing libraries following manufacturer’s instructions. Products were purified (AMPure XP system), and library quality was assessed on the Agilent Bioanalyzer 2100 system. With the recommendation of manufacturer, the clustering of the index-coded samples was conducted on a cBot Cluster Generation System with TruSeq PE Cluster Kit v3-cBot-HS (Illumina). The libraries were sequenced on an Illumina Hiseq X platform, and 150-bp paired-end reads were generated after cluster generation.
Approximately 3 μg of total RNA per sample as input material was applied for the small RNA library. On the basis of the manufacturer’s instructions, NEBNextR Multiplex Small RNA Library Prep Set for IlluminaR (NEB, USA) was conducted to generate sequencing libraries and index codes were added to attribute sequences to every sample. Library quality was assessed on the Agilent Bioanalyzer 2100 system with DNA High Sensitivity Chips. The clustering of the index-coded samples was conducted on a cBot Cluster Generation System using TruSeq SR Cluster Kit v3-cBot-HS (Illumina) following the manufacturer’s instructions. The libraries were sequenced on an Illumina HiSeq 2500 platform, and 50-bp single-end reads were generated following cluster generation.
RNA-Seq data analysis
Quality control of the sequencing raw data was conducted by Fast QC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/), including adapter check, average quality of reads, and base quality distribution. Trimmomatic (Bolger et al. 2014) was applied to remove reads including adapter and low-quality reads from raw data and discard the sequences with N base before alignment. Meanwhile, Q20, Q30, and GC contents of the clean data were calculated. Therefore, the whole downstream analyses were on the basis of clean data with high quality.
The reference genome (OryCun2.0) and gene annotation files were directly downloaded from the Ensembl genome browser (ftp://ftp.ensembl.org/pub/release-92/fasta/oryctolagus_cuniculus/). The clean reads of mRNA were aligned to OryCun2.0 with HISAT2 program (Kim et al. 2015), and the mapped reads were assembled by using StringTie (Pertea et al. 2016). Since there was no lncRNA-related information in the annotated information, lncRNAs in this study were all predicted. To predict the transcripts assembled by stringtie are lncRNAs, the criteria as follows: (a) The transcript does not have the coding potential (PLEK, CNCI) (Li et al. 2014; Sun et al. 2013); (b) the length of the transcript ≥ 200 bp; (c) exon number ≥ 2, filtering a large number of single exon transcripts with low expression level and low confidence in the splicing result of the transcriptome; (d) the transcripts are not indicated in the existing database as protein-coding transcripts. And then, transcripts with FPKM less than 0.5 in six samples were removed. For miRNA reads, alignment was performed with Bowtie. Meantime, miRDeep2 (Friedlander et al. 2012) was used to find novel miRNA with default parameters. Transcripts with count less than 2 in six samples were discarded. The aberrantly-expressed RNAs between experimental and control groups (mRNA and lncRNA) (p value ≤ 0.01 and fold change (FC > 2) were screened by the “edgeR” package with default parameters.
Identification of miRNA-gene interactions
IntaRNA (version 2.0) (Mann et al. 2017) and miRanda (version 3.3a) (John et al. 2004) were applied to predict the interaction of miRNA-lncRNA and miRNA-mRNA in present study. IntaRNA was a program for the fast and accurately predicting interactions between two RNA modules, and miRanda was used as an algorithm for discovering genomic targets for miRNAs. The outMaxEnergy = − 15 of IntaRNA and threshold Energy = − 15 of miRanda were set to screen the potential miRNA-targets. We extracted the intersection set of the corresponding target regulations of both tools to improve the reliability of the predicted target of miRNA.
Identification of AS-related mRNA-lncRNA interactions
Hypergenometric test was performed to extract competing lncRNA-mRNA interactions, which can assess the significance of the shared miRNAs between mRNA and lncRNA. We mapped DE-mRNA and DE-lncRNA into triplets network and then extracted genes and their associated miRNAs to a different triplet (p value < 0.01 was recognized statistically significant). The p value was calculated as follows:
where k was the number of miRNA shared between mRNA and lncRNA and n represented the whole number of human genome miRNA. The number of miRNA interacting with lncRNA and mRNA was t and r respectively.
Topological analysis
In this study, Cytoscape software (version 3.4.0) (Li et al. 2018) was used to construct and visualize the network. The built-in NetworkAnalyzer tool was applied to analyze topological properties such as the closeness, betweenness, and node degree. Closeness centrality (Ci) was the average mean path from one node to the others in a network, betweenness was a measurement of the centrality of a node in a network, and degree is a measure of how many edges connect with the nodes. We applied the Wilcoxon rank-sum test to test for significant differences in topological properties and conducted the statistical calculations with R software version 3.3.1 (p value < 0.05).
Functional enrichment analysis
Functional enrichment analysis at the GO (gene ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) levels were performed using DAVID bioinformatics resources (https://david.ncifcrf.gov/summary.jsp, version 6.8), which provided a comprehensive set of functional annotation and enrichment tools to understand underlying biological mechanisms of an interesting gene set (da Huang et al. 2009). In this study, the biological process (BP) terms and KEGG pathway enrichment analysis with the thresholds of p value < 0.01 were based on the whole rabbit genome.
Results
Successfully establishment of atherosclerotic rabbit carotid arteries models
With the combination of a high-fat diet and an endothelial injury, we can effortlessly detect atherosclerotic plaques in rabbit carotid arteries using the two-dimensional ultrasound at 12th week. Images showed thicken and rough intima and multiple irregular hypoechoic plaques in the right common carotid artery (Fig. 1a–c). In contrast, carotid arteries intima of rabbits treated with normal chow for 12 weeks still maintain clear, smooth, and continuous with no obvious plaque (Fig. 1d–f). Additionally, in AS group, HE-stained vascular on the place with obvious plagues through two-dimensional-ultrasound showed thicken intima, foam cell deposition, smooth muscle cell proliferation, and varying degrees of atherosclerotic plaques (Fig. 2a–c). However, in the control group, we did not identify significant abnormalities (Fig. 2d–f). Furthermore, CD31 immunohistochemistry showed multiple neovascularization in AS group but little even no neovascularization in the control group (Fig. 3a–f).
Images of rabbit carotid arteries in two-dimensional-ultrasound at 12th week. a–c two-dimensional- ultrasound images reveal that obvious atherosclerotic plaques formed on rabbit carotid arterial intima as the arrows show. d–f two-dimensional- ultrasound images demonstrate that carotid arteries intima of rabbits treated with normal chow still maintain smooth as the arrows show
Identification of lncRNAs in rabbit carotid arteries
In our present research, we used StringTie to assemble transcripts, predict results, and filter conditions based on PLEK and CNCI. We totally identified 2855 lncRNAs from two rabbit groups containing 2608 lincRNAs (90.4%) and 247 anti-sense lncRNAs (9.6%) with isoforms that were not reported in previous studies. Additionally, the lncRNA transcripts were widely distributed in all chromosomes (Fig. 4a), approximately 8.27%, 7.81%, 7.53%, and 7.43% coming from chr13, chr1, chr2, and chr12, respectively, while only less than 7% from any other chromosomes. We also identified 2651 lncRNAs in AS group, 2730 lncRNAs in control group and 2526 lncRNAs in both groups (Fig. 4b). The analysis of sequence length demonstrated that lncRNA transcripts were mostly ranged in length from 200 to 1000 bp (Fig. 4c). Furthermore, the length of ORF in lncRNA transcripts was mainly ranged from 38 to 545 and the number of exons was mostly ranged from 2 to 10.
The lncRNAs in rabbit carotid arteries. a Circos plot showing lncRNAs on rabbit chromosome. The outmost layer of ring represents chromosome map of the rabbit genome. The larger inner green ring is all lncRNAs detected by RNA-seq. The smaller inner ring indicates the differentially expressed lncRNAs with fold change > 2 and p value < 0.01, the up- and downregulation lncRNAs were marked in red and blue bars. b Among detected lncRNAs, 2855 common lncRNAs and 2651 specific lncRNAs in AS group and 2730 specific lncRNAs in control group. c Distribution of the sequence length of lncRNAs
Identification of AS-associated aberrant expressed lncRNAs and mRNAs
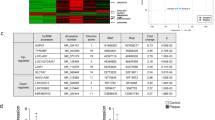
In order to explore the probable biological roles of lncRNAs in atherosclerosis, we studied and analyzed expression profile data of lncRNA, mRNA, and miRNA from atherosclerosis samples and controls. RNAs (lncRNAs and mRNAs) with p value ≤ 0.01 and a fold change > 2 were recognized as significantly aberrant expressed. A total of 255 differentially expressed lncRNAs were detected in AS by RNA-seq analysis, including 87 upregulated and 168 downregulated lncRNAs (Fig. 5a) (Supplementary Table S1). At the same time, 168 upregulated and 253 downregulated mRNAs were identified respectively in atherosclerosis sample group compared to control group (Fig. 5b) (Supplementary Table S2). Top ten DE-mRNAs and DE-lncRNAs based on p value are shown in Tables 1 and 2, respectively. Hierarchical clustering showed expression patterns of differentially expressed lncRNAs and mRNAs among samples were distinguishable (Fig. 6). Among differential expression genes, 84 genes were well-known atherosclerosis-related genes, which were mostly dysregulated expression genes, for example, MMP12, CHI3L1, SOAT1, EDNRA, and CAPG. The aberrantly expressed RNAs were identified as underlying key modulators in the initiation and progression of atherosclerosis.
Volcano plots showing expression profiles of lncRNAs and mRNAs. a, b The negative log of p value (base 10) is plotted on the X-axis, and the log of the fold change (base 2) is plotted on the Y-axis. The green points on the graphs represent differentially expressed lncRNAs and mRNAs in AS compared with normal samples, respectively (fold change > 2 and p value < 0.01). The red points on the graphs represent non-differentially expressed lncRNAs and mRNAs in AS compared with normal samples, respectively (fold change > 2 and p value < 0.01). a A total of 255 differentially expressed lncRNAs were detected in AS by RNA-seq analysis, including 87 upregulated and 168 downregulated lncRNAs. b A total of 168 upregulated and 253 downregulated mRNAs were identified respectively in atherosclerosis sample group compared to control group
Hierarchical cluster analysis using a heatmap. a, b Hierarchical cluster analysis was conducted to assess the significantly different expressions of lncRNA and mRNA, respectively (fold change > 2 and p value < 0.01). a Custer analysis of differentially expressed lncRNAs. b Cluster analysis of differentially expressed mRNAs. Red and green represented increased and decreased expression, respectively. Each sample was indicated by a single column, and each RNA was denoted by a single row of colored boxes
Construction of atherosclerosis-associated lncRNA-mRNA network
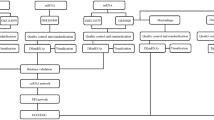
To construct global lncRNA-miRNA-mRNA network and identify putative miRNA-target interactions, according to the analysis of mRNA/lncRNA/miRNA sequence information obtained, we applied miRanda and IntaRNA tools in that a target predicted by two programs at the same time would be more reliable. Totally, we obtained 55,162 miRNA-lncRNA, and 176,684 miRNA-mRNA interaction pairs. Next, all miRNA-mRNA and miRNA-lncRNA pairs were merged to construct the lncRNA-miRNA-mRNA ternary interaction network, including 231,846 edges, 398 miRNAs, 2557 lncRNAs, and 6364 mRNAs. LncRNA, mRNA, and miRNA were defined as the nodes of this ternary interaction network. Additionally, an edge would be added between them if they have interaction with each other; then, the ternary interaction network can be constructed.
In order to investigate the triplet regulation in the AS and discover the AS-related lncRNAs, we extract 421 aberrant expressed mRNAs and 255 aberrant expressed lncRNAs and their linked miRNAs from global lncRNA-miRNA-mRNA ternary interaction network to set up a different DE lncRNA-miRNA-DE mRNA ternary interaction network. The result showed that new network has 385 miRNAs, 226 lncRNAs and 367 mRNAs, 5972 miRNA-lncRNA, and 10,467 miRNA-mRNA interaction pairs. Then, we used hypergeometric test to conduct statistical significance test of the amount of shared miRNAs between mRNA and lncRNA pairs. Finally, we set up a new AS-associated lncRNA-mRNA interaction network (ASLMN) (Fig. 7a) through adding an edge between mRNA and lncRNA if they shared the same miRNA when p value < 0.01. The new AS-associated lncRNA-mRNA network (ASLMN) comprised 22,324 edges, 209 lncRNA nodes, and 362 mRNA nodes. The degree distribution of nodes in the network was analyzed (Supplementary Table S3). The average degree of lncRNAs was 106.81 and significantly higher than the average degree of mRNAs (61.67) (p = 1.62e−09, Wilcoxon rank-sum test). In addition, we discovered that lncRNA nodes exhibited more betweenness centrality compared with mRNA nodes (p = 1.673e−07). These results suggest that although the lncRNAs were non-coding RNA, they showed more specific topological properties compared with mRNA in the network. This result revealed that a part of lncRNA nodes linked many mRNA nodes in ASLMN act as hubs.
The layout of lncRNA-related ceRNA network (ASLMN) and its structural characteristics. a The view of ASLMN. The ASLMN was comprised of 22,324 edges between 362 mRNAs and 209 lncRNAs. The green circle nodes represent mRNAs, and red triangle nodes represent lncRNAs. b The node distribution of mRNA. X-axis was the degrees of the nodes, and the Y-axis was the number of nodes. c The node distribution of lncRNA. X-axis was the degrees of the nodes, and the Y-axis was the number of nodes
Topological analysis of dysregulated lncRNA in AS-associated lncRNA-mRNA network
Firstly, we analyzed and calculated three topological properties of the ceRNA network which were the degree, closeness and betweenness. Generally, a higher degree suggested that the node was acted as hub involved in much more ceRNA interactions. A higher betweenness indicated that the node was bridge connecting different network modules, and bigger closeness suggested that the node tend to be at the center of the network. Figure 7b and c show the mRNA and lncRNA node distribution in the ceRNA network. We then ranked topological characteristics of all nodes in the network and found that six lncRNAs (MSTRG.10603.16, 5258.4, 12799.3, 5352.12, 12022.1, 12250.4) were common nodes in top 10 of each index (Table 3). By mapping these lncRNAs into a ceRNA network, we found that these lncRNAs and their neighbors formed a key module (Fig. 8a). Surprisingly, 331 genes were detected in this key module, and some dysregulated genes in the module were associated with atherosclerosis. In order to explore the underlying roles of the 6 lncRNAs, we conducted GO and KEGG which is the functional enrichment analysis for their first near mRNA neighbors in ASLMN, respectively. About the lncRNA MSTRG.5258.4, we discovered that “cell adhesion,” “immune response,” and “adaptive immune response” were contained in the biological processes, and the KEGG pathways were enriched including “T cell receptor signaling pathway” and “ECM-receptor interaction” (Fig. 8b) (Supplementary Tables S4 and S5). The other five lncRNAs were performed with similar annotations (Supplementary Figures 1–5.). The results demonstrated that the six lncRNAs with higher betweenness degree and closeness were pivot nodes in the ASLMN and performed essential and key effects in atherosclerosis-related biological processes contributing to progress of atherosclerosis. What is more, the six lncRNAs may be used as potential diagnostic factors and novel therapeutic targets of atherosclerosis. However, further specific studies are required to confirm the potential values of these lncRNAs.
The key module of ASLMN. a The module network of 6 lncRNAs with largest degree, betweenness and closeness in ASLMN. The blue nodes represented mRNA and the red nodes represented lncRNAs. b GO enrichment analysis of the module about the lncRNA MSTRG.5258.4. The enriched GO terms were ranked by gene count
Discussion
AS is a complicated disorder that afflicts millions of persons in the worldwide. The pathogenesis and process of the disorder are complex, thus investigating the mechanisms of AS and its various complications could demand a global perspective. LncRNAs are emerging functional modulators of complicated and extensive biological processes that influence cell development, differentiation, and metabolism (Ponting et al. 2009; Zhao et al. 2019), whose novel features have lately been focus of attention. Millions of lncRNAs have been discovered in the mammalian genome with new techniques of RNA sequencing reaching unprecedented levels. Nevertheless, a comprehensive database of experimentally validated lncRNA functions still lacks (Mo et al. 2019). At present, the integration of lncRNAs and coding gene expression profiles has become one of the most common research methods for studying the role of lncRNAs in different biological processes (Zhao et al. 2019). High-throughput screening assay helps us to identify whole transcriptomes on a single slide at the same time. In present study, we successfully induced AS in rabbits (n = 3) by a high-fat diet and a balloon injury as case group and age-matched rabbits (n = 3) were treated with normal chow as the control group. We performed RNA-seq analysis on rabbit carotids (n = 6) with or without atherosclerotic plaques formation. Attractively, the lncRNA transcripts were broadly distributed in all chromosomes. We discovered that 244 lncRNAs were significantly aberrant expressed including 87 upregulated and 168 downregulated lncRNAs. A total of 421 aberrantly expressed mRNAs were simultaneously indentified, containing 168 upregulated and 253 downregulated mRNAs respectively. In addition, hierarchical clustering based on the expression value of DE-lncRNAs showed that the samples from atherosclerosis clustered together. Also, we demonstrated that the DE-lncRNAs were generally distributed on each chromosome.
Among differential expression genes, many were well-known atherosclerosis-related genes, which were mostly dysregulated expression genes. Hu et al. demonstrated that MMP-12 can increase instability of plaques and promote the transformation from stable to vulnerable plaques to accelerate the occurrence and development of carotid atherosclerosis (Hu et al. 2018). Researches demonstrated that APPsw can increase CHI3L1 expression through downregulating miR-342-3p expression in the arterial endothelium, which promote AS (Jung et al. 2018). Nan Wu et al. indicated that deficiency of SOAT1 can inhibit cholesterol transportation from preventing progression of AS in Apo E/− mice (Wu et al. 2018). EDNRA rs1878406 have been revealed as the significant signals associated with presence of carotid plaques (Bis et al. 2011). Burillo et al. reported that rs6886 polymorphism in CAPG was associated with carotid IMT and could modulate response of macrophages to ox- LDL in AS (Burillo et al. 2009).
In our study, we used miRanda-3.3a (− en = − 15) and IntaRNA-2.0 (− out Max E = − 15) tools for mRNA, miRNA, and lncRNA on the basis of ceRNA theory of miRNA sharing lncRNA and mRNA in one triplet to establish a global network of interactions and describe the development of atherosclerosis. From the ternary interaction network, we obtained the AS-related lncRNA-mRNA interaction network (ASLMN) by mapping the DE genes into the ternary interaction network, so as to analyze the function of lncRNAs. ASLMN contained 209 lncRNA nodes and 362 mRNA nodes with a total of 22,324 lncRNA-mRNA edges. Then, we applied the topological properties analysis method to study the ASLMN, conducted a cluster analysis of ASLMN, and further discovered 6 lncRNAs (MSTRG.10603.16, 5258.4, 12799.3, 5352.1, 12022.1, and 12250.4) were highly associated with the development of AS. The lncRNAs connecting many mRNAs were highly associated with the AS by GO enrichment analysis and KEGG pathway analysis. About the lncRNA MSTRG.5258.4, we discovered that cell adhesion, immune response, and adaptive immune response were contained in the biological processes, and the KEGG pathways were enriched including T cell receptor signaling pathway and ECM-receptor interaction. Previous researches had demonstrated that these biological processes and pathways played a critical role in AS. Accumulative studies supported the involvement of the immune response in AS from initiation to thrombotic complications. Furthermore, atherosclerosis is involved in adaptive and innate immune response that regulates lesion occurrence, development, and possible thrombotic complications (Miteva et al. 2018; Vieceli Dalla Sega et al. 2019). T cells with antigen reactivity have a proatherogenic function; therefore, their response could be regulated. Signals or co-stimulatory molecules are required for T cells to proliferate and further differentiate into effector cells (Singh et al. 2014). Co-stimulatory molecules are primarily generated through engagement of the CD28 receptor on T cells with its ligands CD86 and CD80 on antigen-presenting cells (Huang et al. 2016). Inducible co-stimulator (ICOS) is the third member of the CD28 family. Studies indicated that ICOS co-stimulatory signaling pathway is evident in human atherosclerosis and researches on mice suggested that it was potentially associated with protection against lesion initiation and development (Gerdes and Zirlik 2011). While in the KEGG pathway T cell receptor signaling pathway in our study, we enriched “ICOS” gene. Thus, it is suggested that ICOS may perform a downregulated effect in atherosclerosis through T cell receptor signaling pathway, which is rarely reported in the past. Cell adhesion molecules (CAMs) as transmembrane proteins can modulate adhesion between cells or cell and extra-cellular matrix and leukocytes traffic, which may play essential parts in the early stage of atherogenesis. CAMs were focal expressed in atherosclerotic lesions (Ling et al. 2012), which precedes leukocyte infiltration and seems to be modulated partly by modified lipoproteins or their constituents (Schmidt et al. 2016). sVCAM-1 and sICAM-1 are closely relative to carotid intima-media thickness (IMT), an index of AS in the early stage (Figueroa-Vega et al. 2015). Thrombospondin-1 (THBS1), a matricellular protein involved in the maintenance of vascular structure through cell proliferation, apoptosis, and adhesion, was released from activated platelets and accumulated in sites of vascular injury (Khosravi et al. 2019). Immunohistochemical experiment has shown that THBS1 was high-expressed in injured arteries in hypercholesterolemic atherosclerotic rabbits (Ma et al. 2013); our study supports this result and THBS1 is the significant differentially expressed gene in the network. Furthermore, THBS1 can induce expression of CAM, including vascular CAM-1 (VCAM-1) and intracellular CAM-1 (ICAM-1), result in an increase of monocyte recruitment to sites of injury, and involve in atherosclerotic plaque development (Narizhneva et al. 2005). We enriched the gene “THBS1” in the biological process of cell adhesion and KEGG pathway ECM-receptor interaction, and suggested that THBS1 may upregulate AS through ECM-receptor interaction pathway, which is rare reported in the past. According to the results of enrichment, we suggested that lncRNA MSTRG.5258.4 may downregulate ICOS to perform a downregulated role in AS through T cell receptor signaling pathway, and THBS1 to conduct a upregulated function in AS through ECM-receptor interaction pathway with the modulation of lncRNA MSTRG.5258.4, which is rare reported in the past and further reveals the mechanisms of AS.
However, the results of the present study have not been experimentally validated, which is a limiting factor that requires resolution and study in the future.
In present research, we pioneered to comprehensively investigate the differentially expressed lncRNA-associated ceRNA network in AS. Six lncRNAs related to AS were successfully detected based on the theory of ceRNA. We have a different strategy that studies AS or other diseases. Meanwhile, lncRNA performs an important function in AS and may provide a new therapeutic target for AS mechanism research. Despite this progress, the discovery of experimental atherosclerosis and the leap in clinical use of laboratory animals to humans are also challenging. More detailed functional studies of mRNA and lncRNA are necessary.
References
Bernstein E, Allis CD (2005) RNA meets chromatin. Genes Dev 19:1635–1655. https://doi.org/10.1101/gad.1324305
Bis JC et al (2011) Meta-analysis of genome-wide association studies from the CHARGE consortium identifies common variants associated with carotid intima media thickness and plaque. Nat Genet 43:940–947. https://doi.org/10.1038/ng.920
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Burillo E et al (2009) Proteomic study of macrophages exposed to oxLDL identifies a CAPG polymorphism associated with carotid atherosclerosis. Atherosclerosis 207:32–37. https://doi.org/10.1016/j.atherosclerosis.2009.03.054
Chen L et al (2016) Global transcriptomic study of atherosclerosis development in rats. Gene 592:43–48. https://doi.org/10.1016/j.gene.2016.07.023
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Figueroa-Vega N, Moreno-Frias C, Malacara JM (2015) Alterations in adhesion molecules, pro-inflammatory cytokines and cell-derived microparticles contribute to intima-media thickness and symptoms in postmenopausal women. PLoS One 10:e0120990. https://doi.org/10.1371/journal.pone.0120990
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N (2012) miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 40:37–52. https://doi.org/10.1093/nar/gkr688
Gerdes N, Zirlik A (2011) Co-stimulatory molecules in and beyond co-stimulation - tipping the balance in atherosclerosis? Thromb Haemost 106:804–813. https://doi.org/10.1160/th11-09-0605
Guttman M et al (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227. https://doi.org/10.1038/nature07672
Hu W, Wei R, Wang L, Lu J, Liu H, Zhang W (2018) Correlations of MMP-1, MMP-3, and MMP-12 with the degree of atherosclerosis, plaque stability and cardiovascular and cerebrovascular events. Exp Ther Med 15:1994–1998. https://doi.org/10.3892/etm.2017.5623
Huang X et al (2016) Detection of CD28/CD86 co-stimulatory molecules and surface properties of T and dendritic cells: an AFM study. Scanning 38:365–375. https://doi.org/10.1002/sca.21279
Husain K, Hernandez W, Ansari RA, Ferder L (2015) Inflammation, oxidative stress and renin angiotensin system in atherosclerosis. World J Biol Chem 6:209–217. https://doi.org/10.4331/wjbc.v6.i3.209
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human microRNA targets. PLoS Biol 2:e363. https://doi.org/10.1371/journal.pbio.0020363
Jung YY et al (2018) Atherosclerosis is exacerbated by chitinase-3-like-1 in amyloid precursor protein transgenic mice. Theranostics 8:749–766. https://doi.org/10.7150/thno.20183
Khosravi M, Najafi M, Amirfarhangi A, Karimi M, Fattahi F, Shabani M (2019) The increase of pFAK and THBS1 protein and gene expression levels in vascular smooth muscle cells by histamine-treated M1 macrophages. Iran J Allergy Asthma Immunol 18:72–79
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Koenen RR, Weber C (2010) Therapeutic targeting of chemokine interactions in atherosclerosis. Nat Rev Drug Discov 9:141–153. https://doi.org/10.1038/nrd3048
Li A, Zhang J, Zhou Z (2014) PLEK: a tool for predicting long non-coding RNAs and messenger RNAs based on an improved k-mer scheme. BMC Bioinformatics 15:311. https://doi.org/10.1186/1471-2105-15-311
Li H, Zhu H, Ge J (2016) Long noncoding RNA: recent updates in atherosclerosis. Int J Biol Sci 12:898–910. https://doi.org/10.7150/ijbs.14430
Li M, Yang J, Wu FX, Pan Y, Wang J (2018) DyNetViewer: a Cytoscape app for dynamic network construction, analysis and visualization. Bioinformatics 34:1597–1599. https://doi.org/10.1093/bioinformatics/btx821
Ling S, Nheu L, Komesaroff PA (2012) Cell adhesion molecules as pharmaceutical target in atherosclerosis. Mini-Rev Med Chem 12:175–183
Ma Y, Yabluchanskiy A, Lindsey ML (2013) Thrombospondin-1: the good, the bad, and the complicated. Circ Res 113:1272–1274. https://doi.org/10.1161/circresaha.113.302749
Mann M, Wright PR, Backofen R (2017) IntaRNA 2.0: enhanced and customizable prediction of RNA-RNA interactions. Nucleic Acids Res 45:W435–w439. https://doi.org/10.1093/nar/gkx279
Meng L, Lv B, Zhang S, Yv B (2008) In vivo optical coherence tomography of experimental thrombosis in a rabbit carotid model. Heart 94:777–780. https://doi.org/10.1136/hrt.2007.117382
Miteva K, Madonna R, De Caterina R, Van Linthout S (2018) Innate and adaptive immunity in atherosclerosis. Vasc Pharmacol. https://doi.org/10.1016/j.vph.2018.04.006
Mo XB et al (2019) Detection of lncRNA-mRNA interaction modules by integrating eQTL with weighted gene co-expression network analysis. Funct Integr Genomics 19:217–225. https://doi.org/10.1007/s10142-018-0638-4
Narizhneva NV et al (2005) Thrombospondin-1 up-regulates expression of cell adhesion molecules and promotes monocyte binding to endothelium. FASEB J 19:1158–1160. https://doi.org/10.1096/fj.04-3310fje
Pandey RR et al (2008) Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell 32:232–246. https://doi.org/10.1016/j.molcel.2008.08.022
Parsons C, Agasthi P, Mookadam F, Arsanjani R (2018) Reversal of coronary atherosclerosis: role of life style and medical management. Trends Cardiovasc Med 28:524–531. https://doi.org/10.1016/j.tcm.2018.05.002
Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL (2016) Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc 11:1650–1667. https://doi.org/10.1038/nprot.2016.095
Pilyugin M, Irminger-Finger I (2014) Long non-coding RNA and microRNAs might act in regulating the expression of BARD1 mRNAs. Int J Biochem Cell Biol 54:356–367. https://doi.org/10.1016/j.biocel.2014.06.018
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. https://doi.org/10.1016/j.cell.2009.02.006
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146:353–358. https://doi.org/10.1016/j.cell.2011.07.014
Schmidt EP, Kuebler WM, Lee WL, Downey GP (2016) Adhesion molecules: master controllers of the circulatory system. Compr Physiol 6:945–973. https://doi.org/10.1002/cphy.c150020
Schramm G, Surmann EM, Wiesberg S, Oswald M, Reinelt G, Eils R, Konig R (2010) Analyzing the regulation of metabolic pathways in human breast cancer. BMC Med Genet 3:39. https://doi.org/10.1186/1755-8794-3-39
Singh AK, Gaur P, Das SN (2014) Natural killer T cell anergy, co-stimulatory molecules and immunotherapeutic interventions. Hum Immunol 75:250–260. https://doi.org/10.1016/j.humimm.2013.12.004
Song C et al (2016) Construction and analysis of cardiac hypertrophy-associated lncRNA-mRNA network based on competitive endogenous RNA reveal functional lncRNAs in cardiac hypertrophy. Oncotarget 7:10827–10840. https://doi.org/10.18632/oncotarget.7312
Sun L, Luo H, Bu D, Zhao G, Yu K, Zhang C, Liu Y, Chen R, Zhao Y (2013) Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res 41:e166. https://doi.org/10.1093/nar/gkt646
Tian Y et al (2019) Comprehensive analysis of microarray expression profiles of circRNAs and lncRNAs with associated co-expression networks in human colorectal cancer. Funct Integr Genomics 19:311–327. https://doi.org/10.1007/s10142-018-0641-9
Vieceli Dalla Sega F, Fortini F, Aquila G, Campo G, Vaccarezza M, Rizzo P (2019) Notch signaling regulates immune responses in atherosclerosis. Front Immunol 10:1130. https://doi.org/10.3389/fimmu.2019.01130
Weber C, Noels H (2011) Atherosclerosis: current pathogenesis and therapeutic options. Nat Med 17:1410–1422. https://doi.org/10.1038/nm.2538
Wheelock CE et al (2009) Systems biology approaches and pathway tools for investigating cardiovascular disease. Mol BioSyst 5:588–602. https://doi.org/10.1039/b902356a
Wu N, Li RQ, Li L (2018) SOAT1 deficiency attenuates atherosclerosis by regulating inflammation and cholesterol transportation via HO-1 pathway. Biochem Biophys Res Commun 501:343–350. https://doi.org/10.1016/j.bbrc.2018.03.137
Zhao R, Li FQ, Tian LL, Shang DS, Guo Y, Zhang JR, Liu M (2019) Comprehensive analysis of the whole coding and non-coding RNA transcriptome expression profiles and construction of the circRNA-lncRNA co-regulated ceRNA network in laryngeal squamous cell carcinoma. Funct Integr Genomics 19:109–121. https://doi.org/10.1007/s10142-018-0631-y
Funding
Present research was supported by the National Natural Science Foundation of China (No. 81671689) and the Natural Science Foundation of Heilongjiang Province (H2017021).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All performed procedures involved in this study were endorsed by the Medical Ethics Committee on Animal Research of the Second Affiliated Hospital of Harbin Medical University (Ethics No.KY2016-090) and were in compliance with the principles and regulations of laboratory animal care.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Figure 1
(PNG 2050 kb)
Supplementary Figure 2
(PNG 1789 kb)
Supplementary Figure 3
(PNG 2066 kb)
Supplementary Figure 4
(PNG 1717 kb)
Supplementary Figure 5
(PNG 1565 kb)
Supplementary Table S1.
Differentially expressed lncRNA identified by ‘edge R’ R package. (XLSX 34 kb)
Supplementary Table S2.
Differentially expressed mRNA identified by ‘edge R’ R package. (XLSX 36 kb)
Supplementary Table S3.
AS associated lncRNA-mRNA pairs based on hypergeometric test. (XLSX 708 kb)
Supplementary Table S4.
GO enrichment analysis for the first near mRNA neighbors of lncRNA MSTRG.5258.4 in ASLMN. (XLS 25 kb)
Supplementary Table S5.
KEGG enrichment analysis for the first near mRNA neighbors of lncRNA MSTRG.5258.4 in ASLMN. (XLS 27 kb)
Rights and permissions
About this article
Cite this article
Wu, Y., Zhang, F., Li, X. et al. Systematic analysis of lncRNA expression profiles and atherosclerosis-associated lncRNA-mRNA network revealing functional lncRNAs in carotid atherosclerotic rabbit models. Funct Integr Genomics 20, 103–115 (2020). https://doi.org/10.1007/s10142-019-00705-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-019-00705-z