Abstract
Glyoxalase pathway, ubiquitously found in all organisms from prokaryotes to eukaryotes, consists of glyoxalase I (GLY I) and glyoxalase II (GLY II) enzymes, which detoxify a cytotoxic molecule, methylglyoxal (MG). Increase in MG has been correlated with various diseases in humans and different abiotic stresses in plants. We have previously shown that overproduction of GLY I and/or GLY II enzymes in transgenic plants provide tolerance towards salinity and heavy metal stresses. We have identified nineteen potential GLY I and four GLY II proteins in rice and twenty two GLY I and nine GLY II proteins in Arabidopsis. An analysis of complete set of genes coding for the glyoxalase proteins in these two genomes is presented, including classification and chromosomal distribution. Expression profiling of these genes has been performed in response to multiple abiotic stresses, in different tissues and during various stages of vegetative and reproductive development using publicly available databases (massively parallel signature sequencing and microarray). AtGLYI8, OsGLYI3, and OsGLYI10 expresses constitutively high in seeds while AtGLYI4, AtGLYI7, OsGLYI6, and OsGLYI11 are highly stress inducible. To complement this analyses, qRT-PCR is performed in two contrasting rice genotypes, i.e., IR64 and Pokkali where OsGLYI6 and OsGLYI11 are found to be highly stress inducible.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The glyoxalase system has been documented as a major pathway for detoxification of methylglyoxal (MG) in living systems. MG is a cytotoxic α-ketoaldehyde by-product of glycolysis which forms adducts with protein and nucleic acid molecules and is thus deleterious to biological systems (Speer et al. 2003; Li et al. 2008). Irreversible covalent adducts formed by the interaction of MG with proteins are called “advanced glycation end products” or AGEs (Thornalley 2007). The modification of a protein to an AGE frequently eliminates or reduces function of the molecule (Vander Jagt and Hunsaker 2003). MG, a three carbon metabolite, exists ubiquitously across the genera from microorganisms to higher eukaryotes. MG level increases upon exposure to various abiotic stresses such as salinity, drought, and cold in plant systems (Yadav et al. 2005a). Two metalloenzymes, i.e., glyoxalase I (GLY I) and glyoxalase II (GLY II) comprise the glyoxalase system wherein, the reduced glutathione (GSH) participates as a cosubstrate. GLY I uses one molecule of GSH to convert MG to S-D-lactoylglutathione, which is converted to D-lactic acid by GLY II and one molecule of GSH is recycled back into the system (Racker 1951).

Glyoxalase pathway enzymes are well studied in microbial and animal systems (Rhee et al. 1987; Thornalley 1990). However, the presence and role of glyoxalase pathway in plants has been shown only recently where the two enzymes GLY I and GLY II have been purified and characterized from different plant species (Norton et al. 1990; Talesa et al. 1990; Deswal et al. 1993; Paulus et al. 1993; Singla-Pareek et al. 2003; Yadav et al. 2007). The GLY I and GLY II activity has also been shown to be stimulated under various abiotic stresses and developmental cues, which indicates their role in stress response as well as in development (Ramaswamy et al. 1984; Sethi et al. 1988; Deswal et al. 1993; Veena et al. 1999; Yadav et al. 2005a; Lin et al. 2010; Singla-Pareek et al. 2003, 2006, 2008).
In the present manuscript, genome-wide analysis for GLY I and GLY II genes has been carried out in Arabidopsis and rice genomes, which indicated the presence of eleven GLY I genes each in Arabidopsis and rice and five GLY II genes in Arabidopsis and three in rice. Detailed expression profiling of these genes based on publicly available microarray and massively parallel signature sequencing (MPSS) data has also been carried out that indicate glyoxalase family members to be differentially regulated in response to various abiotic stresses and developmental cues. The AtGLYI4, AtGLYI7, OsGLYI6, and OsGLYI11, in particular, are most likely key components of the gene network interaction responsible for detoxifying MG. Further validation of some of these results has also been performed by qRT-PCR analysis using IR64 and Pokkali rice, which show contrasting response towards salinity.
Materials and methods
Search and analysis of glyoxalase proteins in Arabidopsis and rice
The hidden Markov Model profile of glyoxalase (PF00903) and metallo-beta-lactamase (PF00753) domains in GLY I and GLY II proteins, respectively, was retrieved from Pfam (http://www.sanger.ac.uk/Software/Pfam). These profiles were utilized to identify all the putative glyoxalase proteins by searching against the annotated proteins in the whole Arabidopsis (TAIR 9) and rice genomes (TIGR version 6.1) with an e value cutoff of 0.001 as described earlier (Pareek et al. 2006; Kushwaha et al. 2009). All the proteins were checked individually, and rice LOC_Os02g07160 was manually removed as it showed that it matched with proteins other than lactoylglutathione lyase (GLY I). The metallo-beta-lactamase domain was found to be present in several proteins in rice and Arabidopsis genome. However, only three proteins in rice and five proteins in Arabidopsis showed significant similarity with known GLY II (hydroxyacylglutathione hydrolase) proteins and were included in this study. For nomenclature of various glyoxalase proteins, prefix “At” and “Os” were added in Arabidopsis thaliana and Oryza sativa, respectively, followed by GLY I or GLY II and Arabic numbers. For representing alternatively spliced forms, Arabic numbers were added after “.” sign.
Genomic distribution of GLY I and GLY II genes on Arabidopsis and rice chromosomes
Position of each of the glyoxalase genes on Arabidopsis and rice chromosome pseudomolecules available at TAIR version 9 and TIGR version 6.1 was determined by BLASTn search.
Expression analysis using MPSS database
Expression evidence from MPSS tags was determined from Arabidopsis (http://mpss.udel.edu/at) and rice (http://mpss.udel.edu/rice) MPSS project website mapped to Arabidopsis and rice gene models, respectively. The signature was considered to be significant if it uniquely identifies an individual gene and shows perfect match (100% identity over 100% length of the tag). The normalized abundance (tags per million) of these signatures for a given gene in the given library represents a quantitative estimate of expression of that gene.
The description of MPSS libraries in A. thaliana and O. sativa is available at http://mpss.udel.edu/at and http://mpss.udel.edu/rice respectively.
Expression analysis using microarray data
The microarray data for Arabidopsis glyoxalase genes under various abiotic stress conditions, such as salt, drought, osmotic, cold, heat, oxidative, genotoxic, wounding, and UV/B stress (Kilian et al. 2007), and different developmental stages (Schmid et al. 2005) were taken from the AtGenExpress (http://jsp.weigelworld.org/expviz/expviz.jsp). For stress treatments, datasets obtained were corresponding to different time points of stress viz. 0.5, 1, 3, 6, 12, and 24 h for root and shoot tissues. Fold increase in transcript abundance under stress conditions was calculated with respect to their controls. For the developmental stage dataset, the raw Affymetrix values were log10 transformed. Heatmaps were generated, and hierarchical clustering was done using the Institute for Genomic Research MeV software package (Eisen et al. 1998).
For microarray analysis of rice glyoxalase genes, Affymetrix GeneChip Rice Genome Arrays (http://www.ncbi.nlm.nih.gov/geo/; Gene Expression Omnibus platform accession nos. GSE6893 and GSE6901) were used. The values were log2 transformed and heatmaps generated and hierarchical clustering done using the Institute for Genomic Research MeV software package (Eisen et al. 1998). Here, the rice panicle and seed developmental stages are divided into six and five major categories, respectively, based on the studies by Itoh et al. (2005), information from oryzabase (http://www.shigen.nig.ac.jp/rice/oryzabase/top/top.jsp), and some histochemical analysis (Arora et al. 2007). The description of microarray dataset in O. sativa is: ML—mature leaf (collected before pollination), R—roots of 7-day-old seedling, SD—7-day-old seedling, P1 (0–3 cm panicle), P2 (3–5 cm panicle), P3 (5–10 cm panicle), P4 (10–15 cm panicle), P5 (15–22 cm panicle), P6 (22–30 cm panicle), S1 (0–2 DAP), S2 (3–4 DAP), S3 (5–10 DAP), S4 (11–20 DAP), S5 (21–29 DAP), CS—cold stress, DS—dehydration stress, and SS—salt stress.
Plant material and stress treatment for qRT-PCR analysis
Seedlings of Pokkali and IR64 rice cultivars were grown under control conditions in growth chamber at 28 ± 2°C and 16 h photoperiod. The seeds were surface sterilized with 1% Bavistin for 20 min and allowed to germinate in a hydroponic system. Germinated seeds were supplied with modified Yoshida medium (Yoshida et al. 1972). After 7 days, various treatments (200 mM NaCl for salt stress or 5 mM H2O2 for oxidative stress or 10 mM MG for methylglyoxal stress; dissolved in Yoshida medium) were given. After 16 h, shoots were cut, weighed, and packed in aluminum foil; labeled properly, and then frozen in liquid nitrogen. Untreated seedlings were used as control.
Real-time PCR
Total RNA was isolated from the shoot tissues of control and stressed plant samples using RaFlex™ solution I and solution II (GeNei, India) as per the manufacturer’s protocol. Poly A+ RNA was isolated from the total RNA samples as described earlier (Kumari et al. 2009). First-strand cDNA synthesis was done using RevertAid™ RNAse H minus cDNA synthesis kit as per manufacturer’s instructions (Fermentas Life Sciences, USA). The primers for real-time PCR were designed using Primer3 software from the 3′-UTR regions of each of the GLY I and GLY II genes (Table 1). The PCR mixture contained 5 μl of cDNA (50 times diluted), 12.5 μl of 2× SYBR Green PCR Master Mix (Applied Biosystems, USA) and 200 nM of each gene-specific primer in a final volume of 25 μl. The qRT-PCR was performed employing ABI Prism 7500 Sequence Detection System and software (PE Applied Biosystems). All the PCRs were performed: 2 min at 50°C, 10 min at 95°C, and 40 cycles of 15 s at 95°C, 1 min at 55°C, and 30 s at 72°C in 96-well optical reaction plates (Applied Biosystems). The specificity of the amplification was tested by dissociation curve analysis and agarose gel electrophoresis. Three technical replicates were analyzed for each sample. The relative expression ratio of each glyoxalase gene was calculated using delta Ct or comparative Ct value method (Livak and Schmittgen 2001).
Results
Identification of genes for GLY I and GLY II proteins in Arabidopsis and rice genomes
We have identified the glyoxalase proteins within the Arabidopsis and rice genomes. Proteins having putative function as lactoylglutathione lyase and containing glyoxalase domain (PF00903) have been classified as GLY I proteins. Similarly, proteins having putative function as hydroxyacylglutathione hydrolase and containing metallo-beta-lactamase (PF00753) domain have been classified as GLY II proteins. In Arabidopsis, twenty two GLY I proteins are encoded by eleven genes and nine GLY II proteins by five genes. However, in rice, a total of nineteen GLY I proteins are encoded by eleven genes and four GLY II proteins by three genes. This indicated presence of alternatively spliced products for both GLY I and GLY II genes in the two genomes. However, the number of alternative spliced products for both GLY I and GLY II genes are lesser in rice as compared with Arabidopsis. In case of GLY II, even the number of genes is less in rice as compared to Arabidopsis.
In Arabidopsis, six GLY I gene members, i.e., AtGLYI2, AtGLYI3, AtGLYI4, AtGLYI6, AtGLYI7, and AtGLYI9, and four GLY II gene members, i.e., AtGLYII1, AtGLYII2, AtGLYII3, and AtGLYII4, show alternative spliced forms; while OsGLYI5, OsGLYI7, OsGLYI9, OsGLYI11, and OsGLYII1 were found to be alternatively spliced in rice (Tables 2, 3).
Chromosomal distribution of GLY I and GLY II genes
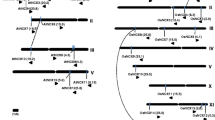
The physical distribution of glyoxalase genes on different chromosomes of Arabidopsis and rice is shown in Fig.1. In Arabidopsis, seven GLY I genes are located on chromosome I and two each on chromosomes II and V (Fig. 1a). In rice, three GLY I genes are located on chromosome V and two each on chromosomes III and VII and one each on chromosomes I, II, IV, and VIII (Fig. 1c). For the chromosomal distribution of GLY II genes, two genes are located on chromosomes I and II each and one on chromosome III in Arabidopsis (Fig. 1b), while in rice, chromosomes I, III, and IX have one GLY II gene each (Fig. 1d).
Chromosomal distribution and segmental duplication events for a, c GLY I and b, d GLY II genes of Arabidopsis and rice, respectively. Only the chromosomes having glyoxalase genes are shown and their number is indicated by Roman numbers. The scale is in megabase (Mb), and the centromeric regions are indicated by ellipses. The chromosomal positions (Mb) and orientation of each of the glyoxalase genes for rice and Arabidopsis are shown as horizontal bars and arrows, respectively. The glyoxalase genes present on duplicated chromosomal segments are connected by broken lines
MPSS analysis of genes encoding glyoxalase proteins
We have used the publicly available 17 and 20 bp MPSS libraries of Arabidopsis and rice and obtained the expression evidence for all the glyoxalase genes identified in this study. Heat maps for GLY I and GLY II genes of Arabidopsis and rice have been generated using TIGR MeV software, which provides information about tissue and stage specificity of the expression of these genes (Fig. 2). Expression analyses of Arabidopsis GLY I genes revealed that AtGLYI6 has higher constitutive expression in almost all the tissues analyzed, thereby indicating its function at all the developmental stages (Fig. 2a). The expression of AtGLYI2 is moderate in tissues other than leaf treated with salicylic acid for 52 h (S52) and in germinating seedling (GSE). Similarly, expression of AtGLYI11 is constitutive in all the tissues except callus (CAF) and untreated root tissues (ROF). The expression of AtGLYI3 is specifically higher in callus, inflorescence, root, and silique, while expression of AtGLYI4 is higher in callus and root than other tissues suggesting their role in specific tissues. Among all the Arabidopsis GLY I genes, expression of AtGLYI1 and AtGLYI10 is lowest in all the tissues.
Expression profile of GLY I and GLY II genes from MPSS data (name of the MPSS libraries are shown below each heat map) in different tissues/organs of a Arabidopsis and b rice. The description of MPSS libraries is given at the website mentioned in the “Materials and methods” section. The heatmaps have been generated using TIGR MeV software package and represent hierarchical clustering of average log signal values of all GLY I and GLY II genes in various tissues/organs (indicated at the top of each lane). The color bar below represents relative expression values; thereby, green color represents lowest expression levels, black represents medium expression levels, and red signifies highest expression level
Based on MPSS data analysis of GLY I genes, OsGLYI11 transcript is found to be the most abundant in various tissues of rice (Fig. 2b). On the other hand, OsGLYI1 and OsGLYI4 show the lowest level of expression in all plant parts. OsGLYI2 and OsGLYI9 genes express more in leaves than other tissues. The OsGLYI6, OsGLYI7, and OsGLYI8 show high expression in seeds.
Based on the analysis of GLY II genes of Arabidopsis, AtGLYII2 and AtGLYII4 are highly expressed in all the tissues other than leaf treated with salicylic acid for 52 h (S52). The AtGLYII1 has higher expression in callus, leaf, and silique than other tissues, while AtGLYII3 is highly expressed in callus, inflorescence, and silique. In rice, OsGLYII1 is highly induced by salt stress in both shoot (NSL) and root (NSR) tissues. OsGLYII2 shows high expression in all plant parts except in seed where its expression is lower. OsGLYII3 is the only GLY II gene in rice which is expressed in seeds (Fig. 2b). This gene also shows high expression under cold (refer NCL lane) and salinity (refer NSL lane) stress. These results are in corroboration to our previous finding where transcript induction for OsGLYII3 has been reported in response to several stresses such as desiccation, salinity, heat, cold, ABA, and salicylic acid (Yadav et al. 2007). OsGLYII3 has also been well characterized by our group and shown to confer tolerance towards abiotic stresses such as salinity, heavy metal (Singla-Pareek et al. 2003, 2006, 2008), and MG (Yadav et al. 2007).
Microarray data analysis of glyoxalase genes in Arabidopsis
Detailed analyses of Arabidopsis and rice microarray data reveal various glyoxalase genes to be differentially regulated at different time points in response to various abiotic stresses and developmental stages. The expression level of GLY I genes has been shown to be affected by exogenous factors like hormone, light, and polyamines (Sethi et al. 1988; Deswal et al. 1993; Chakravarty and Sopory 1998). Previous studies showed that GLY I transcript level increased in response to salinity and osmotic stress (Espartero et al. 1995). The upregulation of Brassica juncea GLY I in response to salt; dehydration and heavy metal like zinc stress has also been documented (Veena et al. 1999). In shoots, AtGLYI7 is shown to be highly upregulated in response to salt, osmotic stress, and wounding; whereas in roots, it is induced under all stresses (Fig. 3a, b). Expression of AtGLYI4 is also induced in response to most of the abiotic stresses at different time points in both shoot and root tissues except under salt stress in roots. AtGLYI10 is induced at early time points (0.5 to 1 h) in shoots in response to all the stresses except salinity. In roots, expression of AtGLYI10 is induced more by heat and cold stress. In shoots, expression of AtGLYI3 is highly induced in response to salt and drought and moderately induced in cold, heat, and wounding stress, while in roots, its expression is moderately induced under all the stresses, except oxidative stress and UV/B. Expression of AtGLYI11 is highly induced under salt and osmotic stress in both shoot and root tissues. In response to heat stress, it is highly induced in root tissues (Fig. 3a, b). Expression of AtGLYI6 is moderately induced by most of the abiotic stresses such as salinity, drought, cold, and heat in both shoot and root tissues.
Heatmap analyses of GLY I and GLY II genes from Arabidopsis in a shoot and b root tissues. The microarray data for expression of glyoxalase genes under various abiotic stress conditions such as salt, drought, osmotic, cold, heat, oxidative, genotoxic, wounding, and UV/B stress has been retrieved from TAIR (ver 9). The datasets obtained are corresponding to shoot and root tissues at different time sets of stress namely 0.5 h, 1 h, 3 h, 6 h, 12 h and 24 h and analyzed with respect to the control. Color bar below represents relative expression values, thereby green color representing lowest expression levels, black medium expression levels and red signifies highest expression level. The hierarchical clustering is performed and heatmaps have been generated using TIGR MeV software package
Amongst the GLY II genes, the expression of AtGLYII2 is highly upregulated in response to osmotic, salt, and drought stress in both shoot and root tissues whereas moderately induced in response to all other stresses (Fig. 3a, b). The AtGLYII1 is induced in all the stresses except cold and wounding in shoots, while in roots, this gene is upregulated in all the stresses except cold stress. AtGLYII4 is induced most profoundly by salt, drought, osmotic, cold, and UV/B stresses in shoot tissues, while in roots, it is moderately induced by all stresses except salt and drought stress (Fig. 3a, b).
Microarray data have also been analyzed with respect to different vegetative and reproductive developmental stages of Arabidopsis. This reveals that AtGLYI2, AtGLYI3, AtGLYI6, AtGLYI9, and AtGLYI11 are highly expressed at all the stages of development. The AtGLYI8 is specifically expressed in seeds, which possibly indicates its important role in seed development (Fig. 4a, b). AtGLYI4 expresses in roots, flower, and seeds and AtGLYI7 expresses in flower and seeds, but not in roots. All the GLY II genes of Arabidopsis express at all the stages of development, though the expression level is higher for AtGLYII2 and AtGLYII5 (Fig. 4).
Expression analysis of GLY I and GLY II genes at different developmental stages of Arabidopsis using microarray data. Heatmap and hierarchical cluster display differential expression profile for glyoxalase genes. Color bar at the left represents log10 expression values, thereby green color representing lower expression level, black represents medium expression level, and red signifies higher expression level. Developmental stages used for expression profiling are mentioned on the right side. Various stages are listed (from top to bottom) in the temporal order of development. In case of 8 weeks old siliques and seeds, stages 3, 4, 5, 6, 7, 8, 9 and 10 represent-mid globular to early heart embryos, early to late heart embryos, late heart to mid torpedo embryos, mid to late torpedo embryos, late torpedo to early walking-stick embryos, walking-stick to early curled cotyledons, curled cotyledons to early green cotyledons, green cotyledons, respectively
Microarray data analysis of glyoxalase genes in rice
The microarray data analyses revealed that among all the GLY I genes, OsGLYI3 and OsGLYI10 are seed specific (Fig. 5). OsGLYI3 is highly expressed at 3–29 days after pollination (DAP; S2–S5), whereas the OsGLYI10 is expressed at 5–29 DAP stage (S3–S5) of the seed. Since the heading stage is one of the most salinity sensitive stages of rice (Lutts et al. 1995), seed specific GLY I genes might have an important role to play under stress conditions. Interestingly, AtGLYI8 in Arabidopsis—a homologue of OsGLYI10, also expresses specifically in the silique/seed. The OsGLYI1 is expressed only in roots and seedlings, while OsGLYI5 is expressed only at the mature panicle stage (P6; 22–29 cm panicle) and in salt stressed seedlings (SS). Expression of OsGLYI2, OsGLYI4, OsGLYI6, OsGLYI7, OsGLYI8, and OsGLYI11 is high at all the developmental stages and stress conditions. Interaction of various developmental processes and environmental cues has been reported previously (Chen et al. 2002; Potocky et al. 2003; Jain et al. 2007). Such an interaction may be responsible for co-regulation of certain genes by both environmental factors and development (Jain et al. 2008). The expression level of OsGLYI9 is negligible in roots and early stages of panicle development. However, it is highly responsive to cold, desiccation, and salt stress. In case of GLY II genes, OsGLYII1 shows high expression preferentially in root tissues (Fig. 5). OsGLYII2 and OsGLYII3 are expressed at all the stages of development and stress conditions. Interestingly, all the GLY II genes also showed high transcript accumulation under cold, desiccation, and salt stress conditions.
Expression analysis of rice GLY I and GLY II genes at different developmental stages and stress conditions using microarray data. Heatmap and hierarchical cluster display of expression profile for 11 GLY I and three GLY II genes shows their different levels of expression. Color bar at the base represents log2 expression values, thereby green color representing lowest expression levels, black medium expression levels and red signifies highest expression level. Developmental stages used for expression profiling are mentioned on top of each column. Panicle and seed stages have been listed in the temporal order of development. The description of these stages is given in the “Materials and methods” section. In seedlings, CS, DS, and SS represent cold, dehydration, and salt stress, respectively
qRT-PCR-based expression analysis of different glyoxalase genes in contrasting genotypes of rice in response to salinity stress, oxidative stress, and exogenous MG
The relative transcript level of all the GLY I and GLY II gene family members of rice is determined in response to salt stress, oxidative stress (H2O2), and MG (the main substrate for the glyoxalase system, whose level increases in response to various abiotic stresses), in two contrasting genotypes of rice with respect to salinity tolerance viz. IR64 (salt sensitive) and Pokkali (salt tolerant; Fig. 6). In this analysis, differential expression of alternatively spliced forms of glyoxalase genes is also quantified based on difference in their 3′ UTR regions that aided the designing of specific primers for each of the spliced forms. The OsGLYI1 and OsGLYI10 could not be amplified from the shoot cDNA used in this study; these are therefore not included in the expression analysis. The EST database of rice (TIGR version 6.1) has also not registered any EST for OsGLYI1 and OsGLYI10 in rice shoot tissues; however, the rice microarray data indicated their expression to be highly root and seed specific (Fig. 5). We are investigating further to establish the root and seed specific nature of these GLY I members.
Histogram depicting relative transcript abundance of all the a GLY I and b GLY II gene members, in contrasting rice cvs. IR64 and Pokkali, based on qRT-PCR analysis. Real time PCR analysis was done with cDNA template generated from shoot tissue of 7-day-old stressed (NaCl, MG, and H2O2) or control seedlings. As OsGLYI1, OsGLYI10, OsGLYI7.2, and OsGLYII1.2 could not be amplified; they are not included in the real-time PCR analysis. Heatmap and hierarchical cluster display of expression profile for c GLY I and d GLY II genes showing different levels of expression in contrasting rice cultivars in response to stress. Color bar at the base represents expression values in terms of fold change, thereby green color representing lowest expression levels, black medium expression levels and red signifies highest expression level
The real-time PCR data reveals several interesting aspects pertaining to expression of glyoxalase family members, which are in good correlation with the microarray and MPSS data analysis. Alternative spliced forms of OsGLYI11 (OsGLYI11/OsGLYI11.2/OsGLYI11.3) and OsGLYI6 (OsGLYI6.3/OsGLYI6.4) showed high level of expression in both sensitive and tolerant rice genotypes under salinity, oxidative, and MG stress (Fig. 6a), thereby indicating their multiple stress responsive nature. Since exposure to abiotic stresses leads to increased accumulation of cytotoxic MG (Yadav et al. 2005a) and these genes have a role in MG detoxification, importance of these genes as potential candidate genes for stress tolerance engineering can be suggested.
Expression of OsGLYI2 and OsGLYI9.2 in IR64 (Fig. 6a, c) and OsGLYI2, OsGLYI8, OsGLYI9.1, and OsGLYI9.2 in Pokkali (Fig. 6a, d) is not induced by any of the stresses applied in the present study. This implies that they might be involved in regulating some other biological process rather than stress response. In Datura and Brassica, GLY I activity has been correlated with development, besides stress response (Ramaswamy et al. 1984; Sethi et al. 1988). Once the genome sequence of these plant species is available, it would be interesting to analyze whether their cognate GLY I gene responsible towards developmental regulation is orthologous to the gene members showing downregulation in this study.
OsGLYI3 shows a stark difference in the expression pattern among the two contrasting genotypes of rice. Pokkali accumulates very high levels of transcript for this gene under all the stresses tested here, while IR64 shows its transcript accumulation only under salinity stress (Fig. 6c, d). The differential accumulation of stress responsive genes has been reported recently in contrasting rice cultivars (Kawasaki et al. 2001; Karan et al. 2009; Kumari et al. 2009). The high abundance of transcript of OsGLYI3 under all stresses indicates a possible cross talk under various stresses in Pokkali and warrants further investigations.
Some GLY I gene members show induction in response to specific but distinct stress in contrasting cultivars of rice. OsGLYI3, OsGLYI4, OsGLYI5, OsGLYI6.1, OsGLYI8, and OsGLYI9.1 are found to be specifically upregulated under salinity stress in IR64 (Fig. 6c). In Pokkali, OsGLYI4 is upregulated under both salinity and MG stress, while OsGLYI5 is upregulated only under MG stress. The OsGLYI9.1 shows induction only in response to oxidative stress (Fig. 6d). OsGLYI7.1 transcript accumulates in response to both salinity and oxidative stress in IR64, while in Pokkali, it accumulates in response to salinity and MG (Fig. 6c, d). Stress specific differential regulation of several genes has been reported in previous studies (Karan et al. 2009; Kumar et al. 2009). These results indicate GLY I gene members to be playing distinct roles in response to specific environmental cues in contrasting rice cultivars.
Expression analysis carried out for GLY II gene family members under salinity, oxidative stress, and MG stress shows their differential stress responsive nature in contrasting cultivars. OsGLYII1 is accumulated under salinity, MG, and oxidative stress in both IR64 and Pokkali (Fig. 6b, c, d). OsGLYII2 and OsGLYII3 show salinity inducible expression in IR64 only while under other stresses; these members are downregulated. The role of OsGLYII3 has already been functionally validated wherein overexpression of this gene provides survival advantage to the plants under salinity stress (Singla-Pareek et al. 2003, 2008).
Discussion
Abiotic stresses lead to the accumulation of higher MG levels in plants (Yadav et al. 2005a). It has been shown earlier that the glyoxalase pathway and its engineering have a direct correlation with tolerance towards abiotic stresses (Veena et al. 1999; Singla-Pareek et al. 2003, 2006, 2008). Overexpression of glyoxalase pathway enzymes can keep a check on the elevation of MG level and also helps maintain a higher “reduced to oxidized” glutathione ratio (Yadav et al. 2005a, b). This may be the basis of improved abiotic stress tolerance of glyoxalase-overexpressing transgenic plants (Singla-Pareek et al. 2003, 2006). One aspect of response to stress is at the transcriptional level, which involves alteration in the expression of genes (Tester and Davenport 2003). For these reasons, we wanted to study if glyoxalase genes are differentially regulated in response to various abiotic stress factors. We have analyzed the expression pattern of GLY I and GLY II genes with publicly available MPSS and microarray data. The microarray data has revealed that many glyoxalase genes in Arabidopsis and rice are strongly stress responsive.
We have previously shown the usefulness of contrasting genotypes viz. IR64 (salt sensitive) and Pokkali (salt tolerant) to understand the molecular basis of differential sensitivity towards salinity stress (Kumari et al. 2009). qRT-PCR analysis revealed that each member is uniquely expressed under a given set of conditions in the two cultivars. OsGLYI3 was found to be one of the most stress responsive GLY I in salt tolerant Pokkali. On the other hand, OsGLYI6.3/6.4 and OsGLYI11.2 were strongly induced under salinity stress only in the sensitive cultivar IR64. Among GLY II genes, OsGLYII1 seems to be the most stress responsive in both genotypes. These genes may thus serve as good candidates for raising stress-tolerant transgenic crop plants. Another interesting observation from the qRT-PCR analysis was that in response to MG stress, all alternative spliced forms of OsGLYI11 were induced, which indicates that OsGLYI11 is a substrate inducible GLY I.
References
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8:242
Chakravarty TN, Sopory SK (1998) Blue light stimulation of cell proliferation and glyoxalase I activity in callus cultures of Amaranthus paniculatus. Plant Sci 132:63–69
Chen W, Provart NJ, Glazebrook J, Katagiri F, Chang HS, Eulgem T, Mauch F, Luan S, Zou G, Whitham SA et al (2002) Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14:559–574
Deswal R, Chakaravarty TN, Sopory SK (1993) The glyoxalase system in higher plants: regulation in growth and differentiation. Biochem Soc Trans 21:527–530
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Espartero J, Sanchez-Aguayo I, Pardo JM (1995) Molecular characterization of glyoxalase-I from a higher plant; upregulation by stress. Plant Mol Biol 29:1223–1233
Itoh J, Nonomura K, Ikeda K, Yamaki S, Inukai Y, Yamagishi H, Kitano H, Nagato Y (2005) Rice plant development: from zygote to spikelet. Plant Cell Physiol 46:23–47
Jain M, Nijhawan A, Arora R, Agarwal P, Ray S, Sharma P, Kapoor S, Tyagi AK, Khurana JP (2007) F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol 143:1467–1483
Jain M, Tyagi AK, Khurana JP (2008) Genome-wide identification, classification, evolutionary expansion and expression analyses of homeobox genes in rice. FEBS J 275:2845–2861
Karan R, Singla-Pareek SL, Pareek A (2009) Histidine kinase and response regulator genes as they relate to salinity tolerance in rice. Funct Integr Genomics 9:411–417
Kawasaki S, Borchert C, Deyholos M, Wang H, Brazille S, Kawai K, Galbraith D, Bohnert HJ (2001) Gene expression profiles during the initial phase of salt stress in rice. Plant Cell 13:889–905
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363
Kumar G, Purty RS, Sharma MP, Singla-Pareek SL, Pareek A (2009) Physiological responses among Brassica species under salinity stress show strong correlation with transcript abundance for SOS pathway-related genes. J Plant Physiol 166:507–520
Kumari S, Sabharwal VP, Kushwaha HR, Sopory SK, Singla-Pareek SL, Pareek A (2009) Transcriptome map for seedling stage specific salinity stress response indicates a specific set of genes as candidate for saline tolerance in Oryza sativa L. Funct Integr Genomics 9:109–123
Kushwaha HR, Singh AK, Sopory SK, Singla-Pareek SL, Pareek A (2009) Genome wide expression analysis of CBS domain containing proteins in Arabidopsis thaliana (L.) Heynh and Oryza sativa L. reveals their developmental and stress regulation. BMC Genomics 10:200
Li Y, Cohenford MA, Dutta U, Dain JA (2008) The structural modification of DNA nucleosides by nonenzymatic glycation: an in vitro study based on the reactions of glyoxal and methylglyoxal with 2′-deoxyguanosine. Anal Bioanal Chem 390:679–688
Lin F, Xu J, Shi J, Li H, Li B (2010) Molecular cloning and characterization of a novel glyoxalase I gene TaGLY I in wheat (Triticum aestivum L.). Mol Biol Rep 37:729–735
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Lutts S, Kinet JM, Bouharmont J (1995) Changes in plant response to NaCl during development of rice (Oryza sativa L.) varieties differing in salinity resistance. J Exp Bot 46:1843–1852
Norton SJ, Talesa V, Yuan WJ, Principato GB (1990) Glyoxalase I and glyoxalase II from Aloe vera: purification, characterization and comparison with animal glyoxalases. Biochem Int 22:411–418
Pareek A, Singh A, Kumar M, Kushwaha HR, Lynn AM, Singla-Pareek SL (2006) Whole-genome analysis of Oryza sativa reveals similar architecture of two-component signaling machinery with Arabidopsis. Plant Physiol 142:380–397
Paulus C, Kollner B, Jacobsen HJ (1993) Physiological and biochemical characterization of glyoxalase I, a general marker for cell proliferation, from a soybean cell suspension. Planta 189:561–566
Potocky M, Elias M, Profotova B, Novotna Z, Valentova O, Zarsky V (2003) Phosphatidic acid produced by phospholipase D is required for tobacco pollen tube growth. Planta 217:122–130
Racker E (1951) The mechanism of action of glyoxalase. J Biol Chem 190:685–696
Ramaswamy O, Pal S, Guha-Mukherjee S, Sopory SK (1984) Correlation of glyoxalase I activity with cell proliferation in Datura callus culture. Plant Cell Rep 3:121–124
Rhee H, Murata K, Kimura A (1987) Molecular cloning of the Pseudomonas putida glyoxalase I gene in Escherichia coli. Biochem Biophys Res Commun 147:831–838
Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Sethi U, Basu A, Guha-Mukherjee S (1988) Control of cell proliferation and differentiation by regulating polyamine biosynthesis in cultures of Brassica and its correlation with glyoxalase-I activity. Plant Sci 56:167–175
Singla-Pareek SL, Reddy MK, Sopory SK (2003) Genetic engineering of the glyoxalase pathway in tobacco leads to enhanced salinity tolerance. Proc Natl Acad Sci USA 100:14672–14677
Singla-Pareek SL, Yadav SK, Pareek A, Reddy MK, Sopory SK (2006) Transgenic tobacco overexpressing glyoxalase pathway enzymes grow and set viable seeds in zinc-spiked soils. Plant Physiol 140:613–623
Singla-Pareek SL, Yadav SK, Pareek A, Reddy MK, Sopory SK (2008) Enhancing salt tolerance in a crop plant by overexpression of glyoxalase II. Transgenic Res 17:171–180
Speer O, Morkunaite-Haimi S, Liobikas J, Franck M, Hensbo L, Linder MD, Kinnunen PKJ, Wallimann T, Eriksson O (2003) Rapid suppression of mitochondrial permeability transition by methylglyoxal. Role of reversible arginine modification. J Biol Chem 278:34757–34763
Talesa V, Rosi G, Contenti S, Mangiabene C, Lupattelli M, Norton SJ, Giovannini E, Principato GB (1990) Presence of glyoxalase II in mitochondria from spinach leaves: comparison with the enzyme from cytosol. Biochem Int 22:1115–1120
Tester M, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot (Lond) 91:503–527
Thornalley PJ (1990) The glyoxalase system: new developments towards functional characterization of a metabolic pathway fundamental to biological life. Biochem J 269:1–11
Thornalley PJ (2007) Dietary AGEs and ALEs and risk to human health by their interaction with the receptor for advanced glycation endproducts (RAGE)—an introduction. Mol Nutr Food Res 51:1107–1110
Vander Jagt DL, Hunsaker LA (2003) Methylglyoxal metabolism and diabetic complications: roles of aldose reductase, glyoxalase-I, betaine aldehyde dehydrogenase and 2-oxoaldehyde dehydrogenase. Chem Biol Interact 143–144:341–351
Veena, Reddy VS, Sopory SK (1999) Glyoxalase I from Brassica juncea: molecular cloning, regulation and its over-expression confer tolerance in transgenic tobacco under stress. Plant J 17:385–395
Yadav SK, Singla-Pareek SL, Ray M, Reddy MK, Sopory SK (2005a) Methylglyoxal levels in plants under salinity stress are dependent on glyoxalase I and glutathione. Biochem Biophys Res Commun 337:61–67
Yadav SK, Singla-Pareek SL, Reddy MK, Sopory SK (2005b) Transgenic tobacco plants overexpressing glyoxalase enzymes resist an increase in methylglyoxal and maintain higher reduced glutathione levels under salinity stress. FEBS Lett 579:6265–6271
Yadav SK, Singla-Pareek SL, Kumar M, Pareek A, Saxena M, Sarin NB, Sopory SK (2007) Characterization and functional validation of glyoxalase II from rice. Protein Expr Purif 51:126–132
Yoshida S, Forno DA, Cock JH, Gomez KA (1972) Laboratory manual for physiological studies of rice. International Rice Research Institute, Manila
Acknowledgments
The authors thankfully acknowledge the research funds received from International Centre for Genetic Engineering and Biotechnology (ICGEB), New Delhi. SLS-P thanks award of Innovative Young Biotechnologist award (IYBA) of Department of Biotechnology (DBT), Government of India. AM thanks ICGEB for providing pre-doctoral fellowship. Authors would also like to thank Dr. Sanjay Kapoor, University of Delhi South Campus, New Delhi, India, for providing help in retrieving rice microarray data and Professor RC Pant for the critical reading of the manuscript.
Conflicts of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mustafiz, A., Singh, A.K., Pareek, A. et al. Genome-wide analysis of rice and Arabidopsis identifies two glyoxalase genes that are highly expressed in abiotic stresses. Funct Integr Genomics 11, 293–305 (2011). https://doi.org/10.1007/s10142-010-0203-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-010-0203-2