Abstract
The gut microbiota of aquaculture species contributes to their food metabolism and regulates their health, which has been shown to vary during aquaculture progression of their hosts. However, limited research has examined the outcomes and mechanisms of these changes in the gut microbiota of hosts. Here, Kuruma shrimps from the beginning, middle, and late stages of aquaculture progression (about a time duration of 2 months between each stage) were collected and variations in the gut microbiota of Kuruma shrimp during the whole aquaculture process were examined. High-throughput sequencing demonstrated increases in the diversity and richness of the shrimp gut microbiota with aquaculture progression. In addition, the gut microbiota composition differed among cultural stages, with enrichment of Firmicutes, RF39, and Megamonas and a reduction in Proteobacteria in the mid-stage. Notably, only very few taxa were persistent in the shrimp gut microbiota during the whole aquaculture progression, while the number of taxa that specific to the end of aquaculture was high. Network analysis revealed increasing complexity of the shrimp gut microbiota during aquaculture progression. Moreover, the shrimp gut microbiota became significantly more stable towards the end of aquaculture. According to the results of neutral community model, contribution of stochastic processes for shaping the shrimp gut microbiota was elevated along the aquaculture progression. This study showed substantial variations in shrimp gut microbiota during aquaculture progression and explored the underlying mechanisms regulating these changes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gut microbiota comprises an extremely intricate microbial aggregate in the intestinal tract of animals, which is critical for immunoregulation, nutrient absorption, physiological responses, and host development (Fan and Pedersen 2021). Homeostasis of the gut microbiota is tightly correlated to the physiology and health of the host (Rinninella et al. 2019). Multiple genetic and physiological factors, including age, sex, and reproductive status, have been reported as the regulators for gut microbiota in diverse animals (Vujkovic-Cvijin et al. 2020). Besides, many non-host factors, including diet, climate, and living conditions, also participate to the variations of gut microbiota among different individuals (Leeming et al. 2019; Tasnim et al. 2017). As an important aquaculture species, associations between shrimp gut microbiota with its growth and health have been investigated previously. For example, the gut microbiota of shrimp with high growth rates showed more interspecies interactions and less phylogenetic clustering (Xiong et al. 2017a). High heterogeneity of the gut microbiota has also been seen in freshwater shrimp, possibly due to both the water-quality factors and surrounding bacterial communities (Zhao et al. 2018). An increase in the complexity of the gut microbiota network was observed in Pacific white shrimp with white feces syndrome, with Rhodobacterales, Vibrionales, and Flavobacteriales recognized as the keystone taxa (Dai et al. 2020). Shrimp aquaculture activities take several months from seeding to harvesting. Although the water quality and diet are relatively stable, the shrimp gut microbiota changes, with possible impacts on their growth and health. In this context, characterizing the gut microbiota dynamics of shrimp during aquaculture process may provide a basis for the improvement of aquaculture techniques and management strategies.
Although it is common knowledge that the gut microbiota of shrimp alters during aquaculture, the consequences and mechanisms of these changes remain unclear. Recent network analyses have unraveled the structure of such complex communities by exploring the co-occurrence arrangements of gut microbiota in some animals (Lee et al. 2021; Berry and Widder 2014; Pascoe et al. 2017). Co-occurrence networks, with nodes representing species and links representing species interactions, can be used to characterize the complexity of the gut microbiota (Hirano and Takemoto 2019). Gut microbiota with higher complexity generally possesses high functional redundancy due to the overlapping metabolic activities of different members (Moya and Ferrer 2016). In addition, a healthy gut microbiota tends to exhibit relative stability despite external disturbances (Yuan et al. 2021), which can also be characterized by some features of co-occurrence networks (Coyte et al. 2015). However, although network analysis can reveal the consequences of gut microbiota changes, the underlying mechanisms remain unclear. Microbial ecological theories suggest that microbial communities are determined by a compensate between deterministic and stochastic processes (Zhao et al. 2022). In communities controlled by deterministic processes, taxa take up specific ecological niches, and the occurrence of which can be predicted (Vanwonterghem et al. 2014). The neutral hypothesis, on the other hand, allows for multiple species to occupy similar or overlapping areas, and for the relative abundance of species within the community to change as a result of random fluctuations (Sloan et al. 2006). Variations in the co-occurrence network for shrimp gut microbiota during aquaculture progression have not yet been investigated. Moreover, the assembly mechanisms shaping the gut microbiota network may provide useful information for ecological aquaculture.
In this study, we addressed the following unresolved issues: (i) how does shrimp aquaculture progression affect the gut microbiota at the aspects of diversity, composition, and function? (ii) do the network complexity and stability of the shrimp gut microbiota vary along the aquaculture process? and (iii) do deterministic or stochastic processes dominate the gut microbiota assembly of shrimps? To answer these questions, we followed the temporal dynamics of the gut microbiota in Kuruma shrimp (Penaeus japonicus) throughout the whole aquaculture progression using high-throughput sequencing and analyzed changes in the shrimp gut microbiota. We also used co-occurrence network analyses to unveil variations in the complexity and stability of the gut microbiota and determined the underlying assembly mechanisms using a neutral community model (NCM). The findings of our research greatly expanded our knowledge of the ecological mechanism affecting the shrimp gut microbiota during aquaculture progression.
Materials and Methods
Aquaculture Progression and Sampling
Kuruma shrimps were obtained from a cultural pond in the coastal area of Dalian, China. The surface area of the pond was about 7 hm2 with a 1.5–2.0 m water depth, with twice water-exchanges per month. Kuruma shrimp seedlings were put into the pond in late May and harvested in late September. The animal concentrations and diets are shown in Table S1. The use of therapeutic or growth-promoting compounds, such as antibiotics, was strictly prohibited during the aquaculture process. Ten healthy individuals were randomly selected from the aquaculture pond on June 3 (initial, several days after releasing seedlings into the ponds), July 27 (Mid), and September 27 (end, the time of harvest) 2021, respectively, and dissected to collect the intestinal tracts. The intestinal contents were gently squeezed out from the intestinal tracts and kept at – 80 °C for further analyses.
DNA Extraction and High-Throughput Sequencing
DNA of gut microbiota was obtained from the contents of the intestinal samples using a QIAamp Power Fecal DNA Kit (Qiagen, CA, USA). Successful DNA extraction was verified by agarose gel electrophoresis (1% concentration). Subsequently, Nanodrop 2000 (ThermoFisher, CA, USA) was applied to estimate the purity and concentration of successfully extracted DNA. And then, all successfully extracted DNA were stored at −20 °C for further application. Gut microbiota of Kuruma shrimps were investigated using the amplicon sequencing based on bacterial 16S rRNA gene using the primers (341F–806R) for V3–V4 regions (Berg et al. 2012). PCR amplification, gel extraction of PCR products, and sequencing library construction were according to a previous study (Hou et al. 2021). Finally, the library of each sample was sequenced at BIOZERON Biotech. Co., Ltd. (Shanghai, China) using an Illumina Novaseq6000 platform based on the 250 bp paired-end strategy.
Sequence Data Processing
The raw reads were assigned to each sample on the basis of their unique barcodes and then the barcode sequences were truncated. Low-quality reads (Phred scores lower than 20, mismatches in the primers, homopolymer runs high than 8, containing ambiguous bases, or read length lower than 250 bp) were eliminated (Bokulich et al. 2013). Then, paired reads were assembled, chimeras were eliminated, and the clean data were aggregated to the amplicon sequence variants (ASVs) using the QIIME2 software with the DADA2 plugin unit (Bokulich et al. 2018). All ASVs were assigned to a taxonomy according to the SILVA database (Release 138) (Yilmaz et al. 2014), and then, singletons (ASVs with only one read) were dropped. Finally, the ASV abundance table was normalized based on the lowest read number among all samples (22,758).
Statistical Analysis
Four alpha diversity indices related to different facets, including richness (Chao1), diversity (Shannon), evenness (Pielou’s evenness index, Pielou_J), and evolution (Faith's phylogenetic diversity, Faith_pd) were estimated. Tukey’s honest significant difference (HSD) test was applied to confirm the variations in alpha diversity indices, major gut bacteria abundances, and functional pathways of shrimp gut microbiota during aquaculture progression (“multcomp” package in R). In addition, according to the Bray–Curtis distance, adonis test and principal coordinate analysis (PCoA) were executed (“vegan” package in R) to evaluate the impacts of shrimp aquaculture progression on the gut microbiota compositions. The specificity and occupancy of each ASV in samples from each aquaculture stage were calculated and projected onto a SPEC-OCCU plot (Gweon et al. 2021) to explore specialists present in different culture stages. Venn diagram analysis (“ggplot2” package in R) was executed to recognize the ASVs shared by all studied shrimps, and their composition was exhibited using a Sankey diagram (“d3Network” package in R). Tukey’s HSD test was also performed to determine if the sum abundances of these ASVs in individuals from different aquaculture stages were significantly different.
Spearman’s rank correlations among all ASVs of samples from each aquaculture stage were respectively calculated using the “WGCNA” package in R to visualize the co-occurrence network. Only ASVs detected in at least six out of 10 samples in each aquaculture stage were used for correlation analysis. Correlations between two ASVs with the |correlation coefficient|> 0.8 and p value adjusted by the Benjamini–Hochberg method < 0.05 were retained. Network graphs were visualized using the Gephi software (Bastian and Jacomy 2009) and the network topological parameters were calculated using the “igraph” package in R. To assess the gut microbiota, stability, robustness, vulnerability, and cohesion were estimated for the co-occurrence networks of samples from each aquaculture stage (Yuan et al. 2021). Changes in these stability indices during shrimp aquaculture progression were analyzed by Tukey’s HSD test. In addition, a NCM was applied to estimate the relative importance of stochastic processes shaping the shrimp gut microbiota (Sloan et al. 2006). The ASVs of samples from each aquaculture stage were then divided into three partitions according to whether they occurred more frequently (above partition), less frequently (below partition), or within (neutral partition) the 95% confidence interval of the NCM.
Results
Shifts in Diversity of Shrimp Gut Microbiota During Aquaculture Progression
The gut microbiota from each of 10 shrimp from the initial, mid, and end aquaculture stages were analyzed, and a total of 2,533,629 high-quality reads were obtained in total (Table S2). These reads were clustered into 7319 ASVs, which were annotated into 58 phyla, 138 classes, 334 orders, 527 families, 1125 genera, and 635 species (Table S3). All the ASVs were annotated at the phylum level, 75.71% were assigned to a genus, but only 10.75% were annotated at the species level (Fig. S1). These results suggested that analyses at the bacterial genus level enabled efficient assessment of relative intact shrimp gut microbiota.
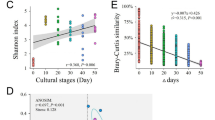
Significantly higher Shannon and Chao1 indices (Tukey’s HSD test, p < 0.05) were found in samples from the end stage than those from the initial and mid-stages (Fig. 1a). In contrast, the Pd_faith indices of samples from the mid and end stages were significantly lower (Tukey’s HSD test, p < 0.05) compared with individuals from the initial stage (Fig. 1). Moreover, the Pielou_J index of shrimp gut microbiota continued to increase during aquaculture progression and was significantly different (Tukey’s HSD test, p < 0.05) among the different aquaculture stages (Fig. 1). These results suggested that the richness, diversity, and evenness of the shrimp gut microbiota increased but the phylogenetic diversity decreased during aquaculture progression. PCoA revealed that the gut microbiota of shrimp from different aquaculture stages clustered separately (Fig. 1b). The adonis test also proved the significant effects (p < 0.05) of aquaculture progression on the shrimp gut microbiota, which could explain 34.2% of the gut microbiota variations (Fig. 1b).
a Variations in alpha diversity of shrimp gut microbiota among different aquaculture stages. Different lowercases letters in the same sub-figure represent significant differences among samples from different aquaculture stages (Tukey’s HSD test, p < 0.05). b PCoA and adonis test based on the Bray–Curtis distance for shrimp gut microbiota samples from different aquaculture stages
Changes in Composition of Shrimp Gut Microbiota During Aquaculture Progression
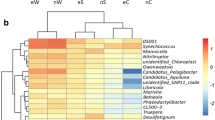
Firmicutes (25.57%) and Proteobacteria (25.18%) were the major bacterial phyla in the shrimp gut microbiota, followed by Desulfobacterota (8.77%), Bacteroidota (7.96%), Chloroflexi (6.80%), and Planctomycetota (5.57%) (Fig. S2). Firmicutes increased while Proteobacteria decreased in shrimps at the mid-stage in contrast to individuals at the initial and end stages (Fig. 2a). Additionally, Acidobacteriota and Verrucomicrobiota increased but Chloroflexi, Desulfobacterota, and Planctomycetota decreased in shrimp gut microbiota in line with aquaculture progression (Fig. 2a). In terms of bacterial genera, RF39 (9.70%) was the most abundant genus in the shrimp gut microbiota, followed by Megamonas (4.95%), OM190 (4.05%), Roseibacterium (3.84%), SBR1031 (2.34%), and Lactobacillus (Fig. S3). Among these, RF39 and Megamonas were significantly enriched (Tukey’s HSD test, p < 0.05) in shrimp from the mid-stage compared to individuals from the initial and end stages (Fig. 2b).
Specialists and Generalists in Shrimp Gut Microbiota During Aquaculture Progression
SPEC-OCCU plots were applied to explore potential specialists in the shrimp gut microbiota from different aquaculture stages. ASVs with specificity and occupancy ≥ 0.7 were identified as specialists, indicating that they were specific to an aquaculture stage and common in samples from this stage. The number of these specialists showed an increasing trend during aquaculture progression, representing 43.35%, 23.33%, and 46.72% of the total taxa in the initial, mid, and end stages, respectively (Fig. 3a). Proteobacteria were specialists in all three aquaculture stages, while notable specialists at the end stage included ASVs from multiple rare bacterial phyla (Others) (Fig. 3b). In contrast, only seven ASVs were shared by all the studied samples (generalists) from different stages of aquaculture progression (Fig. 3c). These generalists belonged to six bacterial phyla, including Firmicutes, Desulfobacterota, Proteobacteria, Actinobacteriota, Gemmatimonadota, and Latescibacterota (Fig. 3d). Among these, ASVs annotated to RF39, Megamonas, Desulfosarcinaceae, and Roseibacterium were more abundant (Fig. 3d). These generalists made up a significant proportion of the total gut microbiota, averaging about 10% in samples from the initial and end stages but > 40% in individuals from the mid-stage (Fig. 3e).
a SPEC-OCCU plot showing amplicon sequence variants (ASVs) in gut microbiota of shrimp from different aquaculture stages. b Pie charts showing the composition of specialists in each aquaculture stage. c Venn diagram exhibiting shared ASVs in shrimp gut microbiota during aquaculture progression. d Average relative abundance and taxonomy of generalists in all samples. e Total proportion of shared ASVs in gut microbiota of samples from different aquaculture stages. Different lowercase letters represent significant differences among samples from different aquaculture stages (Tukey’s HSD test, p < 0.05)
Co-occurrence Networks of Shrimp Gut Microbiota
Co-occurrence networks of the shrimp gut microbiota from the different aquaculture stages were constructed (Fig. 4a–c), and the topological parameters are given in Table 1. Networks for all aquaculture stages obeyed a power-law distribution, suggesting a non-random distribution pattern. In addition, the small-world coefficient was > 1 for all three networks, indicating the small-world characteristic. The networks for the initial and mid-stages had 49 and 47 nodes, 203 and 230 edges, and average degrees of 8.286 and 9.787, respectively. In contrast, the network for samples from the end stage presented 112 nodes and 1367 edges, with average degree of 24.411. Moreover, the modularity of the networks for the samples from the initial, mid, and end stages were 0.455, 0.249, and 0.259, respectively. These results revealed a more complex co-occurrence pattern in the shrimp gut microbiota at the end of aquaculture activities.
Co-occurrence networks of shrimp gut microbiota at the initial (a), mid (b), and end (c) aquaculture stages. Nodes belonging to different modules labeled in different colors. Differences in robustness (d), vulnerability (e), and negative:positive cohesion (f) among networks of different aquaculture stages, respectively. Different lowercase letters in the same sub-figure represent significant differences among samples from different aquaculture stages (Tukey’s HSD test, p < 0.05)
For network indices, significantly higher robustness (Tukey’s HSD test, p < 0.05) was observed in the network from the mid and end stage samples compared with the initial stage (Fig. 4d). In contrast, vulnerability was obviously lower in the network for samples from the end stage than those from the initial and mid-stages (Fig. 4e). Moreover, there was an increasing trend in cohesion in networks in line with aquaculture progression, with a significant difference (Tukey’s HSD test, p < 0.05) between the initial and end stages (Fig. 4f). Taken together, all the network indices demonstrated increasing stability of the network pattern for shrimp gut microbiota in line with aquaculture progression.
Assembly Mechanism of Shrimp Gut Microbiota
Based on the results of NCM, 28.1%, 42.9%, and 55.4% of the ASVs conformed to the neutral-based theory (Fig. 5). These results indicated that the assembly of shrimp gut microbiota at the beginning and ending of the aquaculture activities were managed by the deterministic and stochastic processes, respectively. The ratio of ASVs revealed that the neutral fraction occupied for most of the richness in all three aquaculture stages (Fig. 5). In contrast, the abundance proportions of the above and below partitions occupied more than the richness proportions, with the above fraction dominant in the shrimp gut microbiota in the mid and end stages (Fig. 5).
Distinct fitting distribution, richness, and abundance of shrimp gut microbiota at the initial (a), mid (b), and end (c) stages by neutral community models. The middle and bottom pie charts represent the percentage of the ASVs belong to different parts based on the number and abundance, respectively
Discussion
Numerous studies have examined the gut microbiota associated with the Pacific white shrimp (Litopenaeus vannamei) due to its extensive aquaculture and high production (Holt et al. 2021). The relationships of shrimp diseases and its gut microbiota are important issues, and the gut microbiota has become as an important driver of successful aquaculture (Sha et al. 2022). Kuruma shrimp is a commercial important species cultured in China, but high mortality and slow growth being the main barriers to the prosperity of the Kuruma shrimp industry (Dong et al. 2014). Limited studies of the gut microbiota of Kuruma shrimp investigated the influences of host immune factors or probiotic supplementation (Duan et al. 2017; Yang et al. 2016). To our knowledge, variations in gut microbiota of Kuruma shrimp throughout the whole aquaculture progression have been investigated for the first time in the present study. We sequenced the gut microbiota of Kuruma shrimp and revealed significant variations in its diversity and composition during the whole aquaculture process from putting the seedlings into the pond to harvest.
A broad spectrum of health conditions has been linked to changes in the gut microbiota of animals. Reduction in the richness and diversity of the gut microbiota has been shown to be responsible for the poor health rank of hosts with weaker resistance to potentially harmful factors (Lawley and Walker 2013). The currently observed increasing trends in the diversity and richness of the shrimp gut microbiota during aquaculture progression indicated that the shrimp in the study ponds were in a healthy growth state. This inference consistent to the weighted increase trend of Kuruma shrimps during the aquaculture progression, which from the 0.1 g weight at the initial stage increased to about 15 g at the middle stage and finally reached to 20 g at the end stage. The gut microbiota can also affect the growth of the host by regulating the proportion of functional bacteria (Magne et al. 2020). The enrichment of Firmicutes in the intestines of animals can induce a more capable assimilation of energy from the diet (Estrada-Velasco et al. 2015). In contrast, Proteobacteria were dramatically increased in the shrimp gut microbiota with retarded growth compared with normal individuals (Xiong et al. 2017a). Enriched Firmicutes and deleted Proteobacteria in the gut microbiota of mid-stage shrimp in this study indicated the potential for a period of fast growth of shrimp during the aquaculture progression. In addition, the RF39 genus was the most dominant bacterial genus in the shrimp gut microbiota and significantly enriched in samples from the mid-stage. Previous phylogenomic analysis revealed that RF39 was received with novel clades in Bacilli and probably with acetate and hydrogen producers in the animal gut environment (Wang et al. 2020). Megamonas, as another bacterial genus that was significantly enriched in shrimp at the mid-stage, has been reported to include major propionate producers in the gut microbiota of chickens (Polansky et al. 2016). Propionate and acetate are important short-chain fatty acids (SCFAs) formed by the gut microbiota by breakdown of complex carbohydrates (Macfarlane and Macfarlane 2003). SCFAs play a crucial role in the maintenance of host health through regulating the homeostasis of gut microbiota and immunity (Tan et al. 2014). The high abundance of SCFA producers in the gut microbiota reflected the favorable health status of the shrimp during the aquaculture progression.
Examining the dynamics of the shrimp gut microbiota during aquaculture progression provided key insights into the roles of aquaculture activities in mediating the stability and complexity of the gut microbial network. The richness, specialists, and co-occurrence network results indicated that the aquaculture activities improved the complexity of the shrimp gut microbiota over time, possibly because shrimp growth acts as a deterministic screening agent for the selection of specific intestinal bacteria (Xiong et al. 2017b). Consequently, less responsive neutral taxa could prevail in the shrimp gut microbiota at the beginning of aquaculture (Burns et al. 2016), resulting in less complex network structures. This theory was agreed with the findings of the NCM in this study, indicating that a small number of non-neutral taxa occupied for most of the gut microbiota. In contrast to the complexity, gut microbiota stability could be more critical for host health and well-being by ensuring that beneficial symbionts and the functions associated with them are maintained in a long term (Lozupone et al. 2012). However, the relationship between the complexity and stability of microbial communities is still hotly debated, with controversial research results (Landi et al. 2018). A highly complex network often means that the gut microbiota is more robust to external perturbations, suggesting high stability (Santolini and Barabási 2018). A similar phenomenon was found in the gut microbiota of healthy pigs in relation to age (Ke et al. 2019), fish gut microbiota after exposure to silver nanoparticles (Chen et al. 2021), and shrimp gut microbiota in line with disease progression (Dai et al. 2020). The current results provided clear proof that network stability increased in line with network complexity in the shrimp gut microbiota during aquaculture progression. Cooperating networks of microbes can be efficient but are often unstable, while the introduction of species to enhance competition can stabilize the cooperating network (Coyte et al. 2015). The richness of the shrimp gut microbiota continued to increase with aquaculture progression. Most of the increased richness was previously shown to come from the living environment of the shrimp (Zhang et al. 2021), with microbes colonizing the shrimp gut leading to increased competition.
Previous research showed that larvae often have a stochastic dominated gut microbiota because of their unripe intestine (Vallès et al. 2012), while deterministic processes governed more in adults, due to stronger selection exerted by the host as their age (Stephens et al. 2016). However, the current NCM results showed that the higher contribution of stochastic processes in terms of determining the shrimp gut microbiota during aquaculture progression compared to deterministic processes. In addition to an unstable state in larvae, the stochastic assembly of gut microbiota could also result in a more balanced state in adults, which may enhance the resistance of the microbiota to external stresses (Zhao et al. 2022). The increased stability of the gut microbiota of shrimp at the end of the aquaculture process could be due to the contribution of stochastic processes to gut microbiota assembly (Chase 2010). Moreover, studies of shrimp diseases indicated more-stochastic assembly of the gut microbiota in diseased shrimp in contrast to healthy individuals (Chen et al. 2017; Dai et al. 2019; Zhu et al. 2016). Notably, the gut microbial network in healthy shrimp has been reported to be better connected and more complex than the networks in diseased counterparts (Sha et al. 2022). The occurrence of shrimp diseases was closely related to the gut microbiota dysbiosis, with less complexity and weak stability (Xiong et al. 2017b), while the current study suggested that healthy shrimp showed a more complex and stable gut microbiota. Therefore, in addition to the roles of stochastic and deterministic processes, more information is needed to determine if the gut microbiota is in balance or dysbiosis in order to evaluate the health status of hosts.
Conclusions
The present study provided the first molecular perspective of the roles of aquaculture progression on the gut microbiota of Kuruma shrimp. The richness and diversity of the shrimp gut microbiota increased significantly in line with aquaculture progression. The gut microbiota composition also varied significantly during the aquaculture process, with enrichment of bacteria involved in energy absorption and SCFA production in the mid-stage. Notably, these results revealed increases in the stability and complexity of the shrimp gut microbiota with aquaculture progression. Although deterministic processes dominated the shrimp gut microbiota at the beginning of aquaculture, stochastic assembly had a greater contribution at the end of the process. These findings contributed valuable insights into our knowledge of the shrimp gut microbiota and provided information to help improve management strategies in relation to shrimp health and productivity in aquaculture.
Data Availability
The data that support the findings of this study will be made available when needed and on request.
References
Bastian M, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. In: International AAAI Conference on Weblogs and Social Media: San Jose, CA, USA
Berg J, Brandt KK, Al-Soud WA, Holm PE, Hansen LH, Sørensen SJ, Nybroe O (2012) Selection for Cu-tolerant bacterial communities with altered composition, but unaltered richness, via long-term Cu exposure. Appl Environ Microbiol 78:7438–7446
Berry D, Widder S (2014) Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front Microbiol 5:219
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Huttley GA, Caporaso JG (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with qiime 2’s q2-feature-classifier plugin. Microbiome 6:90
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
Burns AR, Stephens WZ, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJ (2016) Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J 10:655–664
Chase JM (2010) Stochastic community assembly causes higher biodiversity in more productive environments. Science 328:1388–1391
Chen P, Huang J, Rao L, Zhu W, Yu Y, Xiao F, Chen X, Yu H, Wu Y, Xu K, Zheng X, Hu R, He Z, Yan Q (2021) Resistance and resilience of fish gut microbiota to silver nanoparticles. MSystems. 6(5):e00630–21
Chen WY, Ng TH, Wu JH, Chen JW, Wang HC (2017) Microbiome dynamics in a shrimp grow-out pond with possible outbreak of acute hepatopancreatic necrosis disease. Sci Rep 7:1–12
Coyte KZ, Schluter J, Foster KR (2015) The ecology of the microbiome: networks, competition, and stability. Science 350:663–666
Dai W, Qiu Q, Chen J, Xiong J (2019) Gut eukaryotic disease-discriminatory taxa are indicative of Pacific white shrimp (Litopenaeus vannamei) white feces syndrome. Aquaculture 506:154–160
Dai W, Sheng Z, Chen J, Xiong J (2020) Shrimp disease progression increases the gut bacterial network complexity and abundances of keystone taxa. Aquaculture 517:734802
Dong HB, Su YQ, Mao Y, You XX, Ding SX, Wang J (2014) Dietary supplementation with Bacillus can improve the growth and survival of the Kuruma shrimp Marsupenaeus japonicus in high-temperature environments. Aquacult Int 22:607–617
Duan Y, Zhang Y, Dong H, Wang Y, Zhang J (2017) Effect of the dietary probiotic Clostridium butyricum on growth, intestine antioxidant capacity and resistance to high temperature stress in Kuruma shrimp Marsupenaeus japonicus. J Therm Biol 66:93–100
Estrada-Velasco BI, Cruz M, Garcia-Mena J, Valladares Salgado A, Peralta Romero J, Guna Serrano Mde L, Madrid-Marina V, Orbe Orihuela C, López Islas C, Burguete-García AI (2015) Childhood obesity is associated to the interaction between Firmicutes and high energy food consumption. Nutr Hosp 31:1074–1081
Fan Y, Pedersen O (2021) Gut microbiota in human metabolic health and disease. Nat Rev Microbiol 19:55–71
Gweon HS, Bowes MJ, Moorhouse HL, Oliver AE, Bailey MJ, Acreman MC, Read DS (2021) Contrasting community assembly processes structure lotic bacteria metacommunities along the river continuum. Environ Microbiol 23:484–498
Hirano H, Takemoto K (2019) Difficulty in inferring microbial community structure based on co-occurrence network approaches. BMC Bioinform 20:1–14
Holt CC, Bass D, Stentiford GD, van der Giezen M (2021) Understanding the role of the shrimp gut microbiome in health and disease. J Invertebr Pathol 186:107387
Hou Y, Li B, Xu G, Li D, Zhang C, Jia R, Li Q, Zhu J (2021) Dynamic and assembly of benthic bacterial community in an industrial-scale in-pond raceway recirculating culture system. Front Microbiol 12:4083
Ke S, Fang S, He M, Huang X, Yang H, Yang B, Chen C, Huang L (2019) Age-based dynamic changes of phylogenetic composition and interaction networks of health pig gut microbiome feeding in a uniformed condition. BMC Vet Res 15:1–13
Landi P, Minoarivelo HO, Brännström Å, Hui C, Dieckmann U (2018) Complexity and stability of ecological networks: a review of the theory. Popul Ecol 60:319–345
Lawley TD, Walker AW (2013) Intestinal colonization resistance. Immunology 138:1–11
Lee KH, Guo J, Song Y, Ariff A, O’sullivan M, Hales B, Mullins BJ, Zhang G (2021) Dysfunctional gut microbiome networks in childhood IgE-mediated food allergy. Int J Mol Sci 22:2079
Leeming ER, Johnson AJ, Spector TD, Le Roy CI (2019) Effect of diet on the gut microbiota: rethinking intervention duration. Nutrients 11:2862
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R (2012) Diversity, stability and resilience of the human gut microbiota. Nature 489:220–230
Macfarlane S, Macfarlane GT (2003) Regulation of short-chain fatty acid production. P Nutr Soc 62:67–72
Magne F, Gotteland M, Gauthier L, Zazueta A, Pesoa S, Navarrete P, Balamurugan R (2020) The Firmicutes/Bacteroidetes ratio: a relevant marker of gut dysbiosis in obese patients? Nutrients 12:1474
Moya A, Ferrer M (2016) Functional redundancy-induced stability of gut microbiota subjected to disturbance. Trends Microbiol 24:402–413
Pascoe EL, Hauffe HC, Marchesi JR, Perkins SE (2017) Network analysis of gut microbiota literature: an overview of the research landscape in non-human animal studies. ISME J 11:2644–2651
Polansky O, Sekelova Z, Faldynova M, Sebkova A, Sisak F, Rychlik I (2016) Important metabolic pathways and biological processes expressed by chicken cecal microbiota. Appl Environ Microbiol 82:1569–1576
Rinninella E, Raoul P, Cintoni M, Franceschi F, Miggiano GAD, Gasbarrini A, Mele MC (2019) What is the healthy gut microbiota composition? A changing ecosystem across age, environment, diet, and diseases. Microorganisms 7:14
Santolini M, Barabási AL (2018) Predicting perturbation patterns from the topology of biological networks. Proc Natl Acad Sci U S A 115:E6375–E6383
Sha H, Lu J, Chen J, Xiong J (2022) A meta-analysis study of the robustness and universality of gut microbiota-shrimp diseases relationship. Environ Microbiol 24:3924–3938
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP (2006) Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol 8:732–740
Stephens WZ, Burns AR, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJ (2016) The composition of the zebrafish intestinal microbial community varies across development. ISME J 10:644–654
Tan J, McKenzie C, Potamitis M, Thorburn AN, Mackay CR, Macia L (2014) The role of short-chain fatty acids in health and disease. Adv Immunol 121:91–119
Tasnim N, Abulizi N, Pither J, Hart MM, Gibson DL (2017) Linking the gut microbial ecosystem with the environment: does gut health depend on where we live? Front Microbiol 8:1935
Vanwonterghem I, Jensen PD, Dennis PG, Hugenholtz P, Rabaey K, Tyson GW (2014) Deterministic processes guide long-term synchronised population dynamics in replicate anaerobic digesters. ISME J 8:2015–2028
Vallès Y, Gosalbes MJ, de Vries LE, Abellan JJ, Francino MP (2012) Metagenomics and development of the gut microbiota in infants. Clin Microbiol Infec 18:21–26
Vujkovic-Cvijin I, Sklar J, Jiang L, Natarajan L, Knight R, Belkaid Y (2020) Host variables confound gut microbiota studies of human disease. Nature 587:448–454
Wang Y, Huang JM, Zhou YL, Almeida A, Finn RD, Danchin A, He LS (2020) Phylogenomics of expanding uncultured environmental Tenericutes provides insights into their pathogenicity and evolutionary relationship with Bacilli. BMC Genomics 21:1–12
Xiong J, Dai W, Zhu J, Liu K, Dong C, Qiu Q (2017a) The underlying ecological processes of gut microbiota among cohabitating retarded, overgrown and normal shrimp. Microb Ecol 73:988–999
Xiong J, Zhu J, Dai W, Dong C, Qiu Q, Li C (2017b) Integrating gut microbiota immaturity and disease discriminatory taxa to diagnose the initiation and severity of shrimp disease. Environ Microbiol 19:1490–1501
Yang HT, Yang MC, Sun JJ, Shi XZ, Zhao XF, Wang JX (2016) Dual oxidases participate in the regulation of intestinal microbiotic homeostasis in the Kuruma shrimp Marsupenaeus japonicus. Dev Comp Immunol 59:153–163
Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, Schweer T, Peplies J, Ludwig W, Glöckner FO (2014) The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res 42:D643–D648
Yuan MM, Guo X, Wu L, Zhang YA, Xiao N, Ning D, Shi Z, Zhou X, Wu L, Yang Y, Tiedje JM, Zhou J (2021) Climate warming enhances microbial network complexity and stability. Nat Clim Chang 11:343–348
Zhang X, Li X, Lu J, Qiu Q, Chen J, Xiong J (2021) Quantifying the importance of external and internal sources to the gut microbiota in juvenile and adult shrimp. Aquaculture 531:735910
Zhao Y, Duan C, Zhang X, Chen H, Ren H, Yin Y, Ye L (2018) Insights into the gut microbiota of freshwater shrimp and its associations with the surrounding microbiota and environmental factors. J Microbiol Biotechno 28:946–956
Zhao Z, Jiang J, Zheng J, Pan Y, Dong Y, Chen Z, Gao S, Xiao Y, Jiang P, Wang X, Zhang G, Wang B, Yu D, Fu Z, Guan X, Sun H, Zhou Z (2022) Exploiting the gut microbiota to predict the origins and quality traits of cultured sea cucumbers. Environ Microbiol 24:3882–3897
Zhu J, Dai W, Qiu Q, Dong C, Zhang J, Xiong J (2016) Contrasting ecological processes and functional compositions between intestinal bacterial community in healthy and diseased shrimp. Microb Ecol 72:975–985
Funding
This work was supported by the Foundation for Natural Science of Liaoning (2023-MS-350).
Author information
Authors and Affiliations
Contributions
Saisai Zhang: methodology, formal analysis, and writing original draft; Shuang Liu, formal analysis; Hongwei Li: resources; Hui Li: formal analysis; Jun Luo: methodology; Yinpeng Ding, sources; Tongjun Ren: conceptualization, and writing—review and editing; Wenbo Chen: conceptualization, project administration, and writing—review and editing.
Corresponding author
Ethics declarations
Competing Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Liu, S., Liu, H. et al. Stochastic Assembly Increases the Complexity and Stability of Shrimp Gut Microbiota During Aquaculture Progression. Mar Biotechnol 26, 92–102 (2024). https://doi.org/10.1007/s10126-023-10279-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-023-10279-4