Abstract
The possibility of the brine shrimp Artemia to produce dormant embryo (cysts) in diapause is a key feature in its life history. In the present study, we obtained a proteomic reference map for the diapause embryo of Artemia sinica using two-dimensional gel electrophoresis with a pH range of 4–7 and a molecular weight range of 10–100 kDa. Approximately 233 proteins were detected, and 60 of them were analyzed by capillary liquid chromatography tandem mass spectrometry (LC–MS/MS). Of these, 39 spots representing 33 unique proteins were identified, which are categorized into functional groups, including cell defense, cell structure, metabolism, protein synthesis, proteolysis, and other processes. This reference map will contribute toward understanding the state of the diapause embryo and lay the basis and serve as a useful tool for further profound studies in the proteomics of Artemia at different developmental stages and physiological conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brine shrimp (Artemia) is a worldwide-distributed crustacean, widely used as a main food resource in aquaculture and applied in basic research areas ranging from developmental biology to evolution and ecology (Abatzopoulos 2002). It presents interesting features in development and depending on the environmental conditions; two developmental pathways can be taken: ovoviviparous and oviparous development. In the former case, free-swimming nauplius are released from females and in another style encysted gastrulae (cysts) are produced under unfavorable environmental conditions (Bowen 1962). The cysts enter diapause, a state at which development arrests, metabolic activity reduces greatly, and the resistance to severe physiological stress is high (Drinkwater and Crowe 1987; Liang and MacRae 1999). Upon breakage of diapause and placed in suitable conditions, the embryo is activated and resumes its development releasing a swimming nauplius. This is a special strategy of the brine shrimp to cope with harsh environment and to breed offspring. Therefore, the encysted diapause state of the brine shrimp embryo is a crucial feature and of major importance in Artemia’s life history. The initiation of postdiapause development is accompanied by activation of transcription and protein synthesis but not by DNA synthesis and cell division (Abatzopoulos 2002; Olson and Clegg 1978). During the oviparous development process, large degree of internal differentiation and obvious morphological changes occur in the absence of cell division. These characteristics make the cysts of Artemia an ideal system for assessing the role of protein synthesis in the initial developmental events.
Moreover, the Artemia cysts (in diapause or postdiapause) can withstand great physiological stresses compared with other diapause forms in many eukaryotes. They can survive long-term anoxia (Clegg 1997), temperature extremes (Clegg et al. 2001; Liang and MacRae 1999), repeated hydration and dehydration, desiccation (Clegg 2005), and exposure to pollutants (Browne et al. 1991). Therefore, the encysted diapause embryo of brine shrimp is a good model for studies on the mechanism of stress resistance.
Proteome analysis, a highly efficient technology for the systematic analysis of the proteins expressed by a genome, is becoming increasingly important because proteins are directly related to cellular functions (Pandey and Mann 2000). Proteomics allows us to observe the global expression of proteins in various physiological states and to examine the changes in complex biological processes, helping to unravel possible underlying mechanisms. The establishment of a protein reference map is an essential starting point for all physiological studies that may follow. The reference map is usually established from cells grown under “standard conditions,” a physiological point allowing further study on how proteins respond to different genetic and environmental stimuli (Hecker et al. 2003). Recently, several proteomic studies in marine animals have been published, such as dogfish shark (Lee et al. 2006), mussels (Apraiz et al. 2006), and shrimp (Wang et al. 2007). However, a brine shrimp proteome investigation is virtually absent, and only one report on the microtubule proteome of Artemia (O’Connell et al. 2006) has been published.
We report the construction of a reproducible and well-resolved 2-D electrophoresis (2-DE) protein reference map of Artemia sinica cyst and the protein identifications by capillary liquid chromatography tandem mass spectrometry (LC–MS/MS). This map presents the global intracellular protein expression of the diapause embryos of Artemia sinica and will serve as a useful tool to study many important physiological processes, such as dormancy breaking, hatching, and stress resistance. This map could become a platform for further studies on changes in protein expression during development. To our knowledge, this is the first proteome for the dormant embryo with diapause state.
Materials and Methods
Experimental Animals
Cysts of the brine shrimp Artemia sinica were harvested from Yuncheng Salt Lake, Shanxi Province, China in 1998 and stored at 4°C in dark until use. Some of the cysts were incubated in saturated sodium chloride solution for 1 month at 4°C. This treatment causes cysts dehydration, known to terminate diapause (Lavens and Sorgeloos 1987). The hatching of dehydrated cysts and nontreated cysts was performed according to Lavens and Sorgeloos (1996). Briefly, 1.6-g cysts were incubated in 800 ml 33 g/l seawater under continuous illumination (2000 lux) at 28°C in a cilindroconical tube, with medium aeration from bottom as to keep all cysts in suspension. After 48 h, hatching percentage was assessed by counting the larvae individuals, which were transferred to a solid watch glass and observed under a dissection microscope. The number of hatched nauplii was compared with the total number of cysts incubated.
Protein Sample Preparation
Total proteins were extracted from the nontreated cysts with TCA/acetone. Briefly, 8-g cysts were rinsed in 40 mM Tris on ice using a glass homogenizer and then centrifuged at 10,000 g for 30 min at 4°C. The supernatant was resuspended in five volumes of extracting solution [acetone, 10% trichloroacetic acid (TCA), 0.07% dithiothreitol (DTT)] (pre-cooled at −20°C, to prevent protein degradation) immediately and incubated at −20°C overnight. The sample was then centrifuged for 30 min at 10,000 g at 4°C. The result precipitate was then washed twice with pre-cooled (−20°C) acetone and 0.07% DTT to remove the TCA and centrifuged at 10,000 g for 30 min at 4°C. The precipitate was dried overnight at room temperature and then resuspended in rehydration solution (urea 7 M, thiourea 2 M, CHAPS 4% (w/v), DTT 65 mM, biolyte 0.2%, and bromophenol blue 0.001%). Before loading, the sample was centrifuged at 12,000 g for 15 min and the protein concentration in the supernatant was determined by Bradford’s method (1976).
Two-dimensional Gel Electrophoresis
The Immobiline™ DryStrip Gels (IPG) strips (18-cm, linear pH 4–7, Amersham Biosciences) were loaded with 2 mg total protein and were rehydrated for 12 h at 50 V at room temperature. IEF was performed using the PROTEANs IEF Cell (BioRad) at 250 V for 1 h, 500 V for 1 h, 1000 V for 1 h, 8000 V for 5 h, and 8000 V for 60000 Vhour. Afterwards, the focused IPG strips were equilibrated in a buffer comprised of 6 M urea (w/v), 2% SDS (w/v), 0.05 M Tris (pH 8.8), 20% glycerol (v/v), and 2% DTT (w/v) for 15 min at room temperature with gentle shaking, and a second equilibration of 15 min was conducted in the same buffer with 2.5% iodoacetamide (w/v) instead of 2% DTT. The second-dimension SDS-PAGE was performed with a 5% stacking gel and 12% running gel and was carried out at 14°C on a BioRad Protean® II xi Cell System (BioRad, USA) in Tris/Glycine/SDS buffer and the electrophoresis program was 50 V for 10 min, 150 V for stacking gel, and 300 V for running gel. Triplicates were prepared for each sample to confirm overall reproducibility of the protein spots.
Protein Visualization and Image Analysis
Protein spots were visualized by staining with 0.15% Coomassie brilliant blue R-250 in 40% ethanol, 10% acetic acid for 3 h, and destained in 30% ethanol, 10% acetic acid. Gels were scanned with a Gel Doc. Imaging system (BioRad, USA) and analyzed with Imagemaster™ 2-D Software (Amersham Biosciences).
Tryptic in Gel Digestion
The protein spots were excised from replicate 2-DE gels and digested in gel with a mass spectrometry grade trypsin (Promega, Madison, WI), according to Shevchenko’s method (Shevchenko et al. 1996) with minor modifications. Briefly, the protein spots were excised and sliced to small pieces and destained with 100% acetonitrile (ACN) in equal volume of 0.1 M ammonium bicarbonate (NH4HCO3) twice. Then, the gel particles were dehydrated in 100% ACN and dried in a vacuum centrifuge. Twenty microliters of trypsin digestion solution (containing 20 μg/ml trypsin in 50 mM NH4HCO3/10% ACN) was added to pre-incubate the gel at 4°C for 45 min. Then, 20 μl 25 mM NH4HCO3/10% ACN was added to cover the pieces and the digestion was performed at 37°C for 16 h. After centrifugation at 10,000 g for 20 min, the supernatant from each sample was transferred to a new microcentrifuge tube. Then, 50 μl of 50% acetonitrile/5% formic acid was added to tubes to extract the peptides. This supernatant was combined with the previous supernatant and dried in a vacuum centrifuge. The resultant supernatant containing tryptic peptides were subjected to mass spectrometry analysis for protein identification.
Protein Identification
The mass spectrometer used in this work was an ion-trap mass spectrometer model LCQ DECA XPplus MS (ThermoFinnigan, San Jose, CA). In each experiment, a 20-μl enzyme digested sample was injected. Spectra were collected in the positive ion mode. The liquid chromatography separation was performed at a flow-rate of 120 ml per min on a BioBasic-18 column, 150 × 0.18 mm, particle Sz (μ)5 (Thermohypersil-Keystone, No. 72105-100265). The gradient was developed according to 2% A and 98% B for 15 min, 65% A and 35% B for 45 min, 95% A and 5% B for 10 min, and finally 2% A and 98% B for 15 min, where A is acetonitrile and B is water with 0.1% formic acid. The total acquiring time was 90 min. Protein identification was performed using the TurboSEQUEST algorithm in the BioWorks 3.1 software package by searching the database downloaded from the NCBI (http://www.ncbi.nl.nih.gov). Peptides identified were filtered according to their charge state, cross-correlation score (Xcorr, >1.9 for n+1, >2.5 for n+2 and >3.75 for n+3), and Delta Correlation value (Delta Cn >0.1).
Results and Discussions
The Hatching Test
The diapause state of cysts of Artemia sinica could be terminated effectively by dehydration, whereas other factors, such as cold temperature, light, and H2O2 have little effect on this species (Huang et al. 2002). The average hatchability of the nontreated cysts was 12.2 ± 2%, and a significant different result (92.6 ± 1.1% hatch) was observed in the cysts after dehydration treatment, indicating that the embryoic diapause state of the cysts was terminated by the dehydration treatment and the nontreated cysts used in this study were in the diapasue state.
2-D PAGE and Analysis of Protein Expression Profiles
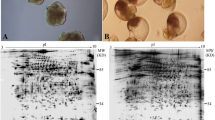
The high resolution and highly reproducible 2-D pattern of the brine shrimp cyst proteome is shown in Fig. 1, which could be used as a reference map. This map is reproduced from replicate gels of three independent biological experiments and comparison of these gels demonstrated that most abundant and visible spots were present on all the gels. Approximately 233 protein spots were detected in the reference gel by the computer 2-D analysis software Imagemaster™.
2-D electrophoretic reference map of proteins from encysted embryos of Artemia sinica. 2-mg proteins were applied in pH 4–7 IPG strip (18-cm) for the first-dimensional electrophoresis and subsequently separated by 12% SDS-PAGE gel. The gel was stained with Coomassie brilliant blue R-250. Spot numbers correspond to those denoted in Table 1
Protein Identification
A total of 60 protein spots with relatively high abundance were excised, 39 of which were identified corresponding to 33 different proteins. These proteins are numbered in the 2-DE reference map shown in Fig. 1. The protein information of the characterized proteins are listed in Table 1 with their function category, protein name, NCBI accession number, number of matched peptides, function description, MW and pI.
Classification of Identified Proteins
The identified proteins were categorized based on their function according to the annotations in the Swiss-Prot database at the ExPASy Molecular Biology Server (http://www.expasy.org). The Swiss-Prot identifiers, employed for linkage to Gene Ontology (GO) Consortium, and providing annotation of each protein with the biological processes, cellular components, molecular functions, and structure, allow us to allocate selected proteins into biologically relevant groups (Tyan et al. 2006). The function description of the identified proteins was presented based on their annotations in the GO database (Table 1). A total of nine functional groups were represented in our result (Fig. 2). Forty percent of the identified protein spots are catalytic enzymes involved in metabolic processes. Approximately 16% of the proteins are associated with protein metabolism, including protein biosynthesis, proteolysis, protein folding, protein binding, and protein phosphatation. Other classes of the identified protein spots include cell defense (10%), cell structure (12%), nucleic acid metabolism (2%), cell adhesion (5%), and cell cycle (2%). In addition, some identified proteins are annotated as hypothetical proteins (5%).
Functional classification of proteins identified in encysted embryos of Artemia. sinica. A total of 39 spots representing 33 different proteins were categorized into 9 groups (Table 1). Assignments were made on the basis of information provided on the Swiss-Prot database at the ExPASy Molecular Biology Server
The diapause cysts (encysted gastrulae) of the brine shrimp are remarkably resistant to physiological stresses. In our result, several spots were identified as cell defense proteins. The most abundant protein spot (spot 1) identified was small heat shock/alpha-crystallin protein precursor, which functions as a molecular chaperone and protects the proteins of encysted embryo of Artemia from stress-induced denaturation (Liang and MacRae 1999; MacRae 2003; Willsie and Clegg 2001). Moreover, it plays a key role during encystment, diapause, and quiescence in developing Artemia and acts on microtubules through p26-dependent disruption of tubulin assembly, which inhibits mitosis and cell division and causes the embryos to stall at gastrulation (Day et al. 2003). Based on our 2-DE result, three protein spots identified as p26 were detected. Compared with the full-sized p26 isoform (spot 1), spots 2 and 3, respectively, have a considerable lower molecular weight and a lower pI value. Such an isoform diversity of p26 in the encysted embryos of Artemia may arise by posttranslational protein modification (Qiu et al. 2004), including ubiquitylation, phosphorylation, glycosylation, acetylation, fragmentation, and so on. The difference in the molecular weight and pI value of p26 isoforms may indicate that they locate in different cells and perform different physiological roles, which need to be answered by more profound research. Heat shock protein 70, an important protein in cytoprotection also was identified (spot 4). It seems to act generally as molecular chaperones for protein folding (Gething and Sambrook 1992) and is involved in repairing protein damage that occurs as a consequence of various stresses. The existence and high abundance of these identified cell defense proteins can partly explain the strong stress resistance of the embryo of Artemia.
The cell structure proteins are the basis of the organization of eukaryotic cells from yeast to human. In our study, several protein spots identified as the cytoskeletal proteins appeared as abundant proteins in the 2-DE gels, including actin and tubulin. Among them, two spots (5 and 6) correspond to actin. Actin is a highly conserved protein in eukaryotic cells, playing important roles in a range of cellular functions including muscle contraction, cell motility, cytoskeletal structure, cell division, intracellular transport, and cell differentiation (Herman 1993). Tublin is a key component of the cytoskeletal microtubules (Howard and Hyman 2003). In accordance with previous study (Langdon et al. 1991), the gastrulation embryo of brine shrimp contains both alpha-tublin and beta-tubulin as shown by Coomassie Blue staining of 2-D gels. The isotubulin composition and the quantity of tubulin do not change during pre-emergence development of Artemia embryos (MacRae and Ludueña 1984) and a mechanism of tubulin synthesis controlling in Artemia may be the limitation of the binding of its mRNA to ribosomes (Langdon et al. 1991). Three identified proteins spots correspond to tublin: alpha-2–tublin (spot 7), beta–tublin (spot 8), and tublin (spot 9). These cytoskeleton proteins constitute complex and highly dynamic network of protein filaments controls a multitude of processes, including cell shape, division, polarity, movement, and intracellular transport (Carballido-Lopez and Errington 2003).
The largest portion of the identified proteins is related to carbohydrate and energy metabolism, representing 36% of the protein identified, including triosephosphate isomerase (spot 15), aldehyde reductase (spot 16), 2-phospho-D-glycerate hydrolase (spots 17 and 18), putative fructose 1,6-bisphosphate aldolase (spot 19), putative chlorohydrolase (spot 20), predicted esterase of the alpha-beta hydrolase superfamily (spot 21) and ENSANGP00000011006 (spot 22), arginine kinase (spots 26 and 27), and ATPase beta-unit (spot 28). This result indicates that the metabolic proteins account for a respectively large proportion of the proteins in the quiescent cyst of Artemia. The metabolic rate depression is an important survival strategy for many animal species and a common element of diapause (Storey and Storey 2004) and the metabolism is severely suppressed in diapause embryos of brine shrimp. However, the cells of the diapause embryo must be able to reverse all the metabolic events quickly to resume its development under favorable conditions (Browne et al. 1991). Our results indicate that the cyst may apply the strategy of storing some of the metabolic proteins to ensure the rapid resumption of its development. ATP has been characterized as a critical component in the early development of Artemia because many important metabolic processes are ATP requiring pathways (Clegg 1964; Conte et al. 1977). Several spots identified were tightly associated with ATP generation and consumption. Arginine kinase (AK) is a type of phosphagen kinase that is the key enzyme for energy metabolism in invertebrates. It functions in the maintenance of ATP level by catalyzing the reversible transfer of a phosphoryl group from a guanidino phosphagen to adenosine diphosphate (ADP), generating a molecule of adenosine triphosphate (ATP) (Ellington 1989). AK is usually present in tissues with high-energy demand or cells experiencing short bursts of energy demand (Dumas 1993; Kotlyar et al. 2000). The ATPsynthase (or F0–F1 complex) regenerates ATP from ADP and Pi in energy-transducing membranes (Boyer 1997). It consists of two parts: a hydrophobic membrane-bound portion called CF0, and a soluble portion that protrudes into the stroma called CF1. CF1 consists of five different subunits: alpha, beta, gamma, delta, and epsilon units. The mRNA of the ATPase subunits of brine shrimp is differentially expressed (de Chaffoy de Courcelles and Kondo 1980): the beta subunit changed slightly but other subunits may rise between the period of early rupture of the shell and complete hatching of the cyst. Only the beta subunit of the ATPase complex was present on our 2D-PAGE gel, indicating the differential expression of the ATPase subunits in the protein level.
Approximately 15% of the proteins are involved in protein metabolism, including 1) protein folding: heat shock protein 70 (spot 4); 2) protein phosphatation: hypothetical protein C17H12.5 (spot 11); 3) protein biosynthesis: glutamyl-tRNA synthetase (spot 30) and elongation factor-2 (spot 31); 4) proteolysis: cathepsin L precursor (spot 32) and cathepsin L-like protease precursor (spot 33), and 5) protein binding: glycine-rich protein (spot 34). Cathepsin L (CL) is a ubiquitous cysteine protease in eukaryotes and essential for development in several organisms, including Xenopus laevis (Miyata and Kubo 1997), Caenorhabditis elegans (Britton and Murray 2002), and Artemia franciscana (Warner et al. 1995). It has been reported that a number of developmental events are dependent on its cysteine protease activity, such as transcription regulation (Hu and Leung 2004), gastrulation (Warner et al. 1995), molting and eggshell remodeling (Guiliano et al. 2004), and yolk metabolism (Fagotto 1990). In embryos of the brine shrimp, the major cysteine protease is a heterodimer composed of a cathepsin L-like polypeptide of 28.5 kDa and a 31.5 kDa polypeptide called the cathepsin L-associated protein or CLAP (Warner et al. 2004). The amount of CL and CLAP remains equivalent level during the embryo development process (Liu and Warner 2006). In our result, besides the protein (spot 32) identified as the cathepsin L precursor, a cathepsin L-like protease precursor (spot 33) also was identified, both of which belong to the CL family. The pI of these two proteins shows remarkable differences, which may associate with the two CLAP isoforms discussed below. The CLAP was shown to be a cell adhesion protein, containing two domains with high similarity to domains in fasciclin I and other cell adhesion proteins. It stabilizes CL at various pH and temperatures (Warner et al. 2004) and plays an important role in targeting and expression regulation of CL during early development of Artemia (Liu and Warner 2006). Herein two protein spots (38 and 39) were identified as CLAP. The molecular weight of spot 38 is close to the calculated molecular mass of 32.3 kDa and the pI of it is slightly lower than that of 8.0 obtained using EXPASY (http://www.expasy.org/). Spot 39 is observed as an isoform with apparently lower molecular weight and pI value. This suggests that variant splicing of the mRNA or further posttranslational modifications of CL may occur. Interestingly, one isoform of CLAP (spot 39) has a similar pI with cathepsin L precursor (spot 32), whereas the pI of another isoform of CLAP (spot 38) shows similarity with cathepsin L-like protease precursor (spot 33). The function and significance of the isoform diversity of CL is worthy of further investigation to clarify whether they are located in a different cell compartment or function at different development stages.
We also identified a protein similar to feminization 1 homolog b (spot 29), which has a DNA-dependent transcription factor activity; Anaphase-promoting complex subunit 7 (spot 35), a component of cell cycle-regulated ubiquitin ligase (anaphase promoting complex/cyclosome) that controls progression through mitosis and the G1 phase of cell cycle. Several hypothetical proteins were identified, the function of which remain unclear.
References
Abatzopoulos TJ (2002) Artemia: basic and applied biology. Kluwer Academic Publishers, The Netherlands
Apraiz I, Mi J, Cristobal S (2006) Identification of proteomic signatures of exposure to marine pollutants in mussels (Mytilus edulis). Mol Cell Proteomics 5:1274–1285
Bowen ST (1962) The genetics of Artemia salina. I. The reproductive cycle. Biol Bull 122:25–32
Boyer PD (1997) The ATP synthase: a splendid molecular machine. Annu Rev Biochem 66:717–749
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Britton C, Murray L (2002) A cathepsin L protease essential for Caenorhabditis elegans embryogenesis is functionally conserved in parasitic nematodes. Mol Biochem Parasit 122:21–33
Browne RA, Trotman CNA, Sorgeloos P (1991) Artemia biology. CRC Press, Boca Raton
Carballido-Lopez R, Errington J (2003) A dynamic bacterial cytoskeleton. Trends Cell Biol 13:577–583
Clegg JS (1964) The control of emergence and metabolism by external osmotic pressure and the role of free glycerol in developing cysts of Artemia salina. J Exp Biol 41:879–892
Clegg JS (1997) Embryos of Artemia franciscana survive four years of continuous anoxia: The case for complete metabolic rate depression. J Exp Biol 200:467–475
Clegg JS (2005) Desiccation tolerance in encysted embryos of the animal extremophile, Artemia. Integ Comp Biol 45:715–724
Clegg JS, Van Hoa N, Sorgeloos P (2001) Thermal tolerance and heat shock proteins in encysted embryos of Artemia from widely different thermal habitats. Hydrobiologia 466:221–229
Conte FP, Droukas PC, Ewing RD (1977) Development of sodium regulation and De Novo synthesis of NA + K-activated ATPase in larval brine shrimp, Artemia salina. J Exp Zool 202:339–362
Day RM, Gupta JS, MacRae TH (2003) A small heat shock/alpha-crystallin protein from encysted Artemia embryos suppresses tubulin denaturation. Cell Stress Chaperon 8:183–193
de Chaffoy de Courcelles D, Kondo M (1980) Lipovitellin from the crustacean, Artemia salina. Biochemical analysis of lipovitellin complex from the yolk granules. J Biol Chem 255:6727–6733
Drinkwater LE, Crowe JH (1987) Regulation of embryonic diapause in Arternia: environmental and physiological signals. J Exp Zool 241:297–307
Dumas C (1993) Cloning and sequence analysis of the gene for arginine kinase of lobster muscle. J Biol Chem 268:21599–21605
Ellington WR (1989) Phosphocreatine represents a thermodynamic and functional improvement over other muscle phosphagens. J Exp Biol 143:177–194
Fagotto F (1990) Yolk degradation in tick eggs: II. Evidence that cathepsin L-like proteinase is stored as a latent, acid-activable proenzyme. Arch Insect Biochem Physiol 14:237–252
Gething MJ, Sambrook J (1992) Protein folding in the cell. Nature 355:33–45
Guiliano DB, Hong XQ, McKerrow JH, Blaxter ML, Oksov Y, Liu J, Ghedin E, Lustigman S (2004) A gene family of cathepsin L-like proteases of filarial nematodes are associated with larval molting and cuticle and eggshell remodeling. Mol Biochem Parasit 136:227–242
Hecker M, Engelmann S, Cordwell SJ (2003) Proteomics of Staphylococcus aureus—current state and future challenges. J Chromatogr B 787:179–195
Herman IM (1993) Actin isoforms. Curr Opin Cell Biol 5:48–55
Howard J, Hyman AA (2003) Dynamics and mechanics of the microtubule plus end. Nature 422:753–758
Hu KJ, Leung PC (2004) Shrimp cathepsin L encoded by an intronless gene has predominant expression in hepatopancreas, and occurs in the nucleus of oocyte. Comp Biochem Phys B-Biochem Mol Biol 138:445
Huang C, Liu G, Zeng L, Wu M (2002) Influence of inducing conditions and specific diapause deactivation methods on hatchability of two species. Artemia cysts produced in lab. Fisheries Sci 21:1–4
Kotlyar S, Weihrauch D, Paulsen RS, Towle DW (2000) Expression of arginine kinase enzymatic activity and mRNA in gills of the euryhaline crabs Carcinus maenas and Callinectes sapidus. J Exp Biol 203:2395–2404
Langdon CM, Rafiee P, Macrae TH (1991) Synthesis of tubulin during early postgastrula development of Artemia - isotubulin generation and translational regulation. Dev Biol 148:138–146
Lavens P, Sorgeloos P (1987) The cryptobiotic state of Artemia cysts, its diapause deactivation and hatching: a review. In: Sorgeloos P et al (eds) Artemia research and its applications: 3. Ecology, culturing, use in aquaculture. In the Second International Symposium on the brine shrimp Artemia, pp 27–63
Lavens P, Sorgeloos P (1996) Manual on the production and use of live food for aquaculture, vol 361. FAO Fisheries Technical Paper, Rome
Lee J, Valkova N, White MP, Kultz D (2006) Proteomic identification of processes and pathways characteristic of osmoregulatory tissues in spiny dogfish shark (Squalus acanthias). Comp Biochem Phys D-Genom Proteom 1:328–343
Liang P, MacRae TH (1999) The synthesis of a small heat shock/[alpha]-crystallin protein in Artemia and its relationship to stress tolerance during development. Dev Biol 207:445–456
Liu LQ, Warner AH (2006) Further characterization of the cathepsin L-associated protein and its gene in two species of the brine shrimp, Artemia. Comp Biochem Phys A-Mol Integ Phys 145:458–467
MacRae TH (2003) Molecular chaperones, stress resistance and development in Artemia franciscana. Semin Cell Dev Biol 14:251–258
MacRae TH, Ludueña RF (1984) Developmental and comparative aspects of brine shrimp tubulin. Biochem J 219:137–148
Miyata S, Kubo T (1997) Inhibition of gastrulation in Xenopus embryos by an antibody against a cathepsin L-like protease. Dev Growth Differ 39:111–115
O’Connell PA, Pinto DM, Chisholm KA, MacRae TH (2006) Characterization of the microtubule proteome during post-diapause development of Artemia franciscana. Biochim Biophys Acta (BBA) Proteins Proteom 1764:920–928
Olson CS, Clegg JS (1978) Cell division during the development of Artemia salina. Dev Genes Evol 184:1–13
Pandey A, Mann M (2000) Proteomics to study genes and genomes. Nature 405:837–846
Qiu ZJ, Viner RI, MacRae TH, Willsie JK, Clegg JS (2004) A small heat shock protein from Artemia franciscana is phosphorylated at serine 50. Biochim Biophys Acta-Proteins Proteom 1700:75–83
Shevchenko A, Jensen, ON, Podtelejnikov AV, Sagliocco F, Wilm M, Vorm O, Mortensen P, Shevchenko A, Boucherie H, Mann M (1996) Linking genome and proteome by mass spectrometry: large-scale identification of yeast proteins from two dimensional gels. Biochemistry 93:14440–14445
Storey KB, Storey JM (2004) Metabolic rate depression in animals: transcriptional and translational controls. Biol Rev 79:207–233
Tyan Y-C, Guo H-R, Liu C-Y, Liao P-C (2006) Proteomic profiling of human urinary proteome using nano-high performance liquid chromatography/electrospray ionization tandem mass spectrometry. Anal Chim Acta 579:158–176
Wang H-C, Wang H-C, Leu J-H, Kou G-H, Wang AHJ, Lo C-F (2007) Protein expression profiling of the shrimp cellular response to white spot syndrome virus infection. Dev Comp Immunol 31:672–686
Warner AH, Pert MJ, Osahan JK, Zielinski BS (1995) Potential role in development of the major cysteine protease in larvae of the brine shrimp Artemia franciscana. Cell Tissue Res 282:21–31
Warner AH, Pullumbi E, Amons R, Liu LQ (2004) Characterization of a cathepsin L-associated protein in Artemia and its relationship to the FAS-I family of cell adhesion proteins. Eur J Biochem 271:4014–4025
Willsie JK, Clegg JS (2001) Nuclear p26, a small heat shock/alpha-crystallin protein, and its relationship to stress resistance in Artemia franciscana embryos. J Exp Biol 204:2339–2350
Acknowledgments
This research was supported by the program of National Natural Science Foundation of China (20060109Z4016) and Natural Science Foundation of Shandong Province for the excellent young researcher (2006BSA02004).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, Q., Wu, C., Dong, B. et al. The Encysted Dormant Embryo Proteome of Artemia sinica . Mar Biotechnol 10, 438–446 (2008). https://doi.org/10.1007/s10126-007-9079-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-007-9079-0