Abstract
A long time has passed since regularly interspaced DNA repeats were discovered in prokaryotes. Today, those enigmatic repetitive elements termed clustered regularly interspaced short palindromic repeats (CRISPR) are acknowledged as an emblematic part of multicomponent CRISPR-Cas (CRISPR associated) systems. These systems are involved in a variety of roles in bacteria and archaea, notably, that of conferring protection against transmissible genetic elements through an adaptive immune-like response. This review summarises the present knowledge on the diversity, molecular mechanisms and biology of CRISPR-Cas. We pay special attention to the most recent findings related to the determinants and consequences of CRISPR-Cas activity. Research on the basic features of these systems illustrates how instrumental the study of prokaryotes is for understanding biology in general, ultimately providing valuable tools for diverse fields and fuelling research beyond the mainstream.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of an RNA-based interference-like mechanism in prokaryotes (Mojica et al. 2005; Makarova et al. 2006), analogous to the adaptive immune system that operates in vertebrates, represented an unanticipated breakthrough in microbiology and immunology. Barrangou and collaborators (Barrangou et al. 2007) validated, in the lactic acid bacterium Streptococcus thermophilus, previous proposals relating clustered regularly interspaced short palindromic repeats (CRISPR) and Cas (CRISPR associated) proteins (Mojica et al. 2000; Jansen et al. 2002) to defence against invasive genetic elements (Mojica et al. 2005; Pourcel et al. 2005). These initial hypotheses were based on the analysis of CRISPR arrays in many Yersinia pestis genomes (Pourcel et al. 2005) and representative strains of the main taxonomic groups of archaea and bacteria (Mojica et al. 2005). The comparison of the CRISPR regions with sequences available in nucleotide databases revealed that repeat intervening spacers matched other sequences (later termed protospacers; Deveau et al. 2008) in mobile genetic elements related to the spacer-carrier strain. Moreover, the presence of a given spacer seemed to be incompatible with the co-occurrence in the cell of perfectly matching protospacers, suggesting the existence in prokaryotes of a CRISPR-based adaptable mechanism of protection (Mojica et al. 2005). Consequently, it was proposed that the immune memory relied on the integration of invading nucleic acids within the CRISPR loci. Indeed, we know now that sequences of foreign origin, either from RNA (Silas et al. 2016) or DNA (Barrangou et al. 2007), can be incorporated into CRISPR arrays during the infection process, resulting in new spacers framed by repeat units. Spacer acquisition is the first step of the CRISPR-Cas mechanism, named ‘adaptation’ or ‘immunisation’. The second stage is referred to as ‘expression’ or ‘CRISPR RNA (crRNA) biogenesis’ and the last one as ‘interference’. During the expression stage, crRNAs are produced after processing the primary transcript (precursor CRISPR RNA or pre-crRNA) generated from the CRISPR locus (Mojica et al. 1993; Brouns et al. 2008). Like the eukaryotic interference RNA (RNAi) system, CRISPR-Cas utilises small guide RNA (crRNA) molecules to recognise complementary sequences (Brouns et al. 2008). However, in addition to RNA (Abudayyeh et al. 2016), CRISPR-Cas binds and cleaves target DNA sequences (Marraffini and Sontheimer 2008; Garneau et al. 2010) during the interference stage.
Apart from the crRNA guides and the Cas proteins that participate in all stages of the CRISPR-Cas mechanism, other main components are needed for CRISPR-based immunity: the leader sequence, the protospacer adjacent motif (PAM) and, in some CRISPR-Cas variants, the trans-activating CRISPR RNA (tracrRNA) or the short-complementarity untranslated RNAs (scoutRNAs). The leader is a sequence conserved at one end of CRISPR arrays that co-evolves with repeat sequences (Bult et al. 1996). The main promoter of the CRISPR locus (Brouns et al. 2008; Pul et al. 2010; Pougach et al. 2010) and motifs related to recognition of the spacer integration site (Rollie et al. 2015; Wei et al. 2015; Nuñez et al. 2016; Yoganand et al. 2017; McGinn and Marraffini 2019) are in the leader. PAMs are short sequences (typically 2 to 5 nt) at the end of the protospacers (Bolotin et al. 2005) of many CRISPR-Cas systems (Mojica et al. 2009; Shah et al. 2013). PAMs are necessary for the efficient recognition of protospacers by Cas proteins during the adaptation and interference stages (Gleditzsch et al. 2019). The requirement for PAMs prevents self-targeting of the CRISPR array as alternative sequences are present in the corresponding location next to the spacers (Weissman et al. 2020). Both tracrRNA and scoutRNA are small RNAs encoded in some CRISPR-Cas types, which partially hybridise with the repeat in CRISPR RNAs, participating in crRNA maturation and target interference (Deltcheva et al. 2011; Jinek et al. 2012; Harrington et al. 2020).
Along with the spacer-matching sequences found in transmissible genetic elements, protospacers were initially located in non-mobile chromosomal regions, suggesting that CRISPR might be playing in-house roles (Mojica et al. 2005). Indeed, as is the case of the immune system in mammals (Sattler 2017), non-canonical functions have been proven for the prokaryotic adaptive system since the initial demonstration of its protective action (Wimmer and Beisel 2020).
The biochemical characterisation of a few CRISPR-Cas systems in the late 2000s and early 2010s enabled easily programmable DNA targeting (Gasiunas et al. 2012; Jinek et al. 2012), providing tools for genome editing, notably those based on Cas9 (Cong et al. 2013; Mali et al. 2013), and for the regulation of gene expression (Bikard et al. 2013). They also allowed the implementation of sequence-specific anti-microbials (Bikard et al. 2014). More recently, mainly thanks to the discovery of novel CRISPR-Cas systems, the spectrum of CRISPR-based devices and applications extended to RNA targeting, molecular diagnostics, epigenetic modification or guided transposition, among others (East-Seletsky et al. 2016; Mojica and Montoliu 2016; Gootenberg et al., 2017; Chavez and Qi 2019; Liu et al. 2020a; Sun et al. 2020b).
Reviews covering CRISPR-Cas have been numerous over the last years in an attempt to capture the growing diversity of CRISPR/Cas configurations and the newly discovered functions and mechanistic peculiarities. At present, CRISPR is a very productive and fast-moving field of research whose updates are followed closely by the large CRISPR community devoted to understanding its biology and, beyond basic researchers, by those interested in applications of CRISPR-based technology. In this manuscript, we summarise the fundamentals of native CRISPR-Cas systems and further elaborate on lesser-known biological aspects, such as the complexities of their regulation and the diverse non-canonical functions they play.
CRISPR-Cas diversity and classification
Although initial identification of CRISPR-Cas components pointed at a limited diversity of these systems and a general mechanism of action (Mojica et al. 2000; Jansen et al. 2002; Haft et al. 2005; Makarova et al. 2006; Barrangou et al. 2007), in-depth analyses of the increasing genomic and metagenomic data have demonstrated a staggering variety in CRISPR-Cas systems (Dwarakanath et al. 2015; Al-Shayeb et al. 2020; Pinilla-Redondo et al. 2020). Early classification schemes, relying mainly on just comparing a subset of cas genes (notably cas1), were gaining in complexity (Haft et al. 2005; Makarova et al. 2006). Cataloguing efforts of CRISPR-Cas elements focused on establishing robust criteria to reflect the phylogeny of the different systems. Thus, Makarova and co-workers recommended in 2011 a polythetic classification strongly supported by evolutionary relationships between CRISPR-Cas components (Makarova et al. 2011). Later, a critical layer was added to the categorisation efforts after defining the functional modular organisation of the CRISPR-cas locus (Makarova et al. 2015). According to the activities carried out by the Cas proteins, four modules of CRISPR-associated genes (sometimes with shared members) were differentiated: (i) adaptation, (ii) expression, (iii) interference or effector and (iv) signal transduction or ancillary (a combination of different accessory genes with unknown or tentatively assigned functions) modules. Subsequently, the comparison among the effector modules became the main classification principle considering Cas sequence similarity, cas locus architecture, the phylogeny of conserved Cas proteins, characteristics of other CRISPR associated elements and, ultimately, available experimental data. This complexity of criteria has resulted in a dynamic classification that must be regularly amended as new data are provided (Lange et al. 2013; Burstein et al. 2017; Harrington et al. 2017; Shmakov et al. 2017a; Yan et al. 2018, 2019; Makarova et al. 2018; Pausch et al. 2020).
The current classification of the CRISPR-Cas systems (Pinilla-Redondo et al. 2020; Makarova et al. 2020; Pausch et al. 2020) comprises two classes (class 1 and class 2), six types (marked with Roman numbers, from Type I to Type VI) and over 30 subtypes (denoted by letters: I-A to I-F, IV-A to IV-E, III-A to III-F, II-A to II-C, V-A to V-K and VI-A to VI-D; provisionally classified systems are labelled with U), some also including multiple recognised variants (indicated with Arabic numerals). Figure 1 shows the typical components of the classified CRISPR-Cas systems.
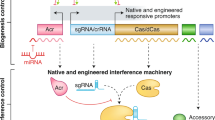
Components of CRISPR-Cas systems. The presence of CRISPR array (CRISPR), trans-activating crRNA (tracrRNA), and genes encoding either core Cas proteins (Core Cas) or accessory proteins/domains (Accessory) involved in ancillary functions (yellow) are shown for class 1 (a) or class 2 (b) CRISPR-Cas subtypes and variants (Sub/Var). Core proteins are coloured based on their role in adaptation (burgundy), crRNA biogenesis (red), target binding (blue) and target cleavage (orange), according to Makarova et al. (2020), Pausch et al. (2020) and Pinilla-Redondo et al. (2020). Constituents that are not invariably present are represented with fainter colours. Genes encoding proteins that contribute multiple functions are depicted with colour schemes consistent with the colour code assigned to each activity. TPR, tetratricopeptide repeat; CHAT, protease domain of the caspase family; RT, reverse transcriptase domain; TnsA-D and TniQ, transposition-related proteins; WYL, protein with the WYL domain
The interference or effector module, responsible for target recognition, encodes for either a multiprotein effector complex (class 1 systems: types I, III and IV; termed Cascade in type I systems and Cmr/Csm in type III, the type IV complex does not have a specific name) or a single effector protein (class 2: types II, V and VI).
The class 1 effector complexes invariably comprise a Cas5 subunit and multiple subunits of Cas7, in addition to a small subunit (collectively denoted Cas11) and a large subunit (Cas8 or Cas10 in type I and III systems, respectively) (Jackson et al. 2014; Osawa et al. 2015). Some type I systems also have a Cas6 homolog associated with the complex (Haurwitz et al. 2010; Sashital et al. 2011). In type I systems, the crRNA-effector complex (surveillance complex) recruits an effector nuclease (Cas3) for target cleavage (Brouns et al. 2008). In other systems (type III and class 2), the effector proteins/complexes themselves are responsible for target cleavage (Jinek et al. 2012; Shmakov et al. 2015; Yan et al. 2019). Cas proteins involved in the interference stage seem to be absent in most type IV systems (Pinilla-Redondo et al. 2020).
CRISPR-Cas types of class 2 are distinguished by the single-protein effector associated with the system, namely, Cas9, Cas12 or Cas13 for type II, type V and type VI, respectively. The three protein families differ in the number, type and architecture of the nuclease(s) domain(s): type II and type VI effector proteins contain two nuclease domains (HNH and RuvC in the case of Cas9, two HEPN domains in Cas13), whereas those of type V have just one (RuvC) (Shmakov et al. 2015; Abudayyeh et al. 2016).
The adaptation module comprises the genes encoding enzymes involved in spacer acquisition, including Cas1 (fused to a reverse transcriptase domain in some type III and type VI systems), Cas2 (Cas1 and Cas2 are present in all the adaptation-proficient, autonomous systems), Cas4 (found in many type I, II and V systems) and Csn2 (exclusive to subtype II-A systems). Nevertheless, some system subtypes, notably within type IV and at a lower proportion in type III and VI systems, are devoid of any genes of the adaptation module (see section below on ‘Adaptation’).
The expression module deals with cleavage of the pre-crRNA and processing into mature crRNAs. Whereas this role is played by a dedicated Cas protein associated with many class 1 systems (see section below on ‘crRNA biogenesis’), class 2 involves a catalytic domain of the effector protein and, at least in type II systems, non-Cas ribonucleases.
Orphan CRISPR arrays and a range of unclassifiable, intermediate and minimal CRISPR-Cas configurations exist (Shmakov et al. 2020a; Pourcel 2020), suggesting degeneration of the CRISPR-Cas systems (Hermans et al. 1991; García-Gutiérrez et al. 2015; Chen et al. 2019). There is evidence that at least some of these apparently incomplete systems are functional (see below).
The general CRISPR-CAS mechanism
Three stages have been identified in the generation of CRISPR-Cas immunity, namely (i) adaptation, (ii) crRNA biogenesis and (iii) interference (Fig. 2). This mechanism involves CRISPR RNAs and core Cas proteins encoded by various gene modules (as discussed above and recently reviewed by Nussenzweig and Marraffini 2020). In addition, the functionality of systems devoid of some of these components, both working autonomously and relying on either auxiliary proteins or CRISPR/Cas activities provided by systems that co-occur in the cell, has also been documented. Most of these atypical systems follow the adaptive, RNA-guided, nucleic-acid targeting and cleavage scheme, thus being considered programmable nuclease systems. Nevertheless, others result in alternative outcomes, such as guided transposition or targeting without cleavage. These alternatives to the canonical process will be covered later in this manuscript.
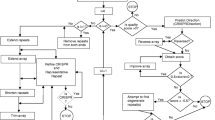
Schematic representation of the Adaptation, crRNA biogenesis and Interference stages of the canonical CRISPR-Cas mechanism. Sequences, typically derived from protospacers located next to a protospacer adjacent motif (PAM), are captured and processed by the adaptation complex (composed of at least Cas1 and Cas2 subunits). Non-Cas proteins may assist pre-spacer processing (not shown). The processed fragments are then preferentially integrated as spacers at the leader-repeat junction of the CRISPR loci by the adaptation complex (Adaptation stage). crRNAs are generated during the crRNA biogenesis stage after cleavage of the CRISPR array’s transcript (pre-crRNA). This cleavage is catalysed by different proteins depending on the system type. In some cases, subsequent maturation of the crRNAs is performed by either Cas or non-Cas exonuclease activities (see text for details). For pre-crRNA and target cleavage, type II systems and some type V subtypes require other CRISPR RNAs (tracrRNA or scoutRNA) that remain hybridised to the partially complementary crRNA. tracrRNA and scoutRNA are drawn with dashed lines when they are involved in only some subtypes of a given CRISPR-Cas type. During the interference stage, mature crRNAs bound to the effector complex (class 1) or protein (class 2), base pair with sequences complementary to the spacer region in RNA (type III, type VI and some type V systems) or DNA (the remaining types) target molecules. The recruitment of endonucleases (i.e. Cas3 in type I) or the activation of nuclease domains in the surveillance complex (other types) after target binding will trigger specific target cleavage. In types III, V and VI, target cleavage or binding leads to collateral unspecific cleavage of RNA (type VI systems), ssDNA (some type V systems) or both ssDNA and RNA (type III systems). Surveillance complex components and substrate identity have not been established for some system types or subtypes (indicated with a question mark)
Adaptation
Adaptation is the first step of the CRISPR-Cas mechanism, where a molecular memory is generated (for a recent review, see Mosterd et al. 2021). During this process, prespacers derived from protospacer-containing sequences are integrated into the CRISPR array as new spacers. Two models of CRISPR adaptation have been described, called primed and naïve (Datsenko et al. 2012; Yosef et al. 2012). Cas proteins of both the adaptation and effector module participate in primed adaptation, leading to a biased acquisition of spacers derived from the genetic element carrying the targets of pre-existing spacers. In contrast, naïve adaptation only requires adaptation machinery, and the selection of protospacers is independent of previous acquisitions. Both naïve and primed adaptations have been experimentally confirmed for the subtype II-A CRISPR-Cas system from Streptococcus mutans and diverse type I systems of Escherichia coli (subtype I-E), Legionella pneumophila, Pseudomonas aeruginosa and Geobacter sulfurreducens (Datsenko et al. 2012; Díez-Villaseñor et al. 2013; Savitskaya et al. 2013; Semenova et al. 2016; Rao et al. 2017; Almendros et al. 2019). In addition, primed but not naïve adaptation was detected in subtype I-B of Haloarcula hispanica (Li et al. 2014a, b) and the subtype I-F of Pectobacterium atrosepticum and E. coli (Richter et al. 2014; Vorontsova et al. 2015). On the contrary, naïve but not primed adaptation has been reported for subtype I-A from Sulfolobus solfataricus and Sulfolobus islandicus, II-A of Streptococcus agalactiae and III-B in S. solfataricus (Deveau et al. 2008; Erdmann et al. 2014; Heler et al. 2015; Shiimori et al. 2017; Nussenzweig et al. 2019; Artamonova et al. 2020).
Cas1 and Cas2 are the essential Cas proteins for adaptation (Makarova et al. 2015; Koonin et al. 2017). The minimal adaptation complex of type I systems comprises two Cas1-dimers joined by a Cas2-dimer (Nuñez et al. 2014, 2015). Nuclease and integrase activities involved in the spacer integration process are provided by Cas1 (Wiedenheft et al. 2009; Babu et al. 2011), while Cas2 has a structural function (Wang et al. 2015). In some CRISPR-Cas systems, additional Cas proteins and activities participate in adaptation: Cas4 in some type I, II and V subtypes (Heler et al. 2015; Hudaiberdiev et al. 2017; Rollie et al. 2018; Kieper et al. 2018; Shiimori et al. 2018; Lee et al. 2019; Almendros et al. 2019), Cas9 in type II, together with Csn2 in subtype II-A systems (Wei et al. 2015) and the reverse transcriptase domains fused to Cas1 (RT-Cas1) in type III and VI variants (Silas et al. 2016; Toro et al. 2019; González-Delgado et al. 2019).
Before their integration, spacer precursors must be recruited by the adaptation complex. Most CRISPR-Cas systems select protospacers after recognising the PAM. Specific motifs located in the PAM region next to the interference target (therefore also referred to as PAM) are likewise required for efficient target recognition and cleavage performed by the effector Cas proteins (Deveau et al. 2008; Datsenko et al. 2012; Swarts et al. 2012; Shah et al. 2013). Although the adaptation and the interference consensus PAMs may differ for a given CRISPR-Cas system (Almendros et al. 2012; Shah et al. 2013), there is a strong preference for acquiring spacers from sequences flanked by a PAM that is compatible with the interference machinery (Yosef et al. 2012, 2013; Díez-Villaseñor et al. 2013; Nuñez et al. 2014; Levy et al. 2015). Bona fide PAMs have not been identified next to protospacers of most type III systems, which is not surprising since type III systems usually display PAM-independent interference. Remarkably, in those type III systems associated with RT-Cas1, ssRNA transcripts are captured and converted into DNA once incorporated into the CRISPR array (Silas et al. 2016; González-Delgado et al. 2019).
In E. coli, although induction of Cas1 and Cas2 of its I-E system triggers naïve acquisition of spacers mainly derived from resident plasmids, chromosomal DNA sequences are also captured (Levy et al. 2015). These chromosomal spacers preferentially derive from the origin of replication and, notably, the terminus region. In this regard, it has been proposed that ssDNA fragments generated upon repair of DNA double-strand breaks, like those produced during replication (Smith 2012), are the primary source of prespacers (Levy et al. 2015; Radovčić et al. 2018). This replication-related origin of naïve spacers would partially explain the prevailing acquisition of sequences from plasmids versus the less often replicated chromosomes. However, it is unknown whether the Cas1-Cas2 complex captures ssDNA or dsDNA molecules, and it remains to be elucidated when and how the dsDNA spacers are generated from repair-derived ssDNA fragments. Concerning primed adaptation, it has been shown that the nuclease activity of Cas3, probably associated with the adaptation machinery in the so-called primed adaptation complex, produces prespacers in I-E and I-F systems (Shiriaeva et al. 2020; Musharova et al. 2021).
Once loaded into the Cas1-Cas2 adaptation complex, the prespacer ends must be trimmed to the size of the spacer. In subtypes I-A and I-C, pruning of the prespacers’ 3′ ends is carried out by the Cas4 protein (Rollie et al. 2018; Lee et al. 2019). However, many CRISPR-Cas systems lack Cas4. Subtype I-E in S. thermophilus is one of the few systems where Cas2 is fused to a DnaQ domain. In vitro studies have shown that this domain has exonuclease activity and that the I-E adaptation complex of this species can process and integrate duplex oligonucleotides with 3′ protruding ends (Drabavicius et al. 2018). In the case of the I-E system of E. coli (devoid of Cas4 and with a Cas2 protein that does not exhibit exonuclease activity), DnaQ and ExoT exonucleases, as well as the proofreading subunit of the DNA polymerase III, have been suggested to be involved in the prespacer trimming (Kim et al. 2020; Ramachandran et al. 2020). According to the model proposed, once the prespacer is loaded into the Cas1-Cas2 complex and the PAM is recognised, host nucleases will degrade the ends of the prespacer until reaching the region protected by the complex (Yoganand et al. 2019).
dsDNA prespacers are preferentially integrated into the CRISPR locus at the leader proximal end of the CRISPR array (Barrangou et al. 2007). The adaptation machinery of type II systems spots this integration site after recognising the leader-anchoring sequence (LAS) within the leader (Wei et al. 2015). Similarly, in subtype I-E, specific sequences in the leader and the repeat are bound by the integration host factor (IHF) protein, which generates a docking site for the Cas1-Cas2 complex upon bending this DNA region at the leader-repeat junction (Nuñez et al. 2016; Yoganand et al. 2017). Once in place, the Cas1-Cas2 complex catalyses direct nucleophilic attack of the 3′-OH ends of the prespacer at the leader-repeat junction and, subsequently, at the repeat-spacer boundary, resulting in a dsDNA spacer flanked by single-stranded repeats that must be repaired and ligated. Neither the ligase nor the DNA polymerase involved in this process has been identified so far. However, E. coli DNA polymerase I mutants cannot acquire spacers, uncovering this protein as a putative candidate (Ivančić-Bace et al. 2015). Meanwhile, a recent study suggested that primase-polymerase homologues associated with some III-A and III-B CRISPR-cas loci might participate in spacer adaptation (Zabrady et al. 2021).
crRNA biogenesis
The biogenesis of crRNAs is a crucial step for target recognition and cleavage. CRISPR arrays are usually transcribed from a single promoter located in the leader, generating the pre-crRNA. Later, this transcript is cleaved into small RNAs. Finally, at least in some systems, these RNA molecules are trimmed to create the mature crRNAs that will become a functional component of the surveillance complex. Occasionally, transcription is observed from promoters within the CRISPR array, either in spacers or in repeats (Lillestøl et al. 2009; Wurtzel et al. 2010; Deng et al. 2012; Zhang et al. 2013).
Class I crRNA maturation is performed by Cas6 protein except for subtypes I-C, III-C and III-D (Carte et al. 2008; Haurwitz et al. 2010; Gesner et al. 2011; Sashital et al. 2011; Nam et al. 2012; Richter et al. 2012; Garside et al. 2012; Özcan et al. 2019). The partially palindromic sequences of the repeats that constitute the CRISPR arrays in types I-D, I-E and I-F adopt a stem-loop structure in the transcribed pre-crRNA. Cas6 recognises this hairpin structure and cleaves the pre-crRNA downstream the loop (Haurwitz et al. 2010; Gesner et al. 2011; Sashital et al. 2011; Nam et al. 2012). The resulting crRNA comprises a spacer flanked by a short repeat-derived sequence at the 5′ end and the stem-loop at the 3′ end. Cas6 remains bond to the crRNA loop and serves as a scaffold for the multimeric effector complex (Jore et al. 2011; Sashital et al. 2011). For subtype I-C, the crRNA maturation occurs similarly, but the function of Cas6 is performed by Cas5d (Nam et al. 2012; Garside et al. 2012). In subtypes I-A and I-B, CRISPR repeats are not palindromic, and in III-A and III-B systems, hairpins are unstable (Koonin et al. 2017). In these cases, dimers of Cas6 generate a conformational change in the pre-crRNA, creating a hairpin-like secondary structure and cleaving within the repeat (Richter et al. 2013; Shao and Li 2013; Reeks et al. 2013; Sefcikova et al. 2017). Later, Cas6 is released from the crRNAs whose 3′ ends are trimmed by a protein not yet identified (Carte et al. 2008, 2010; Hatoum-Aslan et al. 2011). The sequence of type III Cas6 proteins resembles that of types I-A and I-B, and, accordingly, their pre-crRNAs are processed following the same pattern. For subtypes III-C and III-D, as well as III-F, where no homologs to cas6 have been described, Cas5 orthologs could be responsible for pre-crRNA cleavage (Behler and Hess 2020). The maturation of crRNA in type IV has not been studied in detail. Still, for subtype IV-A, it was established that the Cas6 homolog Csf5 protein is responsible for pre-crRNA maturation (Özcan et al. 2019).
Cleavage of pre-crRNA and maturation of crRNA in class II systems relies on the effector protein and, depending on the CRISPR-Cas type, on other genetic elements or proteins. Type II systems require tracrRNA (Deltcheva et al. 2011; Gasiunas et al. 2012; Zhang et al. 2013; Shmakov et al. 2015). Once the tracrRNA anneals with its complementary sequence in the crRNA repeats region, in subtypes II-A and II-B, Cas9 binds and stabilises the crRNA:tracrRNA structure. Later, the homing ribonuclease RNaseIII cleaves the crRNA within the repeat at the 3′ end, and an unknown nuclease processes the 5′ end (Deltcheva et al. 2011). Instead, a tracr-dependent but RNaseIII-independent mechanism was discovered in II-C systems from Campylobacter jejuni, Neisseria meningitidis and Neisseria lactamica (Dugar et al. 2013; Zhang et al. 2013). In these cases, crRNAs are produced individually due to the presence of promoters within the repeats that generate short RNA molecules. In type V, the effector protein Cas12 is responsible for pre-crRNA cleavage. Subtype V-A uses a tracr-independent mechanism to process the pre-crRNA where Cas12, upon recognition of the hairpins formed in the repeat regions, cleaves within them to generate the crRNA. In the other type V subtypes, either tracrRNA or scoutRNAs are needed to efficiently process the crRNA (Yang et al. 2016; Liu et al. 2019a; Harrington et al. 2020). Meanwhile, the mechanism of pre-crRNA cleavage in subtype V-F remains unknown (Behler and Hess 2020). Cas13 RNA-effector protein of type VI can also cut within the pre-crRNA without the tracrRNA, relying on the recognition of the secondary structure of the repeat region in a similar way to type I, cleaving upstream of the hairpin (Shmakov et al. 2015; Abudayyeh et al. 2016; East-Seletsky et al. 2017).
In addition to RNase III, other host proteins may participate in crRNA maturation. In the type III-B of Synechocystis sp. PPC 6803, RNase E is recruited for pre-crRNA processing and cleaves it within the repeat (Behler et al. 2018). Polynucleotide phosphorylase (PNPase) cleaves the pre-crRNA in collaboration with Cas6 in the III-A system of S. epidermidis (Samai et al. 2015; Chou-Zheng and Hatoum-Aslan 2019), and in the I-B system of Haloferax volcanii, RNase Z and RNase P were repurposed to successfully cleave pre-crRNA in a modified strain lacking Cas6 (Maier et al. 2015).
Further investigations will be needed to decipher which yet unknown proteins are involved in the trimming of the crRNA and how the crRNA maturation is achieved in systems lacking Cas6 homologs, RNase III or tracrRNA/scoutRNA.
Interference
Interference is the last stage of the CRISPR-Cas mechanism, where the Cas effector proteins form a surveillance complex with CRISPR RNAs that guide the proteins to a target sequence complementary to the crRNA spacer region (Hille and Charpentier 2016). Firstly, in the case of PAM-dependent dsDNA targeting systems, the surveillance complex scans DNA molecules in search of PAMs. Once a cognate motif is identified, dsDNA is locally unwound. Then, the PAM-proximal positions in the so-called seed sequence (Semenova et al. 2011; Swarts et al. 2017) are probed for complementarity with the crRNA spacer and, subsequently, interrogation of base-pairing proceeds beyond that region. Then, as the RNA:DNA hybrid forms, the non-complementary DNA strand is displaced, forming an R-loop structure (Stella et al. 2017; Xiao et al. 2017). Finally, if sufficient hybridisation is reached, the target becomes fully accessible to the nuclease effector, licensing cleavage.
Even though this can be considered the typical mechanism of interference against dsDNA targets, there are prominent differences among CRISPR-Cas subtypes related to PAM requirement and location, as well as to the nucleic acids that are targeted (dsDNA, ssDNA and/or RNA sequences) and cleaved (only the target or, in addition, unspecific RNA and/or DNA sequences).
For a comprehensive review on class 1 effectors, see Liu and Doudna (2020). Among type I systems, interference has been studied in greater detail for subtype I-E of E. coli. First, the Cascade surveillance complex recognises a downstream (taking the target strand as a reference) PAM in duplex form (Hayes et al. 2016). After R-loop formation, the Cas3 protein is recruited by the complex (Xiao et al. 2017). Then Cas3 nicks within the displaced ssDNA at the R-loop and catalyses subsequent cuts as it translocates along this strand (He et al. 2020). In contrast to type I, the surveillance complex of at least some type III subtypes (III-A, B, C) degrades ssDNA and ssRNA. Through the RNase activity of the Cas7-like subunits in the Cmr/Csm complex, crRNA-complementary sequences in RNA molecules are degraded. The binding of the complex to the RNA target activates the ssDNA nuclease activity of Cas10 (another integral part of the complex). It was revealed in some III subtypes that cyclic oligoadenylates (cOAs) produced by Cas10 activate a separate Csm6 nuclease effector to degrade non-specific RNAs (Kazlauskiene et al. 2017; Niewoehner et al. 2017; Jia et al. 2019). Most type III systems do not require specific motifs flanking the target for efficient recognition, and, accordingly, clear evidence of functional seed sequences has not been found (Marraffini and Sontheimer 2010; Osawa et al. 2015; Estrella et al. 2016). However, the type III-B system from Pyrococcus furiosus recognises a protospacer flanking sequence or PFS (defined as protospacer flanking site by some authors) next to the RNA target (Foster et al. 2020). Interestingly, hybridisation between PFS and the crRNA prevents DNase and cOA production activities but still licences specific RNA degradation. Regarding type IV systems, the first experimental proof of interference in vivo was recently reported by Crowley and co-workers (Crowley et al. 2019). However, the identity of the target (DNA or RNA) and the mechanism involved remain undisclosed (Pinilla-Redondo et al. 2020).
In class 2 systems, type II and some type V subtypes require a hybrid RNA guide composed of tracrRNA and crRNA molecules (Jinek et al. 2012; Shmakov et al. 2015; Liu et al. 2019a; Yan et al. 2019). Subtype V-C and V-D systems, together with crRNA, require scoutRNAs for target cleavage (Harrington et al. 2020). While Cas9 recognises PAMs located downstream of the non-target strand (Gasiunas et al. 2020), PAMs located at the opposite flank are identified on both strands of the DNA target by Cas12 (Shmakov et al. 2015). Type II and some type V systems target dsDNA and cleave the two strands. Other type V subtypes target ssDNA, both dsDNA and ssDNA, or ssRNA (Harrington et al. 2018; Yan et al. 2019; Karvelis et al. 2020; Pausch et al. 2020). Remarkably, after specific target cleavage, at least some Cas12 nucleases develop collateral ssDNA or ssDNA and RNA nuclease activity (Yan et al. 2019). Meanwhile, Cas13 effector proteins of type VI systems degrade non-specific RNAs upon identification of the target RNA (Abudayyeh et al. 2016; Smargon et al. 2017; Liu et al. 2017; Yan et al. 2018). Although no canonical PAM sequence is required for efficient interference by these systems, some Cas13 variants recognise a PFS region downstream of the target (Leenay and Beisel 2017).
Functions of CRISPR-Cas systems
The most apparent benefit for a cell from encoding adaptive immunity machinery such as the one provided by the CRISPR-Cas systems is protection against viruses: the genome of invading viruses and resident proviruses entering a lytic cycle can be specifically degraded, preventing cell damage and the eventual spread of virions. Furthermore, the genetic memory licenced by integrating new spacers derived from the infecting virus will further protect the descendants of the adapted cell for generations, thus perpetuating the anti-virus outcome. Similarly, looking at plasmids as parasitic agents that may place a burden on the cell, the immunity concept also covers interference against these transmissible molecules. Moreover, type IV systems primarily found in plasmids preferentially target sequences in other plasmids, suggesting that this CRISPR-Cas type is specialised in competition between these kinds of molecules (Pinilla-Redondo et al. 2020).
Immunity at the cell and population level is considered the primary purpose of CRISPR-Cas (Edgar and Qimron 2010; Cady et al. 2012; Strotskaya et al. 2017; Watson et al. 2019; Deem 2020). In fact, since 2007 (Barrangou et al. 2007), many studies have proven this defensive role in prokaryotes. Thus, both virus resistance and plasmid cleavage has been used as a recurrent strategy to assess CRISPR-Cas activity (Marraffini and Sontheimer 2008; Garneau et al. 2010; Westra et al. 2013b; Almendros and Mojica 2015; Crowley et al. 2019; Wheatley and MacLean 2020).
Otherwise, invasive mobile genetic elements (iMGEs) such as viruses and plasmids represent an opportunity to acquire foreign DNA. Hence, immunity against these elements has the potential to restrict horizontal gene transfer (HGT). Indeed, it has been shown that CRISPR-Cas systems constitute a barrier to HGT in diverse bacteria and archaea, preventing conjugation, transduction and natural transformation and thereby influencing traits such as bacterial virulence and drug resistance or even microbial speciation (Marraffini and Sontheimer 2008; Mojica and Díez-Villaseñor 2013; Turgeman-Grott et al. 2019; Zhou et al. 2020; Kamruzzaman and Iredell 2020; Wheatley and MacLean, 2020). Interestingly, the inverse has also been reported: recombination between CRISPR spacers in bacterial genomes and their targets in invading bacteriophages facilitates the transfer of CRISPR-Cas systems and adjacent regions through escape transduction particles, favouring, rather than dampening, HGT (Watson et al. 2018; Varble et al. 2019).
However, works assessing the relevance of CRISPR-Cas in the battle against viruses and lateral gene dissemination are scarce. Hence, the actual impact of these systems in natural environments remains to be firmly established (Westra and Levin 2020; Martínez Arbas et al. 2021).
On the other hand, most spacers so far identified in prokaryotic genomes (i.e. chromosomes and resident plasmids) do not match known virus or plasmid sequences (Shmakov et al. 2017b, 2020b). In this context, functions other than the control of iMGEs have been reported for complete and partial CRISPR-Cas systems. These non-canonical activities range from regulatory tasks to guiding transposition events and result in virulence control or genome evolution, among many other outcomes. Some of these roles are just hypothetical, even though well substantiated. For instance, the cOAs synthesised by type III Cas10 (Kazlauskiene et al. 2017) might act as extracellular messengers that enable bacterial communication, integrating Cas proteins within cell signalling pathways. Other non-canonical functions for CRISPR-Cas encoded in prokaryotic genomes have been identified and will be discussed below. Given the peculiarities of the roles played by CRISPR and Cas found in prokaryotic viruses, their functions will be addressed in a dedicated section.
Cytotoxicity, cell dormancy and regulation of gene expression
The indiscriminate degradation of nucleic acids exhibited by some CRISPR-Cas systems after recognising the specific target may result in cell suicide (Hale et al. 2009, 2012; Abudayyeh et al. 2016; Liu et al. 2017).
Furthermore, the observation that many spacers perfectly match fragments within the carrier genome (i.e. self-targeting spacers) (Horvath et al. 2008, 2009; Stern et al. 2010) raised the possibility that CRISPR-Cas activity might have a variety of consequences in non-infected cells (reviewed in Wimmer and Beisel 2020). At present, full-matching self-targeting spacers and spacers with only partial complementarity to resident genomic sequences have been involved in downregulation and upregulation of expression affecting DNA repair, virulence, anti-microbial susceptibility or cell development processes (Newsom et al. 2021). However, in many cases, the underlying regulatory mechanism remains to be established (Table 1). For example, even though endogenous gene regulation unconnected to immunity was predicted as the primary function of the type II-B Cas2 protein in Legionella pneumophila (Gunderson and Cianciotto 2013), and an orphan CRISPR locus in Listeria monocytogenes (Mandin et al. 2007), further studies are necessary to confirm this implication (Bozic et al. 2019).
Better known illustrations of gene regulation executed by CRISPR-Cas systems through DNA or RNA targeting, and their consequences, are discussed below.
Regulation by DNA targeting
The first case of a non-canonical function played by CRISPR-Cas acting on DNA was reported for the type I-F system of P. aeruginosa (Zegans et al. 2009). The Cas3 nuclease, guided by a crRNA partially complementary to a resident prophage sequence, generated minor DNA damage instead of the processive degradation of the target characteristic of full-matching spacers (Xiao et al. 2018). Nevertheless, this DNA insult triggers the SOS response, which leads to de-repression of phage-related lysis genes (Cady and O’Toole 2011; Heussler et al. 2015). In this way, CRISPR-Cas activity indirectly induces the expression of proteins that can kill the cell, dampening dissemination of the carried phage and thus behaving as a population protective mechanism.
Through DNA targeting without cleavage, CRISPR-Cas can also achieve direct regulation of endogenous genes. This action is exemplified by Cas nucleases that bind DNA but are not able to cut it due to only partial complementarity with the crRNA, resulting in transcriptional silencing when the target is located near promoter regions (Ratner et al. 2019; Sampson et al. 2019). Indeed, an early work (Aklujkar and Lovley 2010) suggested that interaction of a crRNA from the I-E system of Pelobacter carbinolicus with a partially complementary sequence in the histidyl-tRNA synthetase gene (hisS) resulted in an attenuated histidyl-tRNA pool. Moreover, estimation of the amount of hisS transcript in a heterologous host carrying the targeting spacer hinted at a cleavage independent regulatory mechanism, ruling out the degradation of the hisS RNA and cleavage of the encoding DNA by the associated Cas nuclease. The type II CRISPR-Cas system in the pathogenic bacterium Francisella novicida typifies another well-substantiated case of gene expression regulation mediated by DNA-binding without cutting. It has been revealed that Cas9 transcriptionally represses endogenous genes through binding to the DNA targets, guided by a tracrRNA-scaRNA (small CRISPR/Cas-associated RNA) hybrid (Ratner et al. 2019; Sampson et al. 2019), rather than through RNA degradation as previously suggested (Sampson et al. 2013). This repression facilitates bacterial evasion of the innate immune system in infected mammalian hosts, enhancing virulence.
Cas proteins can also function as transcriptional activators upon binding to the gene promoters. For instance, the type I-A associated protein Csa3a from S. islandicus activates transcription of the adaptation cas genes and multiple repair genes such as those encoding DNA polymerase II, DNA polymerase IV, the NurA nuclease and the helicase HerA (Liu et al. 2017). Although the precise link between CRISPR and DNA repair has not been established, this case supports the synergy between CRISPR-Cas activity and the DNA repair process (see the ‘DNA repair’ section).
Repression through RNA cleavage
On the other hand, regulation of gene expression by some CRISPR-Cas systems is also achieved through RNA cleavage. In addition to the RNA-targeting CRISPR-Cas, some type II (Louwen et al. 2013; Sampson and Weiss 2013; Sampson et al. 2013; O’Connell et al. 2014; Dugar et al. 2018; Rousseau et al. 2018; Strutt et al. 2018) and type I (Li et al. 2016) systems that typically target DNA possess promiscuous nucleases able to bind and cut within RNA molecules, resulting in regulatory functions. In this context, it has been reported that E. coli subtype I-E Cascade binds ssRNA in vitro (Jore et al. 2011), and Cas3 can degrade ssRNA (Babu et al. 2011). However, evidence of such activities has not been provided in vivo. More recently, Li and co-workers showed that Cas3 protein and Cascade complex of P. aeruginosa PA14 (subtype I-F system) are involved in lasR mRNA degradation requiring just the presence of a PAM-like motif next to the lasR mRNA target and as little as 28% complementarity with the spacer (Li et al. 2016). Surprisingly, both sequence requisites are found in many other mRNAs encoded by the PA14 genome, raising questions such as whether this CRISPR system plays a significant role in regulating the abundance of individual mRNAs in the cell or, if lasR mRNA is the sole target (this question remains to be addressed), how specificity is achieved (Müller-Esparza and Randau 2017). In addition to type I, diverse subtype II-C and S. agalactiae II-A systems have also been reported to cleave RNA and, as in the case of the P. aeruginosa I-F system, efficient degradation of endogenous RNAs occurs despite only partial complementarity to naturally occurring spacers (Dugar et al. 2018; Ma et al. 2018; Rousseau et al. 2018; Strutt et al. 2018).
Genome evolution
Genome evolution is the process of genetic variation in response to environmental variables. In prokaryotes, it occurs through point mutations, horizontal gene transfer and genome rearrangements. These events generate novel genotype variants which are transferred from a source cell to subsequent generations.
Prokaryotic evolution can be affected by CRISPR-Cas activity when it targets foreign genetic elements (e.g. dampening HGT) and when targets the host genome (self-targeting spacers), leading to autointerference. It has been estimated that self-targeting spacers account for 6% of the total pool identified in available sequenced genomes (Shmakov et al. 2017b). Thus, some incomplete, apparently non-functional CRISPR-Cas systems might have emerged because of self-targeting. Otherwise, mutations or rearrangements must occur in the target region to prevent autointerference and cell death, thus pushing genome evolution (Stern et al. 2010; Wimmer and Beisel 2020). This possibility has been experimentally demonstrated in a few cases. For example, induction of the type I-F CRISPR-Cas system in P. atrosepticum carrying self-targeting spacers resulted in deletions involving the target sequence (Vercoe et al. 2013). Likewise, in S. thermophilus, genome rearrangements of a chromosomal locus were also reported to happen at an increased frequency when the resident II-A CRISPR-Cas system targeted sequences located in that region (Selle et al. 2015; Cañez et al. 2019). Similar results have been reported for subtype I-B of Haloferax volcanii (Stachler and Marchfelder 2016) and subtypes I-A and III-B of S. islandicus (Li et al. 2015). Further proof of the concept that CRISPR-Cas immunity can contribute to bacterial diversity has been recently provided (Mo et al. 2021). The authors showed that the III-A CRISPR-Cas system in Staphylococcus species is mutagenic even in the absence of infecting agents. Moreover, increased host mutations occurred upon CRISPR-mediated targeting of lytic phages or plasmids when the associated Cas10 protein was active. These results suggested that the collateral ssDNA cleavage activity of Cas10 and the subsequent DNA repair would be responsible for random mutations in the chromosome, raising the possibility that other systems exhibiting non-specific ssDNase activity (i.e. Type V CRISPR-Cas) could modulate genome evolution in the same way.
Another remarkable case of CRISPR-driven evolution was proposed for the CRISPR-Cas I-E system from P. carbinolicus (Aklujkar and Lovley 2010), involving the previously mentioned (the ‘Cytotoxicity, cell dormancy and regulation of gene expression’ section) spacer matching a sequence within the hisS gene. CRISPR-mediated interference against the histidyl-tRNA-synthetase activity is expected to impair histidine-enriched proteins’ translation. Indeed, in contrast to closely related bacteria lacking such a hisS-interference capacity, P. carbinolicus cannot reduce Fe(III), a catalytic process that involves enzymes with high histidine content. Notably, transcription of hisS in a recombinant, closely related species carrying a hisS-targeting CRISPR-Cas system decreased compared to control strains without targeting capacity. Thus, CRISPR-mediated self-targeting might be responsible for the loss in this species of ancestral genes encoding proteins with high histidine content, having contributed significantly to its metabolic divergence from other members of the Geobacteraceae family.
CRISPR-Cas systems in prokaryotic cells might also accelerate the mutation rate of invasive genetic elements during infection. This effect has been demonstrated for a type II system in heterologous E. coli hosts exposed to a CRISPR-targeted bacteriophage T4 (Tao et al. 2018). Mutation frequencies in the phage genome were several orders of magnitude higher than the frequency observed in the absence of CRISPR-Cas activity. In this way, the CRISPR-Cas systems promote variability of the virus population providing selective advantages to infectious agents in parallel to the protective role played in the host.
DNA repair
Mechanisms of genetic repair that rely on the arrangement of damaged DNA by gap-filling or ligation reactions link to the CRISPR-Cas mode of action.
For example, regarding the adaptation apparatus, the nuclease Cas1 of the I-E system from E. coli interacts with several repair system components (i.e. RecB, RecC, RuvB and RuvC), and it is actively involved in the cell rescue during DNA damage (Babu et al. 2011). Also related to spacer acquisition, it was reported that the II-A CRISPR-Cas associated protein Csn2 can inhibit DNA repair by the non-homologous end-joining (NHEJ) mechanism, explaining the low frequencies at which NHEJ repair and II-A CRISPR-Cas systems coexist within the same microbial genome (Bernheim et al. 2017). Because Csn2 binds the free DNA ends generated after cleavage produced by the adaptation complex at the spacer integration site (Nam et al. 2011; Arslan et al. 2013), it was proposed that repair inhibition is due to competition between Csn2 and the NHEJ-associated Ku protein for binding DNA ends to be repaired.
On another front, it has been shown that the type II effector protein Cas9 triggers the SOS-system response in the heterologous host E. coli as a collateral effect of its DNA-targeting (Cui and Bikard 2016). Interestingly, this SOS response leads to repairing the damaged genetic material through homologous recombination or, in the absence of donor DNA, large deletions due to the action of the RecBCD pathway.
These observations indicate a strong interconnection between CRISPR-Cas and DNA repair machinery, which merits further attention.
Guided transposition
Transposons (Tn) are mobile DNA elements capable of excising and inserting themselves elsewhere in the genome through the activity of transposases assisted by other accessory proteins encoded by the element. The identity of the DNA targeted for integration by the TnsAB transposase complex of the prokaryotic Tn7 transposons is marked by either TnsD (targets the specific attachment site attTn7 in the chromosome) or TnsE protein (targets random sequences in the lagging DNA strand produced during plasmid replication) (Peters et al. 2017; Dimitriu et al. 2019).
Tn7-like transposons associated with I-B and I-F CRISPR-Cas systems were detected in silico in 2017 (Peters et al. 2017). Subsequently, association with similar transposons was also reported for subtype V-K systems (Strecker et al. 2019). These CRISPR-associated transposases (CASTs) consist of core transposase genes, either one (V-K and I-F systems) or two (I-B systems) tniQ genes (homologous to tnsD), a small CRISPR array and the genes encoding Cas effector proteins (Cascade proteins in type I systems or a nuclease-deficient Cas12 in type V systems), lacking adaptation module and effector nuclease (Cas3 in type I) activity (Peters et al. 2017; Faure et al. 2019a; Klompe et al. 2019; Strecker et al. 2019; Saito et al. 2021). A tracrRNA is also present in V-K CRISPR loci. Thus, the capacity to form surveillance complexes with guide RNA molecules is still maintained. Interestingly, TniQ proteins interact in the three subtypes with the surveillance complex to promote transposon integration next to sites targeted by the crRNA (Klompe et al. 2019; Strecker et al. 2019; van der Oost and Mougiakos 2020; Saito et al. 2021). RNA-guided transposition into specific sites of the host genome (homing transposition) involves crRNAs with either a truncated spacer encoded in a CRISPR array located away from the CRISPR-Cas locus (Subtype V-K; Saito et al. 2021) or spacers with low identity to the target, encoded by the cognate CRISPR array but flanked by diverged repeats (I-F systems; Petassi et al. 2020). Noteworthy, in addition to RNA-guided, homing transposition in I-B systems is elicited by a protein-target mechanism independent of crRNA and Cascade (Saito et al. 2021), involving just the larger TniQ protein (the shorter one participates in RNA-guided integration).
CASTs harbouring spacers that target iMGEs could represent a way to bolster gene transfer through crRNA-guided transposition, as opposed to the canonical CRISPR-Cas systems acting as genetic barriers. Once in the recipient cell, homing transposition would facilitate CAST integration into the host genome. However, the limited length of the CASTs CRISPR arrays and the fact that adaptation modules are invariably missing suggest that the expansion of the transposition sites repertoire is restricted. Still, spacers could be integrated de novo by compatible adaptation complexes encoded by CRISPR-Cas systems co-occurring in the cell.
CRISPR/Cas in prokaryotic viruses
Deep in silico analyses have revealed the presence of CRISPR-Cas components in plasmids, viruses and proviruses. Complete CRISPR-Cas systems seem to be very infrequent in viral sequences. Moreover, even though many systems found in chromosomes have been tentatively assigned to provirus regions, CRISPR-Cas components could have been inserted within the viral region after provirus integration. Thus, just two reliable cases of complete CRISPR-Cas systems have been reported so far in viruses: the I-B system of Clostridium botulinum phage D-1873 and the I-F system of Vibrio spp. phages (Seed et al. 2013; Faure et al. 2019b). So far, only the latter has been experimentally validated. Interestingly, on the basis that carried spacers target a host anti-phage island, it has been proposed that the system in Vibrio phages might function as a counter-defence mechanism (Seed et al. 2013; Naser et al. 2017).
In viruses and proviruses, stand-alone CRISPR arrays and sequences like canonical repeat units (named solitary repeat units or SRUs) are significantly more common than complete CRISPR-Cas. SRUs resembling the CRISPR sequences in the host have been tentatively related to anti-CRISPR mechanisms (Faure et al. 2019b). According to the proposal, SRUs might act as dominant-negative inhibitors of the CRISPR-Cas machinery of the host by competing with bona fide crRNAs for binding to the effector proteins. It has also been envisaged that the integration of the viral genome within a host CRISPR-cas locus might occur via homologous recombination between SRUs and similar repeats in resident CRISPR arrays, interrupting their transcription.
Strikingly, some of the larger orphan arrays found in prokaryotic viruses contain spacers that match host genes or intergenic regions, suggesting that they may play regulatory roles (Al-Shayeb et al. 2020). However, most of these CRISPR arrays are very small (mini-CRISPR arrays), composed of a single spacer flanked by either two complete repeats or a complete repeat and a truncated CRISPR-like sequence. The sequences of these repeats in mini-arrays are in most cases identical or very similar to repeats of complete CRISPR-Cas systems present in the respective host genomes (Faure et al. 2019b; Medvedeva et al. 2019). Regarding the spacers, in contrast to the low percentage (roughly 10%) of spacers in prokaryotic genomes that match known sequences, it has been estimated that 67% and 93% of the spacers carried in mini arrays of viruses and proviruses, respectively, have a potential target (Shmakov et al. 2017b). Notably, most of these spacers match sequences in similar viruses or proviruses but not in the respective viral or host genome. Moreover, putative promoters have been tentatively identified upstream most mini arrays detected, suggesting that they are transcribed (Faure et al. 2019b). The absence of cas genes and other CRISPR sequences necessary for interference (e.g. tracrRNA genes are absent in the case of type II mini arrays) implies that the activity of these mini arrays depends on host CRISPR and Cas elements (Faure et al. 2019b; Medvedeva et al. 2019; Iranzo et al. 2020). These observations led to the hypothesis that solitary mini-CRISPR arrays hijack the host CRISPR-Cas systems to tackle virus superinfection. Thus, when a virus carrying a mini array infects the cell, crRNAs produced from the array would guide the host Cas effector proteins to inhibit infection by a targeted competitor virus.
Moreover, the viral mini array could expand its spacer repertoire acquiring new spacers from the second infecting virus through the adaptation machinery of the host. Superinfection inhibition and acquisition of new spacers have been experimentally demonstrated for mini arrays of viruses infecting the archaeon Saccharolobus spp. (Medvedeva et al. 2019).
Finally, a mathematical model on the cost of mini-CRISPR array maintenance and productivity of co-infection events predicted that mini arrays should be more frequent in viruses with a narrower host range, where competition with co-infecting viruses is of prime relevance (Iranzo et al. 2020). According to the theoretical prediction, the mini-CRISPR arrays of viruses that infect the same cell might undergo a rapid co-evolution. As a result, each would be forced to update its spacers pool to re-enlist effective targeting against the respective competitor viruses when CRISPR-evading mutations arise.
Experimental confirmation of these theoretical concepts and the underlying mechanism involved in mini arrays functioning will allow understanding of their role and decipher their regulation, efficiency, and consequences within the microbial population.
CRISPR-Cas control
Having CRISPR-Cas constantly turned on would allow for rapid neutralisation of iMGEs; however, uncontrolled expression may also have several disadvantages. The potential toxic effects of CRISPR-Cas action, notably that of the Cas nucleases acting on self-nucleic acids, and the fitness cost associated with the CRISPR-Cas expression involves the need for checkpoints of the CRISPR mechanism and regulatory strategies to fine-tune repression, induction, activation and inactivation at transcriptional, translational and post-translational levels (Patterson et al. 2017; Leon et al. 2018). Moreover, iMGEs have evolved diverse mechanisms to evade CRISPR-based interference. These aspects of CRISPR-Cas control are summarised in the following sections.
Checkpoints of the CRISPR-Cas mechanism
Determinants of CRISPR-Cas functioning contribute to preventing cell toxicity. Concerning the first stage of CRISPR-Cas immunity, the self-targeting rate is reduced through the preferential uptake of spacers from foreign genetic elements during naïve adaptation (Levy et al. 2015), while primed acquisition leads to a biased integration of spacers derived from pre-targeted regions (Vorontsova et al. 2015). The adaptation complex can capture debris left by DNA repair machinery. In Gram-negative bacteria, upon recognising free DNA ends (Ivančić-Bace et al. 2015; Levy et al. 2015; Radovčić et al. 2018), RecBCD proceeds by degrading DNA until reaching a Chi site (Smith 2012). The AddAB system, a paralog of the RecBCD complex in Gram-positive bacteria, is necessary for efficient spacer acquisition in some of these microorganisms (Modell et al. 2017). Moreover, regions between free DNA ends and Chi sites are more prone to be acquired by the II-A system in Streptococcus pyogenes (Modell et al. 2017). The higher content of Chi sequences in the bacterial chromosome compared to the low frequency of Chi-like sequences usually found in transmissible genetic elements results in fewer spacer-donor regions in self-DNA than in foreign molecules, explaining in part the apparent preference for naïve acquisition of spacers derived from the latter (Levy et al. 2015). Although other repair proteins such as PriA and RecG are involved in primed adaptation in E. coli (Ivančić-Bace et al. 2015; Killelea and Bolt 2017; Radovčić et al. 2018), the precise role played and potential checkpoints remain to be elucidated.
Regarding the crRNA biogenesis stage, Cas ribonuclease-mediated cleavage of RNAs other than pre-crRNAs is impeded due to the recognition of specific sequences and either the stem-loop adopted by palindromic repeats or the structure formed by tracrRNA:pre-crRNA hybrids (see section above on ‘crRNA biogenesis’).
Cleavage by Cas interference nucleases requires the navigation of multiple checkpoints involving sequential conformational rearrangements of the effector protein that occur after binding to the guide RNA, the target or other Cas proteins (reviewed in Jackson et al. 2017). Thus, their nucleolytic capability is only activated during the recognition of guide-complementary targets (Sternberg et al. 2014; Hochstrasser et al. 2014).
Still, the activity of Cas nuclease effector proteins poses a risk to the cell if guided with crRNA targeting sequences that match resident regions. Targeting of the CRISPR locus by Cas nuclease effectors is prevented due to the requirement for a PAM (Westra et al. 2013a; Foster et al. 2020) or mismatches between the target and the crRNA beyond the spacer region (Marraffini and Sontheimer 2010; Meeske and Marraffini 2018; Foster et al. 2020). Finally, the collateral random ssDNA degradation and non-complementary RNA cleavage exhibited by type III systems only occur after binding of the surveillance complex to targeted RNA, thus limiting potential damage of the own nucleic acids to actively transcribed genetic elements (Samai et al. 2015; Jia et al. 2019; Sofos et al. 2020; Foster et al. 2020). Moreover, autoimmunity against the CRISPR locus that might be triggered by anti-sense transcripts generated from some type III CRISPR arrays is prevented by inhibition of Cas10 activities relying on base-pairing between target RNA and crRNA positions flanking the spacer region (Foster et al. 2020; Liu and Doudna 2020).
Indirect parameters such as DNA topology also have an impact on several steps of CRISPR-Cas activity (Westra et al. 2012a, b). For example, R-loop formation after hybridisation between the crRNA and the target dsDNA is influenced by the level of DNA negative supercoiling. Furthermore, DNA bending by hosts factors is required to facilitate recognition of the spacer integration site by the adaptation complex (Dorman and Ní Bhriain 2020).
Regulation of CRISPR-cas expression and Cas activity by cellular regulatory networks
Although constitutive transcription from CRISPR and cas promoters has been observed in diverse systems (Mojica et al. 1993; Hale et al. 2008; Lillestøl et al. 2009; Juranek et al. 2012; Crawley et al. 2018), there is also evidence that transcription from some of these promoters is usually repressed. Typically, these regulated loci are expressed only in certain circumstances, notably when the cell is invaded by potential targets, under stress conditions or when the risk of infection is high (Agari et al. 2010; Quax et al. 2013a; Fusco et al. 2015; León-Sobrino et al. 2016; Patterson et al. 2016; Høyland-Kroghsbo et al. 2017; Yang et al. 2020).
Diverse regulatory proteins and RNAs affecting transcription of CRISPR-Cas components have been identified in bacteria and archaea (reviewed in Patterson et al. 2017). As a reflection of the divergent evolutionary paths on the regulation of CRISPR-Cas expression that each prokaryote can adopt, a given regulatory factor (i.e. cyclic AMP receptor protein) may either repress or activate promoters of homologous cas genes depending on the microorganism (Shinkai et al. 2007; Yang et al. 2014). Moreover, signalling mechanisms (e.g. cellular metabolic sensors, stress-responsive two-component systems, quorum sensing) are involved in CRISPR-Cas regulation. For example, it was shown that quorum sensing signals activate cas gene expression in I-E, I-F and III-A CRISPR-Cas systems of P. aeruginosa and Serratia species (Patterson et al. 2016; Høyland-Kroghsbo et al. 2017). Conceivably, many other bacteria might use this strategy to induce CRISPR immunity at high cell density when the risk of infection increases.
Furthermore, the translation efficiency of cas mRNAs, mainly determined by codon usage biases, has also been related to CRISPR-Cas effective functioning (Quax et al. 2013b), and a CRISPR repeat-binding protein was shown to facilitate transcription of a CRISPR locus (Deng et al. 2012). CRISPR-Cas interference can also be stimulated by Cas protein stabilisation (Yosef et al. 2011), and suppression of both interference and adaptation by cas mRNA-binding regulatory proteins has recently been reported for several CRISPR-Cas types (Campa et al. 2021).
The I-E systems from E. coli and Salmonella enterica strains have been studied in detail, providing an overview of the complexity that the regulation of CRISPR-cas loci may require, involving multiple, complementary and alternative dose-dependent factors. The two systems are tightly regulated at the transcription level by an elaborate regulatory network that involves several transcription factors and nucleoid-associated proteins. Besides, anti-sense RNAs detected in the cas loci of the two species might also be implicated in the regulation of Cas expression. In E. coli, Cas3 protein is stabilised by a chaperon protein induced upon phage infection (Yosef et al. 2011) and the histone-like nucleoid-structuring protein H-NS represses transcription from all cas and CRISPR promoters (Pul et al. 2010; Pougach et al. 2010). As H-NS-mediated silencing is achieved by its cooperative spreading on the promoter regions, DNA topology can have an impact on the activity of the system (Liu et al. 2010). Moreover, transcription from divergent promoters located in one of the intergenic regions of the I-E cas locus of E. coli, where H-NS binds (Pul et al. 2010), is expected to generate a local domain of high negative supercoiling (Mojica and Higgins 1996), therefore facilitating H-NS association and subsequent transcription inhibition. Related to this, H-NS-mediated gene silencing is frequently linked to changes in DNA secondary structure (Mojica and Higgins 1997; Winardhi et al. 2015).
Overexpression of the H-NS antagonist LeuO, a LysR-Type regulator, relieves repression of the I-E Cascade operon in E. coli (Mojica and Díez-Villaseñor 2010; Westra et al. 2010). Based on the preferential binding of H-NS to AT-rich DNA, it has been proposed that H-NS silencing of the cas loci could also be mitigated (Pul et al. 2010; Westra et al. 2010) upon infection by viruses or plasmids with high A-T content (Rocha and Danchin 2002) which would sequester part of the H-NS pool (Doyle et al. 2007; Dillon et al. 2010). The global regulator CRP (cAMP receptor protein) competes with LeuO for binding to the Cascade promoter, preventing LeuO-mediated activation (Yang et al. 2014). However, both strong activation and no significant effect of CRP on the cas3 promoter have been reported, which has tentatively been related to the different growth phases of the E. coli cultures assayed in the two studies (Yang et al. 2014, 2020). Similarly, two recent publications (Mitić et al. 2020; Sun et al. 2020a) documented contradictory results showing that when the gene encoding the H-NS paralog StpA was inactivated or deleted in distinct genetic backgrounds, the cas operon transcription was either increased (in an hns cas1 double mutant) or reduced (hns null mutant). Nevertheless, overexpression of StpA suppressed transcription in both cases. These inconsistencies could be due to the experimental specificities of each study. Meanwhile, in addition to silencing by H-NS and positive regulation by LeuO, transcription of the CRISPR-Cas system from S. enterica serovar Typhi is repressed by the leucine-responsive regulatory protein LRP, and, also in contrast with E. coli, CRP does not participate in its transcriptional control (Medina-Aparicio et al. 2011, 2017).
CRISPR-Cas self-control
There are appealing examples of CRISPR-Cas control executed by canonical Cas and CRISPR arrays, as well as by transcriptional regulators associated with these loci.
In addition to the I-E system, many E. coli strains harbour components of an I-F CRISPR-Cas system (Díez-Villaseñor et al. 2010). However, most evolutionary lineages of E. coli have lost all I-F cas genes and only a small CRISPR array remains. Interestingly, the best matches with the spacers of these orphan arrays correspond almost invariably to sequences of I-F cas genes found in related strains. This observation suggested that the acquisition of cas-targeting spacers might have been responsible for the loss of these genes. Accordingly, it was shown that native orphan arrays can elicit interference against plasmids carrying a complete set of I-F cas genes (Almendros et al. 2016). Therefore, it was proposed that Cas proteins were guided by the constitutively expressed orphan arrays against the targeted cas genes. Moreover, this targeting resulted in the primed acquisition of plasmid-derived spacers that further boosted the degradation of the plasmid. Thus, these orphan arrays behave as a natural anti-cas mechanism that efficiently prevents the establishment in the cell of a cognate CRISPR-Cas system and promotes the destruction of the carrier genetic element.
Within the framework of CRISPR-related regulatory proteins, it has been recently shown that the MM_0565 protein associated with the subtype I-B CRISPR-cas locus of Methanosarcina mazei Gö1 binds to the leader of its I-B CRISPR array and a similar leader region in another CRISPR-Cas system (III-C subtype) present in the genome (Ulbricht et al. 2020). However, expression of the CRISPR-cas loci is not affected by overexpression of this protein, suggesting that it might play a post-transcriptional function influencing spacer integration through direct recruitment of the adaptation complex or affecting the structure of the leader, similarly to the role played by IHF regulatory protein in the I-E CRISPR-Cas system from E. coli (Nuñez et al. 2016).
The I-C system of Myxococcus xanthus offers another example of autoregulation. Its cas locus encodes eight proteins, including DevT (Cas8c), DevR (Cas7) and DevS (Cas5). It has been proposed that DevTRS form a Cascade-like subcomplex that negatively autoregulates the dev transcript (Rajagopalan and Kroos 2017). Even though the mechanism of transcription regulation by DevTRS is unknown, the canonical CRISPR-Cas interference, based on crRNA-guided target-cleavage, is not involved as CRISPR spacers matching the target seem to be absent, and activity of Cas nucleases (Cas3 and Cas6) is not required for this feedback regulation. However, neither non-crRNA guiding nor recruitment of non-Cas nucleases to the RNA or DNA target can be ruled out.
Likewise, a recent work (Workman et al. 2021) demonstrated self-regulation of a CRISPR-Cas system from S. pyogenes. In some subtype II-A systems, an extended tracrRNA (tracr-L) not found in other CRISPR-Cas subtypes is transcribed upstream of the tracrRNA promoter (Deltcheva et al. 2011). Workman and colleagues showed that the tracr-L of S. pyogenes guides Cas9 to repress the promoter of its cas operon. Downstream of this cas promoter, adjacent to the canonical 2-nt PAM of the CRISPR-Cas system, there is an 11-nt sequence complementary to the 5′ end of tracr-L. Such a short match still grants efficient binding of the Cas9:tracr-L repressor complex to the target site within the cas promoter. However, target cleavage appears to be prevented (Workman et al. 2021).
On the contrary, it has been reported that the subtype I-A associated protein Csa3a from S. islandicus activates the expression of its CRISPR adaptation module (Liu et al. 2015, 2017; León-Sobrino et al. 2016). The Csa3b protein encoded in the same locus, on the one hand, represses expression of the I-A interference genes (He et al. 2017), whereas on the other acts as a transcriptional activator of the two III-B effector complex gene cassettes present in the genome (Ye et al. 2020).
How do invasive mobile genetic elements escape from CRISPR-Cas immunity?
Prokaryotic iMGEs have developed diverse strategies to evade CRISPR-Cas, which affect almost every CRISPR-Cas component (recently reviewed in Malone et al. 2021).
Viruses frequently escape CRISPR-defence after deletions (Pyenson et al. 2017; Watson et al. 2019), rearrangements or point mutations in the regions of their genome targeted by CRISPR spacers (Andersson and Banfield 2008). These changes may result in partial or complete removal of the target, mismatches between the crRNA and the target or disruption of the PAM recognised by the effector Cas proteins. Consequently, although effective cleavage may be temporarily avoided, these point mutations could still prompt primed spacer acquisition by some CRISPR-Cas systems, reassembling effective immunity (Semenova et al. 2011; Fineran et al. 2014; Jackson et al. 2019).
Otherwise, DNA chemical modifications of the viral genome can hamper the recognition of sequences targeted by the CRISPR-Cas machinery, although this has only been shown for one phage and host system (Bryson et al. 2015; Vlot et al. 2018).
Viral proteins account for other strategies of counter-defence. Some plasmids and viruses carry genes encoding H-NS homologues (Skennerton et al. 2011; Shintani et al. 2015). It has been proposed that these proteins might repress CRISPR-cas transcription as H-NS and StpA do in the I-E systems of enterobacteria (Dorman and Ní Bhriain 2020). Similarly, it is conceivable that genetic elements targeted by CRISPR-Cas systems relying on activation by quorum sensing could have deployed quorum quenching strategies (Mion et al. 2019). Actually, it has been recently demonstrated that a P. aeruginosa phage encodes a lactonase protein that disrupts the communication pathway responsible for the activation of the CRISPR-Cas systems in the host by inhibiting the receptor of the autoinducer molecule (Shah et al. 2021). In addition, manipulation of the host bacterial quorum sensing pathway has been reported for other viruses (Duddy and Bassleri 2021), suggesting that this might be a procedure frequently used by viruses for CRISPR evasion.
Bondy-Denomy and coworkers discovered in 2013 a CRISPR-Cas evasion strategy based on the so-called anti-CRISPR (Acr) proteins (Bondy-Denomy et al. 2013). Acr encoding genes found in prophages of P. aeruginosa were able to counteract the immunity provided by the host I-F CRISPR-Cas system. Since then, the collection of Acr proteins identified has widely extended, as has an interest in this line of research, due to the potential of Acrs in various CRISPR-Cas technologies (Liu et al. 2020b).
Acr proteins suppress CRISPR-Cas immunity mainly through two mechanisms: obstruction of DNA binding or inhibition of target cleavage (i.e. the surveillance complex binds to the DNA target site, but cleavage is not achieved) (Pawluk et al. 2014; Yang and Patel 2017; Marino et al. 2018; Watters et al. 2018; Knott et al. 2019b; Liu et al. 2019b; Bhoobalan-Chitty et al. 2019; Yin et al. 2019; Lin et al. 2020) In type I CRISPR-Cas systems, some Acrs block target cleavage by recruiting the effector nuclease Cas3 (Wang et al. 2016). Acr can also thwart target binding by type I surveillance complexes via direct interaction with Cascade or via DNA mimicry (i.e. Acr mimics dsDNA). In the first case, upon binding to Cascade, Acr triggers conformational changes that sterically occlude the DNA binding sites (Bondy-Denomy et al. 2015; Maxwell et al. 2016). In the second case, the Acr protein adopts a conformation resembling dsDNA and binds to the target site in the Cascade:crRNA complex instead of the target DNA (Guo et al. 2017; Peng et al. 2017). Acr proteins against type III systems have also been discovered that affect cOAs signalling (Peng et al. 2020; Athukoralage et al. 2020).
In class 2, most Acr proteins inhibit target binding to the surveillance complex using mechanisms equivalent to those exhibited by Acr acting against type I systems. For example, regarding the prevention of target cleavage, it has been observed that some Acrs bind catalytic residues of Cas9 blocking its nuclease activity (Pawluk et al. 2016a, b; Wang et al. 2016; Harrington et al. 2017). However, strategies, leading to (i) dimerisation of the effector protein (thus resulting in the reduction of dsDNA binding), (ii) cleavage of the crRNA spacer sequence in the surveillance complex and (iii) acetylation of specific residues of the Cas protein involved in PAM recognition, have been reported for type II and V systems as well (Marino et al. 2018; Dong et al. 2019; Knott et al. 2019a, b; Zhu et al. 2019). Although Acrs that inhibit the interference activity of type VI systems have been identified, the molecular mechanism involved remains to be elucidated (Smargon et al. 2017; Lin et al. 2020; Peng et al. 2020).
CRISPR and Cas elements encoded in viruses may represent another way to interfere with the host CRISPR-Cas activity (see the “CRISPR/Cas in prokaryotic viruses” section). For instance, regarding Cas proteins, many bacterial and archaeal viruses encode Cas4-like proteins that are thought to participate in counter-CRISPR mechanisms. Their proximity to Cas-inhibiting (i.e. acr) genes supports this hypothesis (Hooton and Connerton 2015; Hooton et al. 2016).
Lastly, Jumbo phages deploy a refined counter-CRISPR mechanism. During infection of the host bacterium, a peculiar nucleus-like structure is formed within which phage DNA replication and transcription takes place. Thus, its DNA is protected from degradation by CRISPR-Cas and restriction-modification systems (Modell et al. 2020; Malone et al. 2020).
Conclusion
Invasive mobile genetic elements put intense selective pressure on bacteria and archaea. This challenge has resulted in numerous prokaryotic defence strategies, notably in the CRISPR-Cas adaptive immune systems. However, the fitness costs associated with CRISPR-Cas may result in stringent regulation of their expression, protein inactivation or, ultimately, loss of the complete system. In addition to immunity, many CRISPR-Cas systems play collateral or alternative roles. These side functions take advantage of the guiding capability of CRISPR RNAs and the multiple roles of Cas proteins in nucleic acid metabolism. Arguably, the associated benefits might account for the maintenance of the CRISPR-Cas systems even when foreign threats could be faced using less-costly protection mechanisms.
The discovery of the innate immunity provided by restriction-modification systems led to the birth of genetic engineering, allowing the cutting and pasting of DNA fragments in vitro almost at will. At present, the versatility of CRISPR-Cas adaptive immune systems has been translated into an arsenal of molecular biology tools that have revolutionised biotechnology, medicine and agriculture. Unfortunately, the research and understanding of the CRISPR-Cas systems that made this scientific revolution possible are often overlooked due to the more attention-grabbing advances in biotechnology and medical applications. Nevertheless, as we have described above, CRISPR-Cas biology is no less impressive. On the contrary, more than three decades after the discovery of the enigmatic repeats in prokaryotes, CRISPR research continues to shed light on the intricacies and complexity of this fascinating system.
Availability of data and material
Not applicable.
Code availability
Not applicable.
References
Abudayyeh OO, Gootenberg JS, Konermann S et al (2016) C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353:aaf5573. https://doi.org/10.1126/science.aaf5573
Agari Y, Sakamoto K, Tamakoshi M et al (2010) Transcription profile of Thermus thermophilus CRISPR systems after phage Infection. J Mol Biol 395:270–281. https://doi.org/10.1016/j.jmb.2009.10.057
Aklujkar M, Lovley DR (2010) Interference with histidyl-tRNA synthetase by a CRISPR spacer sequence as a factor in the evolution of Pelobacter carbinolicus. BMC Evol Biol 10:230. https://doi.org/10.1186/1471-2148-10-230
Almendros C, Guzmán NM, Díez-Villaseñor C et al (2012) Target motifs affecting natural immunity by a constitutive CRISPR-Cas system in Escherichia coli. PLoS ONE 7:e50797. https://doi.org/10.1371/journal.pone.0050797
Almendros C, Mojica FJM (2015) Exploring CRISPR interference by transformation with plasmid mixtures: identification of target interference motifs in Escherichia coli. Methods Mol Biol 1311:161–170. https://doi.org/10.1007/978-1-4939-2687-9
Almendros C, Guzmán NM, García-Martínez J, Mojica FJM (2016) Anti-cas spacers in orphan CRISPR4 arrays prevent uptake of active CRISPR–Cas I-F systems. Nat Microbiol 1:16081. https://doi.org/10.1038/nmicrobiol.2016.81
Almendros C, Nobrega FL, McKenzie RE, Brouns SJJ (2019) Cas4–Cas1 fusions drive efficient PAM selection and control CRISPR adaptation. Nucleic Acids Res 47:5223–5230. https://doi.org/10.1093/nar/gkz217
Al-Shayeb B, Sachdeva R, Chen LX et al (2020) Clades of huge phages from across Earth’s ecosystems. Nature 578:425–431. https://doi.org/10.1038/s41586-020-2007-4
Andersson AF, Banfield JF (2008) Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320:1047–1050. https://doi.org/10.1126/science.1157358
Arslan Z, Wurm R, Brener O et al (2013) Double-strand DNA end-binding and sliding of the toroidal CRISPR-associated protein Csn2. Nucleic Acids Res 41:6347–6359. https://doi.org/10.1093/nar/gkt315
Artamonova D, Karneyeva K, Medvedeva S et al (2020) Spacer acquisition by type III CRISPR–Cas system during bacteriophage infection of Thermus thermophilus. Nucleic Acids Res 48:9787–9803. https://doi.org/10.1093/nar/gkaa685
Athukoralage JS, McMahon SA, Zhang C et al (2020) An anti-CRISPR viral ring nuclease subverts type III CRISPR immunity. Nature 577:572–575. https://doi.org/10.1038/s41586-019-1909-5
Babu M, Beloglazova N, Flick R et al (2011) A dual function of the CRISPR-Cas system in bacterial antivirus immunity and DNA repair. Mol Microbiol 79:484–502. https://doi.org/10.1111/j.1365-2958.2010.07465.x
Barrangou R, Fremaux C, Deveau H et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712. https://doi.org/10.1126/science.1138140
Behler J, Sharma K, Reimann V et al (2018) The host-encoded RNase e endonuclease as the crRNA maturation enzyme in a CRISPR-Cas subtype III-Bv system. Nat Microbiol 3:367–377. https://doi.org/10.1038/s41564-017-0103-5
Behler J, Hess WR (2020) Approaches to study CRISPR RNA biogenesis and the key players involved. Methods 172:12–26
Bernheim A, Calvo-Villamañán A, Basier C et al (2017) Inhibition of NHEJ repair by type II-A CRISPR-Cas systems in bacteria. Nat Commun 8:2094. https://doi.org/10.1038/s41467-017-02350-1
Bikard D, Jiang W, Samai P et al (2013) Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res 41:7429–7437. https://doi.org/10.1093/nar/gkt520
Bikard D, Euler CW, Jiang W et al (2014) Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat Biotechnol 32:1146–1150. https://doi.org/10.1038/nbt.3043
Bhoobalan-Chitty Y, Johansen TB, di Cianni N, Peng X (2019) Inhibition of type III CRISPR-Cas immunity by an archaeal virus-encoded anti-CRISPR protein. Cell 179:448–458. https://doi.org/10.1016/j.cell.2019.09.003
Bolotin A, Quinquis B, Sorokin A, Dusko Ehrlich S (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551–2561. https://doi.org/10.1099/mic.0.28048-0
Bondy-Denomy J, Pawluk A, Maxwell KL, Davidson AR (2013) Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493:429–432. https://doi.org/10.1038/nature11723
Bondy-Denomy J, Garcia B, Strum S et al (2015) Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526:136–139. https://doi.org/10.1038/nature15254
Bozic B, Repac J, Djordjevic M (2019) Endogenous gene regulation as a predicted main function of type I-E CRISPR/Cas system in E. coli. Molecules 24:784. https://doi.org/10.3390/molecules24040784
Brouns SJJ, Jore MM, Lundgren M et al (2008) Small CRISPR RNAs guide antiviral defense in prokaryotes. Science (new York, NY) 321:960–964. https://doi.org/10.1126/science.1159689
Bryson AL, Hwang Y, Sherrill-Mix S et al (2015) Covalent Modification of Bacteriophage T4 DNA Inhibits CRISPR-Cas9. mBio 6:e00648. https://doi.org/10.1128/mBio.00648-15
Bult CJ, White O, Olsen GJ et al (1996) Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273:1058–1073. https://doi.org/10.1126/science.273.5278.1058
Burstein D, Harrington LB, Strutt SC et al (2017) New CRISPR-Cas systems from uncultivated microbes. Nature 542:237–241. https://doi.org/10.1038/nature21059
Cady KC, O’Toole GA (2011) Non-identity-mediated CRISPR-bacteriophage interaction mediated via the Csy and Cas3 proteins. J Bacteriol 193:3433–3445. https://doi.org/10.1128/JB.01411-10
Cady KC, Bondy-Denomy J, Heussler GE et al (2012) The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J Bacteriol 194:5728–5738. https://doi.org/10.1128/JB.01184-12
Campa AR, Smith LM, Hampton HG et al (2021) The Rsm (Csr) post-transcriptional regulatory pathway coordinately controls multiple CRISPR–Cas immune systems. Nucleic Acids Res 388:539–547. https://doi.org/10.1093/nar/gkab704
Cañez C, Selle K, Goh YJ, Barrangou R (2019) Outcomes and characterization of chromosomal self-targeting by native CRISPR-Cas systems in Streptococcus thermophilus. FEMS Microbiology Letters 366:fnz105. https://doi.org/10.1093/femsle/fnz105
Carte J, Wang R, Li H et al (2008) Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev 22:3489–3496. https://doi.org/10.1101/gad.1742908
Carte J, Pfister NT, Compton MM et al (2010) Binding and cleavage of CRISPR RNA by Cas6. RNA 16:2181–2188. https://doi.org/10.1261/rna.2230110
Chavez M, Qi LS (2019) Site-programmable transposition: shifting the paradigm for CRISPR-Cas systems. Mol Cell 75:206–208. https://doi.org/10.1016/j.molcel.2019.07.004
Chen S, Liu H, Liang W et al (2019) Insertion sequences in the CRISPR-Cas system regulate horizontal antimicrobial resistance gene transfer in Shigella strains. Int J Antimicrob Agents 53:109–115. https://doi.org/10.1016/j.ijantimicag.2018.09.020
Chou-Zheng L, Hatoum-Aslan A (2019) A type III-A CRISPR-Cas system employs degradosome nucleases to ensure robust immunity. eLife 8:e45393. https://doi.org/10.7554/eLife.45393
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823. https://doi.org/10.1126/science.1231143
Crawley AB, Henriksen ED, Stout E et al (2018) Characterizing the activity of abundant, diverse and active CRISPR-Cas systems in lactobacilli. Sci Rep 8:11544. https://doi.org/10.1038/s41598-018-29746-3
Crowley VM, Catching A, Taylor HN et al (2019) A type IV-A CRISPR-Cas system in Pseudomonas aeruginosa mediates RNA-guided plasmid interference in vivo. CRISPR J 2:434–440. https://doi.org/10.1089/crispr.2019.0048
Cui L, Bikard D (2016) Consequences of Cas9 cleavage in the chromosome of Escherichia coli. Nucleic Acids Res 44:4243–4251. https://doi.org/10.1093/nar/gkw223
Cui L, Wang X, Huang D et al (2020) CRISPR-cas3 of Salmonella upregulates bacterial biofilm formation and virulence to host cells by targeting quorum-sensing systems. Pathogens 9:53. https://doi.org/10.3390/pathogens9010053
Datsenko KA, Pougach K, Tikhonov A et al (2012) Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nat Commun 3:945. https://doi.org/10.1038/ncomms1937
Deem MW (2020) CRISPR recognizes as many phage types as possible without overwhelming the cas machinery. Proc Natl Acad Sci USA 117:7550–7552. https://doi.org/10.1073/pnas.2002746117
Deltcheva E, Chylinski K, Sharma CM et al (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471:602–607. https://doi.org/10.1038/nature09886
Deng L, Kenchappa CS, Peng X et al (2012) Modulation of CRISPR locus transcription by the repeat-binding protein Cbp1 in Sulfolobus. Nucleic Acids Res 40:2470–2480. https://doi.org/10.1093/nar/gkr1111
Deveau H, Barrangou R, Garneau JE et al (2008) Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J Bacteriol 190:1390–1400. https://doi.org/10.1128/JB.01412-07
Díez-Villaseñor C, Almendros C, García-Martínez J, Mojica FJM (2010) Diversity of CRISPR loci in Escherichia coli. Microbiology 156:1351–1361. https://doi.org/10.1099/mic.0.036046-0
Díez-Villaseñor C, Guzmán NM, Almendros C et al (2013) CRISPR-spacer integration reporter plasmids reveal distinct genuine acquisition specificities among CRISPR-Cas I-E variants of Escherichia coli. RNA Biol 10:792–802. https://doi.org/10.4161/rna.24023
Dillon SC, Cameron ADS, Hokamp K et al (2010) Genome-wide analysis of the H-NS and Sfh regulatory networks in Salmonella Typhimurium identifies a plasmid-encoded transcription silencing mechanism. Mol Microbiol 76:1250–1265. https://doi.org/10.1111/j.1365-2958.2010.07173.x
Dimitriu T, Ashby B, Westra ER (2019) Transposition: a CRISPR way to get around. Curr Biol 29:R886–R889. https://doi.org/10.1016/j.cub.2019.08.010
Dong L, Guan X, Li N et al (2019) An anti-CRISPR protein disables type V Cas12a by acetylation. Nat Struct Mol Biol 26:308–314. https://doi.org/10.1038/s41594-019-0206-1
Dorman CJ, Ní Bhriain N (2020) CRISPR-Cas, DNA supercoiling, and nucleoid-associated proteins. Trends Microbiol 28:19–27. https://doi.org/10.1016/j.tim.2019.08.004
Doyle M, Fookes M, Ivens A et al (2007) An H-NS-like stealth protein aids horizontal DNA transmission in bacteria. Science 315:251–252. https://doi.org/10.1126/science.1137550
Drabavicius G, Sinkunas T, Silanskas A et al (2018) DnaQ exonuclease-like domain of Cas2 promotes spacer integration in a type I-E CRISPR-Cas system. EMBO Rep 19:e45543. https://doi.org/10.15252/embr.201745543
Duddy OP, Bassleri BL (2021) Quorum sensing across bacterial and viral domains. PLoS Pathog 17:e1009074. https://doi.org/10.1371/journal.ppat.1009074
Dugar G, Herbig A, Förstner KU et al (2013) High-resolution transcriptome maps reveal strain-specific regulatory features of multiple Campylobacter jejuni isolates. PLoS Genet 9:e1003495. https://doi.org/10.1371/journal.pgen.1003495
Dugar G, Leenay RT, Eisenbart SK et al (2018) CRISPR RNA-dependent binding and cleavage of endogenous RNAs by the Campylobacter jejuni Cas9. Mol Cell 69:893–905. https://doi.org/10.1016/j.molcel.2018.01.032
Dwarakanath S, Brenzinger S, Gleditzsch D et al (2015) Interference activity of a minimal type I CRISPR–Cas system from Shewanella putrefaciens. Nucleic Acids Res 43:8913–8923. https://doi.org/10.1093/nar/gkv882
East-Seletsky A, O’Connell MR, Knight SC et al (2016) Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538:270–273. https://doi.org/10.1038/nature19802
East-Seletsky A, O’Connell MR, Burstein D et al (2017) RNA targeting by functionally orthogonal type VI-A CRISPR-Cas enzymes. Mol Cell 66:373–383. https://doi.org/10.1016/j.molcel.2017.04.008
Edgar R, Qimron U (2010) The Escherichia coli CRISPR system protects from lysogenization, lysogens, and prophage induction. J Bacteriol 192:6291–6294. https://doi.org/10.1128/JB.00644-10
Erdmann S, le Moine BS, Garrett RA (2014) Inter-viral conflicts that exploit host CRISPR immune systems of Sulfolobus. Mol Microbiol 91:900–917. https://doi.org/10.1111/mmi.12503
Estrella MA, Kuo FT, Bailey S (2016) RNA-activated DNA cleavage by the type III-B CRISPR–Cas effector complex. Genes Dev 30:460–470. https://doi.org/10.1101/gad.273722.115
Faure G, Makarova KS, Koonin Ev (2019a) CRISPR–Cas: complex functional networks and multiple roles beyond adaptive immunity. J Mol Biol 431:3–20. https://doi.org/10.1016/j.jmb.2018.08.030
Faure G, Shmakov SA, Yan WX et al (2019b) CRISPR–Cas in mobile genetic elements: counter-defence and beyond. Nat Rev Microbiol 17:513–525. https://doi.org/10.1038/s41579-019-0204-7
Fineran PC, Gerritzen MJH, Suárez-Diez M et al (2014) Degenerate target sites mediate rapid primed CRISPR adaptation. Proc Natl Acad Sci USA 111:E1629–E1638. https://doi.org/10.1073/pnas.1400071111
Foster K, Grüschow S, Bailey S et al (2020) Regulation of the RNA and DNA nuclease activities required for Pyrococcus furiosus Type III-B CRISPR–Cas immunity. Nucleic Acids Res 48:4418–4434. https://doi.org/10.1093/nar/gkaa176
Fusco S, Liguori R, Limauro D et al (2015) Transcriptome analysis of Sulfolobus solfataricus infected with two related fuselloviruses reveals novel insights into the regulation of CRISPR-Cas system. Biochimie 118:322–332. https://doi.org/10.1016/j.biochi.2015.04.006
Gao NJ, Al-Bassam MM, Poudel S et al (2019) Functional and proteomic analysis of Streptococcus pyogenes virulence upon loss of its native Cas9 nuclease. Front Microbiol 10:1967. https://doi.org/10.3389/fmicb.2019.01967
García-Gutiérrez E, Almendros C, Mojica FJM et al (2015) CRISPR content correlates with the pathogenic potential of Escherichia coli. PLoS ONE 10:e0131935. https://doi.org/10.1371/journal.pone.0131935
Garneau JE, Dupuis M-È, Villion M et al (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468:67–71. https://doi.org/10.1038/nature09523
Garside EL, Schellenberg MJ, Gesner EM et al (2012) Cas5d processes pre-crRNA and is a member of a larger family of CRISPR RNA endonucleases. RNA 18:2020–2028. https://doi.org/10.1261/rna.033100.112
Gasiunas G, Barrangou R, Horvath P, Siksnys V (2012) Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci USA 109:E2579–E2586. https://doi.org/10.1073/pnas.1208507109
Gasiunas G, Young JK, Karvelis T et al (2020) A catalogue of biochemically diverse CRISPR-Cas9 orthologs. Nat Commun 11:5512. https://doi.org/10.1038/s41467-020-19344-1
Gesner EM, Schellenberg MJ, Garside EL et al (2011) Recognition and maturation of effector RNAs in a CRISPR interference pathway. Nat Struct Mol Biol 18:688–692. https://doi.org/10.1038/nsmb.2042
Gleditzsch D, Pausch P, Müller-Esparza H et al (2019) PAM identification by CRISPR-Cas effector complexes: diversified mechanisms and structures. RNA Biol 16:504–517. https://doi.org/10.1080/15476286.2018.1504546
Gootenberg JS, Abudayyeh OO, Lee JW et al (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356:438–442. https://doi.org/10.1126/science.aam932
González-Delgado A, Mestre MR, Martínez-Abarca F, Toro N (2019) Spacer acquisition from RNA mediated by a natural reverse transcriptase-Cas1 fusion protein associated with a type III-D CRISPR-Cas system in Vibrio vulnificus. Nucleic Acids Res 47:10202–10211. https://doi.org/10.1093/nar/gkz746
Gunderson FF, Cianciotto NP (2013) The CRISPR-associated gene cas2 of Legionella pneumophila is required for intracellular infection of amoebae. mBio 4:e00074-13. https://doi.org/10.1128/mBio.00074-13
Guo TW, Bartesaghi A, Yang H et al (2017) Cryo-EM structures reveal mechanism and inhibition of DNA targeting by a CRISPR-Cas surveillance complex. Cell 171:414–426. https://doi.org/10.1016/j.cell.2017.09.006
Haft DH, Selengut J, Mongodin EF, Nelson KE (2005) A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 1:e60. https://doi.org/10.1371/journal.pcbi.0010060
Hale C, Kleppe K, Terns RM, Terns MP (2008) Prokaryotic silencing (psi)RNAs in Pyrococcus furiosus. RNA 14:2572–2579. https://doi.org/10.1261/rna.1246808
Hale CR, Zhao P, Olson S et al (2009) RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 139:945–956. https://doi.org/10.1016/j.cell.2009.07.040
Hale CR, Majumdar S, Elmore J et al (2012) Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol Cell 45:292–302. https://doi.org/10.1016/j.molcel.2011.10.023
Harrington LB, Doxzen KW, Ma E et al (2017) A broad-spectrum inhibitor of CRISPR-Cas9. Cell 170:1224–1233. https://doi.org/10.1016/j.cell.2017.07.037
Harrington LB, Burstein D, Chen JS et al (2018) Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362:839–842. https://doi.org/10.1126/science.aav4294
Harrington LB, Ma E, Chen JS et al (2020) A scoutRNA is required for some Type V CRISPR-Cas systems. Mol Cell 79:416–424. https://doi.org/10.1016/j.molcel.2020.06.022
Hatoum-Aslan A, Maniv I, Marraffini LA (2011) Mature clustered, regularly interspaced, short palindromic repeats RNA (crRNA) length is measured by a ruler mechanism anchored at the precursor processing site. Proc Natl Acad Sci USA 108:21218–21222. https://doi.org/10.1073/pnas.1112832108
Haurwitz RE, Jinek M, Wiedenheft B et al (2010) Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329:1355–1358. https://doi.org/10.1126/science.1192272
Hayes RP, Xiao Y, Ding F et al (2016) Structural basis for promiscuous PAM recognition in type I-E Cascade from E. coli. Nature 530:499–503. https://doi.org/10.1038/nature16995
He F, Vestergaard G, Peng W et al (2017) CRISPR-Cas type I-A Cascade complex couples viral infection surveillance to host transcriptional regulation in the dependence of Csa3b. Nucleic Acids Res 45:1902–1913. https://doi.org/10.1093/nar/gkw1265
He L, St John James M, Radovcic M et al (2020) Cas3 protein—a review of a multi-tasking machine. Genes 11:208. https://doi.org/10.3390/genes11020208
Heidrich N, Hagmann A, Bauriedl S et al (2019) The CRISPR/Cas system in Neisseria meningitidis affects bacterial adhesion to human nasopharyngeal epithelial cells. RNA Biol 16:390–396. https://doi.org/10.1080/15476286.2018.1486660
Heler R, Samai P, Modell JW et al (2015) Cas9 specifies functional viral targets during CRISPR–Cas adaptation. Nature 519:199–202. https://doi.org/10.1038/nature14245
Hermans PWM, van Soolingen D, Bik EM et al (1991) Insertion element IS987 from Mycobacterium bovis BCG is located in a hot-spot integration region for insertion elements in Mycobacterium tuberculosis complex strains. Infect Immun 59:2695–2705. https://doi.org/10.1128/iai.59.8.2695-2705.1991
Heussler GE, Cady KC, Koeppen K et al (2015) Clustered regularly interspaced short palindromic repeat-dependent, biofilm-specific death of Pseudomonas aeruginosa mediated by increased expression of phage-related genes. mBio 6:e00129-15. https://doi.org/10.1128/mBio.00129-15
Hille F, Charpentier E (2016) CRISPR-cas: Biology, mechanisms and relevance. Phil Trans R Soc B 371:20150496. https://doi.org/10.1098/rstb.2015.0496
Hochstrasser ML, Taylor DW, Bhat P et al (2014) CasA mediates Cas3-catalyzed target degradation during CRISPR RNA-guided interference. Proc Natl Acad Sci USA 111:6618–6623. https://doi.org/10.1073/pnas.1405079111
Hooton SPT, Connerton IF (2015) Campylobacter jejuni acquire new host-derived CRISPR spacers when in association with bacteriophages harboring a CRISPR-like Cas4 protein. Front Microbiol 5:744. https://doi.org/10.3389/fmicb.2014.00744
Hooton SPT, Brathwaite KJ, Connerton IF (2016) The bacteriophage carrier state of Campylobacter jejuni features changes in host non-coding RNAs and the acquisition of new host-derived CRISPR spacer sequences. Front Microbiol 7:355. https://doi.org/10.3389/fmicb.2016.00355
Horvath P, Coûté-Monvoisin AC, Romero D, a, et al (2009) Comparative analysis of CRISPR loci in lactic acid bacteria genomes. Int J Food Microbiol 131:62–70. https://doi.org/10.1016/j.ijfoodmicro.2008.05.030
Horvath P, Romero DA, Coûté-Monvoisin AC et al (2008) Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol 190:1401–1412. https://doi.org/10.1128/JB.01415-07
Høyland-Kroghsbo NM, Paczkowski J, Mukherjee S et al (2017) Quorum sensing controls the Pseudomonas aeruginosa CRISPR-Cas adaptive immune system. Proc Natl Acad Sci USA 114:131–135. https://doi.org/10.1073/pnas.1617415113
Hudaiberdiev S, Shmakov S, Wolf YI et al (2017) Phylogenomics of Cas4 family nucleases. BMC Evol Biol 17:232. https://doi.org/10.1186/s12862-017-1081-1
Iranzo J, Faure G, Wolf YI, Koonin Ev (2020) Game-theoretical modeling of interviral conflicts mediated by mini-CRISPR arrays. Front Microbiol 11:381. https://doi.org/10.3389/fmicb.2020.00381
Ivančić-Bace I, Cass SD, Wearne SJ, Bolt EL (2015) Different genome stability proteins underpin primed and Naïve adaptation in E. coli CRISPR-Cas immunity. Nucleic Acids Res 43:10821–10830. https://doi.org/10.1093/nar/gkv1213
Jackson RN, Golden SM, van Erp PBGG et al (2014) Structural biology. Crystal structure of the CRISPR RNA-guided surveillance complex from Escherichia coli. Science 345:1473–1479. https://doi.org/10.1126/science.1256328
Jackson RN, van Erp PB, Sternberg SH, Wiedenheft B (2017) Conformational regulation of CRISPR-associated nucleases. Curr Opin Microbiol 37:110–119. https://doi.org/10.1016/j.mib.2017.05.010
Jackson SA, Birkholz N, Malone LM, Fineran PC (2019) Imprecise spacer acquisition generates CRISPR-Cas immune diversity through primed adaptation. Cell Host Microbe 25:250–260. https://doi.org/10.1016/j.chom.2018.12.014
Jansen R, Embden JD, Gaastra W, Schouls L (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43:1565–1575. https://doi.org/10.1046/j.1365-2958.2002.02839.x
Jia N, Mo CY, Wang C et al (2019) Type III-A CRISPR-Cas Csm complexes: assembly, periodic RNA cleavage, DNase activity regulation, and autoimmunity. Mol Cell 73:264–277. https://doi.org/10.1016/j.molcel.2018.11.007
Jinek M, Chylinski K, Fonfara I et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821. https://doi.org/10.1126/science.1225829
Jore MM, Lundgren M, van Duijn E et al (2011) Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nat Struct Mol Biol 18:529–536. https://doi.org/10.1038/nsmb.2019
Juranek S, Eban T, Altuvia Y et al (2012) A genome-wide view of the expression and processing patterns of Thermus thermophilus HB8 CRISPR RNAs. RNA 18:783–794. https://doi.org/10.1261/rna.031468.111
Kamruzzaman M, Iredell JR (2020) CRISPR-Cas system in antibiotic resistance plasmids in Klebsiella pneumoniae. Front Microbiol 10:2934. https://doi.org/10.3389/fmicb.2019.02934
Karvelis T, Bigelyte G, Young JK et al (2020) PAM recognition by miniature CRISPR-Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res 48:5016–5023. https://doi.org/10.1093/nar/gkaa208
Kazlauskiene M, Kostiuk G, Venclovas Č et al (2017) A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357:605–609. https://doi.org/10.1126/science.aao0100
Kieper SN, Almendros C, Behler J et al (2018) Cas4 facilitates PAM-compatible spacer selection during CRISPR adaptation. Cell Rep 22:3377–3384. https://doi.org/10.1016/j.celrep.2018.02.103
Killelea T, Bolt EL (2017) CRISPR-Cas adaptive immunity and the three Rs. Biosci Rep 37:BSR20170297. https://doi.org/10.1042/BSR20160297
Kim S, Loeff L, Colombo S et al (2020) Selective loading and processing of prespacers for precise CRISPR adaptation. Nature 579:141–145. https://doi.org/10.1038/s41586-020-2018-1
Klompe SE, Vo PLH, Halpin-Healy TS, Sternberg SH (2019) Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 571:219–225. https://doi.org/10.1038/s41586-019-1323-z
Knott GJ, Cress BF, Liu JJ et al (2019) Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a. eLife 8:e49110. https://doi.org/10.7554/eLife.49110
Knott GJ, Thornton BW, Lobba MJ et al (2019b) Broad-spectrum enzymatic inhibition of CRISPR-Cas12a. Nat Struct Mol Biol 26:315–321. https://doi.org/10.1038/s41594-019-0208-z
Koonin E, Makarova KS, Zhang F (2017) Diversity, classification and evolution of CRISPR-Cas systems. Curr Opin Microbiol 37:67–78. https://doi.org/10.1016/j.mib.2017.05.008
Lange SJ, Alkhnbashi OS, Rose D et al (2013) CRISPRmap: an automated classification of repeat conservation in prokaryotic adaptive immune systems. Nucleic Acids Res 41:8034–8044. https://doi.org/10.1093/nar/gkt606
Lee H, Dhingra Y, Sashital DG (2019) The Cas4-Cas1-Cas2 complex mediates precise prespacer processing during CRISPR adaptation. eLife 8:44248. https://doi.org/10.7554/eLife.44248
Leenay RT, Beisel CL (2017) Deciphering, communicating, and engineering the CRISPR PAM. J Mol Biol 429:177–191. https://doi.org/10.1016/j.jmb.2016.11.024
Leon LM, Mendoza SD, Bondy-Denomy J (2018) How bacteria control the CRISPR-Cas arsenal. Curr Opin Microbiol 42:87–95. https://doi.org/10.1016/j.mib.2017.11.005
León-Sobrino C, Kot WP, Garrett RA (2016) Transcriptome changes in STSV2-infected Sulfolobus islandicus REY15A undergoing continuous CRISPR spacer acquisition. Mol Microbiol 99:719–728. https://doi.org/10.1111/mmi.13263
Levy A, Goren MG, Yosef I et al (2015) CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520:505–510. https://doi.org/10.1038/nature14302
Li M, Wang R, Xiang H (2014a) Haloarcula hispanica CRISPR authenticates PAM of a target sequence to prime discriminative adaptation. Nucleic Acids Res 42:7226–7235. https://doi.org/10.1093/nar/gku389
Li M, Wang R, Zhao D, Xiang H (2014b) Adaptation of the Haloarcula hispanica CRISPR-Cas system to a purified virus strictly requires a priming process. Nucleic Acids Res 42:2483–2492. https://doi.org/10.1093/nar/gkt1154
Li R, Fang L, Tan S et al (2016) Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity. Cell Res 26:1273–1287. https://doi.org/10.1038/cr.2016.135
Li Y, Pan S, Zhang Y et al (2015) Harnessing Type I and Type III CRISPR-Cas systems for genome editing. Nucleic Acids Res 44:e34. https://doi.org/10.1093/nar/gkv1044
Lillestøl RK, Shah S, a., Brügger K, et al (2009) CRISPR families of the crenarchaeal genus Sulfolobus: bidirectional transcription and dynamic properties. Mol Microbiol 72:259–272. https://doi.org/10.1111/j.1365-2958.2009.06641.x
Lin P, Qin S, Pu Q et al (2020) CRISPR-Cas13 inhibitors block RNA editing in bacteria and mammalian cells. Mol Cell 78:850–861. https://doi.org/10.1016/j.molcel.2020.03.033
Liu Y, Chen H, Kenney LJ, Yan J (2010) A divalent switch drives H-NS/DNA-binding conformations between stiffening and bridging modes. Genes Dev 24:339–344. https://doi.org/10.1101/gad.1883510
Liu T, Li Y, Wang X et al (2015) Transcriptional regulator-mediated activation of adaptation genes triggers CRISPR de novo spacer acquisition. Nucleic Acids Res 43:1044–1055. https://doi.org/10.1093/nar/gku1383
Liu T, Liu Z, Ye Q et al (2017) Coupling transcriptional activation of CRISPR-Cas system and DNA repair genes by Csa3a in Sulfolobus islandicus. Nucleic Acids Res 45:8978–8992. https://doi.org/10.1093/nar/gkx612
Liu JJ, Orlova N, Oakes BL et al (2019a) CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566:218–223. https://doi.org/10.1038/s41586-019-0908-x
Liu L, Yin M, Wang M, Wang Y (2019b) Phage AcrIIA2 DNA mimicry: structural basis of the CRISPR and anti-CRISPR arms race. Mol Cell 73:611–620. https://doi.org/10.1016/j.molcel.2018.11.011
Liu Z, Dong H, Cui Y et al (2020a) Application of different types of CRISPR/Cas-based systems in bacteria. Microb Cell Fact 19:1–14. https://doi.org/10.1186/s12934-020-01431-z
Liu TY, Doudna JA (2020) Chemistry of class 1 CRISPR-Cas effectors: binding, editing, and regulation. J Biol Chem 295:14473–14487. https://doi.org/10.1074/jbc.REV120.007034
Liu Q, Zhang H, Huang X (2020b) Anti-CRISPR proteins targeting the CRISPR-Cas system enrich the toolkit for genetic engineering. FEBS J 287:626–644. https://doi.org/10.1111/febs.15139
Louwen R, Horst-Kreft D, de Boer AG et al (2013) A novel link between Campylobacter jejuni bacteriophage defence, virulence and Guillain-Barré syndrome. Eur J Clin Microbiol Infect Dis 32:207–226. https://doi.org/10.1007/s10096-012-1733-4
Ma K, Cao Q, Luo S et al (2018) cas9 enhances bacterial virulence by repressing the Regr transcriptional regulator in Streptococcus agalactiae. Infect Immun 86:e00552-e617. https://doi.org/10.1128/IAI.00552-17
Maier LK, Stachler AE, Saunders SJ et al (2015) An active immune defense with a minimal CRISPR (clustered regularly interspaced short palindromic repeats) RNA and without the Cas6 protein. J Biol Chem 290:4192–4201. https://doi.org/10.1074/jbc.M114.617506
Makarova KS, Grishin NV, Shabalina SA et al (2006) A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 1:7. https://doi.org/10.1186/1745-6150-1-7
Makarova KS, Haft DH, Barrangou R et al (2011) Evolution and classification of the CRISPR–Cas systems. Nat Rev Microbiol 9:467–477. https://doi.org/10.1038/nrmicro2577
Makarova KS, Wolf YI, Alkhnbashi OS et al (2015) An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol 13:722–736. https://doi.org/10.1038/nrmicro3569
Makarova KS, Wolf YI, Koonin E (2018) Classification and nomenclature of CRISPR-Cas systems: where from here? CRISPR J 1:325–336. https://doi.org/10.1089/crispr.2018.0033
Makarova KS, Wolf YI, Iranzo J et al (2020) Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat Rev Microbiol 18:67–83. https://doi.org/10.1038/s41579-019-0299-x
Mali P, Yang L, Esvelt K et al (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826. https://doi.org/10.1126/science.1232033
Malone LM, Warring SL, Jackson SA et al (2020) A jumbo phage that forms a nucleus-like structure evades CRISPR–Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat Microbiol 5:48–55. https://doi.org/10.1038/s41564-019-0612-5
Malone LM, Birkholz N, Fineran PC (2021) Conquering CRISPR: how phages overcome bacterial adaptive immunity. Curr Opin Biotechnol 68:30–36. https://doi.org/10.1016/j.copbio.2020.09.008
Mandin P, Geissmann T, Cossart P et al (2007) Identification of new noncoding RNAs in Listeria monocytogenes and prediction of mRNA targets. Nucleic Acids Res 35:962–974. https://doi.org/10.1093/nar/gkl1096
Marino ND, Zhang JY, Borges AL et al (2018) Discovery of widespread type I and type V CRISPR-Cas inhibitors. Science 362:240–242. https://doi.org/10.1126/science.aau5174
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845. https://doi.org/10.1126/science.1165771
Marraffini LA, Sontheimer EJ (2010) Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463:568–571. https://doi.org/10.1038/nature08703
Martínez Arbas S, Narayanasamy S, Herold M et al (2021) Roles of bacteriophages, plasmids and CRISPR immunity in microbial community dynamics revealed using time-series integrated meta-omics. Nat Microbiol 6:123–135. https://doi.org/10.1038/s41564-020-00794-8
Maxwell KL, Garcia B, Bondy-Denomy J et al (2016) The solution structure of an anti-CRISPR protein. Nat Commun 7:13134. https://doi.org/10.1038/ncomms13134
McGinn J, Marraffini LA (2019) Molecular mechanisms of CRISPR–Cas spacer acquisition. Nat Rev Microbiol 17:7–12. https://doi.org/10.1038/s41579-018-0071-7
Medina-Aparicio L, Rebollar-FloresGallego-Hernández JEaL et al (2011) The CRISPR/Cas immune system is an operon regulated by LeuO, H-NS, and leucine-responsive regulatory protein in Salmonella enterica serovar Typhi. J Bacteriol 193:2396–2407. https://doi.org/10.1128/JB.01480-10
Medina-Aparicio L, Rebollar-Flores JE, Beltrán-Luviano AA et al (2017) CRISPR-Cas system presents multiple transcriptional units including antisense RNAs that are expressed in minimal medium and upregulated by pH in Salmonella enterica serovar Typhi. Microbiology 163:253–265. https://doi.org/10.1099/mic.0.000414
Medvedeva S, Liu Y, Koonin EV et al (2019) Virus-borne mini-CRISPR arrays are involved in interviral conflicts. Nat Commun 10:5204. https://doi.org/10.1038/s41467-019-13205-2
Meeske AJ, Marraffini LA (2018) RNA guide complementarity prevents self-targeting in Type VI CRISPR systems. Mol Cell 71:791–801. https://doi.org/10.1016/j.molcel.2018.07.013
Mion S, Plener L, Rémy B et al (2019) Lactonase SsoPox modulates CRISPR-Cas expression in Gram-negative proteobacteria using AHL-based quorum sensing systems. Res Microbiol 170:296–299. https://doi.org/10.1016/j.resmic.2019.06.004
Mitić D, Radovčić M, Markulin D, Ivančić-Baće I (2020) StpA represses CRISPR-Cas immunity in H-NS deficient Escherichia coli. Biochimie 174:136–143. https://doi.org/10.1016/j.biochi.2020.04.020
Mo CY, Mathai J, Rostøl JT et al (2021) Type III-A CRISPR immunity promotes mutagenesis of staphylococci. Nature 592:611–615. https://doi.org/10.1038/s41586-021-03440-3
Modell AE, Siriwardena SU, Choudhary A (2020) A Jumbo phage forms a nucleus-like compartment to evade bacterial defense systems. Biochemistry 59:1869–1870. https://doi.org/10.1021/acs.biochem.0c00273
Modell JW, Jiang W, Marraffini LA (2017) CRISPR-Cas systems exploit viral DNA injection to establish and maintain adaptive immunity. Nature 544:101–104. https://doi.org/10.1038/nature21719
Mojica FJM, Juez G, Rodríguez-Valera F (1993) Transcription at different salinities of Haloferax mediterranei sequences adjacent to partially modified PstI sites. Mol Microbiol 9:613–621. https://doi.org/10.1111/j.1365-2958.1993.tb01721.x
Mojica FJM, Higgins CF (1996) Localized domains of DNA supercoiling: topological coupling between promoters. Mol Microbiol 22:919–928. https://doi.org/10.1046/j.1365-2958.1996.01541.x
Mojica FJM, Higgins CF (1997) In vivo supercoiling of plasmid and chromosomal DNA in an Escherichia coli hns mutant. J Bacteriol 179:3528–3533. https://doi.org/10.1128/jb.179.11.3528-3533
Mojica FJM, Díez-Villaseñor C, Soria E, Juez G (2000) Biological significance of a family of regularly spaced repeats in the genomes of archaea, bacteria and mitochondria. Mol Microbiol 36:244–246. https://doi.org/10.1046/j.1365-2958.2000.01838.x
Mojica FJM, Díez-Villaseñor C, García-Martínez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182. https://doi.org/10.1007/s00239-004-0046-3
Mojica FJM, Díez-Villaseñor C, García-Martínez J, Almendros C (2009) Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155:733–740. https://doi.org/10.1099/mic.0.023960-0
Mojica FJM, Díez-Villaseñor C (2010) The on-off switch of CRISPR immunity against phages in Escherichia coli. Mol Microbiol 77:1341–1345. https://doi.org/10.1111/j.1365-2958.2010.07326.x
Mojica FJM, Díez-Villaseñor C (2013) Right of admission reserved, no matter the path. Trends Microbiol 21:446–448. https://doi.org/10.1016/j.tim.2013.06.003
Mojica FJM, Montoliu L (2016) On the origin of CRISPR-Cas technology: from prokaryotes to mammals. Trends Microbiol 24:811–820. https://doi.org/10.1016/j.tim.2016.06.005
Mosterd C, Rousseau GM, Moineau S (2021) A short overview of the CRISPR-Cas adaptation stage. Can J Microbiol 67:1–12. https://doi.org/10.1139/cjm-2020-0212
Müller-Esparza H, Randau L (2017) Commentary: Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity. Front Microbiol 8:319. https://doi.org/10.3389/fmicb.2017.00319
Musharova O, Medvedeva S, Klimuk E et al (2021) Prespacers formed during primed adaptation associate with the Cas1–Cas2 adaptation complex and the Cas3 interference nuclease–helicase. Proc Natl Acad Sci USA 118:e2021291118. https://doi.org/10.1073/pnas.2021291118
Nam KH, Kurinov I, Ke A (2011) Crystal structure of clustered regularly interspaced short palindromic repeats (CRISPR)-associated Csn2 protein revealed Ca2+-dependent double-stranded DNA binding activity. J Biol Chem 286:30759–30768. https://doi.org/10.1074/jbc.M111.256263
Nam KH, Ding F, Haitjema C et al (2012) Double-stranded endonuclease activity in Bacillus halodurans clustered regularly interspaced short palindromic repeats (CRISPR)-associated Cas2 protein. J Biol Chem 287:35943–35952. https://doi.org/10.1074/jbc.M112.382598
Naser I, Hoque MM, Nahid MA et al (2017) Analysis of the CRISPR-Cas system in bacteriophages active on epidemic strains of Vibrio cholerae in Bangladesh. Sci Rep 7:14880. https://doi.org/10.1038/s41598-017-14839-2
Newsom S, Parameshwaran HP, Martin L, Rajan R (2021) The CRISPR-Cas mechanism for adaptive immunity and alternate bacterial functions fuels diverse biotechnologies. Front Cell Infect Microbiol 10:619763. https://doi.org/10.3389/fcimb.2020.619763
Niewoehner O, Garcia-Doval C, Rostøl JT et al (2017) Type III CRISPR-Cas systems produce cyclic oligoadenylate second messengers. Nature 548:543–548. https://doi.org/10.1038/nature23467
Nuñez JK, Kranzusch PJ, Noeske J et al (2014) Cas1-Cas2 complex formation mediates spacer acquisition during CRISPR-Cas adaptive immunity. Nat Struct Mol Biol 21:528–534. https://doi.org/10.1038/nsmb.2820
Nuñez JK, Harrington LB, Kranzusch PJ et al (2015) Foreign DNA capture during CRISPR–Cas adaptive immunity. Nature 527:535–538. https://doi.org/10.1038/nature15760
Nuñez JK, Bai L, Harrington LB et al (2016) CRISPR immunological memory requires a host factor for specificity. Mol Cell 62:824–833. https://doi.org/10.1016/j.molcel.2016.04.027
Nussenzweig PM, McGinn J, Marraffini LA (2019) Cas9 cleavage of viral genomes primes the acquisition of new immunological memories. Cell Host Microbe 26:515–526. https://doi.org/10.1016/j.chom.2019.09.002
Nussenzweig PM, Marraffini LA (2020) Molecular mechanisms of CRISPR-Cas immunity in bacteria. Annu Rev Genet 54:93–120. https://doi.org/10.1146/annurev-genet-022120-112523
O’Connell MR, Oakes BL, Sternberg SH et al (2014) Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 516:263–266. https://doi.org/10.1038/nature13769
Osawa T, Inanaga H, Sato C, Numata T (2015) Crystal structure of the CRISPR-Cas RNA silencing Cmr complex bound to a target analog. Mol Cell 58:418–430. https://doi.org/10.1016/j.molcel.2015.03.018
Özcan A, Pausch P, Linden A et al (2019) Type IV CRISPR RNA processing and effector complex formation in Aromatoleum aromaticum. Nat Microbiol 4:89–96. https://doi.org/10.1038/s41564-018-0274-8
Patterson AG, Jackson SA, Taylor C et al (2016) Quorum sensing controls adaptive immunity through the regulation of multiple CRISPR-Cas systems. Mol Cell 64:1102–1108. https://doi.org/10.1016/j.molcel.2016.11.012
Patterson AG, Yevstigneyeva MS, Peter C (2017) Regulation of CRISPR–Cas adaptive immune systems. Curr Opin Microbiol 37:1–7. https://doi.org/10.1016/j.mib.2017.02.004
Pausch P, Al-Shayeb B, Bisom-Rapp E et al (2020) CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369:333–337. https://doi.org/10.1126/science.abb1400
Pawluk A, Bondy-Denomy J, Cheung VHW et al (2014) A new group of phage anti-CRISPR genes inhibits the type I-E CRISPR-Cas system of Pseudomonas aeruginosa. mBio 5:e00896. https://doi.org/10.1128/mBio.00896-14
Pawluk A, Amrani N, Zhang Y et al (2016a) Naturally occurring off-switches for CRISPR-Cas9. Cell 167:1829–1838. https://doi.org/10.1016/j.cell.2016.11.017
Pawluk A, Staals RHJ, Taylor C et al (2016b) Inactivation of CRISPR-Cas systems by anti-CRISPR proteins in diverse bacterial species. Nat Microbiol 1:16085. https://doi.org/10.1038/nmicrobiol.2016.85
Peng R, Xu Y, Zhu T et al (2017) Alternate binding modes of anti-CRISPR viral suppressors AcrF1/2 to Csy surveillance complex revealed by cryo-EM structures. Cell Res 27:853–864. https://doi.org/10.1038/cr.2017.79
Peng X, Mayo-Muñoz D, Bhoobalan-Chitty Y, Martínez-Álvarez L (2020) Anti-CRISPR proteins in Archaea. Trends Microbiol 28:913–921
Petassi MT, Hsieh SC, Peters JE (2020) Guide RNA categorization enables target site choice in Tn7-CRISPR-Cas transposons. Cell 183:1757–1771. https://doi.org/10.1016/j.cell.2020.11.005
Peters JE, Makarova KS, Shmakov S, Koonin E, v. (2017) Recruitment of CRISPR-Cas systems by Tn7-like transposons. Proc Natl Acad Sci USA 114:E7358–E7366. https://doi.org/10.1073/pnas.1709035114
Pinilla-Redondo R, Mayo-Muñoz D, Russel J et al (2020) Type IV CRISPR-Cas systems are highly diverse and involved in competition between plasmids. Nucleic Acids Res 48:2000–2012. https://doi.org/10.1093/nar/gkz1197
Pougach K, Semenova E, Bogdanova E et al (2010) Transcription, processing and function of CRISPR cassettes in Escherichia coli. Mol Microbiol 77:1367–1379. https://doi.org/10.1111/j.1365-2958.2010.07265.x
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 151:653–663. https://doi.org/10.1099/mic.0.27437-0
Pourcel C (2020) New insights into CRISPR arrays. CRISPR J 3:422–424. https://doi.org/10.1089/crispr.2020.29111.cpo
Pul Ü, Wurm R, Arslan Z et al (2010) Identification and characterization of E. coli CRISPR-cas promoters and their silencing by H-NS. Mol Microbiol 75:1495–1512. https://doi.org/10.1111/j.1365-2958.2010.07073.x
Pyenson NC, Gayvert K, Varble A et al (2017) Broad targeting specificity during bacterial Type III CRISPR-Cas immunity constrains viral escape. Cell Host Microbe 22:343–353. https://doi.org/10.1016/j.chom.2017.07.016
Quax TEF, Voet M, Sismeiro O et al (2013a) Massive activation of archaeal defense genes during viral infection. J Virol 87:8419–8428. https://doi.org/10.1128/jvi.01020-13
Quax TEF, Wolf YI, Koehorst JJ et al (2013b) Differential translation tunes uneven production of operon-encoded proteins. Cell Rep 4:938–944. https://doi.org/10.1016/j.celrep.2013.07.049
Radovčić M, Killelea T, Savitskaya E et al (2018) CRISPR-Cas adaptation in Escherichia coli requires RecBCD helicase but not nuclease activity, is independent of homologous recombination, and is antagonized by 5’ ssDNA exonucleases. Nucleic Acids Res 46:10173–10183. https://doi.org/10.1093/nar/gky799
Rajagopalan R, Kroos L (2017) The dev operon regulates the timing of sporulation during Myxococcus xanthus development. J Bacteriol 199:e00788-e816. https://doi.org/10.1128/JB.00788-16
Ramachandran A, Summerville L, Learn BA et al (2020) Processing and integration of functionally oriented prespacers in the Escherichia coli CRISPR system depends on bacterial host exonucleases. J Biol Chem 295:3403–3414. https://doi.org/10.1074/jbc.RA119.012196
Rao C, Chin D, Ensminger AW (2017) Priming in a permissive type I-C CRISPR-Cas system reveals distinct dynamics of spacer acquisition and loss. RNA 23:1525–1538. https://doi.org/10.1261/rna.062083.117
Ratner HK, Escalera-Maurer A, le Rhun A et al (2019) Catalytically active Cas9 mediates transcriptional interference to facilitate bacterial virulence. Mol Cell 75:498–510. https://doi.org/10.1016/j.molcel.2019.05.029
Reeks J, Sokolowski RD, Graham S et al (2013) Structure of a dimeric crenarchaeal Cas6 enzyme with an atypical active site for CRISPR RNA processing. Biochem J 452:223–230. https://doi.org/10.1042/BJ20130269
Richter H, Zoephel J, Schermuly J et al (2012) Characterization of CRISPR RNA processing in Clostridium thermocellum and Methanococcus maripaludis. Nucleic Acids Res 40:9887–9896. https://doi.org/10.1093/nar/gks737
Richter H, Lange SJ, Backofen R, Randau L (2013) Comparative analysis of Cas6b processing and CRISPR RNA stability. RNA Biol 10:700–707. https://doi.org/10.4161/rna.23715
Richter C, Dy RL, McKenzie RE et al (2014) Priming in the Type I-F CRISPR-Cas system triggers strand-independent spacer acquisition, bi-directionally from the primed protospacer. Nucleic Acids Res 42:8516–8526. https://doi.org/10.1093/nar/gku527
Rocha EPC, Danchin A (2002) Base composition bias might result from competition for metabolic resources. Trends Genet 18:291–294. https://doi.org/10.1016/S0168-9525(02)02690-2
Rollie C, Schneider S, Brinkmann AS, Bolt EL, White MF (2015) Intrinsic sequence specificity of the Cas1 integrase directs new spacer acquisition. eLife 4:e08716. https://doi.org/10.7554/eLife.08716.001
Rollie C, Graham S, Rouillon C, White MF (2018) Prespacer processing and specific integration in a type I-A CRISPR system. Nucleic Acids Res 46:1007–1020. https://doi.org/10.1093/nar/gkx1232
Rousseau BA, Hou Z, Gramelspacher MJ, Zhang Y (2018) Programmable RNA cleavage and recognition by a natural CRISPR-Cas9 system from Neisseria meningitidis. Mol Cell 69:906–914. https://doi.org/10.1016/j.molcel.2018.01.025
Saito M, Ladha A, Strecker J et al (2021) Dual modes of CRISPR-associated transposon homing. Cell 184:2441–2453. https://doi.org/10.1016/j.cell.2021.03.006
Samai P, Pyenson N, Jiang W et al (2015) Co-transcriptional DNA and RNA cleavage during type III CRISPR-cas immunity. Cell 161:1164–1174. https://doi.org/10.1016/j.cell.2015.04.027
Sampson TR, Saroj SD, Llewellyn AC et al (2013) A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 497:254–257. https://doi.org/10.1038/nature12048
Sampson TR, Weiss DS (2013) Degeneration of a CRISPR/Cas system and its regulatory target during the evolution of a pathogen. RNA Biol 10:1618–1622. https://doi.org/10.4161/rna.26423
Sampson TR, Saroj SD, Llewellyn AC et al (2019) Author correction: A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 570:E30–E31. https://doi.org/10.1038/s41586-019-1253-9
Sashital DG, Jinek M, Doudna JA (2011) An RNA-induced conformational change required for CRISPR RNA cleavage by the endoribonuclease Cse3. Nat Struct Mol Biol 18:680–687. https://doi.org/10.1038/nsmb.2043
Sattler S (2017) The role of the immune system beyond the fight against infection. Adv Exp Med Biol 1003:3–14. https://doi.org/10.1007/978-3-319-57613-8_1
Savitskaya E, Semenova E, Dedkov V et al (2013) High-throughput analysis of type I-E CRISPR/Cas spacer acquisition in E. coli. RNA Biol 10:716–725. https://doi.org/10.4161/rna.24325
Seed KD, Lazinski DW, Calderwood SB, Camilli A (2013) A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 494:489–491. https://doi.org/10.1038/nature11927
Sefcikova J, Roth M, Yu G, Li H (2017) Cas6 processes tight and relaxed repeat RNA via multiple mechanisms: A hypothesis. BioEssays 39:1700019. https://doi.org/10.1002/bies.201700019
Selle K, Klaenhammer TR, Barrangou R (2015) CRISPR-based screening of genomic island excision events in bacteria. Proc Natl Acad Sci USA 112:8076–8081. https://doi.org/10.1073/pnas.1508525112
Semenova E, Jore MM, Datsenko KA et al (2011) Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc Natl Acad Sci USA 108:10098–10103. https://doi.org/10.1073/pnas.1104144108
Semenova E, Savitskaya E, Musharova O et al (2016) Highly efficient primed spacer acquisition from targets destroyed by the Escherichia coli type I-E CRISPR-Cas interfering complex. Proc Natl Acad Sci USA 113:7626–7631. https://doi.org/10.1073/pnas.1602639113
Shabbir MAB, Tang Y, Xu Z et al (2018) The involvement of the Cas9 gene in virulence of Campylobacter jejuni. Front Cell Infect Microbiol 8:285. https://doi.org/10.3389/fcimb.2018.00285
Shah M, Taylor VL, Bona D et al (2021) A phage-encoded anti-activator inhibits quorum sensing in Pseudomonas aeruginosa. Mol Cell 81:571–583. https://doi.org/10.1016/j.molcel.2020.12.011
Shah SA, Erdmann S, Mojica FJM, Garrett RA (2013) Protospacer recognition motifs: Mixed identities and functional diversity. RNA Biol 10:891–899. https://doi.org/10.4161/rna.23764
Shao Y, Li H (2013) Recognition and cleavage of a nonstructured CRISPR RNA by its processing endoribonuclease Cas6. Structure 21:385–393. https://doi.org/10.1016/j.str.2013.01.010
Shiimori M, Garrett SC, Chambers DP et al (2017) Role of free DNA ends and protospacer adjacent motifs for CRISPR DNA uptake in Pyrococcus furiosus. Nucleic Acids Res 45:11281–11294. https://doi.org/10.1093/nar/gkx839
Shiimori M, Garrett SC, Graveley BR, Terns MP (2018) Cas4 nucleases define the PAM, length, and orientation of DNA fragments integrated at CRISPR loci. Mol Cell 70:814–824. https://doi.org/10.1016/j.molcel.2018.05.002
Shinkai A, Kira S, Nakagawa N et al (2007) Transcription activation mediated by a cyclic AMP receptor protein from Thermus thermophilus HB8. J Bacteriol 189:3891–3901. https://doi.org/10.1128/JB.01739-06
Shintani M, Suzuki-Minakuchi C, Nojiri H (2015) Nucleoid-associated proteins encoded on plasmids: occurrence and mode of function. Plasmid 80:32–44. https://doi.org/10.1016/j.plasmid.2015.04.008
Shiriaeva A, Fedorov I, Vyhovskyi D, Severinov K (2020) Detection of CRISPR adaptation. Biochem Soc Trans 48:257–269. https://doi.org/10.1042/BST20190662
Shmakov S, Abudayyeh OO, Makarova KS et al (2015) Discovery and functional characterization of diverse Class 2 CRISPR-Cas systems. Mol Cell 60:385–397. https://doi.org/10.1016/j.molcel.2015.10.008
Shmakov S, Smargon A, Scott D et al (2017a) Diversity and evolution of class 2 CRISPR–Cas systems. Nat Rev Microbiol 15:169–182. https://doi.org/10.1038/nrmicro.2016.184
Shmakov SA, Sitnik V, Makarova KS et al (2017) The CRISPR spacer space is dominated by sequences from the species-specific mobilome. mBio 8:e01397-17. https://doi.org/10.1101/137356
Shmakov SA, Utkina I, Wolf YI et al (2020a) CRISPR arrays away from cas Genes. CRISPR J 3:535–549. https://doi.org/10.1089/crispr.2020.0062
Shmakov SA, Wolf YI, Savitskaya E et al (2020b) Mapping CRISPR spaceromes reveals vast host-specific viromes of prokaryotes. Communi Biol 3:321. https://doi.org/10.1038/s42003-020-1014-1
Silas S, Mohr G, Sidote DJ et al (2016) Direct CRISPR spacer acquisition from RNA by a natural reverse transcriptase-Cas1 fusion protein. Science 351:aad4234. https://doi.org/10.1126/science.aad4234
Skennerton CT, Angly FE, Breitbart M et al (2011) Phage encoded H-NS: a potential Achilles heel in the bacterial defence system. PLoS ONE 6:e20095. https://doi.org/10.1371/journal.pone.0020095
Smargon AA, Cox DBT, Pyzocha NK et al (2017) Cas13b Is a Type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol Cell 65:618–630. https://doi.org/10.1016/j.molcel.2016.12.023
Smith GR (2012) How RecBCD enzyme and Chi promote DNA break repair and recombination: a molecular biologist’s view. Microbiol Mol Biol Rev 76:217–228. https://doi.org/10.1128/mmbr.05026-11
Sofos N, Feng M, Stella S et al (2020) Structures of the Cmr-β complex reveal the regulation of the immunity mechanism of type III-B CRISPR-Cas. Mol Cell 79:741–757. https://doi.org/10.1016/j.molcel.2020.07.008
Solbiati J, Duran-Pinedo A, Godoy Rocha F et al (2020) Virulence of the pathogen Porphyromonas gingivalis is controlled by the CRISPR-Cas protein Cas3. mSystems 5:e00852-20. https://doi.org/10.1128/mSystems.00852-20
Spencer BL, Deng L, Patras KA et al (2019) Cas9 contributes to group B streptococcal colonization and disease. Front Microbiol 10:1930. https://doi.org/10.3389/fmicb.2019.01930
Stachler AE, Marchfelder A (2016) Gene repression in haloarchaea using the CRISPR (clustered regularly interspaced short palindromic repeats)-Cas I-B system. J Biol Chem 291:15226–15242. https://doi.org/10.1074/jbc.M116.724062
Stella S, Alcón P, Montoya G (2017) Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 546:559–563. https://doi.org/10.1038/nature22398
Stern A, Keren L, Wurtzel O et al (2010) Self-targeting by CRISPR: Gene regulation or autoimmunity? Trends Genet 26:335–340. https://doi.org/10.1016/j.tig.2010.05.008
Sternberg SH, Redding S, Jinek M et al (2014) DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507:62–67. https://doi.org/10.1038/nature13011
Strecker J, Ladha A, Gardner Z et al (2019) RNA-guided DNA insertion with CRISPR-associated transposases. Science 365:48–53. https://doi.org/10.1126/science.aax9181
Strotskaya A, Savitskaya E, Metlitskaya A et al (2017) The action of Escherichia coli CRISPR–Cas system on lytic bacteriophages with different lifestyles and development strategies. Nucleic Acids Res 45:1946–1957. https://doi.org/10.1093/nar/gkx042
Strutt SC, Torrez RM, Kaya E et al (2018) RNA-dependent RNA targeting by CRISPR-Cas9. eLife 7:e32724. https://doi.org/10.7554/eLife.32724
Sun D, Mao X, Fei M et al (2020a) Histone-like nucleoid-structuring protein (H-NS) paralogue Stpa activates the type I-E CRISPR-Cas system against natural transformation in Escherichia coli. Appl. Environ. Microbiol. 86:e00731-20. https://doi.org/10.1128/AEM.00731-20
Sun JY, Hu HB, Cheng YX, Lu XJ (2020b) CRISPR in medicine: applications and challenges. Brief Funct Genomics 19:151–153. https://doi.org/10.1093/bfgp/elaa011
Swarts DC, Mosterd C, van Passel MWJ, Brouns SJJ (2012) CRISPR interference directs strand specific spacer acquisition. PLoS ONE 7:e35888. https://doi.org/10.1371/journal.pone.0035888
Swarts DC, van der Oost J, Jinek M (2017) Structural basis for guide RNA processing and seed-dependent DNA targeting by CRISPR-Cas12a. Mol Cell 66:221–233. https://doi.org/10.1016/j.molcel.2017.03.016
Tang B, Gong T, Zhou X et al (2019) Deletion of cas3 gene in Streptococcus mutans affects biofilm formation and increases fluoride sensitivity. Arch Oral Biol 99:190–197. https://doi.org/10.1016/j.archoralbio.2019.01.016
Tao P, Wu X, Rao V (2018) Unexpected evolutionary benefit to phages imparted by bacterial CRISPR-Cas9. Sci Adv 4:aar4134. https://doi.org/10.1126/sciadv.aar4134
Toro N, Mestre MR, Martínez-Abarca F, González-Delgado A (2019) Recruitment of reverse transcriptase-Cas1 fusion proteins by type VI-A CRISPR-Cas systems. Front Microbiol 10:2160. https://doi.org/10.3389/fmicb.2019.02160
Turgeman-Grott I, Joseph S, Marton S et al (2019) Pervasive acquisition of CRISPR memory driven by inter-species mating of archaea can limit gene transfer and influence speciation. Nat Microbiol 4:177–186. https://doi.org/10.1038/s41564-018-0302-8
Ulbricht A, Nickel L, Weidenbach K et al (2020) The CARF protein MM_0565 affects transcription of the casposon-encoded cas1-solo gene in Methanosarcina mazei Gö1. Biomolecules 10:1161. https://doi.org/10.3390/biom10081161
van der Oost J, Mougiakos I (2020) First structural insights into CRISPR-Cas-guided DNA transposition. Cell Res 30:193–194. https://doi.org/10.1038/s41422-020-0284-y
Varble A, Meaden S, Barrangou R et al (2019) Recombination between phages and CRISPR−cas loci facilitates horizontal gene transfer in staphylococci. Nat Microbiol 4:956–963. https://doi.org/10.1038/s41564-019-0400-2
Vercoe RB, Chang JT, Dy RL et al (2013) Cytotoxic chromosomal targeting by CRISPR/Cas systems can reshape bacterial genomes and expel or remodel pathogenicity islands. Plos Genet 9:e1003454. https://doi.org/10.1371/journal.pgen.1003454
Vlot M, Houkes J, Lochs SJA et al (2018) Bacteriophage DNA glucosylation impairs target DNA binding by type I and II but not by type V CRISPR–Cas effector complexes. Nucleic Acids Res 46:873–885. https://doi.org/10.1093/nar/gkx1264
Vorontsova D, Datsenko KA, Medvedeva S et al (2015) Foreign DNA acquisition by the I-F CRISPR-Cas system requires all components of the interference machinery. Nucleic Acids Res 43:10848–10860. https://doi.org/10.1093/nar/gkv1261
Wallace RA, Black WP, Yang X, Yang Z (2014) A CRISPR with roles in Myxococcus xanthus development and exopolysaccharide production. J Bacteriol 196:4036–4043. https://doi.org/10.1128/JB.02035-14
Wang J, Li J, Zhao H et al (2015) Structural and mechanistic basis of PAM-dependent spacer acquisition in CRISPR-Cas systems. Cell 163:840–853. https://doi.org/10.1016/j.cell.2015.10.008
Wang X, Yao D, Xu JJG et al (2016) Structural basis of Cas3 inhibition by the bacteriophage protein AcrF3. Nat Struct Mol Biol 23:868–870. https://doi.org/10.1038/nsmb.3269
Wang Y, Yin X, Zhou Z et al (2019) Cas9 regulated gene expression and pathogenicity in Riemerella anatipestifer. Microb Pathog 136:103706. https://doi.org/10.1016/j.micpath.2019.103706
Watson BNJ, Staals RHJ, Fineran PC (2018) CRISPR-Cas-mediated phage resistance enhances horizontal gene transfer by transduction. mBio 9:e02406-17. https://doi.org/10.1128/mBio.02406-17
Watson BNJ, Vercoe RB, Salmond GPC et al (2019) Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity. Nat Commun 10:5526. https://doi.org/10.1038/s41467-019-13445-2
Watters KE, Fellmann C, Bai HB et al (2018) Systematic discovery of natural CRISPR-Cas12a inhibitors. Science 362:236–239. https://doi.org/10.1126/science.aau5138
Wei Y, Terns RM, Terns MP (2015) Cas9 function and host genome sampling in type II-A CRISPR–Cas adaptation. Genes Dev 29:356–361. https://doi.org/10.1101/gad.257550.114
Weissman JL, Stoltzfus A, Westra ER, Johnson PLF (2020) Avoidance of self during CRISPR immunization. Trends Microbiol 28:543–553
Westra ER, Pul Ü, Heidrich N et al (2010) H-NS-mediated repression of CRISPR-based immunity in Escherichia coli K12 can be relieved by the transcription activator LeuO. Mol Microbiol 77:1380–1393. https://doi.org/10.1111/j.1365-2958.2010.07315.x
Westra ER, Nilges B, van Erp PBG et al (2012a) Cascade-mediated binding and bending of negatively supercoiled DNA. RNA Biol 9:1134–1138. https://doi.org/10.4161/rna.21410
Westra ER, van Erp PBG, Künne T et al (2012b) CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol Cell 46:595–605. https://doi.org/10.1016/j.molcel.2012.03.018
Westra ER, Semenova E, Datsenko KA et al (2013a) Type I-E CRISPR-Cas systems discriminate target from non-target DNA through base pairing-independent PAM recognition. Plos Genet 9:e1003742. https://doi.org/10.1371/journal.pgen.1003742
Westra ER, Staals RHJ, Gort G et al (2013b) CRISPR-Cas systems preferentially target the leading regions of MOB F conjugative plasmids. RNA Biol 10:749–761. https://doi.org/10.4161/rna.24202
Westra ER, Levin BR (2020) It is unclear how important CRISPR-Cas systems are for protecting natural populations of bacteria against infections by mobile genetic elements. Proc Natl Acad Sci USA 117:27777–27785. https://doi.org/10.1073/pnas.1915966117
Wheatley RM, MacLean RC (2020) CRISPR-Cas systems restrict horizontal gene transfer in Pseudomonas aeruginosa. ISME J 15:1420–1433. https://doi.org/10.1038/s41396-020-00860-3
Wiedenheft B, Zhou K, Jinek M et al (2009) Structural basis for DNase activity of a conserved protein implicated in CRISPR-mediated genome defense. Structure 17:904–912. https://doi.org/10.1016/j.str.2009.03.019
Wimmer F, Beisel CL (2020) CRISPR-Cas systems and the paradox of self-targeting spacers. Front Microbiol 10:3078. https://doi.org/10.3389/fmicb.2019.03078
Winardhi RS, Yan J, Kenney LJ (2015) H-NS regulates gene expression and compacts the nucleoid: insights from single-molecule experiments. Biophys J 109:1321–1329. https://doi.org/10.1016/j.bpj.2015.08.016
Workman R, Pammi T, Nguyen B et al (2021) A natural single-guide RNA repurposes Cas9 to autoregulate CRISPR-Cas expression. Cell 184:675–688. https://doi.org/10.1016/j.cell.2020.12.017
Wurtzel O, Sapra R, Chen F et al (2010) A single-base resolution map of an archaeal transcriptome. Genome Res 20:133–141. https://doi.org/10.1101/gr.100396.109
Xiao Y, Luo M, Hayes RP et al (2017) Structure basis for directional R-loop formation and substrate handover mechanisms in Type I CRISPR-Cas system. Cell 170:48–60. https://doi.org/10.1016/j.cell.2017.06.012
Xiao Y, Luo M, Dolan AE et al (2018) Structure basis for RNA-guided DNA degradation by Cascade and Cas3. Science 361:eaat0839. https://doi.org/10.1126/science.aat0839
Yan WX, Chong S, Zhang H et al (2018) Cas13d is a compact RNA-targeting Type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol Cell 70:327–339. https://doi.org/10.1016/j.molcel.2018.02.028
Yan WX, Hunnewell P, Alfonse LE et al (2019) Functionally diverse type V CRISPR-Cas systems. Science 363:88–91. https://doi.org/10.1126/science.aav7271
Yang CD, Chen YH, Huang HY et al (2014) CRP represses the CRISPR/Cas system in Escherichia coli: Evidence that endogenous CRISPR spacers impede phage P1 replication. Mol Microbiol 92:1072–1091. https://doi.org/10.1111/mmi.12614
Yang H, Gao P, Rajashankar KR, Patel DJ (2016) PAM-dependent target DNA recognition and cleavage by C2c1 CRISPR-Cas endonuclease. Cell 167:1814–1828. https://doi.org/10.1016/j.cell.2016.11.053
Yang H, Patel DJ (2017) Inhibition mechanism of an anti-CRISPR suppressor AcrIIA4 targeting SpyCas9. Mol Cell 67:117–127. https://doi.org/10.1016/j.molcel.2017.05.024
Yang D, Wang Z, Ma J et al (2020) Glycine cleavage system and cAMP receptor protein co-regulate CRISPR/cas3 expression to resist bacteriophage. Viruses 12:90. https://doi.org/10.3390/v12010090
Ye Q, Zhao X, Liu J et al (2020) CRISPR-associated factor Csa3b regulates CRISPR adaptation and Cmr-mediated RNA interference in Sulfolobus islandicus. Front Microbiol 11:2038. https://doi.org/10.3389/fmicb.2020.02038
Yin Y, Yang B, Entwistle S (2019) Bioinformatics identification of Anti-CRISPR loci by using homology, guilt-by-association, and CRISPR self-targeting spacer approaches. mSystems 4:e00455-19. https://doi.org/10.1128/mSystems.00455-19
Yoganand KN, Muralidharan M, Nimkar S, Anand B (2019) Fidelity of prespacer capture and processing is governed by the PAM-mediated interactions of Cas1-2 adaptation complex in CRISPR-Cas type I-E system. J Biol Chem 294:20039–20053. https://doi.org/10.1074/jbc.RA119.009438
Yoganand KNR, Sivathanu R, Nimkar S, Anand B (2017) Asymmetric positioning of Cas1-2 complex and integration host factor induced DNA bending guide the unidirectional homing of protospacer in CRISPR-Cas type I-E system. Nucleic Acids Res 45:367–381. https://doi.org/10.1093/nar/gkw1151
Yosef I, Goren MG, Kiro R et al (2011) High-temperature protein G is essential for activity of the Escherichia coli clustered regularly interspaced short palindromic repeats (CRISPR)/Cas system. Proc Natl Acad Sci USA 108:20136–20141. https://doi.org/10.1073/pnas.1113519108
Yosef I, Goren MG, Qimron U (2012) Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res 40:5569–5576. https://doi.org/10.1093/nar/gks216
Yosef I, Shitrit D, Goren MG et al (2013) DNA motifs determining the efficiency of adaptation into the Escherichia coli CRISPR array. Proc Natl Acad Sci USA 110:14396–21401. https://doi.org/10.1073/pnas.1300108110
Zabrady K, Zabrady M, Kolesar P et al (2021) CRISPR-associated primase-polymerases are implicated in prokaryotic CRISPR-Cas adaptation. Nat Commun 12:3690. https://doi.org/10.1038/s41467-021-23535-9
Zegans ME, Wagner JC, Cady KC et al (2009) Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J Bacteriol 191:210–219. https://doi.org/10.1128/JB.00797-08
Zhang A, Chen J, Gong T et al (2020) Deletion of csn2 gene affects acid tolerance and exopolysaccharide synthesis in Streptococcus mutans. Mol Oral Microbiol 35:211–221. https://doi.org/10.1111/omi.12308
Zhang Y, Heidrich N, Ampattu BJ et al (2013) Processing-independent CRISPR RNAs limit natural transformation in Neisseria meningitidis. Mol Cell 50:488–503. https://doi.org/10.1016/j.molcel.2013.05.001
Zhou Y, Tang Y, Fu P et al (2020) The type I-E CRISPR-Cas system influences the acquisition of blaKPC-IncF plasmid in Klebsiella pneumonia. Emerg Microbes Infect 9:1011–1022. https://doi.org/10.1080/22221751.2020.1763209
Zhu Y, Gao A, Zhan Q et al (2019) Diverse mechanisms of CRISPR-Cas9 inhibition by type IIC anti-CRISPR proteins. Mol Cell 74:296–309. https://doi.org/10.1016/j.molcel.2019.01.038
Acknowledgements
Figures were created with BioRender.com.
Funding
The authors are supported by grant PROMETEO/2017/129 (Conselleria d’Educació, Investigació, Cultura i Esport, Generalitat Valenciana, Spain).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guzmán, N.M., Esquerra-Ruvira, B. & Mojica, F.J.M. Digging into the lesser-known aspects of CRISPR biology. Int Microbiol 24, 473–498 (2021). https://doi.org/10.1007/s10123-021-00208-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-021-00208-7