Abstract
The aim of the study was to determine factors associated with spread of linezolid (LNZ)-resistant Staphylococcus epidermidis isolates in a surgical intensive care unit (ICU). A case-control study was conducted in one French adult surgical ICU. From January 2012 to December 2016, patients with at least a single positive LNZ-resistant S. epidermidis blood culture were matched to control with LNZ-susceptible S. epidermidis blood culture in a 1:4 manner. Cases were compared to controls regarding baseline clinical characteristics and LNZ exposure before positive blood culture. Bacterial isolates were genotyped by using pulsed-field gel electrophoresis (PFGE) and MLST. We identified 13 LNZ-resistant S. epidermidis isolates, 1 in 2012, 3 in 2014, 6 in 2015, and 3 in 2016. LNZ use increased steadily from 8 DDDs/100 patient days in 2010 to 19 in 2013 and further decrease by more of 50% in 2015 and 2016. The only independent risk factors associated to LNZ-resistant S. epidermidis isolation were length of stay in ICU before infection (OR 1.45; 95% CI 1.07–1.98), prior exposure to LNZ (OR 109; 95% CI 3.9–3034), and Charlson comorbidities score (OR 3.19; 95% CI 1.11–9.14). PFGE typing showed that all LNZ-resistant isolates were clonal belonging to ST2 and that LNZ-susceptible isolates were highly diverse. We report herein that previous exposure to LNZ substantially increased the risk of occurrence of LNZ resistance in S. epidermidis even in the case of clonal spread of LNZ-resistant isolates. These findings highlight the need for reducing the use of LNZ to preserve its efficacy in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coagulase-negative staphylococci (CoNS) are among the most important pathogens involved in hospital associated with bloodstream infections and infections related to vascular prosthetic devices [1]. A large proportion of nosocomial strains of CoNS are resistant to most of the available antibiotics [2]. Linezolid (LNZ) short-term safety, pharmacokinetics/pharmacodynamics profile, and clinical effectiveness as well could make it more attractive than vancomycin, especially in the ICU setting. However, three LNZ resistance mechanisms have been characterized so far: mutations in the domain V region of 23S rRNA genes, particularly a G2576T substitution; acquisition of the ribosomal methyltransferase gene cfr; and mutations in the ribosomal proteins L3 and L4 [3]. Both vertical and horizontal transmission of LNZ resistance may occur. LNZ-resistant CoNS are increasingly reported worldwide [4,5,6,7,8] .
We therefore conducted a case control study for a 5-year period in the surgical ICU of a French University Hospital to identify risk factors associated with LNZ-resistant S. epidermidis. We also characterized the molecular epidemiology of LNZ-resistant S. epidermidis.
Materials and methods
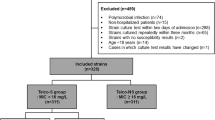
The study was performed in the surgical ICU of the University Hospital of Besancon. To identify risk factors, a case control study was performed. All patients, in whom LNZ-resistant S. epidermidis had been recovered in blood sample between 1 January 2012 and 31 December 2016, were included as case patients. If several episodes occurred, only the isolate from the first episode was considered. They were matched 1:4 with controls that had LNZ-susceptible S. epidermidis in blood sample. For each case, eligible controls were randomly selected.
Infection caused by S. epidermidis was defined when isolates were recovered from multiple blood cultures meeting criteria from the Centers for Disease Control and Prevention for significant bacteremia [9].
Staphylococcus epidermidis isolates were cultured and identified according to routine diagnostic procedures. Antimicrobial susceptibility testing was performed using disk diffusion methods according to EUCAST recommendations and LNZ MIC of LNZ-resistant isolates was determined using Etest®. A MIC > 4 mg/L defined LNZ resistance according to EUCAST guidelines [10].
Demographic and clinical data were collected retrospectively by reviewing the medical charts Administration of antibiotics in the month before admission was also documented. Global LNZ use in the surgical ICU was collected from the pharmacy database and expressed in defined daily doses (DDDs) per 100 patient days.
Bacterial isolates were genotyped by using pulsed-field gel electrophoresis (PFGE) as previously described [11] and clustered in pulsotypes according to international recommendations [12]. LNZ-resistant isolates were further characterized using the MLST scheme developed by Thomas et al. [13].
All variables were examined by univariate analysis using the chi-square or Fisher’s exact test, as appropriate. Continuous variables were analyzed by Student’s t test. All statistical tests were two-tails, and P < 0.05 was considered to be statistically significant. Multivariate analysis was performed by conditional logistic regression. Stepwise selection with an entry and stay level of P = 0.1 was used to build the final multivariate logistic regression model. The adjusted odds ratio (OR) and 95% confidence interval (CI) for the variables selected in the final model are reported. Statistical analyses were computed using the SPSS program version 24 (SPSS Inc., Chicago, USA).
Results
Patient characteristics
During the study period, 13 patients had positive blood cultures with LNZ-resistant S. epidermidis (1 in 2012, 3 in 2014, 6 in 2015, and 3 in 2016). The mean age was 65.7 ± 11 years, 10 (77%) were male, and all patients had received antibiotics within the previous month. Recent exposure to LNZ was reported in 12 (92.3%) patients (Table 1).
Antimicrobial susceptibility
All isolates had a LNZ MIC > 256 mg/L and were co-resistant to methicillin. Co-resistance was also observed with gentamicin (100% of isolates), ofloxacin (100%), rifampicin (31%), and teicoplanin (23%). No vancomycin resistance was observed.
Risk factors and outcome
In multivariate analysis (Table 1), length of stay in ICU before infection (OR 1.45; 95% CI 1.07–1.98), prior exposure to LNZ (OR 109; 95% CI 3.9–3034), and Charlson comorbidities score (OR 3.19; 95% CI 1.11–9.14) were associated with LNZ-resistant S. epidermidis.
There is no difference in mortality in ICU between the 2 groups (p = 0.68), even in the group of patients with blood stream infection (2/9 (22%) for cases vs 3/30 (30%) for controls, p = 0.66).
Linezolid use
The number of LNZ DDDs/100 patient days was 8 in 2010 and increased steadily until it more than doubled to 19 in 2013. LNZ consumption fell by more 50% to 8 DDDs/100 patient days in 2015 and 2016 (Fig. 1).
Molecular typing of bacterial strains
Sixty five isolates of S. epidermidis were genotyped: the 13 LNZ-resistant isolates shared the same PFGE pattern and the 52 LNZ-susceptible isolates showed different PFGE patterns than resistant ones. Globally, LNZ-susceptible isolates displayed a high genomic diversity when considering PFGE results. All LNZ-resistant isolates belonged to ST2.
Discussion
The effectiveness of LNZ against Gram-positive cocci and its favorable short-term safety profile have promoted its widespread use, leading in turn theoretically to the emergence and dissemination of LNZ resistance. In most of published surveillance studies, LNZ-resistant isolates were rare (< 1% in the LEADER surveillance program and 0.4% in the ZAAPS program) [14, 15].
Gu et al. reported a total of 351 LNZ-resistant CoNS cases and the majority from patients in North America (30.8%) and Europe (20%) [16]. LNZ administration is reported to be one of the most important risk factors for LNZ-resistant Gram-positive cocci isolation in hospital outbreaks [17,18,19,20,21,22]. A review of studies that have identified risk factors associated with the isolation of LNZ-resistant CoNS was reported in Table 2.
The emergence of LNZ resistance in S. epidermidis in our ICU was associated with usage of LNZ, which has exerted a high selective pressure. However, it is worth noting that one out of 13 patients did not receive LNZ before resistant S. epidermidis isolated [23]. Other studies showed that being hospitalized near an already colonized patient increased the chances of acquiring such resistant microorganism [24, 25].
PFGE typing showed that all LNZ-resistant isolates belonged to the same pulsotype and that LNZ-susceptible isolates were highly diverse. These findings showed the emergence of a clonal spread of LNZ-resistant S. epidermidis in the ICU that persisted despite a decreased of LNZ consumption.
Beside previous exposure to the drug, we identified the Charlson comorbidities score elevation as an independent risk factor for the isolation of LNZ-resistant S. epidermidis, suggesting that patients with comorbidities were likely to acquire such difficult-to-treat bacteria. One explanation of this finding is that patients with comorbidities require more care, more admission in hospitals, so more opportunities to acquire resistant bacteria. Moreover, increased length of stay was also found to be an important risk factor for LNZ-resistant strains isolated, which confirms our hypothesis. LNZ exposure was probably a factor permitting the emergence of the clone and participate of these diffusion with cross transmission.
Treatment options for LNZ-resistant CoNS are limited and are based on current in vitro susceptibility data. LNZ-resistant CoNS remain universally susceptible to vancomycin, daptomycin, and tigecycline.
Density of LNZ exposure also plays a key role in the emergence of resistance, and it has been suggested that defined daily dose (DDD) longer than 13–15 days generate enough selective pressure to trigger outbreaks [18, 26]. According to our data, the average number of days treated with LNZ was 10.3, and 8 patients (60%) had been treated with LNZ for at least 13 days. Previous use of LNZ before isolation of resistant strain and LNZ DDDs consumption in different studies were reported in Table 3. In France, in 2016, the median [min-max] of LNZ use in 29 intensive care units was 3.2 DDJ/100 patient days [0.1–16.6] (ATB-Raisin surveillance network) [27]. These results showed an important disparity of LNZ use in French ICUs and confirmed a high consumption of these in our hospital.
Taken together our findings suggest that increasing LNZ use produced selective pressure likely to promote resistant strain selection. Patient’s cross-transmission has contributed to the outbreak since patients without prior exposure required a longer time in the ICU before isolation of LNZ-resistant CoNS.
Of note all isolates tested in the present study were clonal by PFGE and belonged to ST2, which is a common S. epidermidis lineage in Europe [17, 28]. There are nine main clonal lineages composing the nosocomial S. epidermidis population worldwide. Also, strains of ST2, ST5, and ST22 are clustered into CC5 which is the most prevalent clonal complex regarding the nosocomial S. epidermidis population [13]. Mihaila et al. reported an outbreak of bloodstream infections with LNZ-resistant S. epidermidis and Staphylococcus pettenkoferi in an ICU in Paris [29]. Their analysis revealed that cross-infection was responsible of the acquisition of the LNZ-resistant CoNS, and they concluded that the emergence of these strains was rather the result of transmission of clonal strains than the selection of resistant mutants under therapeutic treatment. In our study, an overlapping hospital stays were present for 7 patients (54%), which confirm the important role of cross-transmission in the emergence of this clone.
Our study has limitations. Data were collected retrospectively, and the control group was chosen from among patients with non-LNZ-resistant S. epidermidis. However, this design may bias results and lead to an overestimation of the OR, particularly with regard to previous exposure to antibiotics [30].
References
Rogers KL, Fey PD, Rupp ME (2009) Coagulase-negative staphylococcal infections. Infect Dis Clin N Am 23:73–98. https://doi.org/10.1016/j.idc.2008.10.001
Wisplinghoff H, Bischoff T, Tallent SM, Seifert H, Wenzel RP, Edmond MB (2004) Nosocomial bloodstream infections in US hospitals: analysis of 24,179 cases from a prospective nationwide surveillance study. Clin Infect Dis Off Publ Infect Dis Soc Am 1(39):309–317. https://doi.org/10.1086/421946
Long KS, Vester B (2012) Resistance to linezolid caused by modifications at its binding site on the ribosome. Antimicrob Agents Chemother 56:603–612. https://doi.org/10.1128/AAC.05702-11
Dortet L, Glaser P, Kassis-Chikhani N, Girlich D, Ichai P, Boudon M et al (2017) Long-lasting successful dissemination of resistance to oxazolidinones in MDR Staphylococcus epidermidis clinical isolates in a tertiary care hospital in France. J Antimicrob Chemother 30. https://doi.org/10.1093/jac/dkx370/4582307
Tewhey R, Gu B, Kelesidis T, Charlton C, Bobenchik A, Hindler J et al (2014) Mechanisms of linezolid resistance among coagulase-negative staphylococci determined by whole-genome sequencing. mBio 13(5):e00894–e00814. https://doi.org/10.1128/mBio.00894-14
Bonilla H, Huband MD, Seidel J, Schmidt H, Lescoe M, McCurdy SP et al (2010) Multicity outbreak of linezolid-resistant Staphylococcus epidermidis associated with clonal spread of a cfr-containing strain. Clin Infect Dis Off Publ Infect Dis Soc Am 1(51):796–800. https://doi.org/10.1086/656281
Weßels C, Strommenger B, Klare I, Bender J, Messler S, Mattner F et al (2018) Emergence and control of linezolid-resistant Staphylococcus epidermidis in an ICU of a German hospital. J Antimicrob Chemother 1(73):1185–1193. https://doi.org/10.1093/jac/dky010
Li X, Arias CA, Aitken SL, Galloway Peña J, Panesso D, Chang M et al (2018) Clonal emergence of invasive multidrug-resistant Staphylococcus epidermidis deconvoluted via a combination of whole-genome sequencing and microbiome analyses. Clin Infect Dis 18(67):398–406. https://doi.org/10.1093/cid/ciy089
Garner JS, Jarvis WR, Emori TG, Horan TC, Hughes JM (1988) CDC definitions for nosocomial infections, 1988. Am J Infect Control 16:128–140
EUCAST: clinical breakpoints and dosing of antibiotics. 2019 http://www.eucast.org/clinical_breakpoints/, cited 2019 6
Boisson K, Thouverez M, Talon D, Bertrand X (2002) Characterisation of coagulase-negative staphylococci isolated from blood infections: incidence, susceptibility to glycopeptides, and molecular epidemiology. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol 21:660–665. https://doi.org/10.1007/s10096-002-0799-9
Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH et al (1995) Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol 33:2233–2239
Thomas JC, Vargas MR, Miragaia M, Peacock SJ, Archer GL, Enright MC (2007) Improved multilocus sequence typing scheme for Staphylococcus epidermidis. J Clin Microbiol 45:616–619. https://doi.org/10.1128/JCM.01934-06
Mendes RE, Deshpande L, Streit JM, Sader HS, Castanheira M, Hogan PA et al (2018) ZAAPS programme results for 2016: an activity and spectrum analysis of linezolid using clinical isolates from medical centres in 42 countries. J Antimicrob Chemother 1(73):1880–1887. https://doi.org/10.1093/jac/dky099
Pfaller MA, Mendes RE, Streit JM, Hogan PA, Flamm RK (2017) Five-year summary of in vitro activity and resistance mechanisms of linezolid against clinically important Gram-positive cocci in the United States from the LEADER surveillance program (2011 to 2015). Antimicrob. Agents Chemother 61. https://doi.org/10.1128/AAC.00609-17
Gu B, Kelesidis T, Tsiodras S, Hindler J, Humphries RM (2013) The emerging problem of linezolid-resistant Staphylococcus. J Antimicrob Chemother 68:4–11. https://doi.org/10.1093/jac/dks354
Liakopoulos A, Spiliopoulou I, Damani A, Kanellopoulou M, Schoina S, Papafragas E et al (2010) Dissemination of two international linezolid-resistant Staphylococcus epidermidis clones in Greek hospitals. J Antimicrob Chemother 65:1070–1071. https://doi.org/10.1093/jac/dkq065
Treviño M, Martínez-Lamas L, Romero-Jung PA, Giráldez JM, Alvarez-Escudero J, Regueiro BJ (2009) Endemic linezolid-resistant Staphylococcus epidermidis in a critical care unit. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol 28:527–533. https://doi.org/10.1007/s10096-008-0657-5
Kelly S, Collins J, Maguire M, Gowing C, Flanagan M, Donnelly M et al (2008) An outbreak of colonization with linezolid-resistant Staphylococcus epidermidis in an intensive therapy unit. J Antimicrob Chemother 61:901–907. https://doi.org/10.1093/jac/dkn043
Seral C, Sáenz Y, Algarate S, Duran E, Luque P, Torres C et al (2011) Nosocomial outbreak of methicillin- and linezolid-resistant Staphylococcus epidermidis associated with catheter-related infections in intensive care unit patients. Int J Med Microbiol IJMM 301:354–358. https://doi.org/10.1016/j.ijmm.2010.11.001
Potoski BA, Adams J, Clarke L, Shutt K, Linden PK, Baxter C et al (2006) Epidemiological profile of linezolid-resistant coagulase-negative staphylococci. Clin Infect Dis Off Publ Infect Dis Soc Am 15(43):165–171. https://doi.org/10.1086/505114
Bouiller K, Bador J, Bruyère R, Toitot A, Prin S, Quenot J-P et al (2017 http://linkinghub.elsevier.com/retrieve/pii/S0924857917303187 [cited 2017 28]) Recent exposure to linezolid is strongly associated with the isolation of linezolid-resistant coagulase-negative staphylococcus species in patients with related infection or colonization: a case-control study in an intensive care unit. Int J Antimicrob Agents. https://doi.org/10.1016/j.ijantimicag.2017.08.026
Baos E, Candel FJ, Merino P, Pena I, Picazo JJ (2013) Characterization and monitoring of linezolid-resistant clinical isolates of Staphylococcus epidermidis in an intensive care unit 4 years after an outbreak of infection by cfr-mediated linezolid-resistant Staphylococcus aureus. Diagn Microbiol Infect Dis 76:325–329. https://doi.org/10.1016/j.diagmicrobio.2013.04.002
Safdar N, Maki DG (2002) The commonality of risk factors for nosocomial colonization and infection with antimicrobial-resistant Staphylococcus aureus, enterococcus, gram-negative bacilli, Clostridium difficile, and Candida. Ann Intern Med 4(136):834–844
Petinaki E, Kanellopoulou M, Damani A, Foka A, Spiliopoulou I, Skalmoutsou N et al (2009) Linezolid-resistant Staphylococcus cohnii. Greece Emerg Infect Dis 15:116–118. https://doi.org/10.3201/eid1501.080769
Mulanovich VE, Huband MD, McCurdy SP, Lemmon MM, Lescoe M, Jiang Y et al (2010) Emergence of linezolid-resistant coagulase-negative Staphylococcus in a cancer centre linked to increased linezolid utilization. J Antimicrob Chemother 65:2001–2004. https://doi.org/10.1093/jac/dkq238
Surveillance de la consommation des antibiotiques / 2018 / Maladies infectieuses / Rapports et synthèses / Publications et outils / Accueil. [cited 2019 29]; http://invs.santepubliquefrance.fr/Publications-et-outils/Rapports-et-syntheses/Maladies-infectieuses/2018/Surveillance-de-la-consommation-des-antibiotiques
Papadimitriou-Olivgeris M, Giormezis N, Fligou F, Liakopoulos A, Marangos M, Anastassiou ED et al (2013) Factors influencing linezolid-nonsusceptible coagulase-negative staphylococci dissemination among patients in the intensive care unit: a retrospective cohort study. Chemotherapy 59:420–426. https://doi.org/10.1159/000363281
Mihaila L, Defrance G, Levesque E, Ichai P, Garnier F, Derouin V et al (2012) A dual outbreak of bloodstream infections with linezolid-resistant Staphylococcus epidermidis and Staphylococcus pettenkoferi in a liver intensive care unit. Int J Antimicrob Agents 40:472–474. https://doi.org/10.1016/j.ijantimicag.2012.06.014
Harris AD, Samore MH, Lipsitch M, Kaye KS, Perencevich E, Carmeli Y (2002) Control-group selection importance in studies of antimicrobial resistance: examples applied to Pseudomonas aeruginosa, Enterococci, and Escherichia coli, Clin Infect Dis Off Publ Infect Dis Soc Am. 15(34):1558–1563. https://doi.org/10.1086/340533
Author information
Authors and Affiliations
Contributions
K.B. analyzed the data, K.B. wrote the manuscript with support from X.B and C.C. X.B., P.H.W, and C.C conceived the study, P.C performed bacteriological analysis. D.I. and P.H.W collected data. All authors provided critical feedback and helped shape the research, analysis, and manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable. According to French legislation in this period, and because no intervention was performed on patients, no written informed consent was given by the patients.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bouiller, K., Ilic, D., Wicky, P.H. et al. Spread of clonal linezolid-resistant Staphylococcus epidermidis in an intensive care unit associated with linezolid exposure. Eur J Clin Microbiol Infect Dis 39, 1271–1277 (2020). https://doi.org/10.1007/s10096-020-03842-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-020-03842-7