Abstract
Dengue is a rapidly spreading mosquito-borne disease caused by the dengue virus (DENV) and has emerged as a severe public health problem around the world. Guangdong, one of the southern Chinese provinces, experienced a serious outbreak of dengue in 2014, which was believed to be the worst dengue epidemic in China over the last 20 years. To better understand the epidemic, we collected the epidemiological data of the outbreak and analyzed 14,594 clinically suspected dengue patients from 25 hospitals in Guangdong. Dengue cases were then laboratory-confirmed by the detection of DENV non-structural protein 1 (NS1) antigen and/or DENV RNA. Afterwards, clinical manifestations of dengue patients were analyzed and 93 laboratory-positive serum specimens were chosen for the DENV serotyping and molecular analysis. Our data showed that the 2014 dengue outbreak in Guangdong had spread to 20 cities and more than 45 thousand people suffered from dengue fever. Of 14,594 participants, 11,387 were definitively diagnosed. Most manifested with a typical non-severe clinical course, and 1.96 % developed to severe dengue. The strains isolated successfully from the serum samples were identified as DENV-1. Genetic analyses revealed that the strains were classified into genotypes I and V of DENV-1, and the dengue epidemic of Guangdong in 2014 was caused by indigenous cases and imported cases from the neighboring Southeast Asian countries of Malaysia and Singapore. Overall, our study is informative and significant to the 2014 dengue outbreak in Guangdong and will provide crucial implications for dengue prevention and control in China and elsewhere.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dengue is an acute systemic viral disease that affects 40 % of the world’s population. An estimated 50–400 million dengue infections occur worldwide annually [1, 2]. Dengue is prevalent in subtropics and tropics, including Southeast Asia, Africa, South America, the Western Pacific Region, and the Eastern Mediterranean Region [3–5]. Dengue virus (DENV) is the causative agent, and it contains five distinct serotypes (DENV-1 to DENV-5), all of which can be transmitted to humans through the bites of infected Aedes mosquitoes [6–8]. Of these, the DENV-5 strain was only detected in Malaysia in 2007, and it has not been reported in any other countries until now [8].
The illness often begins with high-grade fever and is accompanied by headache, retro-orbital pain, skin erythema, body pain, myalgia, arthralgia, and facial flushing [9, 10]. Most patients recover following a self-limiting non-severe clinical course. However, approximately 5 % of dengue cases may develop into a potentially lethal complication called severe dengue, which includes dengue hemorrhagic fever and dengue shock syndrome [11, 12].
In recent decades, dengue outbreaks have been increasing in size and frequency, which have caused a significant health, economic, and social burden in the affected endemic areas [13–15]. Notably, economic development plays an essential role in the global spread of dengue infections, as viremic travelers carry various dengue serotypes and strains to other parts of the world [16, 17]. Thus, dengue has become a worldwide public health problem.
Since June 2014, DENV has caused dengue infections in Guangdong Province, southern China. The number of dengue cases began to increase rapidly in September and the major outbreak began in October, which was believed to be the worst dengue epidemic in China for the past two decades. Following a rapid increase in the number of cases, the government declared a state of emergency. In order to identify dengue patients earlier, detection of DENV non-structural protein 1 (NS1) antigen and RNA were conducted in multiple clinical laboratories. In this study, the epidemiological data of dengue infections in Guangdong from June to December 2014 were reported. Specifically, a total of 14,594 clinically suspected dengue cases from 25 hospitals in the province were studied. Laboratory diagnosis methods were used to confirm the diagnoses. To identify the etiology of the outbreak, a portion of laboratory-positive serum specimens were chosen stochastically for the further DENV serotyping and bioinformatics analyses.
Methods
Epidemiological investigation
Dengue emerged in June 2014 and caused the explosive epidemic that mainly occurred in September and October of that year. At that time, the government and the Provincial Center for Disease Control and Prevention (CDC) began to report the surveillance data regularly. In order to analyze the epidemiology of the dengue outbreak in Guangdong, we collected the surveillance data of dengue infections from the Provincial CDC (http://www.cdcp.org.cn/gdsjbyfkzzx/index.shtml) during the months of June–December 2014.
Case definitions
According to the Chinese national criteria for dengue diagnosis (WS216-2008) and the 2009 World Health Organization (WHO) guidelines [9], a clinically suspected dengue case was defined as follows: (1) The patient lives in or has traveled to a dengue-endemic area; (2) He/she has had a high fever for 3 days or more, accompanied by two of the following criteria: nausea, vomiting, rash, severe headache, muscle and joint pains, or positive tourniquet test; (3) He/she has low or decreasing white cell counts, and/or has thrombocytopenia. In addition, a confirmed dengue case was the suspected case confirmed by a laboratory-positive diagnostic test, including the identifications of DENV RNA and DENV NS1 antigen that are generally performed in the departments of clinical laboratories in Guangdong.
Laboratory diagnostic assays
To better understand the outbreak, 14,594 clinically suspected dengue cases from 25 hospitals in Guangdong were chosen to identify the causative agent. Serum samples were collected from these patients to determine DENV infection at the acute stage of illness, i.e., 1–7 days from symptom onset. After separation from blood cells, the specimens were used to detect DENV NS1 antigen using enzyme-linked immunosorbent assay (ELISA) and/or DENV RNA using a real-time reverse transcriptase polymerase chain reaction (RT-PCR) assay.
The diagnostic kit for DENV NS1 antigen was provided by Beijing Wantai Biology Pharmacy Company Limited. The method was based on the capture of NS1 antigen using a sandwich-type immunoassay and was carried out following the instructions provided by the manufacturer. The samples were classified as positive or negative in accordance with the cut-off point.
Total RNA was extracted according to the manufacturer’s protocol. The detection kit for DENV RNA was provided by DAAN Gene Company Limited of Sun Yat-sen University. Two degenerate oligonucleotide primers and probe corresponding to a stretch of nucleotides conserved in four serotypes of DENV were as follows: primer 1 (forward), 5′-GARAGACCAGAGATCCTGCTGTCT-3′, primer 2 (reverse), 5′-ACCATTCCATTTTCTGGCGTT-3′, and the TaqMan MGB probe 5′-AGCATCATTCCAGGCAC-3′. The test was considered valid if the positive control showed a CT value ≤32. Samples that had a CT value ≤38 were classified as positive and those with value >38 were considered to be negative.
Clinical characteristics
After having been confirmed by laboratory diagnosis, dengue cases were subjected to the further clinical procedure. Clinical records were analyzed by doctors in charge of the cases. Hematological parameters of dengue cases were performed at clinical laboratories by experienced technicians according to standard operating procedures.
Genetic analysis
Among the serum samples collected from 25 hospitals in Guangdong, 93 specimens with the positive detection of DENV RNA were chosen randomly for genetic analysis. The entire sequence of envelope (E) gene (1485 bp) was amplified and used to identify the dengue virus serotype. Four sets of primers were designed for the gene, based on the reference strains of DENV-1, DENV-2, DENV-3, and DENV-4 (Supplementary Table 1). Afterwards, positive PCR products were sequenced for further analyses.
A phylogenetic tree was constructed using the maximum likelihood (ML) method and Kimura’s two-parameter model in order to evaluate the evolutionary relationship between our strains and the others, as executed in MEGA version 5.0 [18]. One thousand replicates were carried out in a bootstrap test in order to access the reliability of each branch topology. The reference sequences of the DENV envelope gene derived from the GenBank database are exhibited in Supplementary Table 2.
Ethics statement
Informed written consent was obtained from all patients or their guardians. The research protocol was approved by the Ethics Committees of the survey hospitals.
Results
Epidemiology
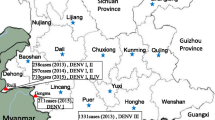
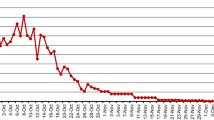
Up to December 15, 2014, epidemic dengue had spread to 20 cities, including Guangzhou, Foshan, Jiangmen, Zhongshan, and Shenzhen (Fig. 1a). From September 4 to October 18, 2014, the infection total increased dramatically, with an average of more than 800 cases being reported daily (Fig. 1b). Additionally, 45,171 cases had been recorded in total by December 15, 2014. Of these, 101 were foreign travelers. Guangzhou contributed 82.7 % (37,354 infections) of confirmed dengue patients. Foshan, a city about 20 km from Guangzhou, bore 7.8 % (3542 infections) of this burden, followed by Zhongshan with 1.5 % (679 infections). In addition, six patients died from the illness, among which five individuals were from Guangzhou and one was from Foshan.
Epidemiological data of the dengue outbreak in Guangdong Province, People’s Republic of China. a Geographic distribution of the dengue outbreak in Guangdong. Epidemic dengue had spread to 20 cities and a total of 45,171 confirmed dengue cases had been recorded in Guangdong (southern China) by December 15, 2014. The number of cases was labeled in the corresponding city and shown using different colors. b The number of confirmed dengue cases by different time points during the dengue outbreak. The dengue epidemic of Guangdong occurred in June 2014 and caused the outbreak in September 2014. The surveillance data has been reported regularly by the Provincial CDC since September 4, 2014
Laboratory diagnostic tests
Serum samples from the 14,594 suspected dengue cases were tested for DENV NS1 antigen and/or viral RNA, respectively. Consequently, 11,387 dengue patients were confirmed by laboratory diagnosis (Fig. S1 and Table 1). In the experiment, 3565 paired serum samples were collected to detect both DENV NS1 and RNA during the acute stage of illness. The concordance between these two tests was 99.69 % (Table 2).
Clinical features
The clinical characteristics of the 11,387 confirmed dengue cases in the acute illness stage are shown in Table 3. All patients typically developed fever in our study. The acute febrile phase lasted 1–15 days, with an average of 5.17 days, with irregular fever (81.30 %) as the most commonly presented symptom. Headache was the second most frequently reported clinical sign (90.88 %). Notably, 4.27 % of the patients needed hospital care and 1.96 % progressed to severe dengue.
Moreover, abnormality in the full blood count was observed in most patients at 1–7 days after the onset of symptoms (Table 4). Over half of the cases (68.5 %) had a decrease in the total white blood cell count, with a minimum of 0.9 × 109/L. Thrombocytopenia was also observed in 84.5 % of the patients. Additionally, the majority of patients suffered from hepatic lesions.
Genetic analysis
Among 93 specimens from 25 hospitals in Guangdong, 91 were amplified and sequenced successfully, all of which were identified as DENV-1. The strains isolated in our study were named Guangzhou14-02 (GZ14-02), Guangzhou14-36 (GZ14-36), and Guangzhou14-55 (GZ14-55). The GenBank accession numbers were KP185303, KT819303, and KT819304, respectively.
Phylogenetic analysis was performed using 67 DENV-1 E gene sequences, including three strains identified in our research and 64 global DENV-1 strains derived from GenBank. Two strains identified in our paper, GZ14-02 and GZ14-36, were subclassified into genotype I, and they were further divided into two related clusters in the phylogenetic ML tree. The strain GZ14-02 was nearly identical to the strains from Guangdong (GZ/25559, GD-D13187, GD-D13012) in the 2013 epidemic, with a bootstrap (BS) value of 99. Furthermore, the strain GZ14-36 was closely related to the strain GZ/ZW-0802 also from Guangdong in the 2014 epidemic, and both of them were clustered together with the isolate of Malaysia in 2008. Moreover, our strain GZ14-55 and the strain CN/GZ37, which was also collected in Guangdong in 2014, were clustered together and fell into genotype V. Additionally, the strain 07354Y14 from Singapore in the same year had strong relevance to these two strains in phylogeny, with a BS value of 98.
Discussion
Guangdong has battled the worst outbreak of dengue over the past 20 years. Notably, Guangzhou contributed more than 80.0 % of confirmed dengue patients. There were several factors contributing to the outbreak in this city. Firstly, climate change plays a critical role in the spatial and temporal distribution of dengue [19–21]. Guangzhou is located the south of the Tropic of Cancer and has a humid subtropical climate influenced by the East Asian monsoon. In September 2014, it had heavy bouts of rainfall alternating with high temperatures, which could have permitted the population of mosquitoes to increase and, thereafter, caused an upsurge in dengue [22]. Secondly, Guangzhou, as the capital and largest city of Guangdong Province, possesses a dense population and a high degree of urbanization. The large-scale population migration may directly influence the epidemiology of dengue infections. Thirdly, Guangzhou is located in the geometrical center of the economic circle of Southeast Asia. In this year, a sharp increase in the incidence of dengue was also reported in most of the Southeast Asian countries. Consequently, travelers may have moved the virus rapidly within and between countries.
In this report, our investigation covered 14,594 clinically suspected dengue patients from Guangzhou and its neighboring cities during a major dengue outbreak in 2014. Among them, 11,387 cases were diagnosed with dengue confirmed by laboratory diagnostic tests. Besides, 93 laboratory-positive sera of dengue patients were chosen to analyze the serotype of DENV. The results showed that 91 out of 93 specimens were identified as DENV-1 (named GZ14-02, GZ14-36, GZ14-55), which elucidated that the dengue epidemic in Guangdong was led by the predominant DENV-1 serotype. During the past three decades, outbreaks of all four serotypes of DENV have been reported in Guangdong Province [23, 24]. Nevertheless, DENV-1 was still the most prevalent serotype in Guangdong in recent years, and led to multiple serious epidemics during the years 1995–2010 [25, 26]. It was reported that a patient infected by DENV-1 had a much lower risk of severe dengue in comparison with the other three serotypes [27, 28]. Our speculation was that DENV-1 was the predominant serotype circulating in Guangdong since 2002 and, therefore, for a new circulating serotype, individuals were more susceptible to infection and progress to a serious condition [29, 30]. Previous studies have revealed that secondary infection with different DENV serotypes carried with it a risk of developing severe dengue symptoms [31–33]. Infection with one serotype can stimulate an effective immunity against reinfection with the same serotype; however, cross-reactive and non-neutralizing antibodies from the previous DENV infection can also bind to a new infecting serotype and facilitate virus entry into susceptible cells. It was a phenomenon called antibody-dependent enhancement of infection (ADE), considered as the most rational explanation for severe dengue [34, 35]. This can also help explain that most of the dengue cases in the present investigation manifested typical dengue fever rather than severe forms of the illness.
In the phylogenetic analysis based on the complete E gene sequences of DENV-1 isolated from different epidemic regions (Fig. 2), our identified strains GZ14-02 and GZ14-36 were subclassified into DENV-1 genotype I, which was one of the most prevalent genotypes in Southeast Asian countries [23, 36]. Additionally, the strain GZ14-55 was grouped into genotype V. Furthermore, the strain GZ14-02 was nearly identical to the isolates from the 2013 epidemic of Guangdong, which indicated that the strain isolated in 2014 possibly originated from the isolates of 2013. That is to say, the virus probably circulated locally and eventually caused the dengue outbreak of 2014. Besides, GZ14-02 and the 2013 Guangdong isolates appeared to share a common ancestral lineage with the 2009 Guangdong endemic strain (GZ11562). Thus, they may evolve from a common ancestor originating from the same place. It was reasonable to propose that endemic infection of dengue circulating locally played a crucial role in causing the dengue epidemic in Guangdong in 2014.
Phylogenetic tree. ML tree of DENV-1 E gene sequences (1485 bp) was constructed with Kimura’s two-parameter model using MEGA 5.0 software. It included three strains collected in this study and 64 global strains. Each strain was named according to the format strain name/country/collection year. The strains identified in this study were labeled with a black dot. Bootstrap values were set for 1000 repetitions. The sequences of reference strains were obtained from GenBank
Notably, the strain GZ14-55 reported here and the other strain CN/GZ37 also collected from the 2014 epidemic in Guangdong formed a close branch, along with the strain from the 2014 epidemic of Singapore. In addition, the isolate GZ14-36 was clustered together with the strain from Malaysia in 2008. These data suggested that the 2014 dengue outbreak in Guangdong might also be caused by imported infections from the neighboring Southeast Asian countries of Singapore and Malaysia. Because of the special geographic location of Guangdong, being surrounded by a large number of dengue-endemic countries, the epidemics from Southeast Asian countries had an important impact on the dengue outbreak in southern China. Wang et al. indicated that the dengue outbreak of Yunnan in 2013 was caused by a newly imported infection from the neighboring country of Myanmar [36]. Wu et al. also found that dengue epidemics in Guangdong were closely associated with those in Southeast Asian countries during the years 1978–2006, such as the Philippines, Indonesia, and Thailand [23].
All of these data revealed that the 2014 dengue epidemic in Guangdong might be triggered by indigenous dengue cases as well as imported cases from the neighboring countries of Singapore and Malaysia. During the past 30 years, imported-cases-induced endemic prevalence and endogenous epidemic outbreak with natural epidemic focuses were believed to be two important modes present in Southern China [23, 25, 37, 38]. They were further validated in the current study.
In addition, we must admit some limitations of our work. Firstly, the number of serum samples used for the genetic analysis may be not sufficient to detect the imported cases and the other serotypes of DENV. Thus, studies with a larger sample size are needed to validate our results. Secondly, the sensitivity of dengue-specific laboratory tests was impossibly 100 % in the acute stage of illness. In other words, the occurrence of omission of the diagnosis of dengue cases was present.
Despite these limitations, our study was still informative and significant to the most current dengue outbreak in Guangdong. Moreover, it may provide crucial implications for dengue prevention and control. Relevant surveillance programs should continue to be strengthened, such as vector control and surveillance, epidemiological surveillance, clinical diagnosis and management, as well as environmental health. Early warnings of an epidemic will help inhabitants increase self-awareness of protection against dengue, and also help clinicians efficiently diagnose and properly treat dengue cases, thereby reducing transmission and improving clinical outcomes.
References
Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL et al (2013) The global distribution and burden of dengue. Nature 496:504–507. doi:10.1038/nature12060
Guzman MG, Halstead SB, Artsob H, Buchy P, Farrar J, Gubler DJ et al (2010) Dengue: a continuing global threat. Nat Rev Microbiol 8:S7–S16. doi:10.1038/nrmicro2460
Whitehead SS, Blaney JE, Durbin AP, Murphy BR (2007) Prospects for a dengue virus vaccine. Nat Rev Microbiol 5:518–528. doi:10.1038/nrmicro1690
Arima Y, Edelstein ZR, Han HK, Matsui T (2013) Epidemiologic update on the dengue situation in the Western Pacific Region, 2011. Western Pac Surveill Response J 4:47–54. doi:10.5365/WPSAR.2012.3.4.019
Mackey TK, Liang BA (2012) Threats from emerging and re-emerging neglected tropical diseases (NTDs). Infect Ecol Epidemiol 2:75–88. doi:10.3402/iee.v2i0.18667
de Alwis R, Williams KL, Schmid MA, Lai C-Y, Patel B, Smith SA et al (2014) Dengue viruses are enhanced by distinct populations of serotype cross-reactive antibodies in human immune sera. PLoS Pathog 10:e1004386. doi:10.1371/journal.ppat.1004386
Holmes EC, Twiddy SS (2003) The origin, emergence and evolutionary genetics of dengue virus. Infect Genet Evol 3:19–28. doi:10.1016/S1567-1348(03)00004-2
Normile D (2013) Tropical medicine. Surprising new dengue virus throws a spanner in disease control efforts. Science 342:415. doi:10.1126/science.342.6157.415
World Health Organization (WHO) (2009) Dengue: guidelines for diagnosis, treatment, prevention and control—new edition. 2009. WHO, Geneva
Senaratne TN, Noordeen F (2014) Diagnosis of dengue in Sri Lanka: improvements to the existing state of the art in the island. Trans R Soc Trop Med Hyg 108:685–691. doi:10.1093/trstmh/tru131
Martina BEE, Koraka P, Osterhaus ADME (2009) Dengue virus pathogenesis: an integrated view. Clin Microbiol Rev 22:564–581. doi:10.1128/CMR.00035-09
Pagliari C, Quaresma JAS, Fernandes ER, Stegun FW, Brasil RA, de Andrade HF et al (2014) Immunopathogenesis of dengue hemorrhagic fever: contribution to the study of human liver lesions. J Med Virol 86:1193–1197. doi:10.1002/jmv.23758
Shepard DS, Undurraga EA, Halasa YA (2013) Economic and disease burden of dengue in Southeast Asia. PLoS Negl Trop Dis 7:e2055. doi:10.1371/journal.pntd.0002055
Young JW, Gibbons RV, Rothman AL, Tannitisupawong D, Srikiatkhachorn A, Jarman RG et al (2014) Dengue severity and blood group. Dengue Bulletin (in press)
Undurraga EA, Betancourt-Cravioto M, Ramos-Castañeda J, Martínez-Vega R, Méndez-Galván J, Gubler DJ et al (2015) Economic and disease burden of dengue in Mexico. PLoS Negl Trop Dis 9:e0003547. doi:10.1371/journal.pntd.0003547
Ninphanomchai S, Chansang C, Hii YL, Rocklöv J, Kittayapong P (2014) Predictiveness of disease risk in a global outreach tourist setting in Thailand using meteorological data and vector-borne disease incidences. Int J Environ Res Public Health 11:10694–10709. doi:10.3390/ijerph111010694
Murray NEA, Quam MB, Wilder-Smith A (2013) Epidemiology of dengue: past, present and future prospects. Clin Epidemiol 5:299–309. doi:10.2147/CLEP.S34440
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Barclay E (2008) Is climate change affecting dengue in the Americas? Lancet 371:973–974
Russell RC, Currie BJ, Lindsay MD, Mackenzie JS, Ritchie SA, Whelan PI (2009) Dengue and climate change in Australia: predictions for the future should incorporate knowledge from the past. Med J Aust 190:265–268
Colón-González FJ, Fezzi C, Lake IR, Hunter PR (2013) The effects of weather and climate change on dengue. PLoS Negl Trop Dis 7:e2503. doi:10.1371/journal.pntd.0002503
Xiong Y, Chen Q (2014) Epidemiology of dengue fever in China since 1978. Nan Fang Yi Ke Da Xue Xue Bao 34:1822–1825
Wu W, Bai Z, Zhou H, Tu Z, Fang M, Tang B et al (2011) Molecular epidemiology of dengue viruses in southern China from 1978 to 2006. Virol J 8:322. doi:10.1186/1743-422X-8-322
Wu Y, Zheng X, Wu Z (2014) Dengue fever in China. In: Treatment of human parasitosis in traditional Chinese medicine. Springer Berlin Heidelberg, Berlin, pp 239–253. doi:10.1007/978-3-642-39824-7_15
Guo R-N, Lin J-Y, Li L-H, Ke C-W, He J-F, Zhong H-J et al (2014) The prevalence and endemic nature of dengue infections in Guangdong, South China: an epidemiological, serological, and etiological study from 2005–2011. PLoS One 9:e85596. doi:10.1371/journal.pone.0085596
Luo L, Liang HY, Hu YS, Liu WJ, Wang YL, Jing QL et al (2012) Epidemiological, virological, and entomological characteristics of dengue from 1978 to 2009 in Guangzhou, China. J Vector Ecol 37:230–240. doi:10.1111/j.1948-7134.2012.00221.x
Liang H, Luo L, Yang Z, Di B, Bai Z, He P et al (2013) Re-emergence of dengue virus type 3 in canton, China, 2009–2010, associated with multiple introductions through different geographical routes. PLoS One 8:e55353. doi:10.1371/journal.pone.0055353
Ocazionez RE, Cortés FM, Villar LA, Gómez SY (2006) Temporal distribution of dengue virus serotypes in Colombian endemic area and dengue incidence. Re-introduction of dengue-3 associated to mild febrile illness and primary infection. Mem Inst Oswaldo Cruz 101:725–731. doi:10.1590/S0074-02762006000700004
Xu G, Dong H, Shi N, Liu S, Zhou A, Cheng Z et al (2007) An outbreak of dengue virus serotype 1 infection in Cixi, Ningbo, People’s Republic of China, 2004, associated with a traveler from Thailand and high density of Aedes albopictus. Am J Trop Med Hyg 76:1182–1188
Wu J-Y, Lun Z-R, James AA, Chen X-G (2010) Dengue fever in mainland China. Am J Trop Med Hyg 83:664–671. doi:10.4269/ajtmh.2010.09-0755
Rothman AL (2004) Dengue: defining protective versus pathologic immunity. J Clin Invest 113:946–951. doi:10.1172/JCI21512
Vaughn DW, Green S, Kalayanarooj S, Innis BL, Nimmannitya S, Suntayakorn S et al (2000) Dengue viremia titer, antibody response pattern, and virus serotype correlate with disease severity. J Infect Dis 181:2–9. doi:10.1086/315215
Kliks SC, Nimmanitya S, Nisalak A, Burke DS (1988) Evidence that maternal dengue antibodies are important in the development of dengue hemorrhagic fever in infants. Am J Trop Med Hyg 38:411–419
Simmons CP, Chau TNB, Thuy TT, Tuan NM, Hoang DM, Thien NT et al (2007) Maternal antibody and viral factors in the pathogenesis of dengue virus in infants. J Infect Dis 196:416–424. doi:10.1086/519170
Wang Y, Si L, Luo Y, Guo X, Zhou J, Fang D et al (2015) Replacement of pr gene with Japanese encephalitis virus pr using reverse genetics reduces antibody-dependent enhancement of dengue virus 2 infection. Appl Microbiol Biotechnol 22:9685–9698. doi:10.1007/s00253-015-6819-3
Wang B, Li Y, Feng Y, Zhou H, Liang Y, Dai J et al (2015) Phylogenetic analysis of dengue virus reveals the high relatedness between imported and local strains during the 2013 dengue outbreak in Yunnan, China: a retrospective analysis. BMC Infect Dis 15:142. doi:10.1186/s12879-015-0908-x
He J-F, Luo H-M, Liang W-J, Zheng K, Kang M, Liu L-P (2007) Epidemic situation of dengue fever in Guangdong province, China, 1990–2005. WHO Regional Office for South-East Asia, New Delhi
Sun J, Wu D, Zhou H, Zhang H, Guan D, He X et al (2015) The epidemiological characteristics and genetic diversity of dengue virus during the third largest historical outbreak of dengue in Guangdong, China, in 2014. J Infect. doi:10.1016/j.jinf.2015.10.007
Acknowledgments
We thank all the participants who consented to take part in this study, as well as the doctors and nurses for recruitment. Especially, we sincerely thank the laboratory technicians from the 25 hospitals who helped us collect and interpret the data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None declared.
Funding
This work was supported by the Guangdong Natural Science Foundation (grant number S2013010014007, 2014A030313070) and Guangdong Province Science & Technology Project Plan & Social Development Foundation (grant number 2010A030400006).
Additional information
Lisi Huang and Xiaohong Luo contributed equally to this work.
Rights and permissions
About this article
Cite this article
Huang, L., Luo, X., Shao, J. et al. Epidemiology and characteristics of the dengue outbreak in Guangdong, Southern China, in 2014. Eur J Clin Microbiol Infect Dis 35, 269–277 (2016). https://doi.org/10.1007/s10096-015-2540-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2540-5