Abstract
Alzheimer disease (AD) is a neurodegenerative process, one of the most common and incident dementia in the population over 60 years. AD manifests the presence of complex biochemical processes involved in neuronal degeneration, such as the formation of senile plaques containing amyloid-β peptides, the development of intracellular neurofibrillary tangles, and the suppression of the acetylcholine neurotransmitter. In this way, we performed a set of theoretical tests of tacrine ligand and acetylcholine neurotransmitter against the human acetylcholinesterase enzyme. Molecular docking was used to understand the most important interactions of these molecules with the enzyme. Computational chemistry calculation was carried out using MP2, DFT, and semi-empirical methods, starting from molecular docking structures. We have also performed studies regarding the non-covalent interactions, electron localization function, molecular electrostatic potential and explicit water molecule influence. For Trp86 residue, we show two main interactions in accordance to the results of the literature for TcAChE. First, intermolecular interactions of the cation-π and sigma-π type were found. Second, close stacking interactions were stablished between THA+ and Trp86 residue on one side and with Tyr337 residue on the other side.
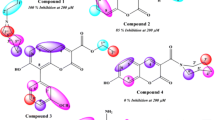
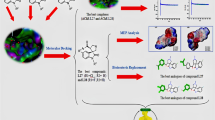
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Besides being one of the most common dementia in the elderly people, Alzheimer disease (AD) is a neurodegenerative process characterized by a progressive loss of cognitive capabilities [1, 2]. However, its cause has not been clearly understood, and it is known that the most alarming risk factor is the presence of the gene mutations in the immunomodulatory glycosylated protein apolipoprotein E (apoE), and considering that only mutations in the ε4 allele have a high probability of developing AD [3,4,5]. Furthermore, environmental factors and health conditions have been related to the manifestation of the disease, only treatable at the palliative level to decrease the harmful effects that cause the cognitive and depressive states of patients [6, 7].
In addition to the progressive loss of cognitive abilities, what is observed in AD patients is the presence of biochemical complex involved in neuronal degeneration, such as the formation of senile plaques containing amyloid-β, the development of intracellular neurofibrillary tangles, yielding to the suppression of the neurotransmitter acetylcholine (ACh). In other words, a decrease in ACh concentration gradient, which is responsible for the transmission of nerve impulses in synaptic regions. This decrease justifies the deleterious effects and induces the patient to massive loss of nervous system capacity and abilities [6, 8].
ACh is a neurotransmitter produced in the central and peripheral nervous systems. Its main physiological effect is the reduction of permeability to potassium ions. ACh binds to subtypes of brain receptors, the nicotinic receptors (nAChRs), composed by transmembrane ion channels, and the muscarinic receptors (mAChRs) that form G-protein receptors. Current studies also suggest that the nicotinic receptor is implicated in the AD evolution and the muscarinic receptors, such as muscarinic M1 postsynaptic and M2 presynaptic [6, 9,10,11,12].
All ACh released from presynaptic vesicles is hydrolyzed, after the synapse process, by the acetylcholinesterase (AChE) enzyme, an allosteric enzyme of the hydrolases class. It is located anchored to the pos-synaptic membrane, with a highly specific biological function, consisting of hydrolyzing the ACh in the synaptic clefts, releasing acetate and choline, in addition to a water molecule regenerating the free enzyme and the acetate [8, 13].
The treatment of patients with AD is a challenge. In this field, the most successful approach for the symptomatic treatment of AD is the cholinergic therapy, which consists in the use of acetylcholinesterase inhibitors (AChEIs). These drugs act competitively and non-competitively with the neurotransmitter and with inhibitory action against the acetylcholinesterase enzyme. This class of inhibitors blocks the hydrolysis of ACh in the synaptic region. Besides AChE and amyloid-β plaque, AD is a multifactorial disease, and the search for drugs capable to modulate more than one target proteins is a prominence [14,15,16,17]. Several recent computational studies, using molecular docking, molecular dynamics, and other techniques, have aided valuable insights to the discovery and development of new potential candidates for these target proteins using natural compounds [18,19,20,21,22,23,24,25,26].
One of the main characteristics of AChEIs is their interaction with the peripheral anionic binding site (PAS), as well as their molecular recognition in the catalytic active site (CAS) of the enzyme composed by the catalytic triad. This triad is formed by one serine residue, commonly used as a nucleophile, which allows the formation of covalent bonds with its hydroxyl, one histidine residue, and one glutamate residue (Ser203, Glu334, and His447) [27]. Some AChEIs present a neuroprotective function by acting negatively on the hyperphosphorylation of amyloid-β protein. Thus, inhibiting its cleavage and consequently forming the amyloid plaques [6, 13]. The elimination of the amyloid plaques has long been a leading strategy in Alzheimer's drug development.
Tacrine (THA) or 1,2,3,4-tetrahydroacridine molecule is a pioneer of AChE inhibitors approved by the FDA (Food and Drug Administration) to treat AD. It is classified as a reversible, non-competitive, and non-selective inhibitor of AChE with moderately long action [27,28,29,30,31]. Several in vitro studies have shown that THA produces an allosteric inhibition of acetylcholinesterase by binding to the anionic site, which is composed by amino acids with hydrophobic side chains [27, 32,33,34]. However, the drug presents high hepatotoxicity, slow pharmacokinetics, and severe collateral effects. Therefore, many efforts have been conducted to develop analogs to THA that can improve the inhibition of the AChE, and presents safer pharmacological properties against inhibition of the AChE mechanism of action [8, 35,36,37,38,39,40,41,42,43]. Some THA ligands presented high selective action against AChE and the aggregation of amyloid-β peptide inside the neurons [44,45,46].
Docking studies of THA with AChE have evidenced that the THA molecule is recognized at the active site of the enzyme from its anionic, peripherical, and catalytic site, through hydrophobic interactions, hydrogen bond with the histidine from the catalytic triad, and the formation of a charge-transfer complex with the Trp86 residue [39, 45]. Electronic structure studies of THA using the frontier molecular orbital (FMO) compared to other AChE inhibitors were applied to understand the main electronic descriptors of this inhibitor [8].
Computational chemistry methods have been widely employed to characterize the AChE enzyme mechanism and the physicochemical parameters that modulate the inhibitory potency of AChE inhibitors [36, 42]. Therefore, this work is based on theoretical chemistry and aims to identify the binding profile of ACh, and tacrine with the human acetylcholinesterase (hAChE) enzyme, to correlate their physicochemical properties, from the electronic structure and molecular docking study of the human AChE enzyme. The study of ligand properties is essential to highlight some important molecular insights through electronic structure associated with receptor-ligand interaction profiles.
Computational details
The geometry structure optimizations of the ligands were performed in vacuum and solvent, by means of implicit integral equation formalism of polarizable continuum model (IEFPCM) solvent method considering the water as solvent. For this purpose, the second-order MP2 perturbational method was carried out with the 6–311+G (d,p) and aug-cc-pVDZ basis sets. MP2 level using the 6–311+G (d,p) and aug-cc-pVDZ basis sets configures the best choice for electronic structure with reasonable computational effort [47,48,49,50,51]. All electronic structure calculations were performed using Gaussian09 computational program [52]. Furthermore, vibrational frequencies were calculated to characterize the equilibrium geometry. Properties as the frontier molecular orbitals, the most negative heteroatom charges, dipole moments, the gap between the highest occupied molecular orbital (HOMO) and the lowest unoccupied molecular orbital (LUMO), the molecular size, molecular electrostatic potential (MEP) map, and distance between its most acid hydrogens were analyzed.
Regarding the 3D structure of the protein used to perform the docking study, we selected the most recent crystallized structure of hAChE, complexed with tacrine molecule. The structure was obtained from the Protein Data Bank (PDB) database, deposited under the code 6O4X with a resolution of 2.30 Å. The pKa and ionization state of carboxyl and amino groups of the catalytic triad residues, Glu202, Glu334, and His447 were analyzed. The protonation state study of the enzyme residues was performed using the ProPka 3.0 program [53].
Two possible charged states were considered for THA. The first state considered the molecule in the neutral state (THA). The second took the tacrine as protonated structure (THA+) while adding a hydrogen atom to the nitrogen (N1) from its pyridine ring. Figure 1 shows the structure of ACh neurotransmitter, THA, and the protonated THA (THA+).
Molecular docking is an in silico simulation procedure used to predict the best likely conformation of a receptor-ligand complex. Each program uses one or more specific search algorithms to predict the conformations of a binary complex, thus identifying potential new inhibitors against a target of interest.
All the water molecules of hAChE of 6O4X structure and the other molecules as stabilizers and ligands were removed. Molecular docking was used to map the main interactions of the amino acids that interact between the active site of the enzyme and its natural ACh, and the THA and THA+ inhibitors. The ligands were docked in the protein considering their conformations obtained with the implicit solvent and vacuum calculations. The simulations were carried out using the AutoDock 4.2.6 program [54] and the Visual Molecular Dynamics (VMD) software [55].
The docked area (binding space) was located in the three-dimensional space of the residues from the active site around the catalytic triad Ser203, Glu334, and His447 residues. Lamarckian Genetic Annealing algorithm (LGA) was used to optimize the docking parameters. The number of genetic algorithm solutions was set to 100, while the number of energy evaluations was set to 5.0 million, and the population size to 150. In the set of 100 solutions for each molecule, the conformation with the most negative energy score and the high number of solutions in the cluster analysis was chosen. This procedure was carried out since these parameters indicate the most stable ligand-receptor complex structures. Furthermore, we used the geometry of the crystallized tacrine inhibitor to validate the results of the docking protocol, which were visualized using the Discovery Studio Visualizer software [56].
To validate our docking procedure, we performed a redocking study using the coordinates of the hAChE coming from the PDB 6O4X, and the THA ligand (THA) included in it. The RMSD between the original conformation and the best pose conformation of the ligand incoming from redocking study was 0.05 Å. The docking and redocking protocols were the same as described previously.
Most of the chemical interactions between a protein and the drugs are dominated by non-covalent interactions, encompassing a wide range of interaction energies, e.g., hydrogen bonding, and van der Waals interactions (dipole–dipole force, and London dispersion). Starting from the optimized three-dimensional structure of the ligands and docking results, we performed the non-covalent interaction (NCI) study. This analysis was developed to study noncovalent interaction in real space, based on the electron density and its derivatives. Comparisons with Quantum Theory of Atoms in Molecule (QTAIM) theory have shown that reduced density gradient (RDG) is an efficient method to support noncovalent interactions study [57]. NCI is based on the concept that non-covalent interactions are characterized by low density and low gradient values and may be localized by generating isosurfaces involving the regions corresponding to the real space [58, 59]. In this perspective, it is possible to analyze the contribution of residues participating in the catalytic triad of hAChE. The Multiwfn program [60] is a multifunctional and flexible wavefunction analyzer, and was used to perform the NCI study from the MP2 wave functions generated in the electronic structure optimization calculations. NCI by means of promolecular density approximation between the main docking residues of the hAChE was studied using the PDB coordinates of the complex. These residues interact with the ligands in order to better understand the interactions between the ligands and the residues of the CAS [60].
The electron localization function (ELF) introduced by Becke and Edgecombe describes how much the Pauli repulsion is efficient at a given point of the molecular space, i.e., it describes the electron density near the atomic nucleus [61]. In this sense, we used the three-dimensional structures of the ligands optimized previously, and through the multifunctional Multiwfn program, the ELF isosurfaces were built. In this way, we achieved a better understanding of the electronic structure of the ligand against the enzyme.
Water molecules play an essential role in biological systems. Therefore, single-point energy calculations were carried out to understand water molecule effects on the CAS activity during the acylation process of ACh and the inhibitory process of THA+. The structures used to study water effects were the following: the PDB 6O4X structure of the best pose of the ligands from docking studies and the main residues shown in docking calculations, including five structural water molecules that are very near the Trp86, Glu202, Ser203, and His447 residues of the CAS. We performed the single-point energy calculations at the B3LYP/6–31+G(d,p) level. For the ACh ad THA+ ligands, a full geometry optimization was performed (with redundant coordinates and frequency calculation, in a vacuum and implicit PCM solvated method using water as solvent) at the B3LYP/6–31+G(d,p).
Results and discussion
Electronic structure
Table 1 shows the structural parameters of ACh, THA, and THA+, where the root mean square deviation (RMSD) was calculated using the PDB structure as the reference. In general, the geometry obtained in solution model does not show a significant difference compared to the vacuum for size, RMSD, and volume properties. The exception is the RMSD of THA+ calculated at MP2/6–311+G(d,p) due to a dihedral angle in relation to the amine group N2 atom (Fig. 1).
THA is a condensed ring molecule, and ACh is a small neurotransmitter produced in the presynaptic myeline neurons. Due to their molecular size and volume, both can access the CAS and interact properly with the hAChE pocket main residues. Despite the small range of molecule size, the distance between the most acidic hydrogens (H1-H2 distance) varied in the range of 1.66 and 8.23 Å, and is justified by the fact that both ligands have distinct hydrogens corresponding to the H–H distance. In the case of THA+, the H–H distance is large due to the most acid hydrogens involved, which are are in both sides of the ring (see Figure S1 of SI for the corresponding distances).
Figure 2 shows the electronic parameters of ACh, THA, and THA+. The gap values show negligible deviation between vacuum and solvent, mainly for THA and THA+ molecules. It is important to note that water molecule only affects the Gorge (the enzyme active site) entrance, but not for its catalysis [62]. In other words, hydrophobic inhibitors are better evaluated for the interaction with the residues of the CAS of the hAChE protein. For the calculations using solvent or vacuum results, the ChelpG charge observed suggests a higher molecule polarization. The carbon atoms of the pyridine ring are more positive and better candidates for the nucleophilic attack, which is necessary for the interaction with the catalytic triad of the hAChE (Figure S1 and S2 of supplementary information). As expected, the ACh structure has a polarization in the acetoxyethyl moiety, where the carbon (C9) atom of the carboxyl group has the most positive ChelpG charge (1.229 in vacuum; 0.959 in solution), which is in accordance with the nucleophilic attack of this carbon center (Figure S2 of supplementary information).
Comparing the charges of the heteroatoms obtained in vacuum and implicit solvent model calculations, the distribution presents a small deviation, decreasing slightly in the absolute value, becoming less negative. Specifically, for ACh molecule, the heteroatom charge becomes less negative when using the solvent implicit model, which is a suggestion of how this ligand has a good affinity with water. The dipole moment presents the same significant variation, its value increasing for the solvated ligand.
Likewise, neutral THA presents acceptor regions close to the nitrogen of the pyridine ring (Figure S2), justifying the addition of a proton in this region to form the THA+ ligand. On the other hand, THA + showed similarity to ACh electrostatic potential map, which presented its two nitrogen atoms as the most acidic hydrogen donors for the nucleophilic attack. In relation to the MEP (Figure S2), it is observed a strong polarization shown by ACh in contrast to THA+, which in consequence made its proton available to interact with His447. In the case of ACh, the positive distribution region turns the hAChE substrate able to be recognized by Asp74 residue in the entrance of the GORGE active site. Furthermore, the same explanation can be applied to THA+, which has a significant positive electronic density distribution and is probably easy to be recognized by the Asp74 residue.
The FMOs are useful tools to study the reactivity of molecules. Figure 3 shows the FMO of optimized structures obtained at MP2 level, with and without solvent. The ligands present the same trend for vacuum and solvent for both basis set used. The ACh and THA+ ligands show almost the same energy values (approximately − 1.20 eV) for the LUMO series without solvent model. However, the analysis of Ach and THA+ using implicit solvent model leads the LUMO energies to higher values close to + 2.00 eV. This trend of higher LUMO energy values is probably assigned relevant for discussing the reactivity of THA+. Solvent model shows no effect on THA HOMO-n (n = 0–2) orbital energy values, while THA \+ has the same trend of ACh with a difference between solvent and vacuum between 2.35 and 3.30 eV for both basis set.
Molecular docking simulation
The molecular docking study was carried out using the human AChE protein (PDB: 6O4X) in a complex with the THA classified as a low nanomolar AChE inhibitor [63, 64]. Figure 4 shows the open active site used to study the interaction of ACh, THA, and THA+.
Table 2 presents the docking results, i.e., the score (binding energy), theoretical Ki, and the most favorable interactions from the best solution obtained based on the LGA study. The THA+ inhibitory activity from the corresponding eeAChE (human AChE from erythrocytes) has the IC50 value of 0.5 ± 0.1 µM [63]. Our theoretical Ki was estimated at 1.62 µM in good accordance with the experimental value.
Considering the best docking conformation of natural substrate ACh, the interaction types between the ACh and the residues of its active site are depicted in Fig. 5. The interaction profile is composed mainly by hydrogen interactions between the ligand and the Trp86 and Trp439 residues, located at the quaternary ammonium binding site (at the distance of 2.10 and 2.15 Å, respectively). The intermolecular interactions of the cation-π and sigma-π were observed with the Trp86 residue. This kind of interaction is equivalent to the cation-π interaction between the methyl group and the condensed aromatic rings of Trp84 residue in the Torpedo Californica (TcAChE) [36]. The attractive charge interaction between the quaternary nitrogen and the Glu202, His447, and Tyr337 residues act as a counterion to stabilize the positive charge of ACh [36].
The neutral form of tacrine (Fig. 6) presented the π-stacked interaction with Trp86 and Tyr337 (4.04 to 4.84 Å) residues. In addition, van der Waals interactions with the residues Gly82 and Thr83 were observed with THA neutral form. These interactions suggest that the THA+ fits better in hAChE inhibition than THA. This is probably due to the protonated conformation of THA+ ligand that takes into account the ligand charge at its physiological pH and has the most significant favorable interactions for the inhibition mechanism.
On the other hand, THA+ (Fig. 7) presents important interactions at hAChE peripheral site residues by interactions of van der Waals type with residues Glu202, Tyr449, Thr83, Ser203, and Gly82. The interaction at the catalytic site of the enzyme was done mainly by hydrogen bonds. First, the His447 residue (2.07 Å) corresponds to the interaction with the proton added to the ligand. The second is between one of the acidic hydrogens and Tyr337 residue. More precisely, binding was observed at the hAChE active site GORGE base, establishing close stacking interactions between the tricyclic aromatic ring and Trp86 residue on one side and with Tyr337 residue on the other side. This interaction is known as the sandwich p-stacking-interactions, in the range of 3.81 to 4.71 Å, respectively [64]. These are similar interactions between the residues Trp84 and Phe330 found in TcAChE [64].
NCI and ELF study
The NCI analysis comprehends the interactions of different binding energies from hydrogen bonding, dipole–dipole, and London dispersion [58]. We studied the non-covalent interactions of the THA, THA+, and ACh, in vacuum and in implicit solvent. Moreover, ELF was used since it allows mapping the attracting sites of electron pairs.
The NCI and ELF results for free ACh molecule at MP2/aug-cc-pVDZ level are shown in Figures S3–S5 of supplementary material. The promolecular NCI isosurfaces and the RDG of ACh, THA, and THA + complexed with residues of protein obtained from the docking study are shown in Figure S5. We observed the presence of attractive interaction regions on ACh between the methyl groups of the quaternary nitrogen and the residues of His447, Trp86, and Tyr337, which correspond to the interactions found in the docking study. The HOMO orbital distribution (Fig. 3) shows an agreement with the docking study, corroborating the electronic contribution observed in the ELF isosurface. These interactions are mainly in the proton attractor sites found in the two oxygen atoms, where the atom O2 (Fig. 1) establishes essential hydrogen bonding with the Trp439 and Trp86 residues (Figure S3 of supplementary material).
The NCI and ELF of THA calculated at MP2/aug-cc-pVDZ level showed the same trend of THA + (Figure S4), and both showed very weak interactions in their most acidic hydrogens. The interpretation of NCI compared to the docking structure is consistent, as it presents essentially the same behavior when comparing THA and THA+, and demonstrates the ability to correctly predict a descriptor for the interaction profile from the free molecules. THA results (Figure S4) show regions with electron density located in the nitrogen of the pyrimidine ring, which qualifies the N in which a proton was added to form the protonated tacrine ligand, in addition to having electron-attracting sites in the most acidic hydrogens.
For protonated tacrine, regions of weak interactions were located around the aromatic rings, and the residues Glu202, Ser203, Trp86, Ty337, and His447, which presented the contribution of van der Waals forces, and the same profile was found for neutral tacrine. We observed the presence of a wide attractive region close to the Trp86 residue corroborating the inhibitor π-stacking interactions found in the molecular docking simulation. Furthermore, regardless of its orientation, the results suggested that the phenyl group may be necessary for the inhibitor complexation step [65]. In this case, it allows for the formation of attractive interactions such as π-stacking of residues Trp86 and Tyr337 and contributing with new interactions, which are responsible for the improved performance of tacrine analogs. Moreover, a strong favorable NCI is observed in the THA + molecule, where there is a hydrogen bond between its most acidic hydrogen linked in the N1 atom, with the oxygen atom of the carbonyl group of the His447 residue.
Influence of water molecules in the catalytic active site
Figure 8 shows the HOMO orbital for the CAS and the complexed CAS with ACh and THA+ in the PDB with water and the excluded water molecules. Glu202 is present in the contribution of HOMO in the CAS and the CAS-ACh complex. HOMO has also contributed to the Ser203 residue of the catalytic triad, close to the water molecule. However, in the case of THA+, the contribution to the HOMO orbital comes from THA+ and Trp86. The system with excluded water has the same profile for the HOMO of CAS and CAS-ACh system. The difference is regarding the HOMO orbital contribution of Tyr449 of CAS-THA+ system. In general, the HOMO orbital energy is stabilized by the same amount from CAS to CAS-ACh in regard to the system with (5.14 eV) and without (5.33 eV) water molecules. In the case of THA+, the stabilization is close to the ACh, with water molecules being 5.18 eV, while without water molecules is 5.37 eV. Inside Fig. 8, ACh and THA+ in vacuum and the PCM solvent model are shown. In both cases, the HOMO orbital distribution has no effect on the solvent, despite the frontier orbital energies. The other occupied molecular orbitals HOMO-n (n = 1,2,3,4), also show contributions from Ser203, Glu202, and Trp86.
HOMO orbital contribution and energies for CAS, CAS-ACh, and CAS-THA+. In the first line, calculations with five nearest explicit structural water molecules. In the second line, calculations without water molecules. In the blue boxes, it is represented ACh and THA+ free molecules in vacuum (top) and PCM solvent (bottom)
The LUMO orbital of complexed CAS-ACh and CAS-THA+ is shown in Fig. 9. In contrast to the HOMO orbital, the LUMO orbital shows a significant difference between the distribution of LUMO orbital comparing the PDB with water and without water. In general, the orbital energies show the same trend with or without water molecules. Therefore, regarding the orbital energies, the influence of explicit water molecules is negligible. However, in regard to the orbital distribution, the water molecules have an influence.
LUMO orbital contribution and energies for CAS, CAS-ACh, and CAS-THA+. In the first line, calculations considered five nearest explicit structural water molecules. In the second line, calculations without the water molecules. In the blue boxes, it is represented ACh and THA+ free molecules in vacuum (top) and PCM solvent (bottom)
Conclusions
Cholinergic therapy through AChE inhibitors (AChEI) has been widely used in AD treatment. In this way, investigating the electronic properties and molecular interactions of these ligands with the residues of the active site is essential. Therefore, we have analyzed frontier molecular orbitals, potential hydrogen acid donor regions, NCI, ELF, molecular docking studies, and interactions with active hAChE enzyme site for ACh, THA, and THA+ ligands. Aug-cc-pVDZ and 6–311 + G(d,p) are suitable basis set, and both shows the same trend for the structural and electronic properties studied. Structurally, ACh and THA molecular size and volume turn possible to access the CAS and interact properly with the hAChE pocket main residues, where the polarization in the acetoxyethyl moiety of ACh structure is in accordance with the nucleophilic attack of this carbon center. Moreover, the nucleophilic attack is related to the LUMO series that has almost the same energies for ACh and THA. Neutral THA presents acceptor regions close to the nitrogen of its pyridine ring, justifying the addition of a proton in this region to form the THA+ ligand. THA+ showed similarity to ACh electrostatic potential map, which presented its two nitrogen atoms as the most acidic hydrogen donors for the nucleophilic attack. ACh interaction profile comprises mainly of hydrogen interactions between the ligand quaternary ammonium binding site and the Trp86 and Trp439 residues. Trp86 residue also shows intermolecular interactions of the cation-π and sigma-π type, equivalent to the cation-π interaction found in the literature in the TcAChE. The sandwich-type interaction was found for THA + binding at the hAChE active site GORGE base, establishing close stacking interactions between the tricyclic aromatic ring and Trp86 residue on one side and with Tyr337 residue on the other side. Similar interactions between the residues Trp84 and Phe330 were found in TcAChE. We found that the presence of a wide attractive region close to the Trp86 residue corroborates the inhibitor π-stacking interactions, and the results suggested that the phenyl group may be necessary for the inhibitor complexation step. NCI and ELF results indicate the presence of attractive interaction regions on ACh between the methyl groups of the quaternary nitrogen and the residues of His447, Trp86, and Tyr337, also found in the molecular docking study. The importance of the solvent is shown by the less negative heteroatom charge, indicating how those ligands have a good affinity with water. In general, the orbital energies have small influence on water molecules, while the water molecules have significative influence concerning the orbital distribution.
References
Hou Y, Dan X, Babbar M et al (2019) (2019) Ageing as a risk factor for neurodegenerative disease. Nat Rev Neurol 1510(15):565–581. https://doi.org/10.1038/s41582-019-0244-7
Weis S, Sonnberger M, Dunzinger A, et al (2019) Neurodegenerative diseases: Alzheimer disease (AD). Imaging Brain Dis 897–931. https://doi.org/10.1007/978-3-7091-1544-2_32
Zhang H-L, Wu J, Zhu J (2010) The immune-modulatory role of apolipoprotein E with emphasis on multiple sclerosis and experimental autoimmune encephalomyelitis. Clin Dev Immunol 2010:186813. https://doi.org/10.1155/2010/186813
Liu C-C, Kanekiyo T, Xu H (2013) Bu G (2013) Apolipoprotein E and Alzheimer disease: risk mechanisms and therapy. Nat Rev Neurol 92(9):106–118. https://doi.org/10.1038/nrneurol.2012.263
Lm B, Ce Y, Td B, DW T, (2010) Genetics of Alzheimer disease. J Geriatr Psychiatry Neurol 23:213–227. https://doi.org/10.1177/0891988710383571
Martorana A, Esposito Z, Koch G (2010) Beyond the cholinergic hypothesis: do current drugs work in Alzheimer’s disease? CNS Neurosci Ther 16:no--no https://doi.org/10.1111/j.1755-5949.2010.00175.x
Cummings JL, Cole G (2002) Alzheimer disease. J Am Med Assoc 287:2335–2338
Nascimento ECM, Martins JBL, dos Santos ML, Gargano R (2008) Theoretical study of classical acetylcholinesterase inhibitors. Chem Phys Lett 458:285–289. https://doi.org/10.1016/j.cplett.2008.05.006
Ohno-Shosaku T, Matsui M, Fukudome Y et al (2003) Postsynaptic M1 and M3 receptors are responsible for the muscarinic enhancement of retrograde endocannabinoid signalling in the hippocampus. Eur J Neurosci 18:109–116. https://doi.org/10.1046/j.1460-9568.2003.02732.x
Yi JH, Whitcomb DJ, Park SJ, et al (2020) M1 muscarinic acetylcholine receptor dysfunction in moderate Alzheimer’s disease pathology. Brain Commun 2 https://doi.org/10.1093/braincomms/fcaa058
Suzuki E, Momiyama T (2021) M1 muscarinic acetylcholine receptor-mediated inhibition of GABA release from striatal medium spiny neurons onto cholinergic interneurons. Eur J Neurosci 53:796–813. https://doi.org/10.1111/ejn.15074
Baxter MG, Crimins JL (2018) Acetylcholine receptor stimulation for cognitive enhancement: better the devil you know? Neuron 98:1064–1066. https://doi.org/10.1016/j.neuron.2018.06.018
Anand P, Singh B (2013) A review on cholinesterase inhibitors for Alzheimer’s disease. Arch Pharm Res 36:375–339. https://doi.org/10.1007/s12272-013-0036-3
Rosini M, Simoni E, Bartolini M et al (2008) Inhibition of acetylcholinesterase, β-amyloid aggregation, and NMDA receptors in Alzheimer’s disease: a promising direction for the multi-target-directed ligands gold rush. J Med Chem 51:4381–4384. https://doi.org/10.1021/jm800577j
Wang Y, Wang H, Chen H (2016) AChE inhibition-based multi-target-directed ligands, a novel pharmacological approach for the symptomatic and disease-modifying therapy of Alzheimer’s disease. Curr Neuropharmacol 14:364–375. https://doi.org/10.2174/1570159x14666160119094820
Han J, Du Z, Lim MH (2021) Mechanistic insight into the design of chemical tools to control multiple pathogenic features in Alzheimer’s disease. Acc Chem Res 54:3930–3940. https://doi.org/10.1021/acs.accounts.1c00457
Kumar N, Gahlawat A, Kumar RN et al (2022) Drug repurposing for Alzheimer’s disease: in silico and in vitro investigation of FDA-approved drugs as acetylcholinesterase inhibitors. J Biomol Struct Dyn 40:2878–2892. https://doi.org/10.1080/07391102.2020.1844054
dos Santos TC, Gomes TM, Pinto BAS et al (2018) Naturally occurring acetylcholinesterase inhibitors and their potential use for Alzheimer’s disease therapy. Front Pharmacol 9:1192. https://doi.org/10.3389/fphar.2018.01192
Abdul Manap AS, Wei Tan AC, Leong WH et al (2019) Synergistic effects of curcumin and piperine as potent acetylcholine and amyloidogenic inhibitors with significant neuroprotective activity in SH-SY5Y cells via computational molecular modeling and in vitro assay. Front Aging Neurosci 11 https://doi.org/10.3389/fnagi.2019.00206
Shaik JB, Kandrakonda YR, Kallubai M et al (2021) Deciphering the AChE-binding mechanism with multifunctional tricyclic coumarin anti-Alzheimer’s agents using biophysical and bioinformatics approaches and evaluation of their modulating effect on Amyloidogenic peptide assembly. Int J Biol Macromol 193:1409–1420. https://doi.org/10.1016/j.ijbiomac.2021.10.204
YudiUtomo R, Asawa Y, Okada S et al (2021) Development of curcumin-based amyloid β aggregation inhibitors for Alzheimer’s disease using the SAR matrix approach. Bioorg Med Chem 46:116357. https://doi.org/10.1016/j.bmc.2021.116357
Liao Y, Hu X, Pan J, Zhang G (2022) Inhibitory mechanism of baicalein on acetylcholinesterase: inhibitory interaction, conformational change, and computational simulation. Foods 11
Viayna E, Sabate R, Muñoz-Torrero D (2013) Dual inhibitors of β-amyloid aggregation and acetylcholinesterase as multi-target anti-Alzheimer drug candidates. Curr Top Med Chem 13:1820–1842. https://doi.org/10.2174/15680266113139990139
Koide da Silva BY, da Neves Lopes A, S, José Sousa Maia P, et al (2021) Chemical and biological evaluation of the aqueous extract of Peumus boldus Molina (Monimiaceae) leaves. Pharmacognosy Res 14:45–52. https://doi.org/10.5530/pres.14.1.8
Meira Menezes T, Assis C, Lacerda Cintra AJ et al (2021) Binding mechanism between acetylcholinesterase and drugs pazopanib and lapatinib: biochemical and biophysical studies. ACS Chem Neurosci 12:4500–4511. https://doi.org/10.1021/acschemneuro.1c00521
de Andrade RG, Souza de Oliveira A, Bartolini M et al (2021) Discovery of sustainable drugs for Alzheimer’s disease: cardanol-derived cholinesterase inhibitors with antioxidant and anti-amyloid properties. RSC Med Chem 12:1154–1163. https://doi.org/10.1039/D1MD00046B
Sussman JL, Silman I (1992) Acetylcholinesterase: structure and use as a model for specific cation—protein interactions. Curr Opin Struct Biol 2:721–729. https://doi.org/10.1016/0959-440X(92)90207-N
Radic Z, Pickering NA, Vellom DC et al (1993) Three distinct domains in the cholinesterase molecule confer selectivity for acetyl- and butyrylcholinesterase inhibitors. Biochemistry 32:12074–12084. https://doi.org/10.1021/bi00096a018
Saxena A, Redman AMG, Jiang X et al (1999) Differences in active-site gorge dimensions of cholinesterases revealed by binding of inhibitors to human butyrylcholinesterase. Chem Biol Interact 119–120:61–69. https://doi.org/10.1016/S0009-2797(99)00014-9
Freeman SE, Dawson RM (1991) Tacrine: a pharmacological review. Prog Neurobiol 36:257–277. https://doi.org/10.1016/0301-0082(91)90002-I
Sussman JL, Harel M, Silman I (1993) Three-dimensional structure of acetylcholinesterase and of its complexes with anticholinesterase drugs. Chem Biol Interact 87:187–197. https://doi.org/10.1016/0009-2797(93)90042-W
Harel M, Schalk I, Ehret-Sabatier L et al (1993) Quaternary ligand binding to aromatic residues in the active-site gorge of acetylcholinesterase. Proc Natl Acad Sci 90(9031):LP-9035. https://doi.org/10.1073/pnas.90.19.9031
Dawson RM (1990) Reversibility of the inhibition of acetylcholinesterase by tacrine. Neurosci Lett 118:85–87. https://doi.org/10.1016/0304-3940(90)90254-7
Summers WK, Tachiki KH, Kling A (1989) Tacrine in the treatment of Alzheimer’s disease. Eur Neurol 29((suppl 3):28–32. https://doi.org/10.1159/000116478
Adem A (1992) Putative mechanisms of action of tacrine in Alzheimer’s disease. Acta Neurol Scand 85:69–74. https://doi.org/10.1111/j.1600-0404.1992.tb04458.x
Borges NM, Sartori GR, Ribeiro JFR et al (2018) Similarity search combined with docking and molecular dynamics for novel hAChE inhibitor scaffolds. J Mol Model 24:1–12. https://doi.org/10.1007/s00894-017-3548-9
Kiametis AS, Gargano R, Martins JBL (2013) Anacardic acid as potential acetylcholinesterase inhibitor. Eur Biophys J with Biophys Lett 42:S160–S160
de Paula AAN, Martins JBL, dos Santos ML et al (2009) New potential AChE inhibitor candidates. Eur J Med Chem 44:3754–3759. https://doi.org/10.1016/j.ejmech.2009.03.045
Nascimento ÉCM, Oliva M, Świderek K et al (2017) Binding analysis of some classical acetylcholinesterase inhibitors: insights for a rational design using free energy perturbation method calculations with QM/MM MD simulations. J Chem Inf Model 57:958–976. https://doi.org/10.1021/acs.jcim.7b00037
Nascimento ECM, Martins JBLJBL, Moreno Nascimento EC et al (2011) Electronic structure and PCA analysis of covalent and non-covalent acetylcholinesterase inhibitors. J Mol Model 17:1371–1379. https://doi.org/10.1007/s00894-010-0838-x
Khan H, Marya Amin S et al (2018) Flavonoids as acetylcholinesterase inhibitors: current therapeutic standing and future prospects. Biomed Pharmacother 101:860–870. https://doi.org/10.1016/j.biopha.2018.03.007
Chufarova N, Czarnecka K, Skibiński R et al (2018) New tacrine–acridine hybrids as promising multifunctional drugs for potential treatment of Alzheimer’s disease. Arch Pharm (Weinheim) 351:1800050. https://doi.org/10.1002/ardp.201800050
Nachon F, Carletti E, Ronco C et al (2013) Crystal structures of human cholinesterases in complex with huprine W and tacrine: elements of specificity for anti-Alzheimer’s drugs targeting acetyl- and butyryl-cholinesterase. Biochem J 453:393–399. https://doi.org/10.1042/BJ20130013
Meena VK, Chaturvedi S, Sharma RK et al (2019) Potent acetylcholinesterase selective and reversible homodimeric agent based on tacrine for theranostics. Mol Pharm 16:2296–2308. https://doi.org/10.1021/ACS.MOLPHARMACEUT.8B01058
Zhou A, Hu J, Wang L et al (2015) Combined 3D-QSAR, molecular docking, and molecular dynamics study of tacrine derivatives as potential acetylcholinesterase (AChE) inhibitors of Alzheimer’s disease. J Mol Model 21:277. https://doi.org/10.1007/s00894-015-2797-8
Simeon S, Anuwongcharoen N, Shoombuatong W et al (2016) Probing the origins of human acetylcholinesterase inhibition via QSAR modeling and molecular docking. PeerJ 4:e2322. https://doi.org/10.7717/PEERJ.2322
Cunha WF, Gargano R, Garcia E et al (2014) Rovibrational energy and spectroscopic constant calculations of CH 4 ⋯ CH4, CH 4 ⋯ H2 O, CH 4 ⋯ CHF3, and H 2O ⋯ CHF3 dimers. J Mol Model 20:2298. https://doi.org/10.1007/s00894-014-2298-1
Pereira WA, Nascimento ÉCM, Martins JBL (2021) Electronic and structural study of T315I mutated form in DFG-out conformation of BCR-ABL inhibitors https://doi.org/10.1080/07391102.2021.1935320
Graef EL, Martins JBL (2019) Analysis of lowest energy transitions at TD-DFT of pyrene in vacuum and solvent. J Mol Model 25:183. https://doi.org/10.1007/s00894-019-4065-9
Rocha KML, Nascimento ÉCM, Martins JBL (2021) Investigation on the interaction behavior of afatinib, dasatinib, and imatinib docked to the BCR-ABL protein. J Mol Model 27:309. https://doi.org/10.1007/s00894-021-04925-8
de Almeida AL, Barbosa LPG, Santos RL, Martins JBL (2016) Chemical reactivity indices of the caffeine molecule. Rev Virtual Química 8:483–492. https://doi.org/10.5935/1984-6835.20160035
Frisch MJ, Trucksr GW, Schlegel HB et al (2009) Gaussian09 Revision D01. Gaussian Inc., Wallingford CT
Olsson MHM, Søndergaard CR, Rostkowski M, Jensen JH (2011) PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7:525–537. https://doi.org/10.1021/ct100578z
Morris GM, Huey R, Lindstrom W et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/JCC.21256
Humphrey W, Dalke A, Schulten K (1996) VMD: Visual molecular dynamics J Mol Graph https://doi.org/10.1016/0263-7855(96)00018-5
Dassault Systèmes BIOVIA (2017) Discovery Studio, 2017. Dassault Systèmes, San Diego, CA
Johnson ER, Keinan S, Mori-Sánchez P et al (2010) Revealing noncovalent interactions. J Am Chem Soc 132:6498–6506. https://doi.org/10.1021/ja100936w
Contreras-García J, Boto RA, Izquierdo-Ruiz F et al (2016) A benchmark for the non-covalent interaction (NCI) index or…is it really all in the geometry? Theor Chem Acc 135:242. https://doi.org/10.1007/s00214-016-1977-7
Contreras-García J, Johnson ER, Keinan S et al (2011) NCIPLOT: a program for plotting noncovalent interaction regions. J Chem Theory Comput 7:625–632. https://doi.org/10.1021/ct100641a
Lu T, Chen F (2012) Multiwfn: a multifunctional wavefunction analyzer. J Comput Chem 33:580–592. https://doi.org/10.1002/jcc.22885
Fuster F, Grabowski SJ (2011) Intramolecular hydrogen bonds: the QTAIM and ELF characteristics. J Phys Chem A 115:10078–10086. https://doi.org/10.1021/jp2056859
Henchman RH, Tai K, Shen T, McCammon JA (2002) Properties of water molecules in the active site gorge of acetylcholinesterase from computer simulation. Biophys J 82:2671–2682. https://doi.org/10.1016/S0006-3495(02)75609-9
Hamulakova S, Janovec L, Hrabinova M et al (2014) Synthesis and biological evaluation of novel tacrine derivatives and tacrine–coumarin hybrids as cholinesterase inhibitors. J Med Chem 57:7073–7084. https://doi.org/10.1021/jm5008648
Gerlits O, Ho KY, Cheng X et al (2019) A new crystal form of human acetylcholinesterase for exploratory room-temperature crystallography studies. Chem Biol Interact 309:108698. https://doi.org/10.1016/j.cbi.2019.06.011
Cheung J, Rudolph MJ, Burshteyn F et al (2012) Structures of human acetylcholinesterase in complex with pharmacologically important ligands. J Med Chem 55:10282–10286. https://doi.org/10.1021/jm300871x
Acknowledgements
The authors acknowledge the computational resource of Laboratory of Computational Chemistry (LQC) of the University of Brasilia.
Funding
This work was supported by the Brazilian National Council for Scientific and Technological Development (CNPq 310071/2018–6, João B. L. Martins), Fundação de Apoio a Pesquisa do Distrito Federal (FAPDF 00193–00000229/2021–21, João B. L. Martins).
Author information
Authors and Affiliations
Contributions
Letícia A. Nascimento: investigation, calculations, writing—original draft. Érica C. M. Nascimento: conceptualization, methodology, formal analysis, methodology. João B. L. Martins: conceptualization, formal analysis, methodology, resources, supervision, writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This paper belongs to Topical Collection XXI—Brazilian Symposium of Theoretical Chemistry (SBQT2021).
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nascimento, L.A., Nascimento, É.C.M. & Martins, J.B.L. In silico study of tacrine and acetylcholine binding profile with human acetylcholinesterase: docking and electronic structure. J Mol Model 28, 252 (2022). https://doi.org/10.1007/s00894-022-05252-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-022-05252-2