Abstract
Cold-adaptive methanogens contribute significantly to methane emission from the cold area, while the cold-adaptive mechanisms used by Archaea remain elusive. Methanolobus psychrophilus R15, a cold-adaptive methanogen isolated from a Tibetan plateau wetland, grows at 0–25 °C and optimally at 18 °C when isolated; however, it grows optimally at 30 °C after culturing at 18 °C for several years. Aiming to gain insights into the protein profiles that are involved in optimal growth and cold adaptation of this methanogen, here, we performed a comparative proteomic study using 2D DIGE on the cultures grown at 30, 18 and 4 °C. 1439 protein spots (3167 ORFs annotated in the R15 genome) were detected, and 202 of 322 differentially expressed protein spots were identified by MALDI-TOF/TOF. The protein abundance of most enzymes involved in methanogenesis, energy conservation and central metabolism were increased at 30 °C, while most ribosome proteins were decreased at 30 °C. Proteasome and ROS scavengers increased expressions at 4 °C, suggesting more aberrant proteins and ROS formed at lower temperatures. Different from the cold-adaptive Methanococcoides burtonii, some chaperones were increased at 4 °C, implying that protein folding was impaired at cold in this psychrophilic archaeon. This study indicates that diverse cold-adaptive mechanisms can be used by different methanogenic Archaea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cold-adaptive methanogens play a major role in the methane emission from cold area, which contributes significantly to the global warming, as about 80 % of the Earth’s biosphere is permanently exposed to temperatures below 5 °C (Rodrigues and Tiedje 2008). However, the cold-adaptive mechanisms used by methanogens are poorly understood, except for an Antarctic methanogen Methanococcoides burtonii (Franzmann et al. 1992), one of the fewer psychrophilic archaeal strains cultured so far.
Cold can exert overall severe physicochemical constraints on living organisms, as temperature is of significance on biochemical reactions, molecular diffusions and hydrogen bond-based macromolecule interactions, etc. (Piette et al. 2011a, b). Extensive studies on the cold-adaptive characteristics of M. burtonii indicated that the following cellular processes are involved in the cold adaptation: increasing proportion of unsaturated fatty acid in cellular membrane so as to keep the membrane fluidity (Nichols et al. 2004); increasing the abundance of ribosomal proteins to keep translation process (Thomas et al. 2001); upregulation of the proteins, such as Dead-box helicase, to prevent RNA forming secondary structure at low temperatures (Williams et al. 2010 and more dihydrouracil modification of tRNA (Noon et al. 2003). Similar cold-adaptive mechanisms are also found in cold-adapted bacteria (De Maayer et al. 2014). However, Bowman (2008), upon comparing 19 cold-adapted prokaryote genomes, has concluded that no common feature was shared by these microorganism genomes. Diverse cold-adaption mechanisms are specially shown by the distinct actions of chaperones. Trigger factor (TF), a ribosome-bound chaperone, acts as a major chaperone working at cold in an Antarctic bacterium Pseudoalteromonas haloplanktis, by remarkable expressing at cold(~40-fold) and reducing most other HSP chaperones (Piette et al. 2011a, b). In contrast, Psychrobacter arcticus elevated the expression of GroEL/ES chaperonins and reduced TF at cold. No chaperones have been found involved in the cold adaptation in M. burtonii (Williams et al. 2011). Therefore, cold-adaptations can be diverse among species.
Methanolobus psychrophilus R15 (CGMCC1.5060T) is another cold-adaptive methanogen recently isolated from a cold wetland, by growing at 0–25 °C with a fastest growth at 18 °C (Zhang et al. 2008). Previous transcriptomic study indicates the distinct cold-responsive cellular processes in addition to the similar cold-adaptive mechanisms found for M. burtonii (Chen et al. 2012). In addition, after a few years subculture at 18 °C, R15 can grow at 30 °C and has a fastest growth at 30 °C. In this study, through a comparative proteomic study by 2D DIGE, revealed a suit of proteins that were involved in optimal and cold-adaptation conditions in this cold-adaptive archaeon.
Materials and methods
Strain and culture
Strain R15 was cultured in basal medium as described by Zhang et al. (2008) with trimethylamine at a final concentration of 20 mM as substrate. R15 were cultured at three temperatures, 4, 18 and 30 °C, representing cold stress (4, 18 °C) and optimal (30 °C) conditions, respectively. Growth was monitored by measuring optical density (OD600) of the cultures with an Ultrospec 10 Cell Density Meter (Amersham Biosciences, Piscataway, NJ, USA).
Methane determination
Methane production was determined by Shimadzu GC-14B gas chromatograph (Shimadzu Corp., Kyoto, Japan).
Soluble protein extraction
Four biological replicates for each culture were prepared. Cells at the early exponential phase (OD600 ≈ 0.2) were collected by centrifuging at 5000×g for 5 min at the same temperatures as the culture temperatures. The collected cells were dissolved in a lysis buffer (7 M urea, 4 % CHAPS, 2 M thiourea, 30 mM Tris, 2 % TritonX-100, 1 % DTT, 1 % IPG buffer) containing protease and phosphatase inhibitor and sonicated for 10 min at 70 W (sonicated 3 s, and then stopped 3 s, repeatedly). The protein concentration of each sample was determined using a 2D Quant kit (GE Healthcare, Piscataway, NJ, USA). Each sample was treated with an Ettan 2-D Clean-up Kit (GE Healthcare) to remove contaminating substances before performing isoelectric focusing (IEF).
Protein labeling and 2D DIGE procedures
Protein labeling with CyDye™ DIGE fluor minimal dyes was performed using an Ettan DIGE system (GE Healthcare) according to the manufacturer’s instructions. To meet the 2D DIGE experimental design requirements, 12 samples were labeled with different CyDye DIGE fluor dyes (GE Healthcare). Fluor dyes were dissolved in 5 µL dimethylformamide (DMF) to reconstitute the CyDye DIGE fluor dye. Each protein sample (50 µg protein) was adjusted to pH 8.5 with 1 M Tris, and then labeled with 0.4 µL fluor dye solution and 0.6 µL DMF for 30 min on ice in the dark. The labeling reaction was stopped by adding 1 µL 10 mM lysine and incubating for 10 min on ice in the dark.
Three samples that were run in the same gel were labeled with Cy2, Cy3, and Cy5 and mixed according to the 2D DIGE experimental design instructions (Table S1). DTT and IPG buffers were added to the mixed samples, which were then centrifuged at 18000×g for 5 min at 20 °C. The supernatant was moved to a Reswelling Tray (GE Healthcare) and applied to 18-cm, pH 3–11 NL Immobiline Dry Strips (GE Healthcare), then rehydrated overnight in the dark before performing the first-dimension IEF.
The IEF was run on an Ettan IPGphor manifold (GE Healthcare) with the following steps: 100 V for 2 h; 200 V for 2 h; 300 V for 1 h; 500 V for 1 h; 1000 V for 2 h; then gradients up to 8000 V for 1 h; 8000 V for 80000 V-h; 500 V for 8 h; and 8000 V for 16000 V-h. Next, the IPG strips were equilibrated for 15 min in the dark in a solution (0.375 M Tris–HCl, pH 8.8, 6 M urea, 30 % glycerol, 2 % SDS, 0.002 % bromophenol blue) containing 2 % DTT, and then equilibrated in a buffer containing 2.5 % iodoacetamide for 15 min. The equilibrated IPG strips were loaded onto acrylamide gels (15 %) for the second dimension separation step. Each slab gel was electrophoresed with a constant current of 12 mA on an Ettan™ DALT six system electrophoresis unit (GE Healthcare) in the dark at 25 °C for 18 h.
Acquisition of images and identification of the differentially expressed proteins
The 2D DIGE gels were scanned using Typhoon Trio+ (GE Healthcare) with the recommended emission filters and laser combinations. Differentially expressed proteins in the R15 cultures grown at 4, 18, and 30 °C were analyzed using the differential in-gel analysis (DIA), biological variation analysis (BVA), and extended data analysis (EDA) modules of the DeCyder 6.5 software (GE Healthcare). Statistical significance was assessed using the Student’s t test and one-way analysis of variance (ANOVA) in the BVA module of DeCyder. Proteins detected in ≥75 % of the experimental samples, Student’s t test <0.05, Av. ratio >1.1 and one-way ANOVA <0.05 were selected as differential expression protein spots. Av. ratio represents fold change.
In-gel tryptic digestion and MALDI-TOF/TOF identification of the differentially expressed proteins
Total proteins (1200 µg) from the same 12 samples which were used in above-labeled experiment were used to cut out the differentially expressed proteins. Four parallel 2D gels were collected and a spot picker (The Gel Company, San Francisco, CA, USA) was used to manually cut out the spots of the differentially expressed proteins. The cut gel pieces containing the proteins were vacuum-dried, destained, and digested with 12.5 ng/μL trypsin (Sigma) in 25 mM ammonium bicarbonate (pH 8.0) for 16 h at 37 °C.
Next, the digested supernatants were collected for matrix-assisted laser desorption/ionization time-of-flight/time-of-flight (MALDI-TOF/TOF). For the analysis, the supernatants were mixed with the MALDI matrix material (5.0 mg/ml of α-cyano-4-hydroxycinnamic acid) at 1:1 ratio and spotted onto a MALDI target. After air drying, crystallized spots were analyzed with an MALDI-TOF/TOF Analyzer (AB SCIEX 5800) (AB Sciex, CA, USA). Peptide mass maps were acquired in the positive reflection mode, averaging 1500 laser shots per MALDI-TOF spectrum and 3000 shots per TOF/TOF spectrum. Calibration mixtures (Applied Biosystems) were used to calibrate the spectrum to a mass tolerance within 0.2 Da. Parent mass peaks with a mass range of 800–3500 Da and the ten most abundant ions from MS analysis were chosen for tandem TOF/TOF analysis.
MS and MS/MS data were analyzed using the MASCOT 2.0 search engine (Matrix Science, London, UK) to search against the SwissProt database (Bairoch and Apweiler 2000). The search parameters were as follows: two missed cleavage sites by trypsin, 0.2 Da peptide mass tolerance, 0.5 Da MS/MS ion tolerance, carbamidomethylation of cysteine as a fixed modification, and methionine oxidation as a variable modification. For all the proteins identified by their peptide mass fingerprint and MS/MS, a mascot score greater than 51 was considered significant (p value <0.05).
Bioinformatics analysis
The differential expressed proteins those with Student’s t test <0.05, Av. ratio >1.1, and one-way ANOVA <0.05 were used for analysis of protein homologs among samples and the confidences of differentially expressed spots. Av. ratio means fold change, it was calculated by the lowest value of the four biological replicates in higher group and highest value of the four biological replicates in lower group, it is more correct than calculated by average value of biological replicates, which may cause false positive. A heat map was clustered with the hierarchical clustering method of the EDA module of Decyder 6.5 and the principal component analysis was performed using the EDA module of Decyder 6.5. The Clusters of Orthologous Groups (COGs) functional categories of proteins (Tatusov et al. 2001) were assigned by BLAST searches against the COG database (http://www.ncbi.nlm.nih.gov/COG/).
Results and discussion
Growth and methanogenesis of R15 at different temperatures
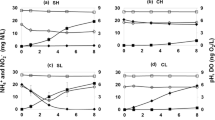
The growth rates of R15 were highest at 30 °C (0.17/h), followed by 18 °C (0.05/h), and lowest at 4 °C (0.005/h), and the generation times were calculated to be 5.8, 20.3 and 198.7 h at the corresponding temperatures (Fig. 1a). At 30 °C, R15 completed the entire growth cycle but the cell appeared lysed when entered the stationary phase. Transient growth at 37 °C was also observed, but it ceased at 24 h during the late exponential phase. Methane production rates showed a similar temperature-related mode (Fig. 1b) as the growth rate, indicating a coupling between the two processes.
Overview of the proteomes of R15 grown at cold and optimal temperatures
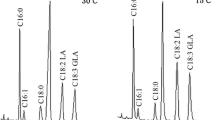
R15 grown at 4, 18 and 30 °C were harvested in their early exponential phases (OD600 ≈ 0.2), and four biological replicates for each sample were used. Each sample was treated using the 2D DIGE protocol (details in “Materials and methods”). Approximately 1439 spots (3167 predicted ORFs in R15 genome) were detected in the 2D DIGE profile (Fig. 2); a total of 322 differentially expressed spots were identified by the DeCyder software, and 202 of them can be identified by MALDI-TOF/TOF and the Mascot database (Table S2).
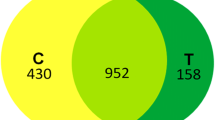
A comparison between the 30- and 18 °C-cultures revealed 135 differently expressed spots; between the 30- and 4 °C-cultures revealed 221 differentially expressed protein spots; and between the 18- and 4 °C-cultures revealed 149 differently expressed spots.
Principal component analysis was performed on the data set containing all the differential expressed proteins (Fig. S1). The score plot showed an overview of the spot maps in which most of the spots were located inside the 95 % confidence interval. As shown in Table S2, 90 % of the differentially expressed proteins in the cells grown at 18 °C compared with the cells grown at 4 °C were in agreement with our transcriptomic data reported previously (Chen et al. 2012). The two studies indicated the high reliability of our analysis results.
The differentially expressed proteins were grouped using hierarchical clustering (details in “Materials and methods”). As shown in Fig. S2, the samples in the four replicate cultures grown at the same temperatures always formed the same clusters; the result also implied high reliability of the experimental data. Compared with the 4 °C-culture, similar profiles of protein abundance were found between 30- and 18 °C-culture. Proteins that increased abundance significantly at 4 °C-cultures (A1 group) appeared at the same amount or decreased at 18- and 30 °C-cultures. Interestingly, the increased expression protein clusters in 30 °C-culture (A2 group) were those of decreased or same expressed proteins in 4- and 18 °C-cultures (Fig. S2). This indicates distinct protein groups involved in the optimal and low temperatures in R15.
All the differentially expressed proteins were assigned to Cluster Orthologous Groups (COG) functional categories, and were found to be involved mainly in energy production and conversion (31–34 %), translation, ribosome structure (9–15 %), protein turnover, chaperones (5–14 %), amino acid transport and metabolism (8–11 %), and carbohydrate transport and metabolism (3–6 %) (Table 1). As shown in Table 1, most of the proteins in energy production category were upregulated at higher temperatures, while those in translation category were downregulated significantly in 30 °C-culture compared with 18 °C-culture, but upregulated significantly in 18 °C-culture than in 4 °C-culture, suggesting that ribosome biogenesis process is sensitive to temperature. Strikingly, most of the chaperones were upregulated in the 4 °C-cultures compared with 18 °C-culture, indicating that the archaeal protein folding can be impaired by cold as reviewed by Piette et al. (2011a, b). Notably, the abundance of two membrane proteins (0505 and 1258) were dramatically increased at 4 °C comparing with at 18 °C (Table S2), suggesting that membrane remodeling can occur in response to cold.
Differentially expressed proteins involved in energy conservation, biomass production and central metabolism pathways
Methanogenesis
R15 is an obligate methylotrophic methanogen that uses trimethylamine (TMA), dimethylamine (DMA), monomethylamine (MMA) and methanol as the only known methanogenic precursors. The methyl groups of these compounds enter a proportional dismutation pathway, by which the oxidation of one methyl group to CO2 generates electrons that reduce up to three other methyl groups to methane (Williams et al. 2010) (Fig. 3). Proteins involved in TMA-derived methanogenesis, such as trimethylaminemethyl transferase (Mpsy_1687, 1688), dimethylaminemethyl transferase (Mpsy_1680), methyl-coenzyme M reductase beta subunit (Mpsy_0832) and F420H2 dehydrogenase subunit FpoB (Mpsy_2567), all showed increased abundance at the higher temperatures with a trend of 30 > 18 > 4 °C (Tables S2). The expression of three corrinoid proteins, trimethylamine corrinoid protein (Mpsy_1686), dimethylamine corrinoid protein (Mpsy_1704) and monomethylamine corrinoid protein (Mpsy_1678) seemed not to be sensitive to temperature increase because they displayed similar abundances at 30 and 18 °C; however, their abundances decreased at 4 °C compared with 18 °C (Table S2). The methyl oxidative branch in the methylotrophic methanogenic pathway produces reducing equivalents (e.g., F420H2) that are used in methane production or other biological processes (Ferry, 1992). Tetrahydromethanopterin S-methyltransferase subunit H (Mpsy_2182) was upregulated at the higher temperatures (30 > 18 > 4 °C). Methyl-coenzyme M reductase alpha subunit (Mpsy_0828), and tetrahydromethanopterin formyltransferase alpha subunit (Mpsy_1734) seemed to be decreased at 30 °C compared with at 18 °C, but increased at 4 °C compared with at 18 °C (Table S2). The expression of formylmethanofuran dehydrogenase subunit E (Mpsy_2441) and methylene tetrahydromethanopterin dehydrogenase (Mpsy_1459) were increased at 4 °C compared with 18 °C. The upregulation of these enzymes at low temperature can be explained by that more enzymes were produced to offset the lower enzymatic activities caused by low temperature. Overall, we proposed the methanogenesis process is more active with temperature increasing, which is consistent with the elevated methylamine-derived methanogenic rate observed for R15 grown at these temperatures. Methanogenesis is the main pathway for energy conservation in a methanogen; therefore, active methanogenic metabolism at increased temperatures will support vigorous growth of R15 at temperatures within a tolerable range.
Differentially expressed proteins involved in energy production and central metabolisms pathways. The temperature-responsive differentially expressed proteins involved in each metabolic pathway are shown. Thick arrows indicate the differentially expressed genes; red arrows, 30 °C-culture- vs. 18 °C-culture; blue arrows, 18 °C-culture vs. 4 °C-culture; green arrows, 30 °C-culture vs. 4 °C-culture. Upward and downward arrows represent upregulation and down-regulation, respectively. Solid thin arrows indicate metabolic flows. Dotted arrows indicate the precursor provider pathways. Arrows of the same color are associated with the same pathway. TMA trimethylamine, CP corrinoid protein, MttP trimethylaminepermease, MttB trimethylamine transferase, Tcp trimethylamine corrinoid protein, MtbB dimethylamine transferase, Dcp dimethylamine corrinoid protein, MtmB monomethylamine transferase, Mcp monomethylamine corrinoid protein, CoM coenzyme M, CoB coenzyme B, MF methanofuran, H 4 MPT tetrahydromethanopterin, PRPP 5-phosphoribosyl 1-pyrophosphate, G1P glucose 1-phosphate, G6P glucose 6-phosphate, F6P fructose 6-phosphate, F1,6BP fructose 1,6-biphosphate, Gap glyceraldehyde 3-phosphate, 1,3BPG 1,3-bisphosphoglycerate, 3PG 3- phosphoglycerate, 2PG 2-phosphoglycerate, PEP phosphoenolpyruvate, Fmd formylmethanofuran dehydrogenase, Ftr formylmethanofuran:H4MPT formyltransferase, Mch methenyl-H4MPT cyclohydrolase, Mtd F420-dependent methylene-H4MPT dehydrogenase, Mer methylene-H4MPT reductase, Mtr methyl-H4MPT:coenzyme M methyltransferase, Mcr methyl-coenzyme M reductase, Hdr heterodisulfide reductase, Fpo F420H2 dehydrogenase, CODH/ACS CO dehydrogenase/acetyl-CoA synthetase complex, Pgm phosphoglucomutase, Gpi glucose-6-phosphate isomerase, Fbp/SuhB inositol-1-monophosphatase/fructose-1,6-bisphosphatase, Fba fructose-bisphosphate aldolase, GAPDH Gap dehydrogenase, GAPOR Gap: ferredoxin oxidoreductase, Gpm phosphoglycerate mutase, Eno enolase, PpsA phosphoenolpyruvate synthase, Por pyruvate ferredoxin oxidoreductase, Phi 3-hexulose-6-phosphate isomerase, Hps 3-hexulose-6-phosphate synthase, Rpi ribose-5-phosphate isomerase, Prs ribose-phosphate pyrophosphokinase, Apt adenine phosphoribosyltransferase, DeoA thymidine phosphorylase, aif2b archaeal translation initiation factor, Rubisco ribulose bisphosphate carboxylase oxygenase, Cdh citrate synthase, Pyc pyruvate carboxylase, Mdh malate dehydrogenase, Fth fumarase, Idh isocitrate dehydrogenase, Acons aconitase, α-KGDH succinate thiokinase, Pfk 6-Phosphofructokinase, Pgk phosphoglycerate kinase, Pyk pyruvate kinase
Interestingly, although strain R15 was grown in trimethylamine in this study, proteins involved in methanol-supported methanogenesis were in abundance in all the samples. Similarly, in M. burtonii, proteins for methylamine-derived methanogenesis were found to be expressed abundantly when it was grown in methanol (Williams et al. 2011). This result suggests that these proteins are expressed constitutively, irrespective of the substrates in the growth medium, and can be categorized as “stand-by” proteins. Remarkably, methanol corrinoidmethyl transferase (Mpsy_0909) and methanol corrinoid protein (Mpsy_0908, 3032) were both upregulated in the 4 °C-cultures (Table S2). This result supports a previous finding that methanol-derived methanogenesis can be a cold-adaptive pathway (Jiang et al. 2010) for which the possible molecular basis was determined in a cold-adaptive Methanosarcina mazei (Cao et al. 2014).
Reductive acetyl-CoA synthesis
Methanogens use the reductive acetyl-CoA pathway for CO2 fixation (Fig. 3) to start the biosynthesis of molecular building blocks (Berg et al. 2010). In the pathway, two CO2 molecules are reduced to generate one acetyl-CoA molecule, one of the CO2 is reduced to CO and bound to a nickel atom in the active center of the CO dehydrogenase/acetyl-CoA synthase complex, and another is reduced to a methyl group and bound to the carrier tetrahydropterin. Subsequently, a methyl-transferring corrinoid protein is involved in methyl transfer, and acetyl-coenzyme A (acetyl-CoA) is synthesized from the CO molecule and the methyl group. Six enzymes are involved in this process (Fig. 3), and some [two subunits of formylmethanofuran dehydrogenase (Mpsy_2758, 2759), formylmethanofuran:tetrahydromethanopterinformyl transferase (Mpsy_1734) and CO dehydrogenase/acetyl-CoA synthase complex (Mpsy_2891)] showed elevated abundance in the proteomes of the cells grown at 30 °C compared with the cells grown at 18 °C. The availability of more acetyl-CoA at the higher temperatures would also be beneficial for increased biomass production, and higher temperature-induced increased methanogenesis would generate more substrate CO2 for acetyl-CoA formation.
Gluconeogenesis
The gluconeogenesis pathway is crucial for biosynthesis, for example, amino acid biosynthesis, which is widely present in all organisms (Ferry 1992). The gluconeogenesis pathway converts acetyl-CoA to pyruvate, 3-phosphoglycerate, and fructose 6-phosphate, all of which are precursors for the synthesis of several types of amino acids. The genes encoding the proteins that are essential for gluconeogenesis are all present in the R15 genome (Fig. 3). The abundances of the alpha (Mpsy_1765) and gamma (Mpsy_1767) subunits of pyruvate synthase, the first enzyme in gluconeogenesis catalyzing the carboxylation of acetyl-CoA by CO2 to pyruvate, display a temperature-dependent trend of 30 > 18 > 4 °C (Table S2). Temperature-responsive proteomics analysis also showed that both glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Mpsy_2902) and fructose-bisphosphate aldolase (Fba) (Mpsy_1418), two enzymes involved in gluconeogenesis, displayed increased abundance in the 30 °C-culture compared with in the 18 °C-culture. Fba also showed elevated abundance in the 30 °C-culture compared with the 4 °C-culture, while another Fba encoding gene Mpsy_1203, was reduced at 30 °C compared with 18 °C. In general, it is predicated that gluconeogenesis will be more active at higher temperatures, so as to meet the higher need for the precursors for amino acid synthesis. Consistently, many enzymes that play a role in amino acid synthesis were upregulated at the higher temperatures, including prolyl-tRNA synthetase (Mpsy_2719) (Table S2).
Glycolysis
Glycolysis pathway is absent in non-heterotrophic Archaea (Verhees et al. 2003), except for the endogenous glycogen-degrading methanogens (Verhees et al. 2001), although the gluconeogenic pathway is found in all Archaea. Glycolysis is a reverse pathway of gluconeogenesis except for three steps: fructose-6-phosphate to fructose-1, 6-biphosphate; glyceraldehyde 3-phosphate (Gap) to 3-phosphoglycerate; and phosphoenolpyruvate to pyruvate. Glycolysis uses 6-phosphofructokinase (Pfk), Gap:ferredoxin oxidoreductase, and pyruvate kinase (Pyk) to complete the three steps (Verhees et al. 2003). Pfk (Mpsy_0617) and Pyk (Mpsy_1041) encoding genes were found in the genome of R15 (Fig. 3). Phosphoglycerate kinase, another enzyme involved in glycolysis, peaked its expression at 4 °C (Table S2), implying that R15 might degrade glycogen generated by the gluconeogenesis at cold temperatures.
Incomplete TCA cycle
Tricarboxylic acid cycle (TCA cycle) is key to energy production as well as for the generation of amino acid biosynthesis precursors. Reductive and oxidative incomplete TCA cycles are found in the Methanococcaceae and Methanosarcinaceae families, respectively (Jones et al. 1987). All the genes required for the oxidative incomplete TCA cycle and two of the genes required for the reductive cycle were found in the R15 genome (Fig. 3). Increased abundance of isocitrate dehydrogenase (IDH) (Mpsy_1029) was found in the 18 °C-culture compared with in the 4 °C-culture, and slightly more IDH was found in the 30 °C- compared with in the 18 °C-culture (Table S2). Pyruvate synthase and pyruvate carboxylase convert acetyl-CoA to oxaloacetate, an intermediate of the incomplete TCA cycle. The two subunits of the former (Mpsy_1765, 1767) were also upregulated at the higher temperature (Table S2). Therefore, we predict that the incomplete TCA cycle in R15 is more active at higher temperatures.
AMP recycling pathway
Ribulose-1,5-bisphosphate carboxylase oxygenase (RuBisCO) is one of the key enzymes in the CBB cycle (Finn and Tabita 2004), a CO2 fixation pathway used by all plants, algae, cyanobacteria, and many other photosynthetic bacteria. The RuBisCO gene has been found in some archaeal genomes, but no homologues of phosphoribulokinase, another important enzyme in the pathway, have been found (Finn and Tabita 2004). Similar pathway in Achaea was named AMP recycling pathway, which has been shown to play a role in AMP recycling in Archaea (Sato et al. 2007). This pathway consisted of (1) conversion of fructose-6-phosphate (F6P) to 5-phosphoribosyl 1-pyrophosphate (PRPP) (shared partial steps of the CBB pathway); (2) conversion of PRPP to 3-phosphoglycerate (3PG), and (3) conversion of 3PG to F6P (shared partial steps of gluconeogenesis) (Fig. 3). By using adenosine 5′-diphosphate (ADP)-dependent (AMP-forming) sugar kinases for glycolysis, such as the ADP-dependent phosphofructokinases and AMP-forming acetyl-CoA synthetase, which can be found in the R15 genome, R15 can produce a larger cellular AMP pool than organisms that possess classical ATP-dependent kinases. ATP can be generated from the AMP recycling pathway product 3-PGA; therefore, this pathway can provide anaerobic archaea with the extra energy through AMP (Sato et al. 2007; Allen et al. 2009). More ribose-5-phosphate isomerase (Rpi, Mpsy_1418), Fba (Mpsy_0045), and GAPDH (Mpsy_2902) (the latter two enzymes are shared by the gluconeogenesis pathway) were produced in the 30 °C-culture compared with the 18 °C-culture. Fba was also found in larger amounts in the 18 °C-culture compared with in the 4 °C-culture (Table S2). 3-PG can also be converted to pyruvate, another important biosynthetic precursor. More energy and biosynthetic precursors can be supplied by an active AMP recycling pathway at higher temperatures, which is consistent with the vigorous growth of R15 at higher temperatures.
Differentially expressed enzymes in methanogenesis and other biosynthesis processes may not respond to temperature simply; their abundance change could be a result of adaptation to different growth rates at different temperatures. In general, methanogenesis, the more active reductive acetyl-CoA pathway, gluconeogenesis, glycolysis and the AMP recycling pathway, which supply ATP, reducing equivalents and biosynthetic precursors for the growth of R15 at higher temperatures, together support R15 to be robust at 30 °C.
RNA transcription and processing
The R15 genome encodes a single gene for TATA box binding protein (TBP, Mpsy_1517). As the first component in transcriptional apparatus that binds to DNA, TBP was most abundant at 4 °C, indicating transcription initiation may be suppressed by cold, so more TBP can compensate for the lower enzymatic activities caused by low temperature. The transcription anti-termination protein NusG had the highest amount at 30 °C; therefore, active transcription is predicted at 30 °C and consistent with the active metabolism at higher temperature. Rrp4, a subunit of the RNA degradation complex exosome, which is responsible for binding the RNA to be degraded, was upregulated at 4 °C compared with 18 °C. RNA turnover at cold can be retarded and more RNase proteins are needed.
Protein synthesis and degradation
Compared with the 18 °C-culture, most of ribosomal proteins and translation initial factors of R15 retained the same amount or decreased expression at 4 °C-culture, only a 50S ribosomal protein L11P (Mpsy_1498) and a translation initiation factor IF-6 (Mpsy_0492) increased expression significantly at lower temperature. This indicates that cold would hinder the ribosome biogenesis though some components are induced by cold, or more such proteins are needed to compensate for lower enzymatic activities at cold. Interestingly, 13 protein spots representing 10 differentially expressed ribosomal proteins (Mpsy_0497, 0881, 0974, 1012, 1013, 1127, 1136, 1141, 1498 and 2863) showed reduced expression at 30 °C compared with their expressions at 18 °C (Table S2). Two translation initiation factors (Mpsy_1579, 0492) were also downregulated at 30 °C compared with at 18 °C, except an elongation factor (Mpsy_1015) increased the abundance as the growth temperatures increased. It is generally believed that more numbers of ribosome proteins support rapid growth at higher temperatures (Maaloe 1979; Neidhardt et al. 1990); however, in E. coli, the peptide chain elongation rate always matches the growth rate at the optimal temperature, while at either lower or higher temperatures the protein synthetic machinery seems to be in excess (Farewell and Neidhardt 1998). The observed decreased expression of ribosome proteins at the optimal temperature suggests that the translation initiation or elongation step may be inhibited at higher temperature, which means that less ribosome proteins are needed. Based on this observation, we postulate that although rapid growth occurs at 30 °C, a mismatch between the initiation and elongation steps can occur, which would lead to a smaller number of ribosomes being required.
Two proteasome subunits (Mpsy_2726, 2881) were found to increase the expression at 4 °C compared with at 18 °C, which may indicate that misfolded proteins or denatured proteins also increased at the lower temperature, so that aberrant protein degradation would be activated; or more proteasome proteins were needed to compensate for the reduced enzymatic activities at low temperature.
Although some proteins can fold spontaneously in vivo, most of them need chaperones to facilitate the folding to a native state in the crowded cytosol. Chaperones consisted of three categories: the ribosome-bound trigger factor interacts with all newly synthesized polypeptides; DnaK assists most small proteins (~70 % of total) and larger proteins (~20 % of total) to fold further; and GroEL/ES help ~10 % of total proteins folding, which is an essential cellular process. Chaperones were also functioned in anti-stresses including heat and cold stresses (Feder and Hofmann 1999; Ting et al. 2010). In accordance with the transcriptomic data, in this study, thermosome, group II chaperonin (Mpsy_1969, 2247, 3167), and other chaperones showed increased abundance in 4 °C-culture compared with the 18 °C-culture, except for chaperone protein ATPase AAA-2 (Mpsy_2749) and prefoldin alpha subunit (Mpsy_0490) (Table S2). Furthermore, most of the chaperones, including thermosome, remained the same or even decreased expression at 30 °C compared with at 18 °C, except for DnaK (Mpsy_2550) and a thermosome subunit (Mpsy_2247) were increased expression significantly(fold changes are 1.5 and 1.3). A thermosome subunit (Mpsy_2247) seemed to have been subjected to post-translational modification, as three spots (spot Master Nos. 229, 252, and 253) (Table S2) were identified to be this protein; spot 229 was induced at 30 °C, while the other two were upregulated at 4 °C. Conversely, thermosomes in M. burtonii increased expression at 23 °C in the comparative transcriptomic and proteomic studies, but not at low-temperature condition (Williams et al. 2011; Campanaro et al. 2011), indicating that the thermosomes of M. burtonii only respond to heat.
Anti-oxidation stress proteins
Reactive oxygen species (ROS), such as O2, H2O2, O2 −, and HO∙, are toxic molecules capable of causing oxidative damage to proteins, DNA and lipids (Suzuki and Mittler 2006). Although R15 is an obligate anaerobe, multiple ROS scavenging ways were found in the genome (Chen et al. 2012). Furthermore, we found that the psychrophilic methanogens (R15 and M. burtonii) contained more genes associated with anti-oxidation stress than other methanogens, a hint that psychrophiles are equipped with more ability to clear oxidants. These genes encode proteins that are involved in three pathways to clear ROS in R15, all the genes in these three pathways were found to be upregulated in 4 °C-culture compared with the 18 °C-culture in the previous temperature-responsive transcriptomic study (Chen et al. 2012). Consistent with the previously reported transcriptomic data, a 3-Cys thioredoxin peroxidase (Mpsy_0762) and a superoxide reductase (Mpsy_2711) were upregulated at 4 °C compared with at 18 °C (Table S2) in this proteomic study. Upregulation of anti-oxidation stress proteins were also reported in M. burtonii and some psychrophilic bacteria (D’Amico et al. 2006). Anaerobic electron-transfer enzymes operate at low potentials that facilitate electron transfer to oxygen (Imlay 2003). At low temperatures, the increased solubility of oxygen and stability of ROS (Piette et al. 2011a, b) aggravate the ROS stress. Proteins associated with anti-oxidation stress were not induced in the 30 °C-culture compared with the 18 °C-culture. Therefore, it is predicted that cellular ROS in this methanogen are increased when growing at low temperatures, but not at higher temperatures.
In conclusion, by comparative proteomic study, this study found that proteins involved in methanogenesis, energy conservation and central metabolism were mostly increased expression at higher temperature. However, ribosome biogenesis is suppressed at higher temperatures. More ROS and aberrant proteins can be formed at cold. In addition, some chaperones including thermosomes were increased the expression at 4 °C in R15, suggesting that protein folding at cold is impaired as well. While the two cultured psychrophilic methanogens showed very similar pattern in upregulation of the enzymes in energy production, transcription and translation at higher temperature, and upregulation of RNase and protein degradation and anti-oxidation proteins at low temperature; the distinct difference between the two is the behavior of chaperones that in R15 they were increased at both high and low temperature, while those in M. burtonii were only increased at high temperature. This study also indicated that, though higher temperature promotes efficient biochemistry cellular processes so that higher the rate of growth and methane formation of R15, the heat-sensitive proteins and protein synthesis process in this cold-adaptive archaeon hinder its growth at further elevated temperature, such as, 37 °C. This study provides another evidence of the species-specific cold-adaptive mechanisms.
References
Allen MA, Lauro FM, Williams TJ, Burg D, Siddiqui KS, De Francisci D et al (2009) The genome sequence of the psychrophilic archaeon, Methanococcoides burtonii: the role of genome evolution in cold adaptation. ISME J 3:1012–1035
Bairoch A, Apweiler R (2000) The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res 28:45–48
Berg A, Kockelkorn D, Ramos-Vera WH, Say RF, Zarzycki J, Hügler M et al (2010) Autotrophic carbon fixation in archaea. Nat Rev Microbiol 8:447–460
Bowman JB (2008) Genomic analysis of psychrophilic prokaryotes. In: Margesin R et al (eds) Psychrophiles: from biodiversity to biotechnology. Springer, Berlin/, pp 265–284
Campanaro S, Williams TJ, Burg DW, De Francisci D, Treu L, Lauro FM, Cavicchioli R (2011) Temperature-dependent global gene expression in the Antarctic archaeon Methanococcoides burtonii. Environ Microbiol 13:2018–2038
Cao Y, Li J, Jiang N, Dong X (2014) Mechanism for stabilizing mRNAs involved in methanol-dependent methanogenesis of cold-adaptive Methanosarcina mazei zm-15. Appl Environ Microbiol 80:1291–1298
Chen Z, Yu H, Li L, Hu S, Dong X (2012) The genome and transcriptome of a newly described psychrophilic archaeon, Methanolobus psychrophilus R15, reveal its cold adaptive characteristics. Environ Microbiol Rep 4:633–641
D’Amico S, Collins T, Marx JC, Feller G, Gerday C (2006) Psychrophilic microorganisms: challenges for life. EMBO Rep 7:385–389
De Maayer P, Anderson D, Cary C, Cowan DA (2014) Some like it cold: understanding the survival strategies of psychrophiles. EMBO Rep 15:508–517
Farewell A, Neidhardt FC (1998) Effect of temperature on in vivo protein synthetic capacity in Escherichia coli. J Bacteriol 180:4704–4710
Feder ME, Hofmann GE (1999) Heat-shock proteins, molecular chaperones, and the stress response: evolutionary and ecological physiology. Annu Rev Physiol 61:243–282
Ferry JG (1992) Biochemistry of methanogenesis. Crit Rev Biochem Mol Biol 27:473–503
Finn MW, Tabita FR (2004) Modified pathway to synthesize ribulose 1, 5-bisphosphate in methanogenic Archaea. J Bacteriol 186:6360–6366
Franzmann PD, Stringer N, Ludwig W, Conway De Macario E, Rohde M (1992) A methanogenic archaeon from Ace Lake, Antarctica: Methanococcoides burtonii sp. nov. Sys Appl Microbiol 15:573–581
Imlay JA (2003) Pathways of oxidative damage. Annu Rev Microbiol 57:395–418
Jiang N, Wang Y, Dong X (2010) Methanol as the primary methanogenic and acetogenic precursor in the cold Zoige wetland at Tibetan plateau. Microb Ecol 60:206–213
Jones WJ, Nagle DP Jr, Whitman WB (1987) Methanogens and the diversity of archaebacteria. Microbiol Rev 51:135–177
Maaloe O (1979) Regulation of the protein synthesizing machinery-ribosomes, tRNA, factors, and so on. Biological regulation and development. Plenum Publishing Corp, New York, pp 487–542
Neidhardt FC, Ingraham JL, Schaechter M (1990) Physiology of the bacterial cell: a molecular approach. Sinauer Associates, Inc, Sunderland, p p507
Nichols DS, Miller MR, Davies NW, Goodchild A, Raftery M, Cavicchioli R (2004) Cold adaptation in the Antarctic Archaeon Methanococcoides burtonii involves membrane lipid unsaturation. J Bacteriol 186:8508–8515
Noon KR, Guymon R, Crain PF, McCloskey JA, Thomm M, Lim J, Cavicchioli R (2003) Influence of temperature on tRNA modification in archaea: Methanococcoides burtonii [optimum growth temperature (Topt), 23°C] and Stetteria hydrogenophila (Topt, 95°C). J Bacteriol 185:5483–5490
Piette F, D’Amico S, Mazzucchelli G, Danchin A, Leprince P, Feller G (2011a) Life in the cold: a proteomic study of cold-repressed proteins in the antarctic bacterium Pseudoalteromonas haloplanktis TAC125. Appl Environ Microbiol 77:3881–3883
Piette F, Struvay C, Feller G (2011b) The protein folding challenge in psychrophiles: facts and current issues. Environ Microbiol 13:1924–1933
Rodrigues DF, Tiedje JM (2008) Coping with our cold planet. Appl Environ Microbiol 74:1677–1686
Sato T, Atomi H, Imanaka T (2007) Archaeal type III RuBisCOs function in a pathway for AMP metabolism. Science 315:1003–1006
Suzuki N, Mittler R (2006) Reactive oxygen species and temperature stresses: a delicate balance between signaling and destruction. Physiol Plant 126:45–51
Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT, Rao BS et al (2001) The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res 29:22–28
Thomas T, Kumar N, Cavicchioli R (2001) Effects of ribosomes and intracellular solutes on activities and stabilities of elongation factor 2 proteins from psychrotolerant and thermophilic methanogens. J Bacteriol 183:1974–1982
Ting L, Williams TJ, Cowley MJ, Lauro FM, Guilhaus M, Raftery MJ, Cavicchioli R (2010) Cold adaptation in the marine bacterium, Sphingopyxis alaskensis, assessed using quantitative proteomics. Environ Microbiol 12:2658–2676
Verhees CH, Tuininga JE, Kengen SW, Stams AJ, van der Oost J, de Vos WM (2001) ADP-dependent phosphofructokinases in mesophilic and thermophilic methanogenic archaea. J Bacteriol 183:7145–7153
Verhees CH, Kengen SW, Tuininga JE, Schut GJ, Stams AJ, de Vos WM, van der Oost J (2003) The unique features of glycolytic pathways in Archaea. Biochem J 375:231–246
Williams TJ, Burg DW, Raftery MJ, Poljak A, Guilhaus M, Pilak O, Cavicchioli R (2010) Global proteomic analysis of the insoluble, soluble, and supernatant fractions of the psychrophilic archaeon Methanococcoides burtonii. Part I: the effect of growth temperature. J Proteome Res 9:640–652
Williams TJ, Lauro FM, Ertan H, Burg DW, Poljak A, Raftery MJ, Cavicchioli R (2011) Defining the response of a microorganism to temperatures that span its complete growth temperature range (−2 to 28 °C) using multiplex quantitative proteomics. Environ Microbiol 13:2186–2203
Zhang G, Jiang N, Liu X, Dong X (2008) Methanogenesis from methanol at low temperature by a novel psychrophilic methanogen, Methanolobus psychrophilus sp. nov., prevalent in Zoige wetland of Tibetan Plateau. Appl Environ Microbiol 74:6114–6161
Acknowledgments
This work was supported by National Natural Science foundation of China under No. 30621005, 30830007 and Y2113B5531.
Conflict of interest
The authors have declared no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Z. Chen and D. Feng contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, Z., Feng, D., Zhang, B. et al. Proteomic insights into the temperature responses of a cold-adaptive archaeon Methanolobus psychrophilus R15. Extremophiles 19, 249–259 (2015). https://doi.org/10.1007/s00792-014-0709-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-014-0709-y