Abstract
It has been suggested that archaea carrying the accA gene, encoding the alpha subunit of the acetyl CoA carboxylase, autotrophically fix CO2 using the 3-hydroxypropionate/4-hydroxybutyrate pathway in low-temperature environments (e.g., soils, oceans). However, little new information has come to light regarding the occurrence of archaeal accA genes in high-temperature ecosystems. In this study, we investigated the abundance and diversity of archaeal accA gene in hot springs in Yunnan Province, China, using DNA- and RNA-based phylogenetic analyses and quantitative polymerase chain reaction. The results showed that archaeal accA genes were present and expressed in the investigated Yunnan hot springs with a wide range of temperatures (66–96 °C) and pH (4.3–9.0). The majority of the amplified archaeal accA gene sequences were affiliated with the ThAOA/HWCG III [thermophilic ammonia-oxidizing archaea (AOA)/hot water crenarchaeotic group III]. The archaeal accA gene abundance was very close to that of AOA amoA gene, encoding the alpha subunit of ammonia monooxygenase. These data suggest that AOA in terrestrial hot springs might acquire energy from ammonia oxidation coupled with CO2 fixation using the 3-hydroxypropionate/4-hydroxybutyrate pathway.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The accA gene encodes the alpha subunit of the Acetyl CoA carboxylase (ACCase), a biotin-dependent enzyme that catalyzes the irreversible carboxylation of acetyl-CoA to produce malonyl-CoA (Brownsey et al. 1997). This gene is present in a wide range of organisms and is known to be involved in fatty acid biosynthesis (Moss Lane 1971). However, ACCase would recently be expected to be specific to carbon fixation in Archaea due to their unique lipids and lack of fatty acids (Berg et al. 2007, 2010a). Specifically, ACCase represents one of the key enzymes responsible for CO2 fixation through the 3-hydroxypropionate/4-hydroxybutyrate cycle (Berg et al. 2007). This pathway was first discovered in a thermophilic chemolithoautotroph Metallosphaera sedula (belonging to the archaeal order Sulfolobales) isolated from a terrestrial hot spring (Berg et al. 2007) and has also been suggested to function in other autotrophic members of Sulfolobales (Berg et al. 2007, 2010b) and mesophilic Crenarchaeota (Thaumarchaeota, which includes all known ammonia-oxidizing archaea, i.e. AOA) as well (Hallam et al. 2006; Berg et al. 2007; Hatzenpichler et al. 2008; Tourna et al. 2011).
Recently, several studies investigated the diversity and abundance of archaeal accA and amoA genes in agricultural soils and ocean waters and showed that the 3-hydroxypropionate/4-hydroxybutyrate pathway has significant correlation with archaeal ammonia oxidation process (Yakimov et al. 2009, 2011; Hu et al. 2011a, b; Pratscher et al. 2011). AOA are widespread in terrestrial geothermal sites in Iceland (Reigstad et al. 2008), Kamchatka (Russia) (Zhang et al. 2008; Zhao et al. 2011), Great Basin and Yellowstone National Park (United States) (de la Torre et al. 2008; Zhang et al. 2008), and Yunnan geothermal zone (China) (Zhang et al. 2008; Jiang et al. 2010). The Yunnan geothermal zone is one of the most active geothermal areas in the world. It possesses thousands of hot springs characteristic of a variety of hydrothermal features, such as hydrothermal explosion craters, geysers, fumaroles, and boiling springs (Song et al. 2013). Previously, two cultivation-independent studies showed that AOA amoA genes (encoding the subunit A of ammonia monooxygenase) were abundant and diverse in Yunnnan hot springs with temperatures higher than 74 °C and up to 94 °C (Zhang et al. 2008; Jiang et al. 2010). However, little is known about whether the AOA found in hot springs also autotrophically fix CO2 through the 3-hydroxypropionate/4-hydroxybutyrate pathway.
In order to fill the above knowledge gap, the abundance and diversity of archaeal accA gene were investigated in the samples collected from the Yunnan hot springs with a range of temperature 66–96 °C and pH 4.3–9.0. An integrated approach was employed, including DNA- and RNA-based phylogenetic analyses and quantitative polymerase chain reaction (qPCR).
Materials and methods
Field measurements and sampling
Eight hot springs were selected in this study and they were located in three thermal areas (Tengchong, Longling, and Eryuan) in the Yunnan geothermal zone (Table 1). At each hot spring, water temperature and pH were determined using a portable meter (PT-10, SARTORIUS, Germany). Concentrations of nitrite, nitrate, ammonium, phosphate, and ferrous iron were measured using HACH colorimeter (model CEL 850/product #: 2687900, Hach Chemical Co., Iowa, USA) according to the manufacturer’s instructions in the field. The measurements of major and trace elements were performed with inductively coupled plasma mass spectroscopy (ICP-MS, Thermo, USA) in laboratory. After chemical measurements, mat-containing sinter or sediments were collected into 50-mL Falcon tubes and immediately stored in liquid nitrogen. The samples were kept in liquid nitrogen in the field and during transportation, and then were stored at −80 °C in the laboratory until further analyses.
Nucleic acid isolation
Both DNA- and RNA-based molecular approaches were employed. Total DNA and RNA were extracted from 1 to 2 g of sediment using E.Z.N.A. ® Soil DNA Kit (Omega Bio-Tek Inc., USA) and E.Z.N.A. ® Soil RNA Kit (Omega Bio-Tek Inc., USA), respectively, according to the manufacturer’s instructions. To remove the residual DNA, the soluble crude RNA was digested with RNase-free DNase I (Fermentas, USA). The DNase-digested RNA samples were checked for potential genomic DNA contamination by PCR amplification with specific primer sets of Arch21F/Arch958R, Bac27F/Univ1492R, and Crena_529F/Crena_981R (Table 2) for archaeal and bacterial 16S rRNA genes and accA gene, respectively.
cDNA synthesis by reverse transcription
The checked RNA samples were reverse-transcribed into cDNA using the Fermentas AMV Reverse Transcriptase (Fermentas, USA) and random hexamer primer according to the manufacturer’s protocol. Absence of contamination from DNA and chemical reagents was verified by conducting the same reactions without the AMV reverse transcriptase and template, respectively.
Quantitative PCR (qPCR)
The archaeal accA and amoA genes and archaeal and bacterial 16S rRNA genes were quantified by qPCR with the primer sets listed in Table 2. Amplification conditions were 95 °C for 10 min, followed by 40 cycles of 15 s at 94 °C, 45 s for annealing at temperatures specified in Table 2, 1 min at 72 °C for extension, and data collection. The reaction volume is 20 μL, containing 20–50 ng DNA, 10 μL of SYBR®PREMIX TaqTM (2×) (TaKaRa, China), and 10 pmol of each primer. Plasmids of an accA gene clone (obtained in this study), one amoA gene clone (obtained in this study), and 16S rRNA genes of Shewanella piezotolerans WP3 and Natronomonas sp. were used as standard templates for the qPCR of archaeal accA and amoA genes and bacterial and archaeal 16S rRNA genes, respectively. Standard templates were made into serial dilutions of 102–108 gene copies per micro-liters. R 2 values of the standard curves were 0.996–0.967 for the targeted genes. The qPCR amplification efficiencies were in the range of 85–95 %. All qPCR reactions were performed in triplicate with an ABI7500 real-time thermal cycler (ABI, USA). Melting curve analysis was performed using the default conditions set in ABI7500 (95 °C). The melting curve had only one peak, indicating that the SYBR green signals were not from primer-dimer artifacts or non-specific PCR amplification.
PCR amplification of archaeal accA gene
Archaeal accA genes from DNA and cDNA extracts were PCR amplified using archaeal accA gene-specific primers Crena_528F and Crena_981R (Yakimov et al. 2009; see Table 2 for detail). PCR reactions were carried out in a total volume of 50 μL containing 1× PCR buffer with 1.5 mM Mg2+, dNTPs (100 μM each), 0.25 μM each primer, 2.5 U of DNA polymerase (Ex-Taq) (TaKaRa, Dalian, China), and 1–10 ng of total DNA/cDNA. Amplification consisted of an initial denaturation at 95 °C for 10 min, followed by 35 cycles of denaturation at 95 °C for 45 s, annealing at 50 °C for 1 min and extension at 72 °C, and a final extension at 72 °C for 20 min. PCR products were purified using an E.Z.N.A. ® Gel Extraction Kit (Omega Bio-Tek Inc., USA) according to the manufacturer’s instructions.
Clone library construction and phylogenetic analysis
The archaeal accA gene clone libraries were constructed according to the procedures previously established (Jiang et al. 2010). Briefly, the purified PCR products were ligated into pMD18-T Vector system (TaKaRa, Dalian, China) and transformed into competent Escherichia coli JM109 cells. The transformed cells were plated on Luria–Bertani (LB) plates containing 100 μg/mL of ampicillin, 80 μg/mL of X-Gal (5-bromo-4-chloro-3-indolyl-b-d-galactopyranoside) and 0.5 mM IPTG (isopropyl-b-d-thiogalactopyranoside), and incubated overnight at 37 °C. A total of 20–40 randomly chosen colonies per sample were analyzed for the insert accA gene sequences by PCR amplification with the accA gene specific primer set (Crena_528F/Crena_981R). Archaeal accA gene clone were sequenced using primer M13+ with the BigDye Terminator version 3.1 chemistry (Applied Biosystems, Foster City, CA, USA). The accA gene sequences were determined with an ABI 3730 automated sequencer. The obtained raw archaeal accA clone sequences were edited using the DNASTAR program v.5.0 and their validity was checked by using TBLASTX (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
Operational taxonomic units (OTUs) of the obtained archaeal accA gene clone sequences were identified using the DOTUR 1.53 program (Schloss and Handelsman 2005) at 98 % nt identity cutoff. Coverage (C) was used to assess the adequacy of clone sampling. Coverage of the clone libraries was calculated as follows: C = 1 − (n1/N), where n1 is the number of OTUs that occurred only once in the clone library and N is the total number of clones analyzed (Jiang et al. 2010). One representative sequence was selected from each OTU for further analysis. The selected representative OTU sequences were translated into amino acid sequences using TBLASTX. The representative sequences and their closest references were aligned using CLUSTALX1.83. Phylogeny was constructed using neighbor-joining and maximum likelihood methods with the MEGA 5.0 program (Tamura et al. 2011). Bootstrap analysis was performed using 1000 replications. The archaeal accA gene sequences obtained in this study were deposited in the GenBank database under accession numbers KC433711–KC433733.
Statistical analysis
The diversity indices of Chao 1, Shannon, and Simpson were calculated using the SPADE software (Chao 2010). Pearson correlation analysis and Mantel test (http://bioinformatics.psb.ugent.be/webtools/zt/) were performed to assess the correlation between the environmental variables and the archaeal accA gene abundance and diversity according to the procedures described previously (Jiang et al. 2009).
Results
Environmental characteristics
Water temperature, pH, and major ions were determined at all the investigated hot springs (Table 1). Temperature ranged from 66 to 96 °C, and pH from 4.3 to 9.0. Most of the examined ions were variable in the investigated sites. For example, the highest concentrations of ammonia and nitrate were 382 and 40 mg/L, respectively, but the lowest concentrations of these two variables were below detection limits.
Abundances of archaeal accA and amoA and archaeal and bacterial 16S rRNA genes
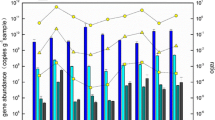
The archaeal accA gene abundance ranged from 2.9 × 104 to 6.8 × 105 copies per gram of dry solids, a little higher than (but was very close to) the archaeal amoA gene abundance, in the investigated hot springs. The abundance of accA gene was markedly lower than that in soils and oceans (Yakimov et al. 2009; Pratscher et al. 2011; Hu et al. 2011b). The abundances of archaeal accA and amoA genes were three to six orders of magnitude lower than that of archaeal 16S rRNA gene copies (Fig. 1).
DNA-based diversity and phylogeny of archaeal accA gene
Eight DNA-based clone libraries were constructed (one for each sample). A total of 224 archaeal accA gene clone sequences were obtained and they could be grouped into 23 OTUs at the cutoff of 98 % identity (Table 3). The coverage ranged from 89 to 100 %, indicating that the sampled clones represent well the AOA accA gene diversity in the studied hot springs. Chao 1, Simpson’s inverse index (D), and Shannon index were 4.5–14.0, 2.1–6.3, and 1.1–2.3, respectively.
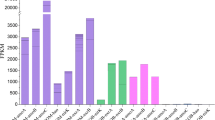
Neighbor-joining and maximum likelihood phylogenetic trees consistently showed that all the obtained archaeal accA gene clone sequences could be divided into three clusters: Clusters A, B, and C (Fig. 2). Clusters A and B were affiliated with AOA lineages. Cluster A was affiliated with Group 1.1a (Dawson et al. 2006) and it was composed of two OTUs (OTU1 and OTU2), representing 6.3 % of the obtained archaeal accA gene clone sequences from hot springs Gmd, Wm3 and Sx4 (temperature 66–84 °C, pH 7.2–9.0) (Fig. 3a). Cluster B consisted of fourteen OTUs (OTU3 to OTU16) and was the dominant group representing of 86 % of the obtained archaeal accA gene clone sequences (Fig. 2). Cluster B has a relatively deep-branching association with all the known lineages of AOA and forms a distinct, well supported sister lineage to Group 1.1a and 1.1b. BLAST analysis showed that clone sequences in Cluster B had 68–71 % amino acid identity with all the closely related sequences known up to date in the NCBI database. Seven OTUs (OTU17 to OTU23) tightly formed Cluster C, which was affiliated with the Desulfurococcales (Fig. 3a). The clone sequences within Cluster C represented a total of 8.5 % of the obtained archaeal accA gene clone sequences.
Neighbor-joining tree showing the phylogenetic relationships of archaeal AccA protein sequences (151 amino acid residues) obtained from the studied samples to closely related sequences from the GenBank database. The sequence of Nitrosotalea devanaterra was provided by Dr Laura Lehtovirta (University of Aberdeen). One clone type within each OTU is shown, and the accession number is shown after the OTU number. The relative abundance of clones represented by each OTU in each of the studied samples is shown after the OTU GenBank accession number. The percentages in the parentheses indicate the abundances of DNA/RNA (two percentages separated by a solidus)-based clones represented by the OTU type, respectively. Asterisk represented that the archaeal accA gene expression was detected. Neighbor-joining- and maximum likelihood-based bootstrap values of >50 % (for 1000 iterations) are indicated above and below the nodes, respectively. Scale bars indicate the Jukes–Cantor distances
cDNA-based diversity and phylogeny of AOA accA gene
cDNA-based clone libraries were successfully constructed on four hot spring samples (Gmd, Wm3, Eynj2, and Sx4). accA gene transcripts were not successfully amplified from the other four samples. The failure was possibly due to the following aspects: (1) the four hot spring samples may contain inhibiting substances affecting RNA harvest; and/or (2) RNA expression of accA gene may be low in failed samples. So the four failed samples were not included in downstream cDNA-based analyses.
A total of 84 cDNA-based archaeal accA gene clone sequences were obtained and they were grouped into seven OTUs at the cutoff of 98 % amino acid similarity. Phylogenetic analyses showed that the cDNA-based OTUs fell into AOA-related Clusters A and B and Desulfurococcales-related Cluster C, and they were affiliated with their DNA-based counterparts (Fig. 2). Clusters A and B include 80 cDNA-based archaeal accA gene clone sequences. The Cluster A-related accA genes were expressed in Gmd and Sx4 hot springs (temperature 66 and 84 °C, respectively), and cluster B-related accA genes were expressed in all the four hot springs (temperature 66 to 84 °C). Desulfurococcales-related Cluster C contains four cDNA-based accA gene clone sequences, indicating that Desulfurococcales-related accA genes were expressed in Gmd and Eynj2 hot springs (temperature 73 and 84 °C, respectively) (Fig. 3b).
Statistical analysis
The simple Mantel tests and Pearson correlation showed that the abundances of the archaeal accA and amoA genes were significantly correlated (r > 0.99, P < 0.005) with each other, and they were significantly positively correlated with concentrations of nitrate (r > 0.8, P < 0.005) and nitrite (r > 0.7, P < 0.005), and were negatively correlated (r < −0.8, P < 0.005) with silicate concentration.
Discussion
Occurrence of archaeal accA genes in Yunnan hot springs
To our knowledge, this study is the first to characterize archaeal accA gene diversity and abundance in terrestrial hot springs. The data presented here proved archaeal accA genes were present in Yunnan hot springs with a wide range of temperature and pH and could be expressed at least in four of the investigated sites with temperatures ranging from 66 to 84 °C (Fig. 3), which was consistent with the temperature range for the distribution and expression of archaeal amoA genes in these hot springs (Zhang et al. 2008; Jiang et al. 2010). Furthermore, the abundances of the archeal accA and amoA genes were almost equal with each other at each investigated site (Fig. 1), which was in agreement with other previous studies in marine and soil environments (Yakimov et al. 2009; Hu et al. 2011b). Taken together, our results suggested that the retrieved archaeal accA genes were mainly derived from AOA. The AOA in the Yunnan hot springs may acquire energy from ammonia oxidation coupled with CO2 fixation using the 3-hydroxypropionate/4-hydroxybutyrate pathway, as has been suggested for their relatives in mesophilic environments (Auguet et al. 2008; Yakimov et al. 2009, 2011; Zhang et al. 2010; Pratscher et al. 2011).
Phylogeny of archaeal accA gene in Yunnan hot spring
The archaeal accA gene-based phylogenetic analysis in this study substantiated the wide distribution of ThAOA in terrestrial geothermal habitats. The cluster B has a relatively deep-branching association with all the known AOA lineages and forms a well-supported lineage distinct from groups 1.1a and 1.1b (Fig. 2). The phylogenitic position of this cluster strongly corresponds to the ThAOA/HWCG III group (Thermophilic Ammonia-Oxidizing Archaea/Hot Water Crenarchaeotic Group III), which represented a high-temperature archaeal amoA gene lineage (Nunoura et al. 2005; de la Torre et al. 2008; Stahl and de la Torre 2012; Hatzenpichler 2012). Cluster B might be related to ThAOA although the accA gene sequence of the representative organism Candidatus Nitrosocaldus yellowstonii is lacking, as the genome sequence is not yet available.
Environmental factors affecting AOA accA genes
Erguder et al. (2009) have reviewed the environmental factors affecting AOA abundance and/or diversity, and such factors include organic carbon, temperature, salinity, DO levels, pH, sulfide levels, and phosphate. Gubry-Rangin et al. (2011) proposed that pH was the major factor governing AOA community structure in global soil ecosystems. However, in this study, the diversity and abundance of archaeal accA genes were not correlated with pH in the investigated hot springs. Our previous study also showed the community structures of Crenarchaeota and Thaumarchaeota in Yunnan hot springs were not affected by pH (Song et al. 2010). In addition, recent studies showed that the “Group 1.1a-associated” AOA preferred acidic or acido-neutral niches in soil ecosystems (Lehtovirta-Morley et al. 2011; Gubry-Rangin et al. 2011); however, AOA accA genes belonging to this group were not detected in acid or acido-neutral soil environments (Lehtovirta-Morley et al. 2011). In contrast, previous archaeal amoA gene-based studies showed that “Group 1.1a-associated” AOA were distributed in Yunnan and Yellowstone hot springs with a wide range of pH (pH 3.4–8.0) (de la Torre et al. 2008; Zhang et al. 2008). Taken together, the AOA populations in hot springs may not be limited by pH only.
In this study, a significant correlation was found between the concentrations of nitrate and nitrite and the abundances of archaeal accA and amoA genes in the studied hot springs. This correlation can be explained by the fact that nitrite and nitrate are the products of ammonia and nitrite oxidation, respectively. Such correlation was also indicative of nitrification activity. However, the concentration of ammonia (serving as the substrate for AOA) was not correlated with the abundances of archaeal accA and amoA genes. This can be explained by the fact that AOA subsisted on a very low level of ammonia (Hatzenpichler 2012), so the concentration of ammonia likely was not limited to AOA present in the investigated hot springs.
Desulfurococcales-related accA genes in Yunnan hot springs
It is notable that a small amount of archaeal accA gene clone sequences obtained in this study were affiliated with Desulfurococcales. Up to date, no research has ever shown that Desulfurococcales-related species autotrophically fix CO2 using the 3-hydroxypropionate/4-hydroxybutyrate pathway; instead, they use another pathway of the dicarboxylate/4-hydroxybutyrate cycle (Huber et al. 2008; Berg et al. 2010a, b; Berg 2011). This inconsistency raises questions about of the function of the accA gene in Desulfurococcales-related organisms in geothermal ecosystems, and awaits further investigations. In summary, our study demonstrates the presence and expression of archaeal accA genes in Yunnan hot springs with a wide range of temperatures and pH. The obtained archaeal accA genes were mainly derived from AOA. The AOA in Yunnan hot springs might acquire energy from ammonia oxidation coupled with CO2 fixation using the 3-hydroxypropionate/4-hydroxybutyrate pathway.
References
Auguet JC, Borrego CM, Bañeras L, Casamayor EO (2008) Fingerprinting the genetic diversity of the biotin carboxylase gene (accC) in aquatic ecosystems as a potential marker for studies of carbon dioxide assimilation in the dark. Environ Microbiol 10:2527–2536
Berg IA (2011) Ecological aspects of the distribution of different autotrophic CO2 fixation pathways. Appl Environ Microbiol 77:1925–1936
Berg IA, Kockelkorn D, Buckel W, Fuchs G (2007) A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea. Science 318:1782–1786
Berg IA, Kockelkorn D, Ramos-Vera WH, Say RF, Zarzycki J, Hügler M, Alber BE, Fuchs G (2010a) Autotrophic carbon fixation in archaea. Nat Rev Microbiol 8:447–460
Berg IA, Ramos-Vera WH, Petri A, Huber H, Fuchs G (2010b) Study of the distribution of autotrophic CO2 fixation cycles in Crenarchaeota. Microbiology 156:256–269
Brownsey RW, Zhande R, Boone AN (1997) Isoforms of acetyl-CoA carboxylase: structures, regulatory properties and metabolic functions. Biochem Soc Trans 25:1232–1238
Chao A (2010) SPADE: species prediction and diversity estimation. URL http://chao.stat.nthu.edu.tw/softwareCE.html
Dawson S, DeLong E, Pace NR (2006) Phylogenetic and ecological perspectives on uncultured crenarchaeota and korarchaeota. In: Dworkin M (ed) The Prokaryotes. Springer-Verlag, Berlin Release 3.7
de la Torre JR, Walker CB, Ingalls AE, Konneke M, Stahl DA (2008) Cultivation of a thermophilic ammonia oxidizing archaeon synthesizing crenarchaeol. Environ Microbiol 10:810–818
Erguder TH, Boon N, Wittebolle L, Marzorati M, Verstraete W (2009) Environmental factors shaping the ecological niches of ammonia oxidizing archaea. FEMS Microbiol Rev 33:855–869
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB (2005) Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc Natl Acad Sci USA 102:14683–14688
Gubry-Rangin C, Hai B, Quince C, Engel M, Thomson BC, James P, Schloter M, Griffiths RI, Prosser JI, Nicol GW (2011) Niche specialization of terrestrial archaeal ammonia oxidizers. Proc Natl Acad Sci USA 108:21206–21211
Hallam SJ, Mincer TJ, Schleper C, Preston CM, Roberts K, Richardson PM, DeLong EF (2006) Pathways of carbon assimilation and ammonia oxidation suggested by environmental genomic analyses of marine Crenarchaeota. PLoS Biol 4:e95
Hatzenpichler R (2012) Diversity, physiology, and niche differentiation of ammonia-oxidizing archaea. Appl Environ Microbiol 78:7501–7510
Hatzenpichler R, Lebedeva EV, Spieck E, Stoecker K, Richter A, Daims H, Wagner M (2008) A moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. Proc Natl Acad Sci USA 105:2134–2139
Hu A, Jiao N, Zhang CL (2011a) Community structure and function of planktonic Crenarchaeota: changes with depth in the South China Sea. Microb Ecol 62:549–563
Hu A, Jiao N, Zhang R, Yang Z (2011b) Niche partitioning of marine group I Crenarchaeota in the euphotic and upper mesopelagic zones of the East China Sea. Appl Environ Microbio 77:7469–7478
Huber H, Gallenberger M, Jahn U, Eylert E, Berg IA, Kockelkorn D, Eisenreich W, Fuchs G (2008) A dicarboxylate/4-hydroxybutyrate autotrophic carbon assimilation cycle in the hyperthermophilic Archaeum Ignicoccus hospitalis. Proc Natl Acad Sci USA 105:7851–7856
Jiang H, Dong H, Deng S, Yu B, Huang Q, Wu Q (2009) Response of archaeal community structure to environmental changes in lakes on the Tibetan Plateau, northwestern China. Geomicrobiol J 26:289–297
Jiang H, Huang Q, Dong H, Wang P, Wang F, Li W, Zhang CL (2010) RNA-based investigation of ammonia-oxidizing archaea in hot springs of Yunnan Province, China. Appl Environ Microbiol 76:2541–4538
Lane DJ (1991) 16S/23S rRNA Sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley Sons, New York, pp 115–175
Lehtovirta-Morley LE, Stoecker K, Vilcinskas A, Prosser JI, Nicol GW (2011) Cultivation of an obligate acidophilic ammonia oxidizer from a nitrifying acid soil. Proc Natl Acad Sci USA 108:15892–15897
Moss J, Lane MD (1971) Biotin-dependent enzymes. Adv Enzymol Relat Areas Mol Biol 35:321–442
Nadkarni M, Martin FE, Jacques NA, Hunter N (2002) Determination of bacterial load by real-time PCR using a broad range (universal) probe and primer set. Microbiology 148:257–266
Nunoura T, Hirayama H, Takami H, Oida H, Nishi S, Shimamura S, Suzuki Y, Inagaki F, Takai K, Nealson KH, Horikoshi K (2005) Genetic and functional properties of uncultivated thermophilic crenarchaeotes from a subsurface gold mine as revealed by analysis of genome fragments. Environ Microbiol 7:1967–1984
Pratscher J, Dumont MG, Conrad R (2011) Ammonia oxidation coupled to CO2 fixation by archaea and bacteria in an agricultural soil. Proc Natl Acad Sci USA 108:4170–4175
Reigstad LJ, Richter A, Daims H, Urich T, Schwark L, Schleper C (2008) Nitrification in terrestrial hot springs of Iceland and Kamchatka. FEMS Microbiol Ecol 64:167–174
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71:1501–1506
Song Z, Chen J, Jiang HC, Zhou EM, Tang SK, Zhi XY, Zhang L, Zhang CL, Li W (2010) Diversity of Crenarchaeota in terrestrial hot springs in Tengchong, China. Extremophiles 14:287–296
Song Z, Wang F, Zhi X, Chen J, Zhou E, Liang F, Xiao X, Tang S, Jiang H, Zhang CL, Dong H, Li W (2013) Bacterial and archaeal diversities in Yunnan and Tibetan hot springs, China. Environ Microbiol 15:1160–1175
Stahl DA, de la Torre JR (2012) Physiology and diversity of ammonia-oxidizing archaea. Annu Rev Microbiol 66:83–101
Takai K, Horikoshi K (2000) Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl Environ Microbiol 66:5066–5072
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tourna M, Stieglmeier M, Spang A, Könneke M, Schintlmeister A, Urich T, Engel M, Schloter M, Wagner M, Richter A, Schleper C (2011) Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci USA 108:8420–8425
Yakimov MM, Conoa VL, Denaroa R (2009) A first insight into the occurrence and expression of functional amoA and accA genes of autotrophic and ammonia-oxidizing bathypelagic Crenarchaeota of Tyrrhenian Sea. Deep Sea Res II 56:748–754
Yakimov MM, Cono VL, Smedile F, DeLuca TH, Juárez S, Ciordia S, Fernández M, Albar JP, Ferrer M, Golyshin PN, Giuliano L (2011) Contribution of crenarchaeal autotrophic ammonia oxidizers to the dark primary production in Tyrrhenian deep waters (Central Mediterranean Sea). ISME J 5:9459–9461
Zhang CL, Ye Q, Huang Z, Li W, Chen J, Song Z, Zhao W, Bagwell C, Inskeep WP, Ross C, Gao L, Wiegel J, Romanek CS, Shock EL, Hedlund BP (2008) Global occurrence of archaeal amoA genes in terrestrial hot springs. Appl Environ Microbiol 74:6417–6426
Zhang LM, Offre PR, He JZ, Verhamme DT, Nicol GW, Prosser JI (2010) Autotrophic ammonia oxidation by soil thaumarchaea. Proc Natl Acad Sci USA 107:17240–17245
Zhao W, Song Z, Jiang H, Li W, Mou X, Christopher SR, Juergen W, Dong H, Zhang CL (2011) Ammonia-oxidizing Archaea in Kamchatka Hot Springs. Geomicrobiol J 28:149–159
Acknowledgments
This work was supported by grants from the National Basic Research Program of China (No. 2010CB833801 and 2012CB822004), the National Natural Science Foundation of China (Nos. 31070007, 40972211, 41030211, 41002123, and 41120124003), the Key Project of International Cooperation of China Ministry of Science and Technology (MOST) (No. 2013DFA31980), Key Project of Yunnan Provincial Natural Science Foundation (2013FA004), Program for New Century Excellent Talents in University, MOE (NCET-12-0954), and the Fundamental Research Funds for the China Central Universities, China University of Geosciences (Wuhan). W-J Li was also supported by ‘Hundred Talents Program’ of the Chinese Academy of Sciences. The authors also thank Chaolei Jiang, Jianxin Fang, Jinchuan Yin, Shaochuan Gong, Jiafu Sheng, and Mr. Ma and the entire staff from Yunnan Tengchong Volcano and Spa Tourist Attraction Development Corporation for sampling support, and Drs JI Prosser and Laura Lehtovirta (Institute of Biological and Environmental Sciences, University of Aberdeen) for providing accA gene of Nitrosotalea devanaterra. The authors are grateful to three anonymous reviewers whose constructive comments significantly improved the quality of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by F. Robb.
Rights and permissions
About this article
Cite this article
Song, ZQ., Wang, L., Wang, FP. et al. Abundance and diversity of archaeal accA gene in hot springs in Yunnan Province, China. Extremophiles 17, 871–879 (2013). https://doi.org/10.1007/s00792-013-0570-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-013-0570-4