Abstract
Thermoplasma acidophilum utilizes l-rhamnose as a sole carbon source. To determine the metabolic pathway of l-rhamnose in Archaea, we identified and characterized l-rhamnose dehydrogenase (RhaD) in T. acidophilum. Ta0747P gene, which encodes the putative T. acidophilum RhaD (Ta_RhaD) enzyme belonging to the short-chain dehydrogenase/reductase family, was expressed in E. coli as an active enzyme catalyzing the oxidation of l-rhamnose to l-rhamnono-1,4-lactone. Analysis of catalytic properties revealed that Ta_RhaD oxidized l-rhamnose, l-lyxose, and l-mannose using only NADP+ as a cofactor, which is different from NAD+/NADP+-specific bacterial RhaDs and NAD+-specific eukaryal RhaDs. Ta_RhaD showed the highest activity toward l-rhamnose at 60 °C and pH 7. The K m and k cat values were 0.46 mM, 1,341.3 min−1 for l-rhamnose and 0.1 mM, 1,027.2 min−1 for NADP+, respectively. Phylogenetic analysis indicated that branched lineages of archaeal RhaD are quite distinct from those of Bacteria and Eukarya. This is the first report on the identification and characterization of NADP+-specific RhaD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thermoacidophilic Archaea, such as Thermoplasma acidophilum and Sulfolobus solfataricus, are thought to metabolize glucose via the branched Entner–Doudoroff (ED) pathway, an ED-like pathway in which hexose intermediates are not phosphorylated or semi-phosphorylated (Ahmed et al. 2004, 2005; Buchanan et al. 1999; Lamble et al. 2003, 2005; Reher et al. 2010; Sato and Atomi 2011). Previously, we identified and characterized several archaeal ED (aED) pathway enzymes, including gluconate dehydratase (Kim and Lee 2005; Lamble et al. 2004), KDG kinase (Kim and Lee 2006a; Potter et al. 2008), glycerate kinase (Noh et al. 2006), and glyceraldehyde dehydrogenase (Jung and Lee 2006). Although the details of the aED pathway of glucose are known and most enzymes in the aED pathway have been identified and characterized, relatively little is known about the metabolism of sugars other than glucose in thermoacidophilic Archaea.
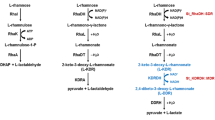
l-Rhamnose is a deoxy sugar that comprises a portion of glycosides in bacteria and plants. In bacteria, l-rhamnose is found in cell surface polysaccharides and in plants l-rhamnose is present as a component of rhamnogalacturonans, which are major pectic polysaccharides of the primary cell walls (de Leder Kremer and Gallo-Rodriguez 2004). l-Rhamnose is also a component of ulvans, the major cell-wall matrix polysaccharides of green seaweeds (Lahaye and Robic 2007). Catabolism of l-rhamnose can be divided into two pathways: phosphorylated and non-phosphorylated metabolic pathways. The phosphorylated pathway for l-rhamnose degradation is found in many bacteria, including E. coli (Power 1967; Wilson and Ajl 1955). In the phosphorylated l-rhamnose pathway in E. coli, l-rhamnose is first isomerized to l-rhamnulose by l-rhamnose isomerase (Takagi and Sawada 1964a; Wilson and Ajl 1957a) and phosphorylated by l-rhamnulose kinase to yield l-rhamnulose 1-phosphate (Takagi and Sawada 1964b; Wilson and Ajl 1957b), which is then converted into l-lactaldehyde and dihydroxyacetone-phosphate by l-rhamnulose 1-phosphate aldolase (Sawada and Takagi 1964). l-Lactaldehyde can be reduced to 1,2-propanediol or oxidized to lactic acid depending on the redox conditions.
The non-phosphorylated pathway for l-rhamnose was identified in the fungus Aureobasidium pullulans (Rigo et al. 1976) and the yeasts Pichia stipitis and Debaryomyces polymorphus (Koivistoinen et al. 2008). This pathway resembles the aED pathway for glucose (Watanabe et al. 2008). l-Rhamnose dehydrogenase (RhaD) oxidizes l-rhamnose into l-rhamno-1,4-lactone in the first step. l-Rhamno-1,4-lactone is subsequently hydrolyzed to l-rhamnonate by l-rhamno-1,4-lactonase. l-Rhamnonate is then converted into l-lactaldehyde and pyruvate via 2-keto-3-deoxy-l-rhamnonate (l-KDR) by l-rhamnonate dehydratase and l-KDR aldolase. Recently, it has been reported that there is another modified non-phosphorylated l-rhamnose pathway in Sphingomonas species (Watanabe and Makino 2009). In the alternative pathway, the l-KDR intermediate is converted to l-lactate and pyruvate via 2,4-diketo-3-deoxy-l-rhamnonate (l-DKDR) by l-KDR 4-dehydrogenase and l-DKDR hydrolase.
We have found that T. acidophilum utilizes l-rhamnose as a sole carbon source. However, the l-rhamnose metabolic pathway has not been reported in any species of the archaeal domain. In the present study, we identified and characterized T. acidophilum l-rhamnose dehydrogenase (Ta_RhaD), catalyzing the oxidation of l-rhamnose to l-rhamnono-1,4-lactone in the non-phosphorylated l-rhamnose pathway. The identification and characterization of l-rhamnose dehydrogenase will contribute to the elucidation and understanding of l-rhamnose catabolism in T. acidophilum.
Materials and methods
Strains and culture conditions
T. acidophilum (JCM9062) was obtained from the Japan Collection of Microorganisms. The composition of the growth medium used was (per L) yeast extract 0.5 g, (NH4)2SO4 1.3 g, KH2PO4 0.3 g, MgSO4·7H2O 0.5 g, CaCl2·2H2O 0.25 g, and carbon source 2.0 g (initial pH 2.0). Cultures were aerobically grown in a 500 mL screw-capped flask at 60 °C and 250 rpm. Escherichia coli TOP10 (Invitrogen) was used to construct recombinant plasmid, and E. coli BL21 CodonPlus (DE3)-RIL (Invitrogen) was selected as expression strain. E. coli BL21 CodonPlus (DE3)-RIL has tRNAs coding argU (AGA, AGG), ileY (AUA), and leuW (CUA).
Construction of recombinant plasmid
The RhaD gene was cloned by PCR using T. acidophilum genomic DNA as a template. The primers were designed: sense direction, 5′-cgggatccgATGCTCGACTTCAAAG-3′; antisense direction, 5′-ggggtaccTTATTGAAGATTTATAAGC-3′. This primer contained restriction sites (underlined) for BamHI and KpnI upstream and downstream, respectively, from the initiation site. PCR was carried out using a PCR Thermal Cycler (Takara, Japan) and Maxime™ PCR premix, (Intron, USA) including Taq polymerase and dNTPs. The PCR fragments were cloned into the corresponding site of the pRSET B vector and then transformed into E. coli TOP10 and BL21 CodonPlus (DE3)-RIL strain.
Expression of recombinant protein
The recombinant protein was expressed in E. coli strain BL21 CodonPlus (DE3)-RIL, which is similarly to our previous studies (Kim and Lee 2006b). Seed culture was carried out in 10 mL of Luria–Bertani medium supplemented with appropriate antibiotics in 50 mL conical tubes at 37 °C and 250 rpm [E. coli BL21-Codon-Plus (DE3)-RIL: 34 μg/mL of chloramphenicol and 100 μg/mL of ampicillin]. Main culture was performed in 1 L of LB medium in 2-L flasks at 37 °C. For induction of recombinant protein, 1 mM (as final concentration) isopropyl-1-thio-β-d-galactopyranoside (IPTG) was added to the culture media when the recombinant cells reached an optical density of 0.4–0.6 at OD600. The recombinant cells were harvested at an optical density of 1.8–2.0 at OD600.
Purification of recombinant dehydrogenase
Harvested cells were concentrated 10-fold by resuspending the cell pellet in 100 mM sodium phosphate, 300 mM NaCl, and 10 mM imidazole (pH 7). Cell lysis was achieved by sonication at 25 % amplification for 1 h on ice with an Ultrasonic Disintegrator, after which the cell debris was removed by centrifugation at 13,000 rpm for 1 h at 4 °C. The cell lysate was subjected to Ni-NTA agarose (QIAGEN) column chromatography to purify His-tagged proteins. The proteins were eluted stepwise by increasing the imidazole concentration (0–250 mM). The Ni-NTA purification fractions containing enzyme activity were then pooled, desalted through a HiPrep 26/10 desalting column, and the purified enzymes were concentrated using a Vivaspin™ concentrator (Vivascience, Lincoln, UK). The enzymes were then loaded into a vial and centrifuged (13,000 rpm × 30 min) to a sufficient concentration. Protein concentration was measured by Bradford assay (Bradford 1976). Standard curve was drawn using bovine serum albumin (BSA) as standard sample, and protein concentration was measured in triplicate.
RhaD assay
RhaD activity was determined spectrophotometrically at A 340. Reaction mixtures (total volume of 1 mL) containing 10 mM l-rhamnose, 1 mM NADP+ as coenzyme, and appropriate amounts of enzyme in 100 mM sodium phosphate (pH 7) were incubated at 60 °C for 4 min. Any increase in absorption at A 340 due to reduction of NADP+ was monitored in a spectrophotometer (Valero and Garcia-Carmona 1996). Enzyme activity was calculated using the molar absorption coefficient of NADPH at A 340 (ε = 5,841 M−1 × cm−1). One unit of RhaD activity was defined as the amount of enzyme required to reduce 1 μmol NADP+ per min at 60 °C and pH 7.
Substrate specificity and kinetic parameters
The substrate specificity of RhaD for sugar groups was determined using NAD+ or NADP+ as a cofactor. The kinetic parameters for RhaD were continuously determined by measuring its enzyme activity at various concentrations of l-rhamnose, l-lyxose, l-mannose, and NADP+. The initial reaction rates were obtained by a linear regression of time-course data. Apparent V max and K m values were calculated by fitting initial rate data to the Lineweaver–Burk plot.
Optimum temperature and optimum pH
The effects of temperature and pH on the enzymatic activity of RhaD were determined using the assay method described above. Temperature profiles were determined between 20 and 90 °C by incubating the purified enzyme at 3 μg/mL in 100 mM sodium phosphate (pH 7). The effect of pH on RhaD activity was determined at 60 °C in 100 mM citrate-NaOH buffer (pH 3–6), 100 mM sodium phosphate (pH 6–9), and 100 mM glycine/NaOH buffer (pH 9–11). All pH values were measured at 60 °C.
Bioinformatic tools
Amino acid sequence homologies and alignment were analyzed using PSI-BLAST database and ClustalX. The phylogenetic trees were built using the neighbor-joining method and the maximum likelihood in MEGA 5.0 software (Tamura et al. 2007).
Results and discussion
Identification of the Ta_RhaD
To examine the existence of metabolic pathways of deoxy-pentoses in T. acidophilum, cells were cultivated in a medium containing l-rhamnose, d-fucose, or l-fucose as a limiting carbon source. For comparison purposes, cells were also cultivated in a medium containing hexoses. Experimental data (Supplementary Fig. S1) showed that all sugars tested could be utilized by T. acidophilum. After 48 h of cultivation, cell densities of T. acidophilum were in the order of d-glucose > d-galactose > l-rhamnose > d-fucose > l-fucose. These results indicate that T. acidophilum was able to utilize deoxy-pentoses. However, no enzymes related to deoxy-pentose metabolism are known in T. acidophilum. In the present study, we identified and characterized T. acidophilum l-rhamnose dehydrogenase (Ta_RhaD), the first enzyme of the non-phosphorylated l-rhamnose pathway.
To determine the genes involved in l-rhamnose utilization, we conducted a proteomic study on cells grown in medium containing l-rhamnose. From the proteomic profiles of T. acidophilum grown, we found that Ta0747 gene product was induced by the presence of l-rhamnose. On the other hand, Ta0747 protein was not detected in the presence of d-glucose. This indicated that the Ta0747 protein is an inducible enzyme responsible for the utilization of l-rhamnose (details will be described elsewhere). In addition, Ta0746, which is located in front of Ta0747, was found to be homologous with the transporter (lp_3596) associated with the l-rhamnose utilization in Lactobacillus plantarum (Beekwilder et al. 2009). The phylogenetic analysis of Ta0746 and other sugar transporter in the major facilitator superfamily (MFS) (Pao et al. 1998) revealed that Ta0746 belongs to a novel family of l-rhamnose transporter in the MFS (Supplementary Fig. S2). This suggested that Ta0746 and Ta0747 are part of a gene cluster associated with the l-rhamnose utilization in T. acidophilum.
Ta0747 gene (NCBI accession number NP_394211), annotated as glucose dehydrogenase homolog, is composed of 245 amino acids with a GTG start codon. However, it has been reported that the Ta0747 gene product is not expressed in E. coli (Nishiya et al. 2004). From the sequence alignment of Ta0747 homologs, we found that the size of the Ta0747 gene is shorter than its homologs, and an ATG start codon exists in front of Ta0747 in the T. acidophilum genome. To distinguish the new extended ORF from the original Ta0747 gene, we named it as Ta0747P (P = POSTECH, JN375693). As shown in Fig. 1a, Ta0747P consists of 254 amino acids. The size of Ta0747P is similar to that of other short-chain dehydrogenase/reductase (SDR) family proteins (Fig. 2). As described below, Ta0747P gene encoded a completely active form of T. acidophilum RhaD.
Amino acid alignment of RhaDs from Bacteria, Eukarya, and Archaea. Cofactor-binding motif (*, GXXXGIG) and catalytic triad (#, S-Y-K) are shown. The arrow (↓) indicates Arg residue at position 16 that is only present on sequences of RhaDs with NADP+ preference. In the alignment, the conserved amino acid residues are displayed by black shading with white letters (identity of 100 %), dark-gray shading with white letters (>80 %), and gray shading with black letters (>60 %). The RhaDs are as follows: Ta_RhaD, Thermoplasma acidophilum (JN375693); Fa_RhaD, Ferroplasma acidarmanus (ZP05570274); Av_RhaD, Azotobacter vinelandii (EAM07804); Sp_RhaD, Sphingomonas sp. SKA58 (EAT09360); Ps_RhaD, Pichia stipitis (ABN68405); Dh_RhaD, Debaryomyces hansenii (CAG87576)
PSI-BLAST searches showed that the amino acid sequence of Ta_RhaD (Ta0747P) is homologous with many bacterial SDRs. Calculated sequence identities were 84 % for the dehydrogenase of Thermoplasma volcanium (NP_110967), 62 % for SDR of Ferroplasma acidarmanus (Fa_RhaD, ZP_05570274), 51 % for RhaD of Sphingomonas sp. (Sp_RhaD, EAT_09360), and 46 % for RhaD of Azotobacter vinelandii (Av_RhaD, EAM_07804). While two highly homologous Ta0747P proteins exist in Euryarchaeota (T. volcanium and F. acidarmanus), relatively few homologs could be found in Crenarchaeota such as Sulfolobus species (sequence identity <30 %).
Sequence alignments of archaeal RhaDs with previously reported bacterial and eukaryal RhaDs (Watanabe et al. 2008; Watanabe and Makino 2009) are shown in Fig. 2. Ta_RhaD, similar to bacterial and eukaryal RhaDs, includes the catalytic triad (Ser144-Tyr157-Lys161) and the cofactor-binding motif (Gly13-X-X-X-Gly17-Ile18-Gly19) (Fujimoto et al. 2001; Kallberg et al. 2002; Oppermann et al. 2003). This suggests that the fundamental catalytic mechanism and cofactor recognition of archaeal RhaD are similar to those of known SDR enzymes. It was reported that most structure-determined SDR enzymes with NADP+ specificity have Arg or Lys in Rossmann fold and that the enhanced population of positively charged Arg or Lys favors the binding of NADP+ because of its negatively charged phosphate (Pletnev et al. 2004; Tanaka et al. 1996; Wermuth and Sciotti 2001). NADP+-specific Ta_RhaD contains a conserved Arg16. Therefore, we assumed that Arg16 residue is important for NADP cofactor specificity. This was also reflected by the phylogenetic relationship of these enzymes, each forming distinct clusters in RhaD subfamilies. From the phylogenetic distribution of RhaD families, we detected three discrete families of known and putative RhaDs in sequenced genomes. As shown in Fig. 3, RhaD subfamilies were separated from other members of the SDR superfamily and branched lineages of archaeal RhaD, which include Ta_RhaD protein, are quite distinct from branches of bacterial RhaDs and eukaryal RhaDs. It is known that Av_RhaD and Sp_RhaD from Bacteria display NAD+/NADP+ dual cofactor specificities and that Dh_RhaD and Ps_RhaD from Eukarya exhibit NAD+ specificities. To our knowledge, this is the first study on the identification and characterization of an NADP+-specific RhaD.
The phylogenetic relationships between RhaDs from Bacteria, Eukarya, and Archaea. Characterized enzymes are marked by closed triangles and by a closed circle for Ta0747P enzyme. The number on each branch indicates the bootstrap value. The RhaDs (GenBank™ accession numbers) are as follows: Ps_RhaD, Pichia stipitis RhaD (ABN68405); Cl_SDR, Clavispora lusitaniae SDR (XP002617482); Dh_RhaD, Debaryomyces hansenii RhaD (CAG87576); Pp_SDR, Pichia pastoris SDR (XP002493760); Mg_SDR, Meyerozyma guilliermondii SDR (XP001484210); Ag_SDR, Arthroderma gypseum SDR (XP003172237); Pc_SDR, Penicillium chrysogenum SDR (XP002567487); Av_RhaD, Azotobacter vinelandii RhaD (EAM07804); He_SDR, Halomonas elongata SDR (YP003899282); Bs_SDR, Brevundimonas subvibrioides SDR (YP003819797); Cs_SDR, Caulobacter segnis SDR (YP003593807); Sp_RhaD, Sphingomonas sp. RhaD (EAT09360); Ae_SDR, Asticcacaulis excentricus SDR (YP004087943); Fa_SDR, putative Ferroplasma acidarmanus SDR (ZP05570274); Tv_SDR, Thermoplasma volcanium SDR (NP110967); Ta_RhaD, Thermoplasma acidophilum RhaD ‘Ta0747P’ (JN375693). Other SDR members are as follows (GenBank™ accession numbers or PDB codes): Ta_GluD, d-glucose dehydrogenase from T. acidophilum (NP393669); Bm_GluD, d-glucose dehydrogenase from Bacillus megaterium (1GCO); Ta_ManD, d-mannose dehydrogenase from T. acidophilum (NP394218); Tt_ManD, d-mannose dehydrogenase from Thermus thermophilus (YP143635); Gf_SorR, l-sorbose reductase from Gluconobacter frateurii (3AI1); Cc_XylD, d-xylose dehydrogenase from Caulobacter crescentus (AAK22854); Ec_Kdg5D, 2-keto-3-deoxy-d-gluconate 5-dehydrogenase from Erwinia chrysanthemi (CAA43989); Go_Gla5D, d-gluconate 5-dehydrogenase from Gluconobacter oxydans (CAA56322)
Characterization of recombinant Ta_RhaD
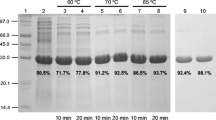
To produce the Ta0747P gene product as an active protein, it was heterologously expressed in E. coli and purified to homogeneity by heat treatment at 55 °C, followed by Ni–NTA affinity chromatography. The purified Ta0747P protein showed a single band in 12 % SDS-PAGE, with a molecular mass of approximately 34 kDa (Fig. 1b). The molecular mass of the single band closely corresponded to the sum of the calculated values of 31,237 Da for Ta0747P and 4,060 Da for His6-tag. To confirm the purified Ta0747P protein, Western blot analysis with anti-His-tag antibody was additionally performed (Fig. 1c).
Twenty-six sugars were tested as possible substrates for Ta_RhaD (Table 1). The highest activity was observed with l-rhamnose. Enzyme activity was assayed routinely in the direction of substrate oxidation by measuring the reduction of NADP+ at A 340. Among the substrates tested, Ta_RhaD was active toward l-rhamnose (100 %), l-lyxose (97 %), and l-mannose (44 %) when NADP+ was used as a cofactor (Table 1). As shown in Table 1, l-rhamnose, l-lyxose, and l-mannose have the same configuration at C1, C2, C3, and C4. Less than 1 % activity was observed for d-mannose, myo-inositol, d-glyceraldehyde, and d-talose. On the other hand, no catalytic activity was detected for the following substrates: d-glucose, d-galactose, l-galactose, d-arabinose, l-arabinose, d-xylose, l-xylose, d-fucose, l-fucose, d-fructose, d-mannitol, d-xylitol, d-sorbitol, d-ribose, d-deoxyribose, d-glucosamine, N-acetyl-d-glucosamine, and d-altrose. When NAD+ was used as a cofactor, Ta_RhaD displayed no activity at all.
Kinetic parameters of Ta_RhaD are summarized in Table 1. The effect of substrate concentration on the activity of purified Ta_RhaD was examined in the range 0–20 mM for sugar substrates and 0–0.5 mM for NADP+ (data are shown in Supplementary Fig. S3). Ta_RhaD enzyme showed highest affinity toward l-rhamnose, with a K m value of 0.46 mM. Turnover number (k cat) was calculated to be 1,341.3 min−1 for l-rhamnose, which yielded a k cat/K m value of 2,946.9 mM−1 × min−1. The k cat/K m value with l-rhamnose was 2.8-fold and 22.2-fold higher than those with l-lyxose and l-mannose, respectively. Figure 4 shows the effects of temperature and pH on Ta_RhaD activity. Purified enzyme displayed optimal activity between 55 and 60 °C, which is similar to the optimal temperature for T. acidophilum growth (Darland et al. 1970). The Ta_RhaD enzyme showed optimal activity at pH 7, which is close to the intracellular pH 6.4–6.9 of T. acidophilum (Hsung and Haug 1975; Searcy 1976).
Effects of temperature and pH on activity of Ta_RhaD. Enzyme activities were assayed at various temperatures (closed symbol) from 20 to 90 °C with intervals of 10 °C. Each temperature point is shown as a closed square shape. Assays were performed as described in the “Materials and methods” section. The enzyme activities at various pHs (open symbol) were assayed. For the pH test, 100 mM citrate-NaOH buffer (pH 3–6; open circles), 100 mM sodium phosphate buffer (pH 6–9; open squares), and 100 mM glycine/NaOH buffer (pH 9–11; open triangles) were used. The dashed line represents l-rhamnose activity while the dotted line indicates l-lyxose activity
Catalytic features of RhaD enzymes
Comparison of certain catalytic properties of RhaDs is summarized in Table 2. All RhaD enzymes were active at pH 7–9 in slightly alkaline conditions. Most RhaDs were mesophilic, except that T. acidophilum RhaD displayed its highest activity at 60 °C, favorable temperature for cell growth. It was reported that thermophilic proteins tend to contain the higher composition of charged residues (Lys, Arg, Glu, Asp) than mesophilic proteins with the difference of approximately 3 % on the composition of charged residues (Szilagyi and Zavodszky 2000). When the average composition of charged residues in RhaD was analyzed, RhaDs of Archaea, Bacteria, and Eukarya in Fig. 3 were found to be composed of 22.5, 18.0, and 19.3 % charged residues, respectively.
Interestingly, Ta_RhaD had excellent affinity for l-rhamnose, as shown in Table 2. The K m value of Ta_RhaD was 0.46 mM, which is 3.7-fold that of P. stipitis RhaD, reported as the RhaD with the lowest K m value among Bacteria and Eukarya. Likely, the affinities of T. acidophilum RhaD for l-lyxose and l-mannose were three to six times higher than those of other RhaDs. Therefore, it can be concluded that the NADP+-specific RhaD from hyperthermophilic Archaea is an attractive catalyst in bioprocessing and biotechnology due to its thermophilicity and strong affinity for l-rhamnose.
References
Ahmed H, Tjaden B, Hensel R, Siebers B (2004) Embden–Meyerhof–Parnas and Entner–Doudoroff pathways in Thermoproteus tenax: metabolic parallelism or specific adaptation? Biochem Soc Trans 32:303–304
Ahmed H, Ettema TJ, Tjaden B, Geerling AC, van der Oost J, Siebers B (2005) The semi-phosphorylative Entner–Doudoroff pathway in hyperthermophilic archaea: a re-evaluation. Biochem J 390:529–540
Beekwilder J, Marcozzi D, Vecchi S, de Vos R, Janssen P, Francke C, van Hylckama VliegJ, Hall RD (2009) Characterization of rhamnosidases from Lactobacillus plantarum and Lactobacillus acidophilus. Appl Environ Microbiol 75:3447–3454
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Buchanan CL, Connaris H, Danson MJ, Reeve CD, Hough DW (1999) An extremely thermostable aldolase from Sulfolobus solfataricus with specificity for non-phosphorylated substrates. Biochem J 343:563–570
Darland G, Brock TD, Samsonoff W, Conti SF (1970) A thermophilic, acidophilic mycoplasma isolated from a coal refuse pile. Science 170:1416–1418
de Leder Kremer RM, Gallo-Rodriguez C (2004) Naturally occurring monosaccharides: properties and synthesis. Adv Carbohydr Chem Biochem 59:9–67
Fujimoto K, Hara M, Yamada H, Sakurai M, Inaba A, Tomomura A, Katoh S (2001) Role of the conserved Ser-Tyr-Lys triad of the SDR family in sepiapterin reductase. Chem Biol Interact 130–132:825–832
Hsung JC, Haug A (1975) Intracellular pH of Thermoplasma acidophila. Biochim Biophys Acta 389:477–482
Jung JH, Lee SB (2006) Identification and characterization of Thermoplasma acidophilum glyceraldehyde dehydrogenase: a new class of NADP+-specific aldehyde dehydrogenase. Biochem J 397:131–138
Kallberg Y, Oppermann U, Jornvall H, Persson B (2002) Short-chain dehydrogenases/reductases (SDRs). Eur J Biochem 269:4409–4417
Kim S, Lee SB (2005) Identification and characterization of Sulfolobus solfataricus d-gluconate dehydratase: a key enzyme in the non-phosphorylated Entner–Doudoroff pathway. Biochem J 387:271–280
Kim S, Lee SB (2006a) Characterization of Sulfolobus solfataricus 2-keto-3-deoxy-d-gluconate kinase in the modified Entner–Doudoroff pathway. Biosci Biotechnol Biochem 70:1308–1316
Kim S, Lee SB (2006b) Rare codon clusters at 5′-end influence heterologous expression of archaeal gene in Escherichia coli. Protein Expr Purif 50:49–57
Koivistoinen OM, Hilditch S, Voutilainen SP, Boer H, Penttila M, Richard P (2008) Identification in the yeast Pichia stipitis of the first l-rhamnose-1-dehydrogenase gene. FEBS J 275:2482–2488
Lahaye M, Robic A (2007) Structure and functional properties of ulvan, a polysaccharide from green seaweeds. Biomacromolecules 8:1765–1774
Lamble HJ, Heyer NI, Bull SD, Hough DW, Danson MJ (2003) Metabolic pathway promiscuity in the archaeon Sulfolobus solfataricus revealed by studies on glucose dehydrogenase and 2-keto-3-deoxygluconate aldolase. J Biol Chem 278:34066–34072
Lamble HJ, Milburn CC, Taylor GL, Hough DW, Danson MJ (2004) Gluconate dehydratase from the promiscuous Entner–Doudoroff pathway in Sulfolobus solfataricus. FEBS Lett 576:133–136
Lamble HJ, Theodossis A, Milburn CC, Taylor GL, Bull SD, Hough DW, Danson MJ (2005) Promiscuity in the part-phosphorylative Entner–Doudoroff pathway of the archaeon Sulfolobus solfataricus. FEBS Lett 579:6865–6869
Nishiya Y, Tamura N, Tamura T (2004) Analysis of bacterial glucose dehydrogenase homologs from thermoacidophilic archaeon Thermoplasma acidophilum: finding and characterization of aldohexose dehydrogenase. Biosci Biotechnol Biochem 68:2451–2456
Noh M, Jung JH, Lee SB (2006) Purification and characterization of glycerate kinase from the thermoacidophilic archaeon Thermoplasma acidophilum: an enzyme belonging to the second glycerate kinase family. Biotechnol Bioprocess Eng 11:344–350
Oppermann U, Filling C, Hult M, Shafqat N, Wu X, Lindh M, Shafqat J, Nordling E, Kallberg Y, Persson B, Jornvall H (2003) Short-chain dehydrogenases/reductases (SDR): the 2002 update. Chem Biol Interact 143–144:247–253
Pao SS, Paulsen IT, Saier MH Jr (1998) Major facilitator superfamily. Microbiol Mol Biol Rev 62:1–34
Pletnev VZ, Weeks CM, Duax WL (2004) Rational proteomics II: electrostatic nature of cofactor preference in the short-chain oxidoreductase (SCOR) enzyme family. Proteins 57:294–301
Potter JA, Kerou M, Lamble HJ, Bull SD, Hough DW, Danson MJ, Taylor GL (2008) The structure of Sulfolobus solfataricus 2-keto-3-deoxygluconate kinase. Acta Crystallogr D Biol Crystallogr 64:1283–1287
Power J (1967) The l-rhamnose genetic system in Escherichia coli K-12. Genetics 55:557–568
Reher M, Fuhrer T, Bott M, Schonheit P (2010) The nonphosphorylative Entner-Doudoroff pathway in the thermoacidophilic euryarchaeon Picrophilus torridus involves a novel 2-keto-3-deoxygluconate-specific aldolase. J Bacteriol 192:964–974
Rigo LU, Nakano M, Veiga LA, Feingold DS (1976) l-Rhamnose dehydrogenase of Pullularia pullulans. Biochim Biophys Acta 445:286–293
Sato T, Atomi H (2011) Novel metabolic pathways in Archaea. Curr Opin Microbiol 14:307–314
Sawada H, Takagi Y (1964) The metabolism of l-rhamnose in Escherichia coli. 3 l-Rhamulose-phosphate aldolase. Biochim Biophys Acta 92:26–32
Searcy DG (1976) Thermoplasma acidophilum: intracellular pH and potassium concentration. Biochim Biophys Acta 451:278–286
Szilagyi A, Zavodszky P (2000) Structural differences between mesophilic, moderately thermophilic and extremely thermophilic protein subunits: results of a comprehensive survey. Structure 8:493–504
Takagi Y, Sawada H (1964a) The metabolism of l-rhamnose in Escherichia coli I. L-Rhamnose isomerase. Biochim Biophys Acta 92:10–17
Takagi Y, Sawada H (1964b) The metabolism of l-rhamnose in Escherichia coli II. l-Rhamnulose kinase. Biochim Biophys Acta 92:18–25
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tanaka N, Nonaka T, Nakanishi M, Deyashiki Y, Hara A, Mitsui Y (1996) Crystal structure of the ternary complex of mouse lung carbonyl reductase at 1.8 angstrom resolution: the structural origin of coenzyme specificity in the short-chain dehydrogenase/reductase family. Structure 4:33–45
Valero E, Garcia-Carmona F (1996) Optimizing enzymatic cycling assays: spectrophotometric determination of low levels of pyruvate and l-lactate. Anal Biochem 239:47–52
Watanabe S, Makino K (2009) Novel modified version of nonphosphorylated sugar metabolism—an alternative l-rhamnose pathway of Sphingomonas sp. FEBS J 276:1554–1567
Watanabe S, Saimura M, Makino K (2008) Eukaryotic and bacterial gene clusters related to an alternative pathway of nonphosphorylated l-rhamnose metabolism. J Biol Chem 283:20372–20382
Wermuth B, Sciotti MA (2001) Coenzyme specificity of human monomeric carbonyl reductase: contribution of Lys-15, Ala-37 and Arg-38. Chem-Biol Interact 130:871–878
Wilson DM, Ajl S (1955) The metabolism of l-rhamnose by Escherichia coli. Biochim Biophys Acta 17:289
Wilson DM, Ajl S (1957a) Metabolism of l-rhamnose by Escherichia coli I. l-rhamnose isomerase. J Bacteriol 73:410–414
Wilson DM, Ajl S (1957b) Metabolism of l-rhamnose by Escherichia coli. II The phosphorylation of l-rhamnulose. J Bacteriol 73:415–420
Acknowledgments
This work was supported by the Marine Biotechnology Program of Korean Ministry of Land, Transport and Maritime Affairs and the 21C Frontier Microbial Genomics and Applications Centre Program of Korean Ministry of Education, Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Albers.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, S.M., Paek, K.H. & Lee, S.B. Characterization of NADP+-specific l-rhamnose dehydrogenase from the thermoacidophilic Archaeon Thermoplasma acidophilum . Extremophiles 16, 447–454 (2012). https://doi.org/10.1007/s00792-012-0444-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-012-0444-1