Abstract
The analysis of the cold-shock domain (CSD)-encoding genes, capB and cspA, by PCR amplification showed presence of capB in all 18 Antarctic Pseudomonas isolates, but the absence of cspA. Nucleotide sequence analysis of capB ORF from a biodegradative Pseudomonas 30/3 and its regulatory sequences including the promoter and 5′-UTR was determined and compared with the other CSD-encoding genes. Expression analysis using translational gene fusion of the putative capB promoter and its flanking sequence from Pseudomonas sp. 30/3 with lacZ′ exhibited a significant increase in β-galactosidase activity at 15 and 6°C. Unlike the expression of E. coli CspA, Pseudomonas sp. 30/3 showed a slow but steady increase of the CapB expression at 6°C. Subcellular localization of CapB at 6°C showed accumulation in and around the nucleoid whereas at 22 or 30°C, it was identified around the nucleoid as well as in the cytosol. Our study attempts to elucidate the detailed structure of capB from Pseudomonas 30/3 and the role of 5′UTR in the transcriptional regulation along with the possible role of CapB in transcription and translation suited for the cold adaptation of this bacterium in Antarctic environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microorganisms inhabiting the Antarctic environment exhibit adaptive features necessary to cope with extreme cold conditions. One of the adaptive responses characterized in Antarctic as well as in some of the mesophilic bacteria is the accumulation of proteins of the cold-shock domain (CSD) family and the regulation of their corresponding genes (Bej et al. 2000; Panicker et al. 2002; Phadtare et al. 1999; Schindler et al. 1999; Wouters et al. 2000; Yamanka et al. 1998). Two classes of small bacterial proteins that consist of a single nucleic acid-binding domain (CSD) have been described: (1) Csps or cold-shock proteins that are expressed immediately after temperature downshift (Goldstein et al. 1990; Graumann and Marahiel, 1996, 1998); and (2) Caps or cold acclimation proteins that are expressed during prolonged growth at cold temperatures (Jiang et al. 1993). CSPs have been extensively described in both Gram-negative and Gram-positive bacterial species (Graumann and Marahiel 1998). A well-studied example is the cspA family of genes in Escherichia coli, which consist of nine homologs to the major cold-shock protein CspA (CS7.4) (Goldstein et al. 1990; Phadtare et al. 1999). And all nine individual members are differentially regulated in response to low temperature stress (Yamanka et al. 1998).
The CSD is an ancient and the evolutionary conserved nucleic acid-binding domain within prokaryotes and eukaryotes (Graumann and Marahiel 1998; Wolffe et al. 1992; Wolffe 1994). In eukaryotes, CSPs exist as a nucleic acid-binding domain within multidomain proteins, called CSD proteins (Somerville, 1999). Among the most widely studied eukaryotic CSD proteins is the Y-box transcription factor that contains an N-terminal domain, a CSD, and a C-terminal auxiliary domain. The functions of Y-box transcription factors have been studied in detail but none of these functions are directly related to cold-shock response or cold adaptation (Somerville, 1999).
The E. coli CspA consists of five β-barrel sheets with two consensus RNA-binding domains (RNP1 and RNP2) placed side by side on separate β-sheets (Newkirk et al. 1994; Schindelin et al. 1994). Similar structures have been observed in other bacterial CSPs and CspB from Bacillus subtilis (Schindelin et al. 1993). It has been reported that CspA functions as a RNA chaperone by minimizing the formation of secondary structures in mRNA and allowing efficient translation at low temperatures (Bae et al. 2000). The cspA promoter is constitutively expressed at 37°C though its activity is enhanced following cold shock of bacterial cultures (Tanabe et al. 1992; Fang et al. 1997). Many other CSPs have been found to have a similar function like that of CspA during the temperature downshift (Bae et al. 2000). In contrast to cspA, which has been identified both in mesophilic and psychrotrophic microorganisms, the caps have so far been identified only in cold-adapted bacteria (Fang et al. 1998; Berger et al. 1997; Gumley and Iniss 1996; Hebraud et al. 1994; Roberts and Inniss 1992). So far, only four caps have been sequenced and characterized: capA and capB in P. fragi (Michel et al. 1997), capA from Arthrobacter globiformis (Berger et al. 1996, 1997) and the capB from Pseudomonas strain 30/3 (Panicker et al. 2002). These genes share 60–70% nucleotide identity with cspA open reading frame.
The expression of cspA is regulated in a complex manner at the transcriptional (Tanabe et al. 1992; Goldenberg et al. 1997; Mitta et al. 1997), mRNA stability (Brandi et al. 1996; Goldenberg et al. 1996; Fang et al. 1997) and translational levels (Mitta et al. 1997). The feature that contributes to the cold-shock induction of cspA and three other genes, cspB, cspG and cspI, is their long, highly homologous 5′ untranslated region (5′-UTR; 159 bases for cspA, 161 bases for cspB, 160 bases for cspG and 145 bases for cspI). They all have similar AT-rich region upstream of the −35 region called the UP element, which plays an important role in their transcription at low temperature (Mitta et al. 1997; Wang et al. 1999). The long 5′-UTRs of cspA, cspB, cspG, and cspI consists of a well-conserved motif, termed the cold box, which is believed to be involved in autoregulation at the end of the acclimation phase when exposed to cold temperatures (Jiang et al.1996; Fang et al. 1998).
In the present study, 35 mesophilic and psychrotrophic bacteria, primarily members of the Enterobactericeae family and 36 Antarctic bacterial isolates were screened for the presence of proteins of the cold-shock domain (CSD) family by PCR method using degenerate primer sets for Enterobacteriaceae and primer sets based on the related genes from P. fragi. Pseudomonas 30/3, a psychrotrophic bacterium belonging to the Pseudomonas syringae cluster (Panicker et al. 2002), was isolated from petroleum hydrocarbon (PAH)-contaminated soil from Wright Valley, Antarctica. In a previous study, we detected elevated expression of an 8-kDa protein in Pseudomonas 30/3 corresponding to the size of the CapB in P. fragi following exposure to 4°C but not at 15, 22, or 37°C (Panicker et al. 2002). In this study, the promoter region of the capB gene was cloned, sequenced and its activity was evaluated at 37, 15, and 6°C to determine its role in capB transcription at cold temperatures.
Materials and methods
Bacterial strains and media requirements
All Antarctic isolates (Table 1) were grown and maintained on R2A agar medium (Becton Dickenson, Franklin Lane, NJ) at 15°C. The mesophilic bacterial strains (Table 1) were cultured on nutrient agar (Becton Dickenson). The growth temperatures for all strains are listed in Table 1. E. coli strain NM522 [F′{proAB+, lacIq, lacZ ΔM15}supE, thi1 Δ(lacpro −AB) Δhsd5(r−m−)λ−] was used for transformation and maintenance of the capB promoters segments cloned on the promoter probe vector pMLB1034.
Analysis of the upstream region of capB ORF
The TOPO-Walker™ kit (Invitrogen, Carlsbad, CA, USA) was used to amplify the upstream sequences of the capB ORF using the oligonucleotide primers GSP1, GSP2 and GSP3 (Table 2). The resulting PCR amplified fragments were cloned onto pCR4-Topo™ using the TOPO-TA™ cloning kit (Invitrogen, Carlsbad, CA). The nucleotide sequence of the cloned gene fragments was determined using M13 forward or reverse primers and an ABI Prism automated DNA sequencer (Perkin Elmer, Norwalk, CT) at the UAB sequencing core facility (http://seqcore.uab.edu/). The sequences were then compared and aligned with the nucleotide sequence database in Genbank (http://www.ncbi.nlm.nih.gov) using the Basic Local Alignment Search Tool (BLAST). The promoter, RBS and ORF sequences were analyzed using Softberry (http://softberry.com/berry.phtml) and Geneious (http://www.geneious.com) software. Multiple sequence alignments were performed by T-Coffee software (Notredame et al. 2000).
PCR amplification
Genomic DNA was purified from all the isolates tested by the method described by Ausubel et al. (1987). The oligonucleotide primers used for the detection of capB and cspA are listed in Table 2. Each PCR amplification was performed in a 25 μl reaction volume consisting of 1 μg of purified genomic DNA; 200 μM of each of the dNTPs; 1 μM of each of the oligonucleotide primers and 2.0 U AmpliTaq (Perkin Elmer, Walham, MA) DNA polymerase; and 1× PCR reaction buffer [10× buffer consisted of 300 mM Tris.Cl (pH 9.0), 75 mM (NH4)2SO4 and 2 mM MgCl2]. All PCR amplifications were performed in a GeneAmp PCR 2400 thermocycler (Perkin Elmer, Norwalk, CT) with 25 cycles of amplification steps each at 94°C for 1 min, 50°C for 1 min and 72°C for 2 min. For those isolates that were PCR negative, an annealing temperature of 45°C was also attempted.
Similarly, PCR was also carried out to amplify different fragments of the promoter region of capB using oligonucleotide primers with appropriate restriction endonuclease recognition sites flanked at the 5′ end to facilitate cloning onto the promoter probe vector pMLB1034 (Table 2). The PCR cycling and reaction conditions were same as above with the annealing temperature being 50°C.
Analysis of the promoter segments of capB on pMLB1034
Segments of the 5′-upstream DNA were PCR amplified using capB-specific L-capBproBamHI and R-capB13BamHI; and L-capB515BamHI and R-capB13BamH1 primer sets generating 136 and 515 bp amplicons, respectively. The amplified DNA fragments were then cloned on a promoter probe vector pMLB1034 to establish two translational fusion plasmids, pBGP136 and pBGP515. E. coli NM522 was transformed with these plasmid constructs and 3 white colonies with putative clones were randomly selected on LB agar plates supplemented with ampicillin (50μg/ml), 3-bromo-4-chloro-β-d-thiogalactosidase (X-gal) and isopropyl-beta-d-thiogalactopyranoside (IPTG). The selected white colonies were inoculated in LB broth and plasmid DNA extracted using the Qiagen mini-prep plasmid purification columns (Qiagen, Valencia, CA). The purified DNA from each putative clone was treated with the respective restriction endonuclease (New England Biolab, Beverly, MA) and separated in a 1% (w/v) agarose gel at 5 V/cm to determine the molecular sizes of the cloned DNA fragments.
β-galactosidase assay
E. coli NM522 harboring pBGP136 or pBGP515 plasmids were grown in 50 ml of LB supplemented with 1 mM IPTG at 37°C until the optical density at 600 nm reached 0.5. Aliquots (15 ml each) of the culture were transferred to 15 and 6°C. β-galactosidase assay (Miller 1972) was performed using 0.5 ml culture at 0 h (time before transfer to different temperatures), 1, 3, and 6 h after temperature downshift. The assay was done in triplicate to ensure the consistency of the results.
Western blot
An aliquot of the overnight culture of Pseudomonas sp. 30/3 grown at 19 ± 2°C (room temperature) was diluted 1:20 with 1:10 Trypticase Soy Broth (TSB) (Becton Dickenson) and grown at 19 ± 2°C (room temperature) on a rotary shaker set at 150 rpm until the optical density at 450 nm reached 0.2. The culture was aliquoted (50 ml) and transferred to 6, 15, or 30°C. At 0, 1, 3, and 6 h of incubation, an aliquot of the culture was removed and the bacteria were harvested by centrifugation (10,000×g for 10 min). The cells were resuspended in PBS (pH 7.2) and the cell membrane disrupted by sonication on ice for 3–5 s. The lysate was resuspended in 50 mM Tris–HCl buffer (pH 6.8) consisting of 2% (w/v) SDS, 10% (v/v) glycerol, 6% (v/v) 2-mercaptoethanol and 0.002% (v/v) bromophenol blue to 0.25 μg total protein/μl, and boiled for 5 min. The concentration of total protein was determined colorimetrically using the BCA Protein Assay Kit (Pierce, Rockford, IL) with bovine serum albumin (Sigma–Aldrich, St. Louis, MO) as a standard. These samples (2.5 μg total protein) were electrophoresed on a 15% (w/v) polyacrylamide gel and electrophoretically transferred onto polyvinyliden difluoride membranes (Millipore, Bedford, MA). For anti-CapB antiserum, first capB from Pseudomonas sp. 30/3 was PCR amplified and cloned into pET24 (b) plasmid (Novagen, EMD Chemicals, Madison, WI). The CapB protein was expressed by IPTG induction and then purified using the His.Tag kit (Novagen, EMD Chemicals). The antiserum for the CapB protein was developed in NZW rabbit with 3 boosts (14, 21 and 49 days) and 2 test-bleeds (35 and 56 days) (Cocalico Biologicals, Inc., Reamstoen, PA). The effective dilution of the anti-CapB rabbit-antiserum for western blot analysis was determined by reaction with the purified CapB protein and serial dilution of the antibody followed by a standard curve analysis (Ausubel et al. 1987). The membrane with immobilized proteins from Pseudomonas sp. 30/3 was incubated with rabbit anti-CapB antiserum (1:1000 dilutions) for 2 h at 22°C. Bound antibodies were detected with goat anti-rabbit peroxidase conjugated IgG (1:5000 dilution, Pierce), and the peroxidase activity was visualized using 0.02% (v/v) of 3, 3′-diaminobenzidine tetrahydrochloride (Pierce) in 0.1 M PBS (pH 6.2) containing 0.06% H2O2 (Thermo Fisher Scientific). The membrane was scanned and the relative amount of CapB expressed was analyzed with Scion Image software (http://www.scopncorp.com). The consistency of the results was determined from four individual assays. Similarly, western blot assays were performed for CapB expression in other Antarctic Pseudomonas sp. at 15°C (Table 1).
In situ localization of CapB by immunofluorescence staining
Immunofluorescence staining was performed by a modified method described by Harry et al. (1995). Pseudomonas 30/3 culture was grown in 1:10 v/v TSB media till exponential phase at 22°C and then aliquots of 50 ml each were subjected to 6 (cold shock), 22, and 30°C, respectively for 1 h.
Fixation and permeabilization of cells
The cells were vortexed to disrupt any clumps of bacteria. A 0.25-ml volume of bacterial culture was fixed with fixative solution at a final concentration of 2.4% (v/v) paraformaldehyde, 0.04% (v/v) glutaraldehyde, in 30 mM Na-PO4 buffer (pH 7.5) for 10 min at room temperature and 50 min on ice. The fixed bacteria were washed thrice in PBS (pH 7.4) at room temperature and then resuspended in 90–125 μl of GTE (50 mM glucose, 20 mM Tris–HCl [pH 7.5], 10 mM EDTA). A freshly prepared lysozyme solution in GTE was added to a final concentration of 2 mg/ml. 10 μl of the fixed cells were dropped on a microscopic slide which was treated with 0.1% (wt/v) poly-l-lysine (Sigma, St. Louis, MO). After 30 s, the liquid was aspirated from the slides, and allowed to dry completely. The slides were dipped in −20°C methanol for 5 min and then in −20°C acetone for 30 s and again allowed to dry. A 10-μl volume of blocking solution (2% bovine serum albumin [BSA] in PBS, pH 7.4 [BSA-PBS]) was then added to the fixed cells, and the slides were then incubated for 15 min at room temperature.
Immunofluorescence staining of cells
The samples on the slides were incubated with rabbit anti-CapB antiserum (1:1500 dilutions in BSA-PBS) for 1 h at 22°C and were washed 10 times with PBS. The slides were then incubated with 7.5 mg/ml solution of the HiLyte Fluor 488-labeled goat anti-rabbit IgG secondary antibody (AnaSpec, San Jose, CA) in BSA–PBS for 1 h at 22°C in the dark. DAPI (Sigma, St. Louis, MO) was added with the secondary antibody at a final concentration of 0.01 mg/ml. The slides were washed 10 times with PBS and then once with mounting medium. Slides were mounted in PBS-glycerol solution and observed under a Leica fluorescence microscope (Bannockburn, IL) or stored at −20°C for several days before observation.
Results
Occurrence of the cspA and the capB genes that code for the CSD proteins
The results of the PCR amplification of cspA and capB in various microbial isolates are presented in Table 1. All 18 Antarctic Pseudomonas soil isolates exhibited 248 bp capB amplicon. Only 1 non-Antarctic strain, P. putida ATCC 17484 exhibited positive amplification. The capB in Pseudomonas sp. 30/3 was used as the positive control in this study (Panicker et al. 2002). These 18 Antarctic Pseudomonas isolates exhibited positive PCR amplification with the capB primers (L-capB515BamH1 or LcapBproBamH1 in combination with RcapB13BamH1), but were negative for cspA when amplified with the cspA gene-specific CSPU5 and CSPU3 universal primers. All Enterobactericeae and a few other mesophilic isolates tested in this study showed positive amplification of a 200-bp cspA ORF with the cspA gene-specific universal primers. An Antarctic marine psychrophile, S. benthica ATCC 43992 was also positive for cspA, but was negative for capB. P. putida ATCC 17484 was the only mesophilic isolate that exhibited positive amplification of capB in this study. All other mesophilic Pseudomonas spp. were negative for both cspA and capB.
Structure of CapB and sequence comparison among other CSD proteins
CapB of Pseudomonas 30/3 is highly identical to CapB of P. fragi (98.6%). When the amino acid sequence of CapB was compared to other known cold-inducible CSPs in different bacteria (i.e., CspA, CspB, CspG and CspI from E.coli and CspB, CspC and CspD from B.subtilis), the amino acid identity/similarity was between 51.4 and 60.9%/60–71% (Fig. 1b). The three-dimensional structure of CspA has already been determined. It consists of five antiparallel β-strands forming a β-barrel structure with two β-sheets (Feng et al. 1998; Newkirk et al. 1994; Schindelin et al. 1994). CapB consists of well-conserved hydrophobic residues V9, I21, V30 (L29 in CapB), V32 (V31 in CapB), and V51 (Fig. 1b) which form a hydrophobic core in CspA. In addition, CapB also consists of two well-conserved RNA binding motifs, RNP1 and RNP2 (Fig. 1b). These facts suggest that like CspA, the CapB may form a similar structure to that of CspA and may also bind to RNA and single-stranded DNA.
a Nucleotide sequence of the 540-bp upstream region of the capB gene along with the 210-bp capB ORF from Pseudomonas sp.30/3. The putative promoter elements –35 and –10 regions as well as the ribosome binding site (RBS), upstream sequence and downstream box (DB) sequences are underlined and labeled. GenBank accession number is AF363392. The putative transcription start site is in bold letters and is marked by an arrow. The translation start codon ATGs are also in bold letters and are underlined. The putative 11-bp cold-box element is labeled and boxed. b Amino acid sequence alignments of CapB-Ps30/3 (database accession. no. AAK35071), CapB-P.fragi (AAC45997), CspA-E.coli (AAN82813), CspB B.subtilis (P32081), CspB-E.coli (AAN81636), CspC-B.subtilis (AAC45646), CspD B.subtilis (AAA96623), CspG-E.coli (AAN79591), CspI-E.coli (AAN81629). Alignments were performed by T-Coffee (Notredame et al. 2000). Their amino acid sequence identities/similarities are shown on the right, with CapB-Ps30/3 set at 100%. The residues forming the hydrophobic core in the β-barrel structure are indicated by asterisks above the sequences. The RNA binding motifs, RNP1 and RNP2, are boxed
Transcriptional regulation of capB expression
Upstream nucleotide sequences of the capB ORF
The BLAST (http://www.ncbi.nlm.nih.gov) nucleotide sequence comparison analysis of the 540-bp region upstream of the ATG of capB of Pseudomonas 30/3 exhibited 93% nucleotide identity with the upstream sequence of the capB gene in P. fragi. The nucleotide sequence information of Pseudomonas 30/3 is elaborated in Fig. 1a and the GenBank accession number is AF363392 (gi:13625472). The putative promoter sequences −35 (5′-TTGGCA-3′), −10 (5′-GGTTAAGGT-3′) were identified. The putative transcription initiation site is 3-bp downstream from the −10 promoter region (Fig. 1a). A typical RBS (5′-AGGA7–9 ATG) and an ORF of 210 bp between ATG and TAA were also identified (Fig. 1a). Thirteen base-pairs downstream from the transcription initiation nucleotide and within the 149 bp long 5′-UTR is a sequence with high identity to eubacterial cold-box elements (Figs. 1a, 2a). In Pseudomonas 30/3 capB, 7 out of 11 nucleotides are identical to the cold-box sequences of E.coli cspA, cspB and cspG (Fig. 2a).The level of identity exceeds that for the cold-box sequence from the E. coli cspI gene, which has only three to five nucleotides in common with other cold-box elements. And the level of identity is less than that for the cold-box sequence from Anabaena crhC and M. burtonii deaD (RNA helicases) which have seven to nine nucleotides in common with other cold-box elements (Fig. 2a). As shown in Fig. 2b, capB also contains downstream box (DB) downstream of translation initiation codon which has 10 out of 14 identical nucleotides to DB of E. coli cspA, the level of identity of DB sequence from Pseudomonas 30/3 capB exceeds that of DB from Anabaena crhC (Fig. 2b). Farther downstream in the 5′-UTR of Pseudomonas 30/3 capB, there is a putative 12-bases upstream sequence, which may be similar to upstream sequence from other cold-shock genes. Interestingly, Pseudomonas 30/3 capB does not have an AT-rich UP element upstream of the −35 promoter sequence, which plays an important role in the transcription at low temperature in E.coli cspA, cspB, cspG and cspI (Mitta et al. 1997; Wang et al. 1999).
a Comparison of cold-box elements from coldshock genes of E. coli, Anabaena and deaD gene from M. burtonii with the putative cold-box element from Pseudomonas 30/3 capB gene. Alignments were performed by T-Coffee (Notredame et al. 2000). 11-bp cold-box element is underlined. b Comparison of the downstream box (DB) sequence from coldshock genes of E. coli and Anabaena with the 14-bp putative DB from Pseudomonas 30/3 capB gene. Alignments were performed by T-Coffee (Notredame et al. 2000)
Cold shock-inducible expression of capB
The pBGP136 construct consisting of a 127-bp DNA fragment excluding the cold box and promoter but including the capB RBS and the codons for the first 13 amino acid residues exhibited nearly insignificant (<30 Miller units) β–galactosidase activity at all temperatures (Fig. 3a, b). However, within this narrow range of β-galactosidase activity, cultures exposed to 6°C showed maximum expression with an increasing trend throughout the 6-h incubation period. The cultures exposed to 15°C showed a slight decline in the activity, whereas cultures at 37°C showed the least activity (<5 Miller unit) and lower than the activity measured at initial time (15 Miller unit).
a Construction of plasmid pBGP136. A translational fusion was constructed with the PCR amplified capB promoter consisting of the −35, −10, and the ribosome binding sequence and the sequences for the first 13 amino acid residues of the capB ORF on pMLB1034. b The results of the β-galactosidase assay exhibited by the pBGP136 construct. See text for the detail description of the results; c Construction of plasmid pBGP515. A translational fusion was constructed with the PCR amplified capB promoter consisting of the 476-bp upstream of the −35 and stretching up to the sequence for the first 13 amino acid residues of the capB ORF on pMLB1034; d The results of the β-galactosidase assay exhibited by the pBGP515 construct. See text for the detail description of the results
The pBGP515 construct consisting of a 517-bp DNA fragment included in pGBP136 along with an additional 390 bp further 5′ DNA sequence upstream including entire 149 bp 5′UTR with cold box and capB promoter, showed significant increase (>1000 fold) in β–galactosidase activity at cold temperatures (Fig. 3c, d). Cultures exposed to 15°C for 1 h showed an increased activity of 3000 Miller units, eventually reaching a maximum activity of >4000 Miller units at 3 h and then declined to approximately 3000 Miller units at 6 h. In contrast, cultures at 6°C exhibited a steady increase in the expression of β-galactosidase ranging from 1000 Miller units after 1 h of incubation to >3000 Miller units at 6 h. However, the cultures exposed to 37°C during the entire 6-h time period exhibited a decline in activity after the initial 1 h of incubation.
CapB expression
Western blot results exhibited elevated expression of CapB in Pseudomonas 30/3 cultures when exposed to 15 and 6°C, whereas the cultures exposed at 30°C exhibited progressively decreased expression from the initial time of incubation (Fig. 4). The expression of CapB continued to increase at a steady level at 6°C, whereas the level decreased slightly for cultures exposed at 15°C. Moreover, all Antarctic Pseudomonas sp. which exhibited positive amplification of capB also showed expression of CapB by western blot (Table 1).
Western blot analysis of the expression of CapB protein in Pseudomonas 30/3 following exposure to various temperatures. Lanes 1–3 Cultures exposed to 6°C for 1, 3, and 6 h following treatment, respectively; Lanes 4–6 cultures exposed to 15°C for 1, 3, and 6 h following treatment, respectively; Lanes 7–9 cultures exposed to 30°C for 1, 3, and 6 h following treatment, respectively; SS Size Standard
In situ immunolocalization of CapB in Pseudomonas 30/3
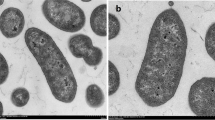
In order to understand the possible cellular role of CapB in Pseudomonas 30/3, immunofluorescence staining was used to localize this protein using the anti-CapB rabbit-antiserum at different temperatures. The cellular location of the nucleoid was confirmed by DAPI staining (Fig. 5a). At 6°C, a dense accumulation of the anti-CapB antibody immunoconjugated with the green Hilyte Fluor 488-labeled goat anti-rabbit IgG secondary antibody was observed around the nucleoid region (Fig. 5b). At 22 or 30°C, the green fluorescence was dispersed in the cytosol as well as in the nucleoid region (Fig. 5c, d, e, f). In addition, cultures exposed at 6°C exhibited compact nucleoid as compared to the cultures exposed to 22 or 30°C.
Immunolocalization of CapB in Pseudomonas 30/3 at various temperatures; a, c, e The DAPI stained (blue fluorescence) nuclei at 6, 22, and 30°C, respectively; b, d, f The CapB bound HiLyte Fluor 488 (green fluorescence) at 6, 22, and 30°C, respectively. CapB can be seen localized around the compact nucleoid region at 6°C (b) and CapB localized both around nucleoid and cytosol at 22 and 30°C (d, f)
Discussion
Psychrotrophic microorganisms demonstrate growth at a wide range of temperatures as high as 25°C or above and remain metabolically active at or below 0°C (Pikuta and Hoover 2007). Cold inducible proteins collectively known as cold-adaptive proteins, and the well-studied cold-shock proteins have been implicated in microbial adaptation to cold temperatures (Hebraud and Potier 1999). The cold adaptive gene, capB is present in cold-tolerant Pseudomonas spp. such as P. fragi K1, P. syringae, P. fluorescens, P. tolaasii, P. solanacearum (Hebraud et al. 1993). Similarly, we have shown in this study that capB is also present in all of the Antarctic biodegradative Pseudomonas spp. tested (Table 1).
We have investigated the role of the 5′-UTR and the cold-box sequences in the regulation of the Pseudomonas 30/3 capB gene. High level of β-galactosidase activity by the pBGP515 construct at cold temperatures suggested that the 515-bp untranslated DNA segment including the entire 149-bp 5′UTR with cold box and capB promoter along with the 13 amino acid residues including the downstream box sequence of the CapB in Pseudomonas 30/3 are required for the regulation of the capB and sustained expression of the CapB at cold temperatures. Unlike the expression of the CspA family of proteins that tends to exhibit transient expression immediately following downshift of the temperature (Etchegaray et al.1996), the CapB exhibited a steady increase following exposure to cold temperatures. This is similar to the previously described characteristic expression of the Caps during prolonged exposure to cold temperatures (Thieringer et al. 1998; Whyte and Inniss 1992). Therefore, it is possible that the unique and different nucleotide sequences on the 5′ upstream region of the capB ORF in Pseudomonas 30/3 and the absence of AT-rich UP element may be contributing to the regulation of this gene leading to the sustained expression of the CapB at cold temperatures. The sustained expression of CapB may be necessary for the adaptation of microorganisms in Antarctic perennially cold temperature environment. It has been shown that there is striking high structural similarity in four csp genes from E. coli and cold induced genes encoding DEAD-box RNA helicases from E. coli, Anabaena and M. burtonii (Lim et al. 2000). And all of the 5′-UTRs are greater than 100 bp in length including Pseudomonas 30/3 capB gene. E. coli cspA, cspB, and cspG, all have a downstream box (DB) located downstream of the translation initiation codon, which has been shown to play an important role in cold-shock induction at the level of translation (Mitta et al. 1997). Pseudomonas 30/3 capB gene also have similar DB sequence (Fig. 2b) and a construct pGP476 consisting of entire 149 bp 5′UTR with cold box and capB promoter but without codons for the first 13 amino acid residues, i.e., without the DB sequence, did not show any β-galactosidase activity at 37 or 15°C and showed a low level of expression after the culture was exposed to 6°C for 3 h (data not shown). In this respect, it is similar to the cspA promoter that also requires the first 13 amino acid residues containing the DB sequence to enhance transcription of this gene and translation of the cspA mRNA at cold temperatures (Mitta et al. 1997). The cold-box sequences are well conserved, and except E. coli cspI sequence, at least 6 of the 11 nucleotides are in common between any one cold-box sequences (Fig. 2a). It has also been observed that the optimal temperature ranges for induction of the four E. coli cold-shock induced csp genes (cspA, B, G and I) are not same. Upon cold shock, CspA can be induced for broader range of temperature than that of CspB and CspG, which are restricted to lower and narrower temperature range (Etchegaray et al.1996). It has been suggested by the authors that specific sequence differences in the 5′-UTR and cold-box elements resulting in different mRNA secondary structures may play important roles in regulation. But authors from the same laboratory concluded that deleting the cold-box had little effect on cold-shock induction of β-galactosidase activity, and that instead a region 11 bp upstream of the ribosome binding site was important for translational efficiency of gene expression (Yamanka et al. 1999). Although all these reports indicate involvement of 5′-UTR in the regulation, the exact genetic structures and precise mechanisms for the function of the CSD-encoding genes at cold temperatures are rather complex and yet to be defined.
Also, we have investigated the possible in situ function of CapB by intracellular localization study. The amino acid sequence alignment among the Pseudomonas 30/3 CapB and other CSPs from different bacteria exhibited a relatively high sequence similarity in the conserved RNA binding motifs (RNP1 and RNP2) suggesting that Pseudomonas 30/3 CapB may have the same cellular function (Fig. 1b). In spite of the conserved RNP1 and RNP2 motifs, our study reveals that there are differences in the subcellular localization of Pseudomonas 30/3 CapB and CspA in E. coli. CspA in E. coli occupies a polar position away from the nucleoid at 37°C and maintains its position when subjected to cold shock (Giangrossi et al. 2001), whereas the CapB was localized in and around the nucleoid during cold shock. Pseudomonas 30/3 cells exhibited a more compact nucleoid at 6°C, which was with the zone of localization of CapB in the cytosolic spaces surrounding the nucleoid region. It has been reported that during cold shock, there is a decrease in the transcriptional and translational capacity of the cells leading to chromosome compaction in B. subtilis (Weber et al. 2001). Also, previous studies in both E. coli and B. subtilis have shown that the ribosomal proteins localize in a manner similar to CSPs, while RNA polymerase subunits and the primary sigma factor localize mainly in nucleoids (Azam et al. 2000; Lewis et al. 2000). During cold shock, CapB was localized mostly in the nucleoid region, which suggests similar localization as RNA polymerase and it implies a possible role for CapB in transcription. Whereas, at higher temperatures (22 and 30°C), CapB localizes as found for the ribosomal proteins suggesting that it functions at the same cellular location as ribosomes during translation. Therefore, the results from this study suggest that CapB has a possible role in both transcription and translation in Pseudomonas 30/3.
We have shown that the capB gene is conserved in Antarctic biodegradative Pseudomonas sp. Although the capB and CSD-encoding genes share common genetic features, the unique regulatory segment of a biodegradative Pseudomonas 30/3 capB gene could be responsible for the sustained expression of CapB protein. Moreover, the in situ localization of the CapB indicated that this protein has both transcriptional and translational regulatory role in this bacterium. The continuous expression of the CapB protein and its regulatory role in the transcription and translation of the essential genes may be necessary for this bacterium and possibly other Antarctic Pseudomonas sp. tested in this study for survival in Antarctic perennially cold temperature environment.
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Smith JG, Sideman JG, Struhl K (eds) (1987) Current protocols in molecular biology. John Wiley & Sons, Inc., New York, pp 2.10–2.11
Azam TA, Hiraga S, Ishihama A (2000) Two types of localization of the DNA-binding proteins within the Escherichia coli nucleoid. Genes Cells 5:613–626
Bae W, Xia B, Inouye M, Severinov K (2000) Escherichia coli CspA-family RNA chaperones are transcription antiterminators. Proc Natl Acad Sci USA 97:7784–7789
Bej AK, Saul D, Aislabie J (2000) Cold tolerant alkane-degrading Rhodococcus species from Antartica. Polar Biol 23:100–105
Berger F, Morellet N, Menu F, Potier P (1996) Cold shock and cold acclimation proteins in the psychrotrophic bacterium Arthrobacter globiformis S155. J Bacteriol 178:2999–3007
Berger F, Normand P, Potier P (1997) capA, a cspA-like gene that encodes a cold acclimation protein in the psychrotrophic bacterium Arthrobacter globiformis SI55. J Bacteriol 179:5670–5676
Brandi A, Pietroni P, Gulazeri CO, Pon CL (1996) Post-transcriptional regulation of CspA expression in Escherichia coli. Mol Microbiol 19:231–240
Etchegaray JP, Jones PG, Inouye M (1996) Differential thermoregulation of two highly homologous cold-shock genes, cspA and cspB, of Escherichia coli. Genes Cells 1:171–178
Fang L, Jiang W, Bae W, Inouye M (1997) Promoter-independent cold-shock induction of cspA and its derepression at 37°C by mRNA stabilization. Mol Microbiol 23:355–364
Fang L, Hou Y, Inouye M (1998) Role of the cold-box region in the 5′ untranslated region of the cspA mRNA in its transient expression at low temperature in Escherichia coli. J Bacteriol 180:90–95
Feng W, Tejero R, Zimmerman DE, Inouye M, Montelione GT (1998) Solution NMR structure and backbone dynamics of the major cold shock protein (CspA) from Escherichia coli: evidence for conformational dynamics in the single-stranded RNA-binding site. Biochemistry 37:10881–10896
Francis KP, Stewart GS (1997) Detection and speciation of bacteria through PCR using universal major cold-shock protein primer oligomers. J Indust Microbiol Biotechnol 19:286–293
Giangrossi M, Exley RM, Le Hegarat F, Pon CL (2001) Different in vivo localization of the Escherichia coli proteins CspD and CspA. FEMS Microbiol Lett 202(2):171–176
Goldenberg D, Azar I, Oppenheim AB (1996) Differential mRNA stability of the cspA gene in the cold-shock response of Escherichia coli. Mol Microbiol 19:241–248
Goldenberg D, Azar I, Oppenheim AB, Brandi A, Pon CL, Gualerzi CO (1997) Role of Escherichia coli cspA promoter sequences and adaptation of translational apparatus in the cold shock response. Mol Gen Genet 256:282–290
Goldstein J, Pollit NS, Inouye M (1990) Major cold shock protein of Escherichia coli. Proc Natl Acad Sci USA 87:283–287
Graumann P, Marahiel MA (1996) Some like it cold: response of microorganisms to cold shock. Arch Microbiol 166:293–300
Graumann PL, Marahiel MA (1998) A superfamily of proteins that contain the cold-shock domain. Trends Biochem Sci 23:286–290
Gumley AW, Iniss WE (1996) Cold shock proteins and cold acclimation proteins in the psychrotrophic bacterium Pseudomonas putida Q5 and its transconjugants. Can J Microbiol 42:798–803
Harry EJ, Pogliano K, Losick R (1995) Use of immunofluorescence to visualize cell-specific gene expression during sporulation in Bacillus subtilis. J Bacteriol 177(12):3386–3389
Hebraud M, Potier P (1999) Cold shock response and low temperature adaptation in psychrotrophic bacteria. J Mol Microbiol Biotechnol 1:211–219
Hebraud M, Garry P, Labadie J (1993) Ubiquity of low molecular mass cold-shock proteins. Abstracts of the fourth international symposium on pseudomonas, p 59
Hebraud M, Dubois E, Potier P, Labadie J (1994) Effect of growth temperatures on the protein levels in a psychrotrophic bacterium, Pseudomonas fragi. J Bacteriol 176:4017–4024
Jiang W, Jones P, Inouye M (1993) Chloramphenicol induced the transcription of the major cold-shock gene of Escherichia coli, cspA. J Bacteriol 175:5824–5828
Jiang W, Fang L, Inouye M (1996) The role of the 5′-end untranslated region of the mRNA for CspA, the major cold-shock protein of Escherichia coli, in cold-shock adaptation. J Bacteriol 178(16):4919–4925
Lewis PJ, Thaker SD, Errington J (2000) Compartmentalization of transcription and translation in Bacillus subtilis. EMBO J 19:710–718
Lim J, Thomas T, Cavicchioli R (2000) Low temperature regulated DEAD-box RNA helicase from the Antarctic archaeon, Methanococcoides burtonii. J Mol Biol 297:553–567
Michel V, Lehoux I, Depret G, Anglade P, Labadie J, Hebraud M (1997) The cold shock response of the psychrotrophic bacterium Pseudomonas fragi involves four low-molecular-mass nucleic acid-binding proteins. J Bacteriol 179:7331–7342
Miller JH (1972) Experiments in molecular genetics. Cold Spring Laboratory, Cold Spring Harbor, NY, pp 352–355
Mitta M, Fang L, Inouye M (1997) Deletion analysis of cspA of Escherichia coli: requirement of the AT-rich UP element for cspA transcription and the downstream box in the coding region for its cold shock induction. Mol Microbiol 26:321–335
Newkirk K, Feng W, Jiang W, Tejero R, Emerson SD, Inouye M, Montelione GT (1994) Solution NMR structure of the major cold shock protein (CspA) from Escherichia coli: identification of a binding epitope for DNA. Proc Natl Acad Sci USA 91:5114–5118
Notredame C, Higgins DG, Heringa J (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Panicker G, Aislabie J, Saul D, Bej A (2002) Cold tolerance of Pseudomonas sp. 30/3 isolated from oil-contaminated soil, Antarctica. Pol Biol 225:5–11
Phadtare S, Alsina J, Inouye M (1999) Cold-shock response and cold-shock proteins. Curr Opin Microbiol 2(2):175–180
Pikuta EV, Hoover RB (2007) Microbial extremophiles at the limits of life. Crit Rev Microbiol 33:183–209
Roberts ME, Inniss WE (1992) The synthesis of cold shock proteins and cold acclimation proteins in the psychrophilic bacterium Aquaspirillum articum. Curr Microbiol 25:275–278
Schindelin H, Marahiel MA, Heinemann U (1993) Universal nucleic acid-binding domain revealed by crystal structure of the B. subtilis major cold-shock protein. Nature 364:164–168
Schindelin H, Jiang W, Inouye M, Heinemann U (1994) Crystal structure of CspA, the major cold shock protein of Escherichia coli. Proc Natl Acad Sci USA 91:5119–5123
Schindler T, Graumann PL, Perl D, Ma S, Schmid FX, Marahiel MA (1999) The family of cold shock proteins of Bacillus subtilis. Stability and dynamics in vitro and in vivo. J Biol Chem 274:3407–3413
Somerville J (1999) Activities of cold-shock domain proteins in translation control. Bioessays 21:319–325
Tanabe H, Goldstein J, Yang M, Inouye M (1992) Identification of the promoter region of the Escherichia coli major cold shock gene, cspA. J Bacteriol 174:3867–3873
Thieringer HA, Jones PG, Inouye M (1998) Cold shock adaptation. BioEssays 20:49–57
Wang N, Yamanka K, Inouye M (1999) CspI, the ninth member of the CspA family of Escherichia coli, is induced upon cold shock. J Bacteriol 181:1603–1609
Weber MHW, Volkov AV, Fricke I, Marahiel MA, Graumann PL (2001) Localization of cold shock proteins to cytosolic spaces surrounding nucleoids in Bacillus subtilis depends on active transcription. J Bacteriol 183(21):6435–6443
Whyte LG, Inniss WE (1992) Cold shock proteins and cold acclimation proteins in a psychrotrophic bacterium. Can J Microbiol 38:1281–1285
Wolffe AP (1994) Structural and functional properties of the evolutionarily ancient Y-box family of nucleic acid binding proteins. Bioessays 16:245–251
Wolffe AP, Tafuri S, Ranjan M, Familari M (1992) The Y-box factors: a family of nucleic acid binding proteins conserved from Escherichia coli to man. New Biol 4:290–298
Wouters JA, Rombouts FM, Kuipers OP, de Vos WM, Abee T (2000) The role of cold-shock proteins in low-temperature adaptation of food-related bacteria. Syst Appl Microbiol 23:165–173
Yamanka K, Fang L, Inouye M (1998) The CspA family in Escherichia coli: multiple gene duplication for stress adaptation. Mol Microbiol 27:247–255
Yamanka K, Mitta M, Inouye M (1999) Mutation analysis of the 5′ untranslated region of the cold shock cspA mRNA of Escherichia coli. J Bacteriol 181:6284–6291
Acknowledgments
This study was supported in part the UAB Faculty Development award and the Department of Biology, and the Foundation of Research, Science and Technology, New Zealand (C09X0018). The logistic support was provided by Antarctica New Zealand; 2008 Tawani International Scientific Expedition (Tawani Foundation, Chicago, IL); and Antarctic Maitri (NCAOR, India) and Novolazarevskaya (Russia) stations. We thank Col (Ret) James Pritzker for supporting the Tawani Expedition; Rasik Ravindra (Director, NCAOR, India), Cdr. Arun Chaturvedi, Cdr. Pradip Malhotra and Ashit Swain for field support. We thank Edward Phillips at UAB High Resolution Imaging Shared Facility for the fluorescent microscopy study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Oren.
G. Panicker and N. Mojib contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Panicker, G., Mojib, N., Nakatsuji, T. et al. Occurrence and distribution of capB in Antarctic microorganisms and study of its structure and regulation in the Antarctic biodegradative Pseudomonas sp. 30/3. Extremophiles 14, 171–183 (2010). https://doi.org/10.1007/s00792-009-0296-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-009-0296-5