Abstract
Objective
Head and neck squamous cell carcinoma (HNSCC) is a prevalent cancer that originates from the squamous cells. The role of the replication factor C subunit 3 (RFC3) in HNSCC progression remains elusive. The aim of this study was to uncover RFC3 significance in HNSCC.
Methods
The Cancer Genome Atlas (TCGA-HNSCC) dataset was initially used to assess RFC3 expression and its association with HNSCC clinical features. Subsequently, quantitative reverse transcription PCR (RT-qPCR) confirmed RFC3 mRNA expression in oral squamous cell carcinoma (OSCC), a primary HNSCC type. Survival rates were evaluated using the Kaplan–Meier plot, while the Tumor Immune Estimation Resource (TIMER) database probed RFC3-immune cell interaction. Additionally, in silico tools were used to examine the RFC3 protein network and engagement in HNSCC pathways.
Results
RFC3 expression is significantly upregulated in HNSCC, including OSCC. Upregulated RFC3 expression was significantly correlated with the clinicopathological features of HNSCC, including tumor stage, grade, metastasis, and patient survival. RFC3 is also associated with immune cell infiltration. Functional analysis has highlighted its involvement in DNA replication, mismatch repair, and cell cycle pathways. Interestingly, RFC3 high expression is linked to well-known oncogenic signaling pathways, such as MYC/MYCN, HIPPO, and mTOR.
Conclusions
In conclusion, RFC3 can be considered a novel prognostic biomarker for HNSCC, and further studies on its functional mechanisms may help to use RFC3 as a therapeutic target for HNSCC.
Clinical relevance
The clinical relevance of this study lies in identifying RFC3 as a novel biomarker and prognostic indicator for HNSCC, offering insights that could impact future clinical approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Head and neck squamous cell carcinoma (HNSCC) ranks among the most prevalent cancers worldwide. Notably, oral squamous cell carcinomas (OSCCs) make up the majority (around 80–90%) of malignancies within the oral cavity, a significant subtype of HNSCC. The prevalence of oral cavity cancers, ranging from the 6th to the 9th most common anatomical site for cancer, varies by country and gender [1]. The primary causative factors for HNSCC encompass smoking, alcohol consumption, and ultraviolet radiation exposure. Additionally, several other factors, including human papillomavirus (HPV) and Candida infections, nutritional deficiencies, and genetic predispositions, have been associated with HNSCC [2]. Clinically, HNSCC typically presents with a central necrotic area surrounded by raised margins. This condition predominantly affects adults and the elderly [3]. Despite advances in surgical resection, radiotherapy, and chemotherapy, the high recurrence rate (20–40%) in HNSCC leads to lower survival rates [4,5,6]. Hence, a comprehensive understanding of the molecular pathways driving HNSCC proliferation and metastasis is imperative for developing novel therapeutic strategies to curb HNSCC progression. Furthermore, maintaining the accuracy of DNA replication is widely recognized as a crucial aspect of cancer progression and the cell cycle. When DNA damage repair mechanisms and cell cycle checkpoints malfunction, they become significant contributors to genomic instability [7].
One such molecular player is replication factor C (RFC), a heteropentameric protein complex pivotal in DNA replication, DNA damage repair, and cell cycle checkpoint control throughout cell cycle progression [8,9,10]. RFC is a component of eukaryotic DNA polymerase [11]. It operates by loading a ring-shaped homotrimer called proliferating cell nuclear antigen (PCNA) onto DNA in an ATP-dependent manner to act as a sliding clamp for various proteins involved in DNA replication processes [12]. Furthermore, RFCs have the capability to interact with proteins involved in cell cycle checkpoints, initiating signal cascades downstream of DNA damage checkpoints. This involvement extends to participating in the repair processes of mismatched DNA and excision repair in cases of damaged DNA [13, 14]. RFC consists of a large subunit, replication factor C subunit 1 (RFC1), and four small subunits, replication factor C subunits 2 through 5 (RFC2–5). Particularly, RFC2 and RFC3 have garnered attention for their involvement in tumorigenesis due to their interaction with the c-MYC oncogene, which promotes cell division and proliferation [15]. It is noteworthy that, except for RFC1, the other four subunits of RFC (RFC2–5) exhibit elevated expression levels in various malignant tumors [16,17,18,19,20]. RFC3, in particular, has been extensively studied in the context of cancer. It forms a complex with proliferating cell nuclear antigen (PCNA), and a decrease in RFC3 expression restricts cancer cell multiplication [21].
Previous investigations have linked RFC3 to liver, breast, esophageal, and ovarian cancers, highlighting its critical role in cellular proliferation, invasion, and metastasis [21,22,23,24]. An interesting observation is the inhibitory effect of 9-cis retinoic acid–activated retinoid X receptor α on cancer growth by disrupting the RFC3-PCNA complex, leading to S phase entry arrest [25]. These findings collectively suggest RFC3 potential significance as a cancer antigen. In esophageal adenocarcinoma, RFC3 was found to be overexpressed, and its knockdown exhibited an anti-proliferative effect. This effect appears to stem from the induction of cell cycle arrest at the S phase, a critical checkpoint governed by DNA replication complex formation and function [16]. The knockdown of RFC3 resulted in elevated levels of S phase–associated proteins, including p21, p53, and p57, while reducing the expression of cyclin A. In the cell cycle, the CDK2/cyclin A complex drives progression through the G1-S phase transition, a tightly regulated step in cell proliferation. CDK inhibitors p21 and p51 bind to CDK2, inhibiting its activity and causing cells to remain in the G1/S phase, effectively halting cell cycle progression. Moreover, p53, known to activate p21 expression, likely plays a role in RFC3-mediated upregulation of p21 and/or p51, leading to the inhibition of G1-S phase transition [22]. However, further research is needed to unveil the intricate mechanisms underlying RFC3 regulation of cell cycle–related proteins, and its role in HNSCC remains largely unexplored. In this study, we comprehensively analyze RFC3 expression in OSCC and HNSCC tumor samples. Our investigation includes examining its clinicopathological correlations, tumor infiltration, patient prognosis, and functional pathways in HNSCC. By shedding light on RFC3 role in HNSCC, we aim to contribute valuable insights into the molecular underpinnings of this challenging cancer and potentially identify new avenues for therapeutic intervention.
Materials and methods
RFC3 expression analysis using online database
The Cancer Genome Atlas-Head-Neck Squamous Cell Carcinoma (TCGA-HNSCC) dataset (n = 520) and normal tissues (n = 44) were identified and used to measure RFC3 gene expression. The expression of RFC3 and the clinicopathological features of HNSCC were investigated using the UALCAN database (http://ualcan.path.uab.edu) [26].
Patient and sample collection
To validate RFC3 expression, we collected tumor and adjacent normal tissue samples from 30 OSCC patients. During surgery, samples of the tumor and surrounding normal tissues were collected and immediately stored at − 80 °C until further processing. Patient with primary OSCC only is included, and patients with systemic disease and recurrence were excluded. Patient clinicopathological data were collected such as stage, grade, site, and pattern of invasion which are listed in Table 1. The Institutional Ethical Committee approved the study and followed the principles of the Declaration of Helsinki. Informed consent was obtained from the patient or guardian.
RNA extraction and RT-qPCR analysis
Total RNA was extracted from the tumor and adjacent normal tissues using TRIzol reagent (Thermo Fisher Scientific, Waltham, MA, USA), according to the manufacturer’s instructions. RNA quality and quantity were analyzed using a Nanodrop One (Thermo Scientific, USA). RNA was converted into cDNA using the Takara 1st-strand cDNA synthesis kit (Takara, Tokyo, Japan) according to the manufacturer’s instructions. For RT-qPCR analysis, cDNA was used as a template, and gene expression was analyzed. The primer sequence was synthesized (Eurofins, Bangalore, India) and collected from the literature. The sequence of the primer RFC3 forward primer 5′-GCC TGC AGA GTG CAA CAA TA-3′ and reverse primer 5′-TCA AGG AGC CTT TGT GGA GT-3′; GAPDH forward primer 5′-TCC AAA ATC AAG TGG GGC GA-3′ and reverse primer 5′-TGA TGA CCC TTT TGG CTC CC-3′. The following components were used to create a total of 25 µl of PCR reactions: 2 µl of cDNA template, 50 mM forward and reverse primers, 12.5 µl of 2 × SYBR, and DDH2O. The following protocol was used for RT-qPCR provided by Bio-Rad CFX Opus 96 (Bio-Rad, Hercules, CA, USA). Initial denaturation at 95 °C for 3 min, followed by 40 cycles of denaturation at 95 °C for 10 s, and annealing at 58 °C for 30 s. GAPDH was used as a reference gene. The 2−ΔΔCt method was used to calculate the relative gene expression.
Survival analysis by Kaplan–Meier plotter
The Kaplan–Meier plot (https://kmplot.com/) [27] contains TCGA cancer patient data on gene expression to analyze patient survival. In this study, the prognostic value of RFC3 mRNA levels in HNSCC was analyzed using Kaplan–Meier survival analysis.
In silico functional analyses
The Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) (https://string-db.org/) [28] is an online database used to analyze protein and protein interaction networks. RFC3 protein interactions were analyzed using the STRING database. Metascape is another database used for gene annotation, network visualization, and functional enrichment analysis. The biological significance of gene sets or clusters can be inferred by using this information to help researchers interpret large-scale omics data. The data were obtained from STRING and processed in Metascape (https://metascape.org/) [29] for functional enrichment analysis of RFC3.
cBioPortal, TIMER, and immunohistochemistry analysis
cBioPortal (https://www.cbioportal.org/) [30] is a platform for the exploratory analysis of massive cancer genomic datasets, hosting data from important consortium projects such as TCGA and TARGET, as well as individual lab studies. To investigate the RFC3 connection between other genes in our study, cBioPortal was used. TCGA or user-provided tumor profiles can be used with TIMER2.0 (http://timer.cistrome.org) [31], which uses six cutting-edge algorithms to provide a more reliable estimation of immune infiltration levels. In our study, we used TIMER to analyze the correlation between RFC3 expression and tumor immune cell infiltration. The most popular method for identifying proteins in tissues is immunohistochemistry (IHC), which is used for histopathological diagnosis and research. In contemporary pathology, antibodies are crucial in determining the presence and location of particular proteins in surgical specimens. We analyzed RFC3 expression in HNSCC samples using the Protein Atlas immunohistochemistry tool (https://www.proteinatlas.org/) [32].
Statistical analysis
SPSS software version 25 (SPSS Inc., Chicago, IL, USA) and GraphPad Prism version 9.4.0 were used for statistical analysis; Student’s t-test or one-way ANOVA and multivariate COX regression were performed. The cutoff for statistical significance was set at P < 0.05.
Results
RFC3 gene and protein expression was significantly upregulated in HNSCC tumors
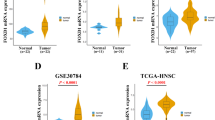
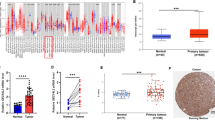
In this study, we used various databases to analyze RFC3 expression in various tumor and normal samples. Similarly, all sources revealed that RFC3 expression was significantly increased in a variety of cancer samples compared to the control (Fig. 1A). UALCAN results suggest that RFC3 expression was significantly higher in HNSCC tissues than in normal tissues (Fig. 1B), and our validation results using RT-qPCR were similar to the UALCAN results (Fig. 1C). Matched OSCC tumor tissues and adjacent normal tissues showed significant differences in RFC3 expression (Fig. 1D). RFC3 protein expression was analyzed using the UALCAN database (Fig. 1E), and immunohistochemistry (Fig. 1F) showed that RFC3 is highly expressed in HNSCC tissues.
UALCAN database showed that RFC3 mRNA is highly expressed in various cancers (A) including in HNSCC (B). RT-qPCR results are also similar to UALCAN database results, the mRNA level significantly upregulated (C) and RFC3 mRNA level significantly increased in matched OSCC tissues (D). RFC3 protein was significantly increased in HNSCC samples (E). Immunohistochemistry results showed that RFC3 protein stained at a medium level in the HNSCC sample (F). ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05
RFC3 expression is associated with clinicopathological features, prognosis, and tumor infiltration
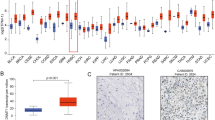
The UALCAN database was used to examine RFC3 expression and clinicopathological features of HNSCC. The UALCAN database results revealed that high expression of RFC3 mRNA was substantially linked to tumor stage, tumor grade, nodal metastasis, and HPV status (Fig. 2A–D). Similarly, high expression of the RFC3 protein was associated with tumor stage and grade in HNSCC (Fig. 2E, F). Further analysis of the proteomics data in the UALCAN database suggested that the high expression of RFC3 was linked to HNSCC via well-known oncogenic signaling pathways, such as MYC/MYCN, HIPPO, and mTOR (Fig. 3A–C). We examined the survival rate using Kaplan–Meier plots. High RFC3 expression was associated with poor overall and relapse-free HNSCC survival rates (Fig. 4A, B). TIMER2.0 database was used to explore the association between RFC3 expression and the infiltration of different immune cells. RFC3 expression was positively correlated with B cells and dendritic cells, whereas it was negatively correlated with CD8 + T cells and macrophages in HNSCC (Fig. 4C).
Functional analysis revealed RFC3 and its network is strongly associated with HNSCC
The protein network of RFC3 was analyzed using the STRING database, which provides details about protein–protein interactions. Several well-known oncogenic proteins, including PCNA, RFC5, RFC2, MCM2, and DSCC1, mainly interacted with RFC3 (Fig. 5A). Metascape analysis of the RFC3 network revealed that it is significantly involved in the cell cycle, DNA replication, and mismatch repair (Fig. 5B). Based on the RFC3 network and pathways, we identified an association between the RFC3 network and HNSCC (Fig. 5C), suggesting that RFC3 plays a crucial role in the pathogenesis of HNSCC. We used cBioPortal to analyze the correlation between RFC3 and oncogene expression and found that RFC3 expression positively correlated with proliferating cell nuclear antigen (PCNA) and Mini chromosome maintenance complex component genes (MCM2) (Fig. 5D, E) in HNSCC. Thus, our results suggested that targeting RFC3 or its network is a promising approach for HNSCC treatment.
STRING database shows the RFC3 protein and its partner protein interaction network (A) and functional enrichment analysis of RFC3 gene associated with cell cycle, mismatch repair (B), and also it shows RFC3 network is associated with HNSCC (C). RFC3 expression positively correlated with PCNA (D) and MCM2 (E) expression in HNSCC
Discussion
In this study, we conducted a comprehensive analysis of RFC3 gene and protein expression in the context of HNSCC. Our findings shed light on the significance of RFC3 in this particular cancer type and its potential implications. RFC3 expression has also been linked to different cancers, including liver, breast, ovarian, and esophageal cancers. Previous studies have also shown that RFC3 is involved in cellular invasion, progression, and metastasis and predicts poor prognosis [21,22,23,24, 33].
In a study conducted by Gong et al., an examination of RFC3 expression in lung squamous cell carcinoma using immunohistology analysis revealed a significant overexpression of RFC3 in major lung cancer tissues when compared to their corresponding normal lung tissue counterparts. This overexpression of RFC3 was found to be significantly associated with nodal metastasis status and had implications for prognosis. Multivariate analysis further underscored RFC3 expression as an independent risk factor in this context [34]. Similarly, in another study by Yu-He et al., RFC3 was found to be highly expressed in breast cancer tissue, and this heightened expression was significantly associated with metastasis and poor prognosis [21]. Additionally, investigations into RFC3 expression in hepatocellular and ovarian carcinoma have also suggested a potential link between RFC3 expression and cancer. Functional knockdown experiments in cell lines and animal models have shown that reducing RFC3 levels leads to decreased cell proliferation and migration, as well as increased apoptosis [22, 23]. Our current research aligns with these findings, as we observed high RFC3 expression in HNSCC tissues. This elevated expression was found to be associated with advanced stage, higher grade, metastasis, and poorer prognosis in HNSCC patients. Multivariate analysis of our data similarly supports RFC3 overexpression as an independent risk factor in the development of HNSCC. These collective results emphasize the significance of RFC3 in various cancer types and its potential as a target for further therapeutic exploration. Because RFC3 is important in the creation of the DNA replication complex, it has been proposed that RFC3 overexpression is responsible for tumor formation.
Previously, a study by Li et al. analyzed the activity of RFC family members using the STRING database and found that RFC is mainly involved in telomere maintenance, DNA mismatch repair, DNA replication, and nucleotide excision repair [28, 35]. RFC can attach DNA polymerase and PCNA to a primer-bound DNA template to create the DNA-RFC-PCNA-DNA polymerase complex. This complex is then extended along the DNA template by human single-stranded DNA-binding protein (SSB) in the presence of deoxynucleotides (dNTPs). RFC can also connect to cell cycle checkpoint proteins to initiate signal transduction after DNA damage checkpoints, allowing it to participate in mismatch and excision repair of damaged DNA [13, 14].
The extensive body of research into RFC3 sheds light on its multifaceted role in cancer, suggesting its potential as a significant biomarker, a key player in cancer cell proliferation and survival, a determinant of cancer cell invasion and metastasis, and a participant in various regulatory networks. RFC3 influence on cancer cell proliferation extends beyond its individual actions. It interacts with various factors to orchestrate cancer cell proliferation in vivo. For instance, RFC3 interaction with retinoid X receptor α (RXRα) is implicated in the suppression of retinoic acid–sensitive breast cancer cell growth [25]. Moreover, in cervical cancer tissues, upregulated SIX homeobox 1 (SIX1) expression led to the significant upregulation of several DNA replication initiation–related genes, including RFC3, RFC4, and RFC5 [36]. Additionally, E2F and cyclic AMP response element-binding protein (CREB) were suggested as regulators of RFC3 expression in certain cancer cell lines [37]. These interactions underline the complexity of RFC3 role in cancer cell proliferation and its integration into broader regulatory networks. Our study’s proteomic analysis of RFC3 through UALCAN also found that high expression of RFC3 is linked to HNSCC via well-known oncogenic signaling pathways, such as MYC/MYCN, HIPPO, and mTOR. Furthermore, STRING and functional enrichment analysis showed that RFC3 dysregulation is associated with HNSCC pathogenesis via cell cycle, DNA replication, and activation of the pre-replication complex pathways. Therefore, RFC3 might be involved in tumorigenesis through regulating cancer signaling pathways and other oncogenes.
Furthermore, we have analyzed RFC3 interacting genes PCNA and MCM2 in HNSCC. PCNA is a nuclear protein that serves as a marker of cell proliferation. Most solid tumor types, including colorectal cancer and breast cancer, show a substantial correlation between PCNA expression and prognosis, and survival [38, 39]. PCNA is significantly overexpressed in HNSCC and can be used as a biomarker for HNSCC [40, 41]. Minichromosome maintenance proteins (MCMs) perform crucial functions in regulating replication time throughout the cell cycle and demonstrate helicase activity during replication. Overexpression of MCMs has been observed in several malignant tissues, including HNSCC and cancer cell lines [42, 43]. Our results also showed that RFC3 expression was positively correlated with PCNA and MCM2 in HNSCC pathogenesis. From these results, we suggest that RFC3 is associated with PCNA and MCM2, and may be involved in cancer cell proliferation and the cell cycle pathway in HNSCC.
In summary, the diverse body of evidence surrounding RFC3 emphasizes its multifaceted involvement in cancer, ranging from its potential as a diagnostic biomarker to its critical roles in cancer cell proliferation, survival, invasion, and interaction within complex regulatory networks. Understanding RFC3 precise mechanisms in different cancer types could pave the way for innovative therapeutic strategies and diagnostic approaches. However, the current study is limited by its small sample size, gene expression, and in silico analysis. Further large-scale studies, protein-level expression analysis, and functional analysis using in vitro and in vivo models will help reveal the molecular mechanism of RFC3 in HNSCC pathogenesis.
Conclusion
In conclusion, our findings suggest that RFC3 expression is significantly increased in HNSCC and that increased expression is important for HNSCC tumorigenesis. Furthermore, findings also suggest that RFC3 is a potential prognostic biomarker and therapeutic target in HNSCC. Nevertheless, further studies are required to understand the crucial role of RFC3 in HNSCC tumorigenesis.
Data Availability
All data generated or analyzed during this study are downloaded from TCGA datasets.
References
Johnson NW, Jayasekara P (2000) Amarasinghe AAHK (2011) Squamous cell carcinoma and precursor lesions of the oral cavity: epidemiology and aetiology. Periodontol 57:19–37
Marur S, D’Souza G, Westra WH, Forastiere AA (2010) HPV-associated head and neck cancer: a virus-related cancer epidemic. Lancet Oncol 11:781–789
Neville BW, Day TA (2002) Oral cancer and precancerous lesions. CA Cancer J Clin 52:195–215
Anderson CR, Sisson K, Moncrieff M (2015) A meta-analysis of margin size and local recurrence in oral squamous cell carcinoma. Oral Oncol 51:464–469
Wang B, Zhang S, Yue K, Wang X-D (2013) The recurrence and survival of oral squamous cell carcinoma: a report of 275 cases. Chin J Cancer 32:614–618
Panzarella V, Pizzo G, Calvino F et al (2014) Diagnostic delay in oral squamous cell carcinoma: the role of cognitive and psychological variables. Int J Oral Sci 6:39–45
Torgovnick A, Schumacher B (2015) DNA repair mechanisms in cancer development and therapy. Front Genet 6:157
Pascucci B, Stucki M, Jónsson ZO et al (1999) Long patch base excision repair with purified human proteins. DNA ligase I as patch size mediator for DNA polymerases delta and epsilon. J Biol Chem 274:33696–33702
Shimada M, Okuzaki D, Tanaka S et al (1999) Replication factor C3 of Schizosaccharomyces pombe, a small subunit of replication factor C complex, plays a role in both replication and damage checkpoints. Mol Biol Cell 10:3991–4003
Sancar A, Lindsey-Boltz LA, Unsal-Kaçmaz K, Linn S (2004) Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem 73:39–85
Uhlmann F, Cai J, Flores-Rozas H et al (1996) In vitro reconstitution of human replication factor C from its five subunits. Proc Natl Acad Sci U S A 93:6521–6526
Mossi R, Hübscher U (1998) Clamping down on clamps and clamp loaders–the eukaryotic replication factor C. Eur J Biochem 254:209–216
Majka J, Burgers PMJ (2004) The PCNA–RFC families of DNA clamps and clamp loaders. In: Progress in nucleic acid research and molecular biology. Academic Press 227–260
Sakato M, O’Donnell M, Hingorani MM (2012) A central swivel point in the RFC clamp loader controls PCNA opening and loading on DNA. J Mol Biol 416:163–175
Koch HB, Zhang R, Verdoodt B et al (2007) Large-scale identification of c-MYC-associated proteins using a combined TAP/MudPIT approach. Cell Cycle 6:205–217
Xiong S, Wang Q, Zheng L et al (2011) Identification of candidate molecular markers of nasopharyngeal carcinoma by tissue microarray and in situ hybridization. Med Oncol 28(Suppl 1):S341–S348
Arai M, Kondoh N, Imazeki N et al (2009) The knockdown of endogenous replication factor C4 decreases the growth and enhances the chemosensitivity of hepatocellular carcinoma cells. Liver Int 29:55–62
Niu G, Wang D, Pei Y, Sun L (2017) Systematic identification of key genes and pathways in the development of invasive cervical cancer. Gene 618:28–41
Srihari S, Kalimutho M, Lal S et al (2016) Understanding the functional impact of copy number alterations in breast cancer using a network modeling approach. Mol Biosyst 12:963–972
Martinez I, Wang J, Hobson KF et al (2007) Identification of differentially expressed genes in HPV-positive and HPV-negative oropharyngeal squamous cell carcinomas. Eur J Cancer 43:415–432
He Z-Y, Wu S-G, Peng F et al (2017) Up-regulation of RFC3 promotes triple negative breast cancer metastasis and is associated with poor prognosis via EMT. Transl Oncol 10:1–9
Yao Z, Hu K, Huang H et al (2015) shRNA-mediated silencing of the RFC3 gene suppresses hepatocellular carcinoma cell proliferation. Int J Mol Med 36:1393–1399
Shen H, Xu J, Zhao S et al (2015) ShRNA-mediated silencing of the RFC3 gene suppress ovarian tumor cells proliferation. Int J Clin Exp Pathol 8:8968–8975
Lockwood WW, Thu KL, Lin L et al (2012) Integrative genomics identified RFC3 as an amplified candidate oncogene in esophageal adenocarcinoma. Clin Cancer Res 18:1936–1946
Maeng S, Kim GJ, Choi EJ et al (2012) 9-Cis-retinoic acid induces growth inhibition in retinoid-sensitive breast cancer and sea urchin embryonic cells via retinoid X receptor α and replication factor C3. Mol Endocrinol 26:1821–1835
Chandrashekar DS, Karthikeyan SK, Korla PK et al (2022) UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25:18–27
Nagy Á, Munkácsy G, Győrffy B (2021) Pancancer survival analysis of cancer hallmark genes. Sci Rep 11:6047
Szklarczyk D, Kirsch R, Koutrouli M et al (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 51:D638–D646
Zhou Y, Zhou B, Pache L et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404
Li T, Fu J, Zeng Z et al (2020) TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48:W509–W514
Uhlén M, Fagerberg L, Hallström BM et al (2015) Proteomics Tissue-based map of the human proteome. Science 347:1260419
Shen H, Cai M, Zhao S et al (2014) Overexpression of RFC3 is correlated with ovarian tumor development and poor prognosis. Tumour Biol 35:10259–10266
Gong S, Qu X, Yang S et al (2019) RFC3 induces epithelial-mesenchymal transition in lung adenocarcinoma cells through the Wnt/β-catenin pathway and possesses prognostic value in lung adenocarcinoma. Int J Mol Med 44:2276–2288
Li Y, Gan S, Ren L et al (2018) Multifaceted regulation and functions of replication factor C family in human cancers. Am J Cancer Res 8:1343–1355
Liu D, Zhang X-X, Xi B-X et al (2014) Sine oculis homeobox homolog 1 promotes DNA replication and cell proliferation in cervical cancer. Int J Oncol 45:1232–1240
Chae H-D, Mitton B, Lacayo NJ, Sakamoto KM (2015) Replication factor C3 is a CREB target gene that regulates cell cycle progression through the modulation of chromatin loading of PCNA. Leukemia 29:1379–1389
al-Sheneber IF, Shibata HR, Sampalis J, Jothy S (1993) Prognostic significance of proliferating cell nuclear antigen expression in colorectal cancer. Cancer 71:1954–1959
Shrestha P, Yamada K, Wada T et al (1992) Proliferating cell nuclear antigen in breast lesions: correlation of c-erbB-2 oncoprotein and EGF receptor and its clinicopathological significance in breast cancer. Virchows Arch A Pathol Anat Histopathol 421:193–202
Lörz M, Meyer-Breiting E, Bettinger R (1994) Proliferating cell nuclear antigen counts as markers of cell proliferation in head and neck cancer. Eur Arch Otorhinolaryngol 251:91–94
Lu EM-C, Ratnayake J, Rich AM (2019) Assessment of proliferating cell nuclear antigen (PCNA) expression at the invading front of oral squamous cell carcinoma. BMC Oral Health 19:233
Yu S, Wang G, Shi Y et al (2020) MCMs in cancer: prognostic potential and mechanisms. Anal Cell Pathol 2020:3750294
Zhi X, Lamperska K, Golusinski P et al (2015) Gene expression analysis of head and neck squamous cell carcinoma survival and recurrence. Oncotarget 6:547–555
Acknowledgements
We extend our sincere gratitude to all individuals and organizations who contributed to this study. We appreciate the valuable resources provided by The Cancer Genome Atlas (TCGA) and the Tumor Immune Estimation Resource (TIMER) database, which significantly facilitated our data analysis. We are also thankful for the technical support and guidance offered by our research team. Furthermore, we would like to acknowledge the participants who generously provided samples for this research. Our acknowledgments also go to the reviewers and editors for their insightful comments and suggestions, which improved the quality of our work. This research would not have been possible without the collaborative efforts of everyone involved, and we are deeply grateful for their contributions.
Funding
The study was self-funded by the authors’ institution.
Author information
Authors and Affiliations
Contributions
K.R.A., B.K., and P.A. contributed to study conception and design. K.R.A., C.P., and B.K. data curation from databases, recruited participants and collected the data. B.K and A.P. performed formal analysis and supervision. K.R.A., V.P.J., and P.A. analyzed and interpreted the data. K.R.A., C.P., and B.K. drafted the manuscript. All authors critically revised and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
This study was approved by the Institutional Ethical Committee of the Saveetha Dental College and Hospital. All participants signed an informed consent form.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
AVS, K.R., Pandi, C., Kannan, B. et al. RFC3 serves as a novel prognostic biomarker and target for head and neck squamous cell carcinoma. Clin Oral Invest 27, 6961–6969 (2023). https://doi.org/10.1007/s00784-023-05316-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-023-05316-4