Abstract
Objectives
The aim of this study was the analysis of WNT10A variants in seven families of probands with various forms of tooth agenesis and self-reported family history of cancer.
Materials and methods
We enrolled 60 young subjects (aged 13 to 17) from the Czech Republic with various forms of tooth agenesis. Dental phenotypes were assessed using Planmeca ProMax 3D (Planmeca Oy, Finland) with Planmeca Romexis software (version 2.9.2) together with oral examinations. After screening PAX9, MSX1, EDA, EDAR, AXIN2 and WNT10A genes on the Illumina MiSeq platform (Illumina, USA), we further analyzed the evolutionarily highly conserved WNT10A gene by capillary sequencing in the seven families.
Results
All the detected variants were heterozygous or compound heterozygous with various levels of phenotypic expression. The most severe phenotype (oligodontia) was found in a proband who was compound heterozygous for the previously identified WNT10A variant p.Phe228Ile and a newly discovered c.748G > A variant (p.Gly250Arg) of WNT10A. The newly identified variant causes substitution of hydrophobic glycine for hydrophilic arginine.
Conclusions
We suggest that the amino acid changes in otherwise highly conserved sequences significantly affect the dental phenotype. No relationship between the presence of WNT10A variants and a risk of cancer has been found.
Clinical relevance
Screening of PAX9, MSX1, EDA, EDAR, AXIN2 and WNT10A genes in hope to elucidate the pattern of inheritance in families.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tooth development is controlled by a set of highly conserved regulatory genes with similar functions in different animal species [1]. It is of crucial importance for all the components of signaling pathways to be present at the right place, at the right time and at the required quantity [2, 3]. Any deviation in this process may lead to developmental malfunctions [1]. Tooth agenesis is one of the most prevalent craniofacial anomalies [4]. According to the number of missing teeth, tooth agenesis is classified as hypodontia (missing 1–5 teeth), oligodontia (missing 6 or more teeth) or anodontia (complete absence of dentition) [5]. The prevalence of tooth agenesis (excluding the third molars) depends somewhat on the ethnicity of the populations under study. Generally, more severe phenotypes occur less frequently than the milder ones. Females have been found to have higher prevalence of tooth agenesis than males [6,7,8].
Tooth development is a dynamic and complex process involving more than 350 genes [9, 10]. Among these, PAX9, MSX1, EDA, EDAR, AXIN2 and WNT10A are the most studied ones [11,12,13,14,15]. Most of the variants are found in patients with tooth agenesis [16]. The WNT10A gene belongs to the WNT family (wingless type MMTV integration site) which is apparently highly conserved in the animal kingdom, particularly in the vertebrates [17, 18]. The family consists of structurally related genes encoding secreted signaling proteins [19]. WNT proteins bind to Frizzled receptors (G-protein coupled receptors) and to low-density lipoprotein receptor-related protein (LRP) [20]. They are involved in the differentiation of tissues and organs during embryonic development [21,22,23]. The human WNT family consists of 19 genes, among which WNT10A shows the strongest association with odontogenesis [24, 25]. WNT10A encodes a signaling protein, which plays an essential role at multiple stages of tooth morphogenesis including the induction and maintenance of primary and secondary enamel knots [26, 27]. This gene is 13.4 kb long, located on the long arm of 2q35 and has 4 exons. The first and the last exons contain untranslated regions. The transcribed protein is 417 amino acids long with a 23–24 cysteine residue motif conserved among all WNT proteins [19]. WNT proteins share 27–83% amino acid sequence identity [28, 29] and contain a signal peptide (residues 1–35) and a WNT domain ranging from 60–417 amino acids [30]. WNT10A is constitutively expressed in the skin and is particularly abundant during the formation of ectodermal appendages, including hair follicles [31,32,33,34]. Tooth agenesis may occur as an isolated form without further syndromes or as part of one or more syndromes [10, 35]. In addition to tooth agenesis, the most common syndromes caused by WNT10A variants are a wide range of ectodermal dysplasias, including hypohidrotic ectodermal dysplasia, odonto-onycho-dermal dysplasia (OODD) and Shop-Schulz-Passarge syndrome (SSPS) [17, 30]. These conditions show a high degree of phenotypic variability with unclear genotype–phenotype correlation, suggesting a role of other, as yet unknown variants [30, 36, 37]. The occurrence of WNT10A variants in patients with tooth agenesis is between 44 and 62% [38,39,40,41], although one Swedish study reported 27.7% [42]. The number of missing teeth correlates with the proportional presence of variants; in patients missing 1–3 teeth, WNT10A variants were found in 15.8% of cases while in those missing 4 or more teeth the proportion of the variants was 51.6% [41].
WNT signaling has also been associated with oncogenesis [43, 44]. The simultaneous occurrence of tooth agenesis and cancer including in patients carrying particular variants of components of the WNT signaling cascade has recently been reviewed [45]. Nevertheless, details of putative interactions between the mutations associated with tooth agenesis and those involved in mechanisms of cancer are yet to be elucidated [46, 47].
In this study, we discuss WNT10A variants that were detected in seven families of probands with various forms of tooth agenesis and self-reported family history of cancer.
Materials and methods
From among the patients with dental anomalies who presented at the Department of Stomatology of St. Anne's Faculty Hospital and Faculty of Medicine in Brno and at the Institute of Dentistry and Oral Sciences, Palacky University, Olomouc, we selected a sample of 60 individuals for further study. The selection criteria were as follows: age between 13 and 17 years, absence of any major known genetic diseases, no history of cancer and willingness (informed consent) to participate in the project. All participants were fully screened by next generation sequencing for exon mutations in PAX9, MSX1, AXIN2, EDA, EDAR and WNT10A genes. The focus was on WNT10A gene since the mutations in this gene were those found to be most often associated with the patterns of missing teeth observed in the patients. In patients where exon mutations were found, the first-degree relatives (parents and siblings) were then contacted and, if they agreed (and consented), they were added to the sample, clinically examined and NGS sequenced.
To assess dental phenotypes, Planmeca ProMax 3D (Planmeca Oy, Finland) in mode orthopantomogram with Planmeca Romexis software (version 2.9.2) was used together with oral examinations. The diagnosis of agenesis was determined based on the absence of (a) permanent tooth/teeth followed by a radiological examination using orthopantomogram. Additionally, we ruled out the possibility that the missing teeth may have resulted from earlier extractions by questioning the patients and checking their medical records when available. The ageneses were categorized as follows: absence of up to five permanent teeth except third molars was classified as hypodontia, absence of six or more permanent teeth was classified as oligodontia; the total agenesis of all permanent teeth was not encountered in the group. Information on the incidence of oncological diseases was obtained by interviewing the probands, their relatives (or, when possible, interviewing additional family members).
Genomic DNA was obtained from 200 µL of whole blood taken from the fingers by EDTA coated capillaries Microvette® 200 µL, K3 EDTA (Sarstedt, Germany). DNA isolation was performed using the Zephyrus Magneto (Elisabeth Pharmacon, Czech Republic) and Prepito® NA Body Fluid Kit (PerkinElmer, Germany). DNA quality and concentration were verified spectrophotometrically. DNA was diluted to a concentration of 150 ng in 50 µL PCR water and sheared to a size of 200 bp using the Covaris S220 Sonicator (Covaris, USA) (treatment time 180 s, volume 130 µL). Next, size and quantity of DNA fragments were verified on 2200 TapeStation Instrument (Agilent Technologies, USA). NGS library was prepared by enrichment of genomic regions for PAX9, MSX1, AXIN2, EDA, EDAR and WNT10A genes according to the protocol for SeqCap EZ System (Roche NimbleGen, USA). Concentrations of prepared libraries before pooling and of the final library were determined by KAPA qPCR KAPA Library Quantification Kit (Kapa Biosystems, USA). The final library was sequenced on the Illumina MiSeq platform (Illumina, USA) using MiSeq Reagent Kit V2 (300 cycles) according to manufacturer recommendations for 14 pM libraries.
NGS data were analyzed according to Roche NimbleGen workflow [48]. As a reference genome, Hg38 was used. BWA 0.7.13 software package [49] was used for indexing the reference genome and alignment of reads. Adapter trimming was performed using Trimmomatic 0.32 software [50]. PCR duplicates were removed using Picard Tools 1.110 software. Variant calling and filtering were performed using SAM tools 1.3 and BCF tools 1.3 software [51, 52]. Using software R, the depth of coverage was calculated for each position on the reference genome corresponding to the selected genes (PAX9, MSX1, AXIN2, EDA, EDAR and WNT10A) [53]. Sequence data were browsed through using Integrative Genomics Viewer 2.3 (IGV) [54, 55].
Selected variants in WNT10A were further analyzed by capillary sequencing. DNA isolation followed the same protocol as described above for NGS (Table 1).
PCR was carried out using the EliZyme™ HS FAST MIX (Elisabeth Pharmacon, Czech Republic) on a Veriti® thermal cycler (Applied Biosystems, USA). The PCR conditions were as follows: activation/denaturation at 95 °C for 2 min; 40 cycles at 95 °C for 15 s and 62 °C (for exon 2) or 63 °C (for exon 3) for 15 s; and 72 °C for 15 s. Primer sequences are given in Table 1. Amplicons were purified by ExoI‐FastAP (Fermentas, USA), incubated at 37 °C for 15 min and subsequently at 85 °C for 15 min for enzyme deactivation. Sequencing was performed using BigDye® Terminator v.3.1 (Life Technologies, USA). Sequencing reactions/products were purified using EDTA/ethanol precipitation, resuspended in 10 μl of Hi‐Di formamide (Life Technologies, USA) and finally sequenced on the automated ABI 3130 Genetic Analyzer. Sequences were edited and compared with reference sequence (accession NG_012179.1) of the WNT10A gene from the GenBank Database (NCBI).
Results
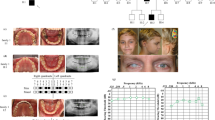
Family histories provided by the probands and available members of their families focused on tooth agenesis and cancer (Fig. 1). Initial (NGS) screening of their DNA samples singled out WNT10A as the gene with the greatest variability in both phenotypes (as indicated by the family histories) and variants causing an amino acid change. Furthermore, the NGS screen pointed to exons 2 and 3 as the locus of potential variants of interest, prompting additional, more focused, analyses (capillary sequencing; Table1). In addition to previously known variants, we found a new missense variant c.748G > A in exon 3, causing Gly250Arg amino acid change (Fig. 2). Based on the above data, we created seven family pedigrees, as displayed in Fig. 1A–G. We should add that the NGS screen failed to reveal any new variants of relevance for the present study in PAX9, MSX1, AXIN2, EDA or EDAR genes; note that, for clarity, ectodermal dysplasia and other hereditary syndromes often associated with WNT10A variants are omitted in Fig. 1.
The phenotypic description of studied families
Family A
The female proband (Z410) has hypodontia of lower central incisors and several extracted teeth. Her mother (Z410B) has fully developed dentition. Her stepbrother and aunt have tooth agenesis which was reported only by family records. The proband is a carrier of heterozygous variant c.208C > T causing p.Arg70Trp.
Family B
The male proband (Z450) has hypodontia of five permanent teeth, including lower central incisors, upper right lateral incisor and upper canines. Neither the mother (Z450B) nor the brother (Z450D) suffer from tooth agenesis. The deceased father had a complete dentition (information about third molars is not available). Grandfather from the father's side had liver cancer. The p.Phe228Ile amino acid change was identified in all three family members.
Family C
The female proband (Z451) has hypodontia of upper second molars and lower right second molar. Her mother (Z451B) is missing only second right upper incisor while neither her father (Z451C) nor her brother (Z451D) have hypodontia. Grandmother from the father's side had breast cancer. DNA sequencing uncovered compound heterozygous variants c.649G > A and c.682 T > A resulting in p.Asp217Asn and p.Phe228Ile in both siblings. Their mother and father are carriers of heterozygous variants c.649G > A and c.682 T > A, respectively.
Family D
The female proband (Z605) has 8 teeth missing: upper left lateral incisor, upper canines, upper second premolars, lower central incisors and lower left second premolar. No other family members have tooth agenesis. Grandfather from the mother's side had liver cancer and grandmother from the father's side had breast cancer. A heterozygous variant c.682 T > A (p.Phe228Ile) was found in the proband, her mother (Z605B) and one brother. No other variants were found in the selected exons.
Family E
The male proband (Z620) has oligodontia, missing 7 teeth: upper right lateral incisor, all upper premolars and lower second premolars. No other family members had missing teeth. This case, as well as that of the previous family D, can be classified as sporadic form of oligodontia. The same p.Phe228Ile change was found in the proband and his father (Z620C).
Family F
The male proband (Z623) suffers from severe oligodontia, missing 15 teeth: upper lateral incisors, upper right canine, upper premolars, lower incisors and canines and lower second premolars. Mother's sister had leukemia. The proband carries a compound heterozygous c.682 T > A and newly found c.748G > A variant resulting in p.Phe228Ile and p.Gly250Arg amino acid changes. Both originate from his parents—the mother (p.Gly250Arg), the father (p.Phe228Ile). The A allele of the novel variant c.748G > A was detected using NGS with average depth of coverage 47 for Z623. The position was read 54 times, 23 times (43%) for G allele and 31 times (57%) for A allele. GGG coding triplet is changed to AGG leading to arginine instead of glycine.
Family G
The male proband (Z624) has severe oligodontia, missing 13 teeth: upper left canine and second premolar, upper second molars, all lower incisors and left canine, lower second premolars and second molars. His aunt (Z458) lacks lower second molars. A compound heterozygous variant c.321C > A together with c.337C > T variant ([p.Cys107*], [p.Arg113Cys]) was identified in the proband. The stop codon variant was inherited from the mother (Z624B), who inherited it from her mother (Z624E). p.Arg113Cys was inherited from the father (Z624C). The proband’s aunt is also a carrier of p.Cys107* (Table 2).
Discussion
The main subject of the present study was the mode of inheritance of selected gene variants in seven families who have members with oligodontia. Comparison of family histories and genetic (DNA) analyses revealed six known variants and one newly identified WNT10A variant, all causing amino acid substitutions, as associated with apparently inherited moderate to severe forms of oligodontia. These substitutions are, respectively, p.Arg70Trp, p.Cys107Ter, p.Arg113Cys, p.Asp217Asn, p.Phe228Ile and (for the newly discovered one) p.Gly250Arg. Only heterozygous and compound heterozygous variants have been identified in the study. Despite PAX9 and MSX1 being among the most studied genes in tooth agenesis (in addition to AXIN2, EDA and EDAR), we have not been able to find any relevant variants of these genes in the seven families. On the face of it, this may sound surprising but, perhaps, it is not: WNT10A carries more variants associated with defective dentition than all the five genes mentioned above (see Supporting Information, Table S1) [15, 38, 39].
The most intriguing feature of our findings is the relationship between genotype and phenotype. Although some family members carry the same WNT10A variants, the phenotypes differ. This is the case in families B, D and E, where probands carry the same c.682 T > A (p.Phe228Ile) variant but are missing a variable number (5–8) of teeth. This corresponds with the mean of 7 from a Swedish study [42]. In contrast, members of their families developed a full dentition (Table 3). The mother (Family E, Z620B) does not carry any variants in the studied gene but may be missing upper left second premolar. The presence of p.Phe228Ile carries the risk of oligodontia but the precise magnitude of the risk remains to be determined. In one study, Mostowska et al. reported a ninefold increase [39] and an even higher risk (30-fold) was subsequently reported by the same group [16]. This is the most prevalent variant [42] and p.Phe228Ile is a frequent amino acid substitution associated with either autosomal dominant or autosomal recessive form of isolated hypodontia. This variant is present both in people with tooth agenesis and in the general population of European origin [24, 30, 36, 38], but is absent in the Chinese population [41]. In controls, the allele frequency has been reported up to 2.3% [30, 38, 39]. It has been suggested that heterozygous c.682 T > A (p.Phe228Ile) variant might provide “some kind of survival advantage” which could, however, depend on the ethnicity of the populations under study [25].
The most severe phenotype belongs to the male proband (Z623) of family F with 15 missing teeth (third molars excluded). He is the only family member suffering from severe oligodontia. According to family records, a female cousin of his mother appears to have had oligodontia, but it is not known how many teeth he had missing. This proband carries a heterozygous c.682 T > A (p.Phe228Ile) variant together with the novel heterozygous c.748G > A variant causing p.Gly250Arg substitution, changing hydrophobic glycine to hydrophilic arginine. Previous in silico analysis indicated that the substitution would be damaging [56]; the variant rs200387103 (c.748G > C) has been assigned at the same position in WNT10A resulting in a similar amino acid exchange. In our case, the c.748G > A variant was inherited from the mother who had no tooth agenesis. As far as we are aware, this variant has not been associated with tooth agenesis. It is located near c.682 T > A (p.Phe228Ile).
The male proband (Z624) from family G has a similarly high number of missing teeth (13). In his case, compound heterozygous (p.Cys107*) and c.337C > T (p.Arg113Cys) are present (see also [39] for a very similar observation). His aunt (Z458) is heterozygous for the nonsense variant, with only the lower second molars being absent. The variant c.321C > A (p.Cys107*) has a shorter protein missing a large part of the WNT domain. Homozygotes for this variant had very severe oligodontia, missing nearly all permanent teeth. Together with p.Phe228Ile, this variant should be considered as common in European populations [41]. Arg113 is a highly conserved amino acid and a change to cysteine is predicted to be potentially damaging [39, 57].
The variant p.Arg70Trp was detected in only one proband (Z410) in a heterozygous state. It has previously been shown to be associated with a particularly severe phenotype, possibly because of a damaging effect on protein structure [57].
Heterozygous p.Asp217Asn was detected in several members of family C, the female proband (Z451) being affected the most with 3 missing teeth. She is also heterozygous for p.Phe228Ile together with her brother (Z451D) who interestingly is not missing teeth (except for the lower right third molar). The inheritance of alleles from parents follows the same pattern as described by Kantaputra and Sripathomsawat [58]. The second aim of the present study has been to look for indications that oligodontia may carry an increased risk of cancer [45]. Of particular relevance here is an earlier suggestion that WNT10A plays a role in tumorigenesis by stimulating extracellular matrix (ECM) through mechanisms driven by WNT signaling [59, 60]. Together with AXIN2 it encodes components of the canonical Wnt pathway [27]. Polymorphism of AXIN2 gene has recently been linked to cancer, particularly that of bowel (colorectal), liver, prostate, ovarian or lung [61]. Studying such correlations could lead to novel ways in predicting the risk of serious malignancies, based on both genetics and specific cell-biological mechanisms.
In the present study, family members (grandparents and an aunt) of several probands were reported as diagnosed with cancer. In Fig. 1B, proband’s Z450 paternal grandfather had liver cancer. However, Z450 inherited his WNT10A variant (heterozygous p.Phe228Ile) from his mother. Thus, the case does not imply any association between p.Phe228Ile and hepatic cancer. As Z450 had none of the other variants of WNT10A under study, no case for an association of WNT10A gene-linked oligodontia with cancer could be made in his family. In Fig. 1C, paternal grandmother of Z451 had breast cancer (Z451B). Z451 could, however, have inherited p.Phe228Ile from either of her parents (Z451B and Z451C). Because this substitution occurs often in patients with tooth agenesis but also in controls, it is unlikely that it plays an important role in this case. The same is probably true in the case of the liver cancer in the maternal grandfather of Z605 (Fig. 1D) and the breast cancer in her paternal grandmother. No variant was found in the father (Z605C), so there is no connection to the breast cancer of his mother (paternal grandmother of Z605). In Fig. 1F, sister of the mother of proband Z623 had leukemia. The mother is a carrier of p.Gly250Arg. As there are, to the best of our knowledge, no publications on p.Gly250Arg in WNT10A (the present study is the first to identify it as having significance in a human condition), the putative association with leukemia would seem to be, at best, speculative.
Looking at the sex influenced expression of heterozygotes, the results suggest an even distribution. Although Bohring et al. proposed a slightly higher incidence in males [30], other studies did not find any distribution pattern [38, 40].
Heterozygosity is responsible for less severe cases of dental agenesis with incomplete penetrance [25, 38, 62]. According to Arzoo et al. [42], bi-allelic WNT10A variants were associated with absence of upper and lower molars as well as lower central incisors. In the present study, we were able to observe similar missing teeth also in the heterozygous state. Probands with bi-allelic variants in WNT10A had a mean number of 14.4 missing teeth [42]. Recorded missing teeth (third molars, premolars, lateral incisors, lower central incisors) correspond to those in other publications [25, 41, 42]. All our probands and family members with WNT10A variants had intact maxillary central incisors, suggesting that these teeth follow a developmental program independent from WNT10A, as shown in mice [63]. Bi-allelic variants are reported to have a significant influence on the number of missing teeth because of their quantitative effect.
To elucidate the peculiar genotype–phenotype interactions of WNT10A variants, we need to figure out the protein’s structure and its interactions. No crystal structure of the human WNT10A protein is currently available [18]. Arte et al. [55] suggested that all missense variants affect the N-terminal domain, which consists of an alpha-helical bundle. Their proposal is based upon alignment with WNT8, whose three-dimensional structure and interactions with the Frizzled receptors were uncovered by X-ray crystallography [64]. There is still uncertainty about the structure, particularly the number of α-helices. He et al. suggested 10 α-helices and 7 β-strands [62], while Nawaz et al. predicted 11 α-helices and 7 β-strands [65]. WNT proteins have a high number of disulfide bonds; therefore, protein conformation may be important for the correct interaction with their receptors [18]. Variants in exons 2 and 3 are responsible for severe oligodontia affecting a highly conserved region and structural elements of the protein [66].
Conclusions
We followed the inheritance of WNT10A variants in seven families of European ethnicity (Czech Republic). The actual inheritance of the alleles appears simple enough, yet their phenotypic expression remains unclear. All identified variants are heterozygous, with some family members carrying compound heterozygous variants. The most abundant variant is p.Phe228Ile. In one of the families, we discovered a genetic variant (causing p.Gly250Arg) never previously associated with any pathology. Together with the p.Phe228Ile variant, it caused a severe oligodontia (19 missing teeth, third molars included). All the sequenced nucleic acid substitutions in the present study code for evolutionarily highly conserved amino acid positions. Differences in phenotypic expression, especially in the heterozygous subjects, suggest possible involvement of additional genetic and/or environmental factors capable of perturbing the highly organized development of human dentition and resulting in tooth agenesis. The main strength of the study is the inclusion of almost complete sets of living first degree relatives and the use of whole gene analysis by NGS. Possible weakness is a rather small number of patients, but this is, in part, compensated by a very detailed genotyping of WNT10A.
References
Thesleff I (2006) The genetic basis of tooth development and dental defects. Am J Med Genet A 140:2530–2535. https://doi.org/10.1002/ajmg.a.31360
Tucker A, Sharpe PT (1999) Molecular genetics of tooth morphogenesis and patterning: the right shape in the right place. J Dent Res 78:826–834. https://doi.org/10.1177/00220345990780040201
Tucker A, Sharpe P (2004) The cutting-edge of mammalian development; how the embryo makes teeth. Nat Rev Genet 5:499–508. https://doi.org/10.1038/nrg1380
Vieira AR, Meira R, Modesto A, Murray JC (2004) MSX1, PAX9, and TGFA contribute to tooth agenesis in humans. J Dent Res 83:723–727. https://doi.org/10.1177/154405910408300913
van der Schalk-Weide Y, Beemer FA (1994) Faber Ja, Bosman F Symptomatology of patients with oligodontia. J Oral Rehabil 21:247–261. https://doi.org/10.1111/j.1365-2842.1994.tb01141.x
Khalaf K, Miskelly J, Voge E, Macfarlane TV (2014) Prevalence of hypodontia and associated factors: a systematic review and meta-analysis. J Orthod 41:299–316. https://doi.org/10.1179/1465313314Y.0000000116
Nieminen P (2009) Genetic basis of tooth agenesis. J Exp Zool B Mol Dev Evol 312B:320–342. https://doi.org/10.1002/jez.b.21277
Polder BJ, Van’t Hof MA, Van Der Linden FPGA, Kuijpers-Jagtman AM (2004) A meta-analysis of the prevalence of dental agenesis of permanent teeth Community. Dent Oral Epidemiol 32:217–226. https://doi.org/10.1111/j.1600-0528.2004.00158.x
Jussila M, Thesleff I (2012) Signaling networks regulating tooth organogenesis and regeneration, and the specification of dental mesenchymal and epithelial cell lineages. Cold Spring Harb Perspect Biol 4:a008425. https://doi.org/10.1101/cshperspect.a008425
Bailleul-Forestier I, Molla M, Verloes A, Berdal A (2008) The genetic basis of inherited anomalies of the teeth. Part 1: clinical and molecular aspects of non-syndromic dental disorders. Eur J Med Genet 51:273–291. https://doi.org/10.1016/j.ejmg.2008.02.009
Vastardis H, Karimbux N, Guthua SW, Seidman JG, Seidman CE (1996) A human MSX1 homeodomain missense mutation causes selective tooth agenesis. Nat Genet 13:417–421. https://doi.org/10.1038/ng0896-417
Šerý O, Bonczek O, Hloušková A, Černochová P, Vaněk J, Míšek I, Krejčí P, Izakovičová Hollá L (2015) A screen of a large Czech cohort of oligodontia patients implicates a novel mutation in the PAX9 gene. Eur J Oral Sci 123:65–71. https://doi.org/10.1111/eos.12170
Bonczek O, Balcar VJ, Šerý O (2017) PAX9 gene mutations and tooth agenesis: A review. Clin Genet 92:467–476. https://doi.org/10.1111/cge.12986
Song S, Han D, Qu H, Gong Y, Wu H, Zhang X, Zhong N, Feng H (2009) EDA gene mutations underlie non-syndromic oligodontia. J Dent Res 88:126–131. https://doi.org/10.1177/0022034508328627
Bergendal B, Klar J, Stecksén-Blicks C, Norderyd J, Dahl N (2011) Isolated oligodontia associated with mutations in EDARADD, AXIN2, MSX1, and PAX9 genes. Am J Med Genet A 155A:1616–1622. https://doi.org/10.1002/ajmg.a.34045
Mostowska A, Biedziak B, Zadurska M, Matuszewska-Trojan S, Jagodziński PP (2015) WNT10A coding variants and maxillary lateral incisor agenesis with associated dental anomalies. Eur J Oral Sci 123:1–8. https://doi.org/10.1111/eos.12165
Peifer M, Polakis P (2000) Wnt signaling in oncogenesis and embryogenesis - a look outside the nucleus. Science 287:1606–1609. https://doi.org/10.1126/science.287.5458.1606
Yuan Q, Zhao M, Tandon B, Maili L, Liu X, Zhang A, Baugh EH, Tran T, Rm S, Hecht JT, Swindell EC, Wagner DS, Letra A (2017) Role of WNT10A in failure of tooth development in humans and zebrafish. Mol Genet Genomic Med 5:730–741. https://doi.org/10.1002/mgg3.332
Miller JR (2002) The Wnts. Genome Biol 3(1):reviews3001.1–reviews3001.15. https://doi.org/10.1186/gb-2001-3-1-reviews3001
Bodine PVN, Komm BS (2006) Wnt signaling and osteoblastogenesis. Rev Endocr Metab Disord 7:33–39. https://doi.org/10.1007/s11154-006-9002-4
Van Amerongen R, Nusse R (2009) Towards an integrated view of Wnt signaling in development. Development 136:3205–3214. https://doi.org/10.1242/dev.033910
Moon RT, Shah K (2002) Developmental biology: signalling polarity. Nature 417:239–240. https://doi.org/10.1038/417239a
Zhang Y, Tomann P, Andl T, Gallant NM, Huelsken J, Jerchow B, Birchmeier W, Paus R, Piccolo S, Mikkola ML, Morrisey EE, Overbeek PA, Scheidereit C, Millar SE, Schmidt-Ullrich R (2009) Reciprocal requirements for EDA/EDAR/NF-kappaB and Wnt/beta-catenin signaling pathways in hair follicle induction. Dev Cell 17:49–61. https://doi.org/10.1016/j.devcel.2009.05.011
Mostowska A, Hozyasz KK, Biedziak B, Wojcicki P, Lianeri M, Jagodzinski PP (2012) Genotype and haplotype analysis of WNT genes in non-syndromic cleft lip with or without cleft palate. Eur J Oral Sci 120:1–8. https://doi.org/10.1111/j.1600-0722.2011.00938.x
Mues G, Bonds J, Xiang L, Vieira AR, Seymen F, Klein O, D’souza RN (2014) The WNT10A gene in ectodermal dysplasias and selective tooth agenesis. Am J Med Genet A 164A:2455–2460. https://doi.org/10.1002/ajmg.a.36520
Liu F, Millar SE (2010) Wnt/beta-catenin signaling in oral tissue development and disease. J Dent Res 89:318–330. https://doi.org/10.1177/0022034510363373
Tamura M, Nemoto E, Sato MM, Nakashima A, Shimauchi H (2010) Role of the Wnt signaling pathway in bone and tooth. Front Biosci (Elite Ed) 2:1405–1413. https://doi.org/10.2741/e201
Cadigan KM, Nusse R (1997) Wnt signaling: a common theme in animal development. Genes Dev 11:3286–3305. https://doi.org/10.1101/gad.11.24.3286
Wodarz A, Nusse R (1998) Mechanisms of Wnt signaling in development. Annu Rev Cell Dev Biol 14:59–88. https://doi.org/10.1146/annurev.cellbio.14.1.59
Bohring A, Stamm T, Spaich C, Haase C, Spree K, Hehr U, Hoffmann M, Ledig S, Sel S, Wieacker P, Röpke A (2009) WNT10A mutations are a frequent cause of a broad spectrum of ectodermal dysplasias with sex-biased manifestation pattern in heterozygotes. Am J Hum Genet 85:97–105. https://doi.org/10.1016/j.ajhg.2009.06.001
Wang J, Shackleford GM (1996) Murine Wnt10a and Wnt10b: cloning and expression in developing limbs, face and skin of embryos and in adults. Oncogene 13:1537–1544
Dassule HR, Mcmahon AP (1998) Analysis of epithelial-mesenchymal interactions in the initial morphogenesis of the mammalian tooth. Dev Biol 202:215–227. https://doi.org/10.1006/dbio.1998.8992
Millar SE, Willert K, Salinas PC, Roelink H, Nusse R, Sussman DJ, Barsh GS (1999) WNT signaling in the control of hair growth and structure. Dev Biol 207:133–149. https://doi.org/10.1006/dbio.1998.9140
Andl T, Reddy ST, Gaddapara T, Millar SE (2002) WNT signals are required for the initiation of hair follicle development. Dev Cell 2:643–653. https://doi.org/10.1016/s1534-5807(02)00167-3
Bailleul-Forestier I, Berdal A, Vinckier F, De Ravel T, Fryns JP, Verloes A (2008) The genetic basis of inherited anomalies of the teeth. Part 2: syndromes with significant dental involvement. Eur J Med Genet 51:383–408. https://doi.org/10.1016/j.ejmg.2008.05.003
Cluzeau C, Hadj-Rabia S, Jambou M, Mansour S, Guigue P, Masmoudi S, Bal E, Chassaing N, Vincent MC, Viot G, Clauss F, Manière MC, Toupenay S, Le Merrer M, Lyonnet S, Cormier-Daire V, Amiel J, Faivre L, de Prost Y, Munnich A, Bonnefont JP, Bodemer C, Smahi A (2011) Only four genes (EDA1, EDAR, EDARADD, and WNT10A) account for 90% of hypohidrotic/anhidrotic ectodermal dysplasia cases. Hum Mutat 32:70–72. https://doi.org/10.1002/humu.21384
Wedgeworth EK, Nagy N, White JML, Pembroke AC, Mcgrath JA (2011) Intra-familial variability of ectodermal defects associated with WNT10A mutations. Acta Derm Venereol 91:346–347. https://doi.org/10.2340/00015555-1028
Van Den Boogaard MJ, Créton M, Bronkhorst Y, Van Der Hout A, Hennekam E, Lindhout D, Cune M, Van Amstel HKP (2012) Mutations in WNT10A are present in more than half of isolated hypodontia cases. J Med Genet 49:327–331. https://doi.org/10.1136/jmedgenet-2012-100750
Mostowska A, Biedziak B, Zadurska M, Dunin-Wilczynska I, Lianeri M, Jagodzinski PP (2013) Nucleotide variants of genes encoding components of the Wnt signalling pathway and the risk of non-syndromic tooth agenesis. Clin Genet 84:429–440. https://doi.org/10.1111/cge.12061
Plaisancié J, Bailleul-Forestier I, Gaston V, Vaysse F, Lacombe D, Holder-Espinasse M, Abramowicz M, Coubes C, Plessis G, Faivre L, Demeer B, Vincent-Delorme C, Dollfus H, Sigaudy S, Guillén-Navarro E, Verloes A, Jonveaux P, Martin-Coignard D, Colin E, Bieth E, Calvas P, Chassaing N (2013) Mutations in WNT10A are frequently involved in oligodontia associated with minor signs of ectodermal dysplasia. Am J Med Genet A 161A:671–678. https://doi.org/10.1002/ajmg.a.35747
Song S, Zhao R, He H, Zhang J, Feng H, Lin L (2014) WNT10A variants are associated with non-syndromic tooth agenesis in the general population. Hum Genet 133:117–124. https://doi.org/10.1007/s00439-013-1360-x
Arzoo PS, Klar J, Bergendal B, Norderyd J, Dahl N (2014) WNT10A mutations account for ¼ of population-based isolated oligodontia and show phenotypic correlations. Am J Med Genet A 164A:353–359. https://doi.org/10.1002/ajmg.a.36243
Clevers H (2006) Wnt/beta-catenin signaling in development and disease. Cell 127:469–480. https://doi.org/10.1016/j.cell.2006.10.018
Li J, Zhang Z, Wang L, Zhang Y (2019) The oncogenic role of Wnt10a in colorectal cancer through activation of canonical Wnt/β-catenin signaling. Oncol Lett 17:3657–3664. https://doi.org/10.3892/ol.2019.10035
Bonczek O, Krejci P, Izakovicova-Holla L, Cernochova P, Kiss I, Vojtesek B (2021) Tooth agenesis: What do we know and is there a connection to cancer? Clin Genet 99:493–502. https://doi.org/10.1111/cge.13892
Jia S, Zhou J, Fanelli C, Wee Y, Bonds J, Schneider P, Mues G, D’Souza RN (2017) Small-molecule Wnt agonists correct cleft palates in Pax9 mutant mice in utero. Development 144:3819–3828. https://doi.org/10.1242/dev.157750
Yu M, Wong SW, Han D, Cai T (2019) Genetic analysis: Wnt and other pathways in nonsyndromic tooth agenesis. Oral Dis 25:646–651. https://doi.org/10.1111/odi.12931
Roche: Sequencing Solutions Technical Note: How To Evaluate NimbleGen SeqCap EZ Target Enrichment Data Roche Diagnostics: Mannheim. http://netdocs.roche.com/DDM/Effective/07187009001_RNG_SeqCap-EZ_TchNote_Eval-data_v2.1.pdf Accessed 2 October 2020
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–17660. https://doi.org/10.1093/bioinformatics/btp324
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
R. CoreTeam (2015) R:A language and environment for statistical computing. Available: https://www.R-project.org/. Accessed 28 September 2020
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26. https://doi.org/10.1038/nbt.1754
Thorvaldsdóttir H, Robinson JT, Mesirov JP (2013) Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14:178–192. https://doi.org/10.1093/bib/bbs017
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249. https://doi.org/10.1038/nmeth0410-248
Arte S, Parmanen S, Pirinen S, Alaluusua S, Nieminen P (2013) Candidate gene analysis of tooth agenesis identifies novel mutations in six genes and suggests significant role for WNT and EDA signaling and allele combinations. PLoS One 8:e73705. https://doi.org/10.1371/journal.pone.0073705
Kantaputra P, Sripathomsawat W (2011) WNT10A and isolated hypodontia. Am J Med Genet A 155A:1119–1122. https://doi.org/10.1002/ajmg.a.33840
Zhang J, Tian XJ, Xing J (2016) Signal Transduction Pathways of EMT Induced by TGF-β, SHH, and WNT and Their Crosstalks. J Clin Med 5:41. https://doi.org/10.3390/jcm5040041
Heise RL, Stober V, Cheluvaraju C, Hollingsworth JW, Garantziotis S (2011) Mechanical stretch induces epithelial-mesenchymal transition in alveolar epithelia via hyaluronan activation of innate immunity. J Biol Chem 286:17435–17444. https://doi.org/10.1074/jbc.M110.137273
Hlouskova A, Bielik P, Bonczek O, Balcar VJ, Šerý O (2017) Mutations in AXIN2 gene as a risk factor for tooth agenesis and cancer: A review. Neuro Endocrinol Lett 38:131–137
He H, Han D, Feng H, Qu H, Song S, Bai B, Zhang Z (2013) Involvement of and interaction between WNT10A and EDA mutations in tooth agenesis cases in the Chinese population. PLoS One 8:e80393. https://doi.org/10.1371/journal.pone.0080393
Suomalainen M, Thesleff I (2010) Patterns of Wnt pathway activity in the mouse incisor indicate absence of Wnt/beta-catenin signaling in the epithelial stem cells. Dev Dyn 239:364–372. https://doi.org/10.1002/dvdy.22106
Janda CY, Waghray D, Levin AM, Thomas C, Garcia KC (2012) Structural basis of Wnt recognition by Frizzled. Science 337:59–64. https://doi.org/10.1126/science.1222879
Nawaz S, Klar J, Wajid M, Aslam M, Tariq M, Schuster J, Baig SM, Dahl N (2009) WNT10A missense mutation associated with a complete odonto-onycho-dermal dysplasia syndrome. Eur J Hum Genet 17:1600–1605. https://doi.org/10.1038/ejhg.2009.81
Tardieu C, Jung S, Niederreither K, Prasad M, Hadj-Rabia S, Philip N, Mallet A, Consolino E, Sfeir E, Noueiri B, Chassaing N, Dollfus H, Manière MC, Bloch-Zupan A, Clauss F (2017) Dental and extra-oral clinical features in 41 patients with WNT10A gene mutations: A multicentric genotype-phenotype study. Clin Genet 92:477–486. https://doi.org/10.1111/cge.12972
Acknowledgements
We would like to thank all the subjects who participated in the research. The authors thank Dr. Philip J. Coates (RECAMO, MMCI, Brno, Czech Republic) for English language editing.
Funding
This work was funded by grants of the IGA MH CZ n.: NT/11420–6/2010, the European Regional Development Fund-Project ENOCH (No. CZ.02.1.01/0.0/0.0/16_019/0000868), MH CZ-DRO (MMCI, 00209805) and AZV CR NU20-06–00189.
Author information
Authors and Affiliations
Contributions
All authors made substantial contribution to the conception and design of the manuscript. P.B. and O.B. drafted manuscript, T.Z. analyzed data from NGS sequencing, P.B. and J.L. performed NGS sequencing, P.K., J.S. and J.V. performed sampling and clinical investigations, J.V., L.I.H. and O.S. conceptualized research and obtained fundings, B.V. and V.J.B. edited and supervised manuscript, and O.S. supervised laboratory analyses and final manuscript. All authors agree to be accountable for all aspects of the study design and its content. All authors approved the final submitted version.
Corresponding author
Ethics declarations
Ethical approval
The study protocol and informed consents were reviewed and approved by the ethical committee of Faculty of Medicine and Dentistry, Olomouc and Faculty of Medicine, Brno. The study was performed in accordance with the Declaration of Helsinki. Care was taken to follow the letter and spirit of the Declaration of Helsinki—ethical principles for medical research involving human subjects.
Informed consent
Written informed consent (No: Fm-L009-001-ZUBNI-014) was obtained from all participating human subjects.
Conflict of interest
All the authors declare that they have no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bielik, P., Bonczek, O., Krejčí, P. et al. WNT10A variants: following the pattern of inheritance in tooth agenesis and self-reported family history of cancer. Clin Oral Invest 26, 7045–7055 (2022). https://doi.org/10.1007/s00784-022-04664-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00784-022-04664-x