Abstract
Hydroxylamine oxidoreductase (HAO) from the ammonia-oxidizing bacterium Nitrosomonas europaea normally catalyzes the four-electron oxidation of hydroxylamine to nitrite, which is the second step in ammonia-dependent respiration. Here we show that, in the presence of methyl viologen monocation radical (MVred), HAO can catalyze the reduction of nitric oxide to ammonia. The process is analogous to that catalyzed by cytochrome c nitrite reductase, an enzyme found in some bacteria that use nitrite as a terminal electron acceptor during anaerobic respiration. The availability of a reduction pathway to ammonia is an important factor to consider when designing in vitro studies of HAO, and may also have some physiological relevance. The reduction of nitric oxide to ammonia proceeds in two kinetically distinct steps: nitric oxide is first reduced to hydroxylamine, and then hydroxylamine is reduced to ammonia at a tenfold slower rate. The second step was investigated independently in solutions initially containing hydroxylamine, MVred, and HAO. Both steps show first-order dependence on nitric oxide and HAO concentrations, and zero-order dependence on MVred concentration. The rate constants governing each reduction step were found to have values of (4.7 ± 0.3) × 105 and (2.06 ± 0.04) × 104 M−1 s−1, respectively. A second reduction pathway, with second-order dependence on nitric oxide, may become available as the concentration of nitric oxide is increased. Such a pathway might lead to production of nitrous oxide. We estimate a maximum value of (1.5 ± 0.05) × 1010 M−2 s−1 for the rate constant of the alternative pathway, which is small and suggests that the pathway is not physiologically important.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Living organisms can extract energy from an astonishing range of oxidation–reduction reactions, and in the process they change their surroundings significantly. In particular prokaryotic respiratory processes play a critical role in shaping the chemical environment of the biosphere, affecting everything from the levels of greenhouse gases and composition of soils to the speciation of toxic metals such as arsenic and mercury [1, 2]. Nitrosomonas europaea is one of a group of bacteria that gain energy primarily by aerobically oxidizing ammonia, and are thus referred to as “ammonia-oxidizing bacteria” (AOB) [3]. These bacteria form an integral part of the biological nitrogen cycle, and thus have great economic and ecological importance [4–11]. In recent years it has become apparent that AOB such as N. europaea, once thought to be obligate aerobes, can in fact respire aerobically or anaerobically. The fate of the nitrogen originating from ammonia will thus depend critically on the specific growth conditions [3, 12]. How the transition from aerobic to anaerobic growth is managed poses an interesting question, because of the potential for toxic intermediates NH2OH and NO· to accumulate as the O2 supply diminishes. Our research group has a longstanding interest in hydroxylamine oxidoreductase (HAO), a Nitrosomonas enzyme that catalyzes the four-electron oxidation of NH2OH to nitrite under aerobic conditions [13–15]. This paper elucidates the pathways by which HAO reacts with NO· and NH2OH under reducing conditions, and discusses the implications of such pathways for AOB metabolism, as organisms switch from aerobic to anaerobic growth.
HAO catalyzes the second of two steps by which NH4 + is oxidized to NO2 − by O2 in N. europaea; the first step, oxidation of NH4 + to NH2OH, is catalyzed by the enzyme ammonia mono-oxygenase [6, 15]. HAO is a highly complex enzyme, consisting of three identical subunits that contain eight hemes each (Fig. 1) [16]. NH2OH oxidation by HAO takes place on three novel hemes known as P460s, which have vacant coordination sites at which NH2OH can bind (Fig. 1) [16–20]. The remaining 21 HAO hemes are of c-type and six-coordinate, with two axial His ligands each [16, 21]. These hemes are closely spaced, and hence can act as electron transfer agents, accepting the electrons that P460 abstracts from NH2OH [16, 21]. Moreover, heme 8 from one subunit lies near heme 2 of the adjacent subunit, so electron transfer between subunits should be facile. Indeed, the 18-heme circle seen in the HAO trimer (Fig. 1b) appears to be designed precisely to allow electrons entering at a single P460 to be rapidly distributed throughout the trimer. Heme 1 (Fig. 1) is the only heme apart from P460 that is solvent-exposed, and appears to be the in vivo exit point for the electrons extracted from NH2OH oxidation. The electrons go from heme 1 to cytochrome c-554, a soluble 26.1-kDa tetraheme protein [22, 23] that is quite abundant in the periplasm of N. europaea [24], and is now generally recognized to be the acceptor of the electrons from HAO in vivo.
a Arrangement of the eight hemes within one subunit of hydroxylamine oxidoreductase (HAO). b Arrangement of all the hemes within the HAO trimer; primes are used to distinguish subunits. The hemes are labeled according to the format used in [16], except for P460, which was labeled heme 4 in the original structure article
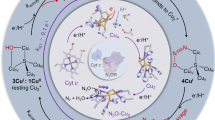
Figure 2 presents a possible mechanism for NH2OH oxidation at HAO that is based on extensive literature precedent [25–32], and provides a useful working hypothesis with which to proceed. The figure also introduces a slight modification of the convenient Enemark–Feltham notation, {P460(NO)}n, to describe intermediates containing Fe–NO fragments [33]. On the basis of this mechanism, NH2OH binds to P460 and is oxidized by two electrons to give a {P460(NO)}7 moiety at the active site. The electrons end up on hemes 3 and 8, which have midpoint potentials of approximately 0 mV and can temporarily store electrons [34, 35]. No other heme on HAO has a midpoint potential suitable for accepting electrons from {P460(NO)}7; however, once electrons have been transferred from hemes 3 and 8 to the physiological acceptor cytochrome c-554, then {P460(NO)}7 can be further oxidized to {P460(NO)}6, which can either lose NO· or undergo reductive nitrosylation to give NO2 −. Either option returns the active site to its resting state.
Putative changes at the P460 active site during catalytic NH2OH oxidation to NO2 −. The electrons leaving the active site will be passed on to the HAO c-hemes (see Fig. 1). The dashed arrows show the reverse pathway that might be expected under reducing conditions, as well as the pathway for generation of the side-product N2O (see text). Active-site Fe–NO fragments are each represented in terms of two resonance structures, and of the corresponding Enemark–Feltham description {Fe(NO)}n. In this notation the superscript n is the sum of the d electrons that would be counted on Fe if the ligand were actually NO· and the π* electron from the NO· [33]. Note that “P460” is used in place of “Fe” in the notation, to emphasize that it is the P460 heme that is nitrosylated
Figure 2 outlines, by way of dashed arrows, the possibility that under appropriate conditions the catalytic cycle could run in reverse, leading to NH4 + and N2O formation. To understand why this possibility must be considered we turn to Fig. 3, which is a Frost diagram for nitrogen at pH 7, showing the energies of NH4 +, NH2OH, and NO2 −, in relation to those of other species with intermediate oxidation states [36, 37]. Several features stand out in this diagram which have implications for understanding Nitrosomonas behavior as it switches from aerobic to anaerobic metabolism. First, both N2 and N2O sit well below NH2OH, NO2 −, and NO· in energy. Thus, conversion of NH2OH, as well as NO2 − and NO·, to the low-energy species N2O and N2 is one obvious pathway by which potentially toxic products could be removed if these accumulated owing to a sudden shortage of O2. Indeed, it is known that N2O and traces of N2 are released both in vivo and in vitro during NH2OH oxidation by HAO [24, 38], and Fig. 2 shows a pathway by which N2O could be generated. That being said, Fig. 3 shows that NH4 + also sits well below NH2OH, NO2 −, and NO·, so reversing the physiological reaction is a second pathway by which potentially toxic products could be removed if they accumulated to unacceptable levels. Furthermore, NH2OH is unstable with respect to disproportionation into NH4 + and NO2 − or NO·. The architecture of HAO (Fig. 1) [16], which allows facile electron transfer between active sites on different subunits of the trimeric enzyme, could in principle make HAO an ideal disproportionation catalyst. To our knowledge, neither of the last two possibilities has been previously addressed in the literature. Herein we report on the HAO-catalyzed reduction of NO· and NH2OH by the methyl viologen monocation radical (MVred), and demonstrate that HAO can reduce NO· to NH4 +, in competition with reduction to N2O.
Materials and methods
Materials
HAO was purified as described in [39], and the photoactive NO·-releasing species N,N′-bis(carboxymethyl)-N,N′-dinitroso-p-phenylenediamine (species 1, Scheme 1) was synthesized according to the procedure described in [40]. Methyl viologen dichloride was obtained from Acros, sodium dithionite from Aldrich, and hydroxylamine hydrochloride from Fisher. Myoglobin (crystallized and lyophilized horse skeletal muscle) was obtained from Sigma in the fully oxidized ferric form, known as metmyoglobin. For experiments requiring ferromyoglobin (Mb), the ferric protein was reduced by titrating it with exactly 0.5 equiv of sodium dithionite. In experiments involving methyl viologen, the total concentration of this species was kept constant at 300 μM. The ratio of one-electron-reduced MVred to fully oxidized methyl viologen dication (MVox) was then adjusted by adding an appropriate amount of sodium dithionite. The concentration of MVred was typically 25% or less than the total methyl viologen concentration. All experiments were performed in solutions buffered with phosphate (μ = 50 mM, pH 7.4), and all solutions were prepared and manipulated in a nitrogen-filled glovebox. Stock solutions were prepared daily, and stored in a fridge at 4 °C until needed. Hydroxylamine hydrochloride stock solutions were prepared in nanopure water; all other stock solutions were made up in phosphate buffer (μ = 50 mM, pH 7.4).
Simplified reaction pathway for generation of NO· from N,N′-bis(carboxymethyl)-N,N′-dinitroso-p-phenylenediamine (1), in the presence of reduced methyl viologen. A more detailed analysis of this reaction can be found in [43]
Data collection and instrumentation
Routine UV/vis spectra were obtained using a Cary 50 spectrophotometer (Varian) that was installed in the glovebox. This spectrometer was also used for collecting the kinetic data in the HAO-catalyzed reduction of hydroxylamine by MVred. Nitric oxide was photogenerated in situ using a previously described method [40–43], and the process is summarized in Scheme 1. Briefly, photochemical fragmentation of species 1 into species 2 was initiated with a 10-ns, 308-nm, 6.0-mJ pulse from a XeCl excimer laser (TUI, Existar 200). In the presence of MVred, 2 was subsequently reduced to stable species 3, trapping an equivalent of NO· for future reactions (Scheme 1, step 2). An Olis RSM-1000 spectrophotometer was used to monitor the absorbance changes induced by the laser pulse. The configuration of the laser and spectrophotometric equipment has been described in general terms elsewhere [40, 43]. All data were collected with the Olis RSM-1000 spectrophotometer in rapid-scanning mode (monochromator entrance slit width 0.6 mm, scanning slit width 0.2 mm, exit slit width 0.12 mm), which allows complete spectra to be obtained in 1 ms. For the laser experiments, 150 μL test solutions were held in 3 mm × 3 mm fluorescence cuvettes. With this configuration, the laser pulse uniformly irradiated the full width of the test solution. This in turn allowed subsequent photochemically induced reactions to be monitored for tens of seconds, without needing to worry about diffusion-induced concentration changes within the region of the cuvette being monitored [43].
Data analysis
Data were analyzed using the commercially available software packages Microcal Origin Version 6.0 (Microcal Software), and Mathcad 13 (Mathsoft Engineering and Education). Complete spectra obtained using the Olis RSM-1000 spectrophotometer or the Cary 50 spectrophotometer were first subjected to singular value decomposition (SVD) to determine the number of colored species, and to decrease the noise in the absorbance matrices [44]. A matrix form of Beer’s law (Eq. 1) was then used to calculate the concentrations of all species in solution as a function of time [44, 45]:
In Eq. 1 A is the SVD-processed absorbance matrix, in which each row corresponds to a spectrum, and each column to a time trace at a fixed wavelength; ε is the matrix of extinction coefficients, in which each row corresponds to a unique species, and each column to a wavelength; C is the matrix of concentrations, in which each column corresponds to a unique species, and each row to a specific time; and l is a scalar representing the pathlength. In the laser experiments, a value of l = 2.55 mm was used in conjunction with 3 mm × 3 mm cuvettes, as described in [43]. The extinction coefficient difference spectra required in Eq. 1, as well as absolute extinction coefficient spectra used to calculate species concentrations in the cuvettes prior to laser photoexcitation, were obtained as follows. The extinction coefficient spectra for Mb and nitrosomyoglobin (MbNO) were obtained as described previously [43]. For MVox, solutions containing 0–100 μM known concentrations were prepared from the solid reagent, using standard gravimetric and volumetric techniques, and spectra were collected at the various concentrations. The ε vector for MVox was then obtained from a rearranged form of Eq. 1. A solution of MVox with known concentration was then titrated with dithionite until it had been fully reduced, at which point the ε vector for MVred could be obtained, after correcting for dilution by added reductant. The extinction coefficient for the NO·-generating species 1 is 13,500 M−1 cm−1 at 300 nm [41], and this value was used to obtain the concentration of a solution of 1, which in turn was used to calculate the complete ε value spectrum for species 1. An identical procedure was used to obtain the full ε value spectrum of oxidized HAO, which has a known extinction coefficient value of 7 × 105 M−1 cm−1 at 408 nm [19]. A solution of HAO was then titrated with titanium(III) citrate [46] to generate spectra at various stages of reduction. After correcting for dilution by added reductant, ε value spectra were generated for all the stages of reduction, and stored in an electronic “lookup table.” A Mathcad routine could then be used to determine the HAO concentration from the spectrum of any solution containing the enzyme, regardless of the enzyme’s oxidation state. An example of the procedure is given in the supplementary material.
Results
Figure 4a shows the spectral changes observed after a solution initially containing 26 μM 1, 65 μM MVred, and 0.8 μM HAO was irradiated with a 308-nm, 10-ns laser pulse. Analysis of the difference spectra by SVD showed that there was only one spectral component, which could be attributed to oxidation of MVred to MVox. The dashed red lines in Fig. 4a represent the least-squares fits to the data, using the independently obtained \( {\mathbf{\varepsilon }}_{{{\text{MV}}_{{{\text{ox}}}} }} - {\mathbf{\varepsilon }}_{{{\text{MV}}_{{{\text{red}}}} }} ({\mathbf{\Updelta \varepsilon }}_{{{\text{MV}}}} ) \) difference spectrum. Figure 4b shows how the concentration of MVox varies as a function of time. The y-intercept in Fig. 4b is not zero, indicating that some MVred was oxidized on the microsecond timescale, within the dead time of the current experiment. Figure 5 shows what happened when the experiment represented by the plots in Fig. 4 was repeated in the presence of 19 μM Mb. This protein is well known to bind NO· rapidly and irreversibly, effectively taking it out of circulation [25, 43, 47]. Here SVD revealed one major and one very minor spectral component. Subsequent least-squares fitting of the spectra with the appropriate Δε components showed that in this case two processes had occurred: oxidation of MVred, and nitrosylation of Mb (Fig. 5a). The concentration versus time trace (Fig. 5b) shows that the initial concentration of MVox is approximately the same as that seen in the experiment represented by the plots in Fig. 4, but thereafter remains virtually unchanged (the very slight increase in MVox of about 6 nM s−1 was not readily reproducible, and is of unknown origin). The MbNO is generated in the first few milliseconds after the laser pulse, and also changes little thereafter.
a Spectral changes (blue lines) observed after a solution initially containing 26 μM 1, 65 μM methyl viologen monocation radical (MVred), and 0.8 μM HAO, was irradiated with a 308-nm, 10-ns laser pulse. The representative difference spectra shown here were collected 0.016, 1.6, 4.8, 8.0, 48, and 88 s after the laser pulse. Theoretical traces (dashed red lines) were calculated from the known extinction coefficients of MVred and methyl viologen dication (MVox), as described in “Materials and methods.” b MVox concentration versus time trace corresponding to the data shown in a
a Spectral change (blue line) observed 90 s after a solution initially containing 26 μM 1, 19 μM ferromyoglobin (Mb), 1.1 μM HAO, and 42 μM MVred was irradiated with a 308-nm, 10-ns laser pulse. The theoretical trace (dashed red line) was calculated from the known extinction coefficients of MVred, MVox, Mb, MbNO, as described in “Materials and methods.” b MVox concentration versus time trace corresponding to the data shown in a
The fast (submillisecond) oxidation phase observed both in the presence and in the absence of Mb is attributed to the reaction of MVred with 2, the highly reactive denitrosated product of photolysis of 1 (Scheme 1) [42, 43]. This phase was also observed in experiments where mixtures of 1 and MVred were irradiated in the presence and absence of Mb, but with no added HAO. The subsequent slow oxidation seen in Fig. 4 must be effected by NO·, since removal of the NO· by reaction with Mb eliminates it. Some slow oxidation was also observed when mixtures of 1 and MVred were irradiated in the absence of both Mb and HAO, but the reaction profiles were substantively different from those observed in the presence of HAO (see below and the supplementary material).
As Mb binds NO· quantitatively, the concentration of MbNO present at a given time in Fig. 5b is a direct measure of the amount of NO· generated up until that time [43]. Thus, Fig. 5b shows that over the timescale of the experiment, 0.85 equiv of NO· was generated for every equivalent of MVox. Further experiments, performed with different initial concentrations of MVred and 1, both in the presence and in the absence of HAO, showed that on average 0.85 ± 0.02 equiv of NO· was generated for every equivalent of MVred oxidized by 2 (see the supplementary material). Knowing this, we were able to determine the amount of photogenerated NO· from the initial concentration of MVox, even in the absence of Mb. Thus, in the experiment represented by the plots in Fig. 4, the concentration of NO· immediately after the laser pulse was estimated to be 5.9 μM. The data in Fig. 4b were empirically fitted to two exponentials (Fig. 4b, red trace), in order to obtain an estimate of the final MVox concentration. From this fit, the final MVox concentration was estimated to be 38.2 μM, of which 31 μM was oxidized by NO·; thus, each NO· was able to oxidize about 5 equiv of MVred. It should be noted that in the absence of HAO, only 1–2 equiv of MVred was oxidized per equivalent of NO· generated, showing that the reaction pathways are different in the absence of enzyme (see the supplementary material).
To act as a five-electron oxidant as indicated in Fig. 4, NO· must be being reduced all the way to ammonia (Scheme 2), demonstrating that HAO can behave like the enzyme cytochrome c nitrite reductase [31, 48–50], at least when exposed to a potent reductant such as MVred. The reaction proceeds in two kinetically distinct phases, the first of which consumes about 3 equiv of MVred, and the second 2 equiv (Fig. 4b). This is consistent with a process in which NO· is converted to NH2OH in the first fast phase, and the NH2OH is then reduced to NH4 + in the second, 10 times slower phase (Scheme 2). The second phase was studied independently by reacting NH2OH with MVred in the presence of HAO (see below), and for this phase the production of ammonia was verified independently (see the supplementary material).
An analysis of the initial rate of MVox formation \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right) \) as a function of HAO, initial MVred, and initial NO· concentrations revealed the rate law shown in Eq. 2:
The complete analysis, which was complicated by technical difficulties encountered in trying to keep two out of three concentrations constant within each given series of experiments, is described in the supplementary material. Here, we present a brief summary of the process. Figure 6 shows a plot of \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right)/[{\text{NO}} \cdot ] \) as a function of HAO concentration, in experiments where the initial concentrations of NO· and MVred were kept approximately constant. Dividing \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) by the NO· concentration virtually eliminated systematic errors arising from minor variations in NO· concentration from one data set to the next. The procedure did not eliminate the contribution from the factor k 2app[NO·] (Eq. 2), but this proves to be exceedingly small (see below). The plot in Fig. 6 is linear, with a nonzero y-intercept k uncat[MVred]0 that arises because NO· reacts with MVred even in the absence of HAO. We estimate the uncertainty in HAO concentration to be ±2 × 10−8 M for each data point (horizontal error bars), and believe this to be the major source of scatter in the data in Fig. 6. It was also possible to estimate the systematic errors propagated into \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right)/[{\text{NO}} \cdot ] \) by the known variations in the initial MVred concentrations (see the supplementary material), but these proved to be very small compared to the uncertainties produced by random variations in HAO concentration. The systematic contributions from MVred concentrations are depicted by vertical half error bars in Fig. 6.
Variation in \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} /[{\text{NO}} \cdot ]_{0} \) as a function of HAO concentration, at roughly constant initial NO· and MVred concentrations (3.6 ± 0.4 and 19 ± 4 μM, respectively), where \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \equiv {\text{d}}[{\text{MV}}_{{{\text{ox}}}} ]/{\text{d}}t|_{{t = 0}} . \) Dividing \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) by the initial NO· concentration virtually eliminates the systematic errors introduced by variations in the initial NO· concentration from one run to the next. The horizontal error bars (±2 × 10−8 M) reflect the estimated uncertainty in the HAO concentration for each data point, which we believe is the major source of scatter in the data. Barely visible are vertical half error bars that reflect the calculated error propagation into \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) arising from known variations in the initial MVred concentration (the error propagation calculations are given in the supplementary material). Least-squares fitting of the data to a straight line yielded a slope of (1.5 ± 0.1) × 106 M−1 s−1, and an intercept of 0.14 ± 0.07 s−1. On the basis of the rate law of Eq. 2, the slope represents the value of 3k 1 + k 2app[NO·]0, and the intercept k uncat[MVred]0
In experiments carried out at roughly constant initial NO· and HAO concentrations, \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right)/[{\text{NO}} \cdot ] \) was found to vary linearly with initial MVred concentration (Fig. 7). Once again the line has nonzero intercept, which in this case shows that consumption of electrons proceeds even as the limit of no electron donor is approached. Thus, the HAO-catalyzed reduction of NO· must have a zero-order dependence on MVred. The slope of the line in Fig. 7 reflects the increase in \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) due to the uncatalyzed reaction, and provides a direct measure of k uncat (Eq. 2). The half error bars in Fig. 7 reflect the systematic error propagated into \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right)/[{\text{NO}} \cdot ] \) by the known variations in the HAO concentration from one data point to the next (see the supplementary material). The dashed line in Fig. 7 is the least-squares fit to the data after the systematic error has been corrected (the corrected data reside at the tips of the half error bars). The dashed line comes substantially closer to the corrected data than the solid line does to the uncorrected data, showing that the variations in HAO concentration account for most of the apparent scatter in the original data. From the slope of the dashed line in Fig. 7, the value of k uncat was estimated to be (3.64 ± 0.08) × 103 M−1 s−1, which is consistent with the less precise value, (7 ± 4) × 103 M−1 s−1, obtained from the intercept of the line in Fig. 6. The value of k uncat was also obtained independently by analyzing the dependence of \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) on the concentrations of MVred and NO· in the absence of HAO (see the supplementary material).
Variation in \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} /[{\text{NO}} \cdot ]_{0} \) as a function of initial MVred concentration, at roughly constant initial NO· concentration and HAO concentration (3.1 ± 0.3 and 0.37 ± 0.02 μM, respectively). The half error bars reflect the systematic error propagated into \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} } \right)/[{\text{NO}} \cdot ] \) by the known variations in HAO concentration (the error propagation calculations are given in the supplementary material). The dashed line is the least-squares fit to the data after the systematic error has been corrected (the corrected data reside at the tips of the half error bars) and has a slope of (3.64 ± 0.08) × 103 M−1 s−1, and an intercept of 0.438 ± 0.003 s−1. On the basis of the rate law of Eq. 2, the slope represents the value of k uncat, and the intercept (3k 1 + k 2app[NO·])[HAO]
Using the calculated value of k uncat and the known initial concentrations of MVred and NO·, one can determine and then subtract out the uncatalyzed contribution to \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) for any given experiment. This tactic proved useful for analyzing the dependence of \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) on NO· concentrations, because in such experiments it was especially difficult to keep MVred truly constant. Figure 8 shows the results of experiments in which the NO· concentration was varied while keeping the HAO and MVred concentrations as constant as possible. A direct plot of \( V_{{{\text{MV}}_{{{\text{ox0}}}} }} \) versus NO· concentration showed a slight but distinct deviation from linearity (see the supplementary material). Such a deviation is less apparent in the corrected data of Fig. 8, but cannot be completely ruled out. Thus, in Fig. 8, the solid trace is a linear fit to the data, whereas the dashed trace is a fit to the parabolic equation y = ax + bx 2. On the basis of the mechanism of Eq. 2, the parameter a from the parabolic fit would represent 3k 1, while the parameter b would represent k 2app.
Variation in \( \left( {V_{{{\text{MV}}_{{{\text{ox0}}}} }} - k_{{{\text{uncat}}}} [{\text{MV}}_{{{\text{red}}}} ]_{0} [{\text{NO}} \cdot ]_{0} } \right)/[{\text{HAO}}] \) as a function of initial NO· concentration. The linear fit (solid line) has a slope of (1.90 ± 0.06) × 106 M−1 s−1, and intercept of −1.2 ± 0.5 s−1. The dashed curve is a fit to the equation y = ax + bx 2, where a = (1.4 ± 0.1) × 106 M−1 s−1, and b = (3 ± 1) × 1010 M−2 s−1
Assuming for the moment that there is a real curvature in Fig. 8, then k 2app could be governing the production of N2O, which is known to be released by N. europaea during respiration [24]. Such a process would require 2 equiv of NO· to coincide at the HAO active site (though probably not simultaneously), which might lead to a second-order dependence on NO· concentration. On the basis of the stoichiometry for the net reaction of NO· with MVred to produce N2O (Scheme 3), a true rate constant k 2 governing the process would be equal to 0.5k 2app. Continuing with the assumption that the curvature in Fig. 8 is real, and adding the additional assumption that the curvature reflects production of N2O at higher NO· concentrations, one can calculate values of (4.7 ± 0.3) × 105 M−1 s−1 for k 1, and (1.5 ± 0.5) × 1010 M−2 s−1 for k 2, from the values of the Fig. 8 parameters a and b, respectively. Figure 9 then shows the predicted relative rates for the production of NH2OH and N2O as a function of NO·. One can see from this figure that even if the second-order dependence on NO· is real, the relative rate of N2O formation should stay well below that of NH2OH formation at NO· concentrations below 20 μM.
The reduction of NH2OH to NH4 +, the putative second step in the reduction of NO· to NH4 +, was studied independently using conventional UV/vis spectral methods. Also, for this step, the production of ammonia was verified independently using a commercially available enzymatic ammonia assay. The ammonia assay is fully described in the supplementary material; here we discuss the UV/vis spectral analysis. As an example, Fig. 10 shows how the concentration MVox increases with time in a solution initially containing 0.22 μM HAO, 38.6 μM MVred, and roughly 4.5 μM NH2OH. The data are well modeled using a single exponential (Fig. 10, red trace), which has an amplitude equal to 2 times the initial NH2OH concentration. Repeating the experiment of Fig. 10 under different HAO concentrations, while keeping all other parameters constant, yielded exponential plots in every case. A plot of the apparent first-order rate constant (k 3app) versus the HAO concentration is linear, with a zero intercept (Fig. 11a). The rate dependence on concentrations of NH2OH and MVred was analyzed using the method of initial rates. The initial rates were found to vary linearly with NH2OH out to 100 μM substrate concentration (Fig. 11b), and to be independent of MVred concentration (data not shown). In the absence of HAO, the rate of MVred oxidation was negligible, even with initial NH2OH concentrations of 100 μM and initial MVred concentrations of approximately 50 μM. The HAO-dependent and NH2OH-dependent data yield values of k 3 within 2% of each other, with the average value being k 3 = (2.06 ± 0.04) × 104 M−1 s−1.
Variation in MVox concentration as a function of time, in a solution initially containing 0.22 μM HAO, 38.6 μM MVred, and roughly 4.5 μM NH2OH. The red curve is a fit to the function y = A 0 + amp[1 − exp(−k 3obs)t], where the rate constant is labeled in accordance with Scheme 2
a Dependence of k 3obs on HAO concentration. The initial MVred concentration was 37 ± 3 μM, and the initial NH2OH concentration was 4.9 ± 0.4 μM. The value of k 3obs was obtained as described in the legend to Fig. 8. b Dependence of \( V_{{{\text{MV}}_{{{\text{red0}}}} }} /[{\text{HAO}}] \) on initial NH2OH concentration. The initial MVred concentration was 46 ± 3 μM, and the HAO concentration was (9 ± 3) × 10−8 M. From these data, the value of k 3 is determined to be (2.06 ± 0.04) × 104 M−1 s−1, where k 3 is labeled in accordance with Scheme 2
Discussion
The results presented herein demonstrate that HAO can catalyze the reduction of NO· and NH2OH to NH4 +. This is an important factor to consider when designing in vitro studies of HAO, and it may also have some physiological relevance. For example, HAO could play a role in limiting the accumulation of NO· and NH2OH if the O2 concentration transiently diminishes in the cells. It has long been known that HAO can generate NO2 − only aerobically, and that in the absence of O2 free NO· is liberated instead [51]. Thus, if the O2 supply were to drop within a cell, NO· and perhaps NH2OH would accumulate. In the presence of a suitable electron donor though, both NO· and NH2OH would then be converted back to NH4 + by HAO. This would be energetically wasteful, but would also slow down the accumulation of potentially toxic NO· and NH2OH. It is not difficult to suggest a suitable redox partner for HAO-catalyzed reduction of NO· to NH4 +. The physiological electron acceptor for HAO, cytochrome c-554 [22, 52–55], could suffice. Cytochrome c-554 has two physiologically relevant electron-accepting hemes with midpoint potentials of 47 mV [56, 57]. Combining this value with the reduction potential for the NO·/NH4 + couple (339 mV, based on Fig. 3), one calculates a favorable cell potential (292 mV) for the reduction of NO· to NH4 + by reduced cytochrome c-554. Even in the absence of an extra electron donor such as cytochrome c-554, NH2OH accumulation would be curtailed by disproportionation, as shown in Scheme 4. Hydroxylamine disproportionates slowly in alkaline solutions, and the process can be accelerated by transition metal complexes such as [Fe(CN)5H2O]3− [58]. The architecture of HAO (Fig. 1) [16] allows facile electron transfer between active sites on different subunits of the trimeric enzyme, which could in principle make HAO an ideal disproportionation catalyst: the electrons needed to reduce NH2OH to NH4 + at one active site would be provided by oxidation of NH2OH to NO· at another active site. Disproportionation would not prevent the accumulation of excess NO·.
HAO-catalyzed reduction of NH2OH and NO· is less likely to be physiologically relevant if the O2 concentration drops for a prolonged period. There is now evidence that, under such conditions, N. europaea switches to anaerobic respiration mode, for which it has specialized proteins. Indeed, it appears that the switch from aerobic to anaerobic respiration is triggered by both low O2 concentration and high NO· concentration [3, 24]. Schmidt et al. [3] recently demonstrated that at concentrations greater than 30 ppm, NO· can act as a signal within N. europaea, which causes the bacteria to cease motile and motile-planktonic growth, and instead form biofilms. At the same time, expression of the protein nitrosocyanin, which may be involved in electron transfer from HAO to nitric oxide reductase, increases significantly [3]. Expression of nitric oxide reductase, which would catalyze the reduction of NO· to N2O, does not appear to be affected by NO· concentration [3]; however, the gene for this protein is present in HAO, and may be turned on at low O2 concentrations [24].
Our results show that HAO-catalyzed reduction of NO· to N2O, if it occurs at all, is negligible at micromolar concentrations. Above approximately 30 ppm, where HAO might begin to contribute significantly to N2O production, the specialized proteins of anaerobic respiration would be turned on as described above. Thus, HAO-catalyzed reduction of NO· to N2O is not expected to play a significant physiological role in Nitrosomonas. This is the same conclusion reached by previous researchers [24]. Our results do not rule out in vitro reduction of NO· to N2O by HAO in experiments where the NO· concentration is very high; such reduction was proposed to explain the rapid oxidation of fully reduced HAO when exposed to 1 atm of NO· [59].
An interesting feature of the HAO-catalyzed reduction of NH2OH by MVred is its linear dependence on NH2OH concentration. The HAO-catalyzed oxidation of NH2OH by mammalian cytochrome c exhibits hyperbolic dependence on NH2OH concentration, for which a K m value of 3.6 μM has been reported [13]. It is not immediately clear why there should be such a dramatic difference in NH2OH dependence between the oxidation and reduction processes, and understanding this difference may ultimately yield some valuable mechanistic insights. One possible explanation would be if the affinity of HAO for NH2OH is dependent on the enzyme’s extent of reduction. To test this hypothesis, we plan to investigate the HAO-catalyzed reduction of NH2OH by a series of electron donors with different midpoint potentials.
Abbreviations
- AOB:
-
Ammonia-oxidizing bacteria
- HAO:
-
Hydroxylamine oxidoreductase
- Mb:
-
Ferromyoglobin
- MbNO:
-
Nitrosomyoglobin
- MVox :
-
Methyl viologen dication
- MVred :
-
Methyl viologen monocation radical
- SVD:
-
Singular value decomposition
References
Ehrlich HL (2002) Geomicrobiology. Marcel Dekker Inc., New York
Fenchel T, King GM, Blackburn TH (1998) Bacterial biogeochemistry, 2nd edn. Academic Press, London
Schmidt I, Steenbakkers PJM, Huub JM, Schmidt K, Jetten MSM (2004) J Bacteriol 186:2781–2788
Kroneck PMH, Beuerle J, Schumacher W (1992) In: Sigel H, Sigel A (eds) Metal ions in biological systems. Marcel Dekker Inc., New York, pp 455–505
Wood PM (1988) In: Cole JA, Ferguson SJ (eds) The nitrogen and sulfur cycles. Cambridge University Press, New York, pp 219–243
Hooper AB (1989) In: Schlegel HG, Bowien B (eds) Autotrophic bacteria. Science Tech Publishers, Madison, pp 239–265
Bock E, Koops HP, Harms H, Ahlers B (1991) In: Shively JM, Barton LL (eds) Variations in autotrophic life. Academic Press, San Diego, pp 171–200
Schlegel HG (1981) In: Bothe H, Trebst A (eds) Biology of inorganic nitrogen and sulfur. Springer, New York, pp 3–12
(1994) Biological nitrogen fixation. National Academy Press, Washington, pp 6–32
Burns RC, Hardy RWF (1975) Nitrogen fixation in bacteria and higher plants. Springer, New York
Prince RC, George GN (1997) Nat Struct Biol 4:247–250
Kampschreur MJ, Tan NCG, Picioreanu C, Jetten MSM, Schmidt I, van Loosdrecht MCM (2006) Biochem Soc Trans 34:179–181
Hooper AB, Nason A (1965) J Biol Chem 240:4044–4057
Andersson KK, Hooper AB (1983) FEBS Lett 164:236–240
Ferguson SJ (1998) Curr Opin Chem Biol 2:182–193
Igarashi N, Moriyama H, Fujiwara T, Fukumori Y, Tanaka N (1997) Nat Struct Biol 4:276–284
Hooper AB, Terry KR (1977) Biochemistry 16:455–459
Hendrich MP, Logan M, Andersson KK, Arciero DM, Lipscomb JD, Hooper AB (1994) J Am Chem Soc 116:11961–11968
Logan MSP, Balny C, Hooper AB (1995) Biochemistry 34:9028–9037
Logan MSP, Hooper AB (1995) Biochemistry 34:9257–9264
Arciero DM, Hooper AB (1993) J Biol Chem 268:14645–14654
Iverson TM, Arciero DM, Hsu BT, Logan MSP, Hooper AB, Rees DC (1998) Nat Struct Biol 5:1005–1012
Andersson KK, Lipscomb JD, Valentine M, Munck E, Hooper AB (1986) J Biol Chem 261:1126–1138
Whittaker M, Bergman D, Arciero DM, Hooper AB (2000) Biochim Biophys Acta 1459:346–355
Hoshino M, Maeda M, Konishi R, Seki H, Ford PC (1996) J Am Chem Soc 118:5702–5707
Ford PC, Lorkovic IM (2002) Chem Rev 102:993–1017
Wolak M, van Eldik R (2002) Coord Chem Rev 230:263–282
Fernandez BO, Ford PC (2003) J Am Chem Soc 125:10510–10511
Choi IK, Liu Y, Wei Z, Ryan MD (1997) Inorg Chem 36:3113–3118
Feng D, Ryan MD (1987) Inorg Chem 26:2480–2483
Einsle O, Messerschmidt A, Huber R, Kroneck PMH, Neese F (2002) J Am Chem Soc 124:11737–11745
Cabail MZ, Kostera J, Pacheco AA (2005) Inorg Chem 44:225–231
Enemark JH, Feltham RD (1974) Coord Chem Rev 13:339–406
Kurnikov IV, Ratner MA, Pacheco AA (2005) Biochemistry 44:1856–1863
Hendrich MP, Petasis D, Arciero DM, Hooper AB (2001) J Am Chem Soc 123:2997–3005
Shriver D, Atkins P (2006) Inorganic chemistry, 4th edn. WH Freeman and Co, San Francisco, p 775
Koppenol WH, Moreno JJ, Pryor WA, Ischiropoulus H, Beckman JS (1992) Chem Res Toxicol 5:834–842
Hooper AB, Tran VM, Balny C (1984) Eur J Biochem 141:565–571
Cabail MZ, Pacheco AA (2003) Inorg Chem 42:270–272
Cabail MZ, Lace PJ, Uselding J, Pacheco AA (2002) J Photochem Photobiol A Chem 152:109–121
Namiki S, Arai T, Fujimori K (1997) J Am Chem Soc 119:3840–3841
Bodemer G, Ellis LM, Lace PJ, Mooren PE, Patel NK, Ver Haag M, Pacheco AA (2004) J Photochem Photobiol A Chem 163:53–60
Cabail MZ, Moua V, Bae E, Meyer A, Pacheco AA (2007) J Phys Chem A 111:1207–1213
Press WH, Teukolsky SA, Vetterling WT, Flannery BP (1992) Numerical recipes in C the art of scientific computing, chap 15, 2nd edn. Cambridge University Press, Cambridge, pp 59–70
Strang G (1988) Linear algebra and its applications, 3rd edn. Harcourt Brace Jovanivich Inc., San Diego, pp 153–350
Codd R, Astashkin AV, Pacheco A, Raitsimring AM, Enemark JH (2002) J Biol Inorg Chem 7:338–350
Hoshino M, Ozawa K, Seki H, Ford PC (1993) J Am Chem Soc 115:9568–9575
Cunha CA, Macieira S, Dias JM, Almeida G, Goncalves LL, Costa C, Lampreia J, Huber R, Moura JJG, Moura I, Romao MJ (2003) J Biol Chem 278:17455–17465
Angove HC, Cole JA, Richardson DJ, Butt JN (2002) J Biol Chem 277:23374–23381
Einsle O, Messerschmidt A, Stach P, Bourenkov GP, Bartunik HD, Huber R, Kroneck PMH (1999) Nature 400:476–480
Hooper AB, Terry KR (1979) Biochim Biophys Acta 571:12–20
Yamanaka T, Shinra M (1974) J Biochem 75:1265–1273
Hooper AB, Maxwell PC, Terry KR (1978) Biochemistry 17:2984–2989
Iverson TM, Arciero DM, Hooper AB, Rees DC (2001) J. Biol Inorg Chem 6:390–397
Arciero DM, Balny C, Hooper AB (1991) Biochemistry 30:11466–11472
Arciero DM, Collins MJ, Haladjian J, Bianco P, Hooper AB (1991) Biochemistry 30:11459–11465
Upadhyay AK, Petasis DT, Arciero DM, Hooper AB, Hendrich MP (2003) J Am Chem Soc 125:1738–1747
Alluisetti GE, Almaraz AE, Amorebieta VT, Doctorovich F, Olabe JA (2004) J Am Chem Soc 126:13432–13442
Hendrich MP, Upadhyay AK, Riga J, Arciero DM, Hooper AB (2002) Biochemistry 41:4603–4611
Acknowledgments
This publication was made possible by support from the National Institutes of Environmental Health Sciences (NIEHS, grant no. 1 R15 ES013955-01), and from the University of Wisconsin-Milwaukee’s Research Growth Initiative (101X076).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kostera, J., Youngblut, M.D., Slosarczyk, J.M. et al. Kinetic and product distribution analysis of NO· reductase activity in Nitrosomonas europaea hydroxylamine oxidoreductase. J Biol Inorg Chem 13, 1073–1083 (2008). https://doi.org/10.1007/s00775-008-0393-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-008-0393-4