Abstract
Re-evaluation of the crystallographic data of the molybdenum-containing E. coli formate dehydrogenase H (Boyington et al. Science 275:1305–1308, 1997), reported in two redox states, reveals important structural differences for the formate-reduced form, with large implications for the reaction mechanism proposed in that work. We have re-refined the reduced structure with revised protocols and found substantial rearrangement in some parts of it. The original model is essentially correct but an important loop close to the molybdenum active site was mistraced, and, therefore, catalytic relevant residues were located in wrong positions. In particular selenocysteine-140, a ligand of molybdenum in the original work, and essential for catalysis, is no longer bound to the metal after reduction of the enzyme with formate. These results are incompatible with the originally proposed reaction mechanism. On the basis of our new interpretation, we have revised and proposed a new reaction mechanism, which reconciles the new X-ray model with previous biochemical and extended X-ray absorption fine structure data.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
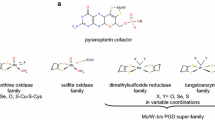
Molybdopterin-containing formate dehydrogenases (Fdhs) are members of the dimethyl sulfoxide reductase family of mononuclear molybdenum enzymes [1–4] and are involved in many different metabolic pathways. In addition to the Mo (or W) catalytic site, they contain Fe,S clusters in variable amount that are responsible for electron transfer. With a few exceptions, Fdhs contain a selenocysteine bound to the Mo atom, which is essential for enzymatic activity.
In Escherichia coli, three selenium-dependent Fdh isoenzymes are involved in formate respiration: Fdh-N, a membrane bound complex, expressed when growing on nitrate [5], Fdh-O, expressed for oxygen metabolism [6], and Fdh-H, part of the hydrogen lyase complex [7]. Crystal structures for Fdh-N and Fdh-H have been determined. While Fdh-N is a large complex of 600 kDa, with an α4β4γ4 structure [8], Fdh-H is an 80-kDa protein, containing one Mo active site and one [4Fe–4S] cluster. The crystal structures of both oxidized (2.8-Å resolution) and formate-reduced (2.3-Å resolution) forms of E. coli Fdh-H were reported in 1997 by Boyington et al. [7]. This was the first crystal structure of a Mo-containing Fdh, and the information provided is crucial for mechanistic interpretation, since it still is the only known structure of a formate-reduced state of the enzyme. The two homologous enzymes for which crystal structures are available were solved only in the oxidized form: E. coli Fdh-N [8] and the Desulfovibrio gigas W-containing Fdh [9, 10].
Extended X-ray absorption fine structure (EXAFS) studies for E. coli Fdh-H were also reported for oxidized and dithionite-reduced forms of the enzyme [11]. However, there are differences in the reduced structure determined by EXAFS that indicate the presence of a Se–S ligation, absent in the crystal structure. As proposed in that work, dithionite-reduced Fdh-H and formate-reduced Fdh-H must correspond to different forms of the enzyme [11]. With dithionite, no break of the Mo–Se bond would be expected. EXAFS experiments on the formate-reduced species, which has been crystallized, would certainly help to clarify this problem. The related Fdh from D. desulfuricans was recently studied by electron paramagnetic resonance (EPR) for as-prepared and inhibited forms of the enzyme (Brondino and Moura, unpublished results) and comparison with the EPR data for E. coli Fdh-H [12] suggests changes in the latter interpretation.
When reanalyzing the original diffraction data (kindly provided by Jeff Boyington) of E. coli Fdh-H, we realized that a loop region close to the active site (residues 138–146) of the formate-reduced E. coli Fdh-H (pdb code 1aa6) was mistraced. This region includes the catalytic relevant active site residues Se-Cys140 and His141, and the observed changes have therefore severe implications for the reaction mechanism proposed in [7] on the basis of the crystal structure. The enzymatic reaction mechanism for Fdhs is now reanalyzed and an alternative mechanism is proposed.
Experimental procedures
The model was rebuilt manually in O [13] and refined with Refmac5 [14] using all available reflections. Hydrogen atoms were added in riding positions, while ARP-wARP [15] was employed for updating the water molecules. The whole protein chain, Fe–S cluster and molybdopterin guanine dinucleotide (MGD) cofactors were defined as a single TLS domain, which basically allows modeling of the anisotropic motion of a rigid body at a cost of only 20 additional refinement parameters. Restraints for the Fe–S cluster and MGD were identical to those used for D. gigas W-containing Fdh [10]. The X-ray weight (vs. geometry) was carefully chosen to minimize R free. Bond lengths to the molybdenum atom were defined (σ = 0.4 Å) to avoid implicit van der Waals restraints.
To distinguish the improvement of our model from the general improvement of refinement programs over time, we have included refinement statistics for the rebuilt model with residues 136–150 in the old position (Table 1). While the global quality indicators R/R free are not very sensitive to local changes, they confirm the better fit of this loop in the new model. The real-space R factor [13]—a local goodness of fit indicator for each atom—showed the same result more clearly (Fig. 1).
Real-space R and correlation coefficients (CC) between the best [2mF o−DF cexp(iϕ)] and the calculated [F c exp(iϕ)] electron density were computed in O [13]. They indicate an improved fit around each of the residues 138–149 in the new interpretation. To avoid showing biologically irrelevant differences caused by the choice of refinement program or resolution cutoff, we used the same hybrid model of Table 1 as the “old” structure to calculate these statistics
The dithiolene function of the MGD cofactor was restrained to be flat (double-bonded dithiolene function) for the MGD which points towards the 4Fe–4S cluster (MGD 802, using common numbering for Fdhs [3]), but restraints for the corresponding reduced form (dithiolate in alternate conformation) gave a better electron density fit for the pterin-group of MGD 801, like in W-Dg Fdh [10]. Anisotropic displacement of the molybdenum atom, its apical ligand and the selenium atom are clearly visible in difference electron density maps. However, anisotropic refinement of these atoms did not improve the R/R free factor. The new model fits well to the electron density, with only two main-chain gaps for residues 410–412 and 658–675.
Results and discussion
When analyzing the crystal structures of E. coli Fdh-H, we found that the deposited model for the formate-reduced form (pdb code 1aa6) revealed a very poor fit of the electron density map for the loop 138RV-SeC-HGPSVA146. Val139, part of the molybdenum ligand Se-Cys140 and His141 were clearly out of density, while other parts showed poor connectivity. To our surprise we found the selenium atom 12 Å away from the molybdenum atom, as indicated by the second highest 2F o−F c electron density peak (8σ above the average) and the highest difference peak (F o−F c) (9σ above the average) (Fig. 2). This implies that, in the formate-reduced Fdh-H, Se-Cys140 is not a Mo ligand.
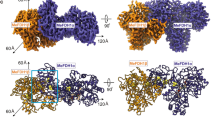
a Stereo picture of the cofactor and the loop 136–149 (for clarity only Cα atoms are shown, except for Se-Cys140, His141, Arg138) from the deposited structure 1aa6 (red) [7] and our interpretation (green). In this picture one can see how different the Se-Cys140 and His141 positions are. The newly traced main chain (138–139) corresponds to an alternate conformation for Arg138 (blue) in the original work. The newly traced loop contains only a single conformation. b Stereo representation of the 2F o−F c electron density map of the same region represented in a, contoured at 1σ, after one round of refinement in refmac5 [14]. The molybdenum, the selenium and the phosphorous atoms are contoured at 5σ (yellow) (the maximum, at the center of the selenium contour, was 8σ). c As in b, zoomed in at the seleniumcysteine and His141. For calculation of the density, residues 130–150 were deleted from the model and all atoms were randomly displaced (0.3 Å r msd) before refinement, to remove model bias. The picture was made with PyMOL [20]. d Cα plot of the deposited structure (1aa6) [7] and the newly refined structure in different shades of gray. The loop 136–149 is shown in red for 1aa6 and in green for the revised structure
We also checked the deposited coordinates of the oxidized form of the enzyme (pdb code 1fdo) using the same approach. The model fits very well to its electron density, leaving no doubt about the selenium being a molybdenum ligand.
Except for the mistraced loop in the reduced form, both forms of the enzyme adopt, however, almost identical structures (Fig. 2d). Unfortunately the mobile 138RV-SeC-HGPSVA146 active-site loop corresponds to a relatively weakly diffracting region. Low-resolution data are important for establishing the connectivity of the electron density. With these data excluded (cf. Table 1), the false double conformation of Arg138b modeled in the original deposited structure was able to hide the true conformation of the active-site loop. In the new interpretation, the selenium atom contacts the conserved Arg333 stabilizing the negative charge of a free selenol (see later). The movement of the polypeptide chain also drags along His141, which now contacts Asp349 and the conserved Arg138.
At the molybdenum site, the apical ligand refined better as a sulfur atom (=S or –SH, not H2O or –OH) as also observed in the related W-Fdh from D. gigas [10]. Although the available resolution does not allow a firm conclusion, it has been reported that E. coli Fdh-H is irreversibly inactivated by cyanide [16], which would be in agreement with the presence of a cyanolysable terminal sulfide, also considered possible in the EXAFS study [11].
On the basis of the new interpretation of the electron density data of formate-reduced E.coli Fdh-H, we can propose a reaction mechanism that fits the experimental data from the group of Stadtman [11, 12, 17, 18] (Fig. 3).
In step I, formate binds directly to Mo, displacing Se-Cys140. The binding of formate is in analogy to that of nitrite, a competitive inhibitor and is shown for the nitrite-bound structure in the oxidized form (pdb code 1fdi) [7]. The 9-Å displacement of the selenium atom from its bound position to the free selenol form places it close (3.5 Å) to the Nε1 of Arg333, which stabilizes the free selenol. Arg333 is highly conserved in Fdhs [3, 4] and is very well ordered in both oxidized and reduced forms, although it adopts different conformations: In the oxidized form, its side chain points towards the active site, in a position to interact and orient the formate molecule (form A). In the formate-reduced form, Arg 333 points away and is in contact with the free selenol from Se-Cys140 (form B).
In step II, the α-proton from formate may be transferred to the nearby His141 that acts as general base, in agreement with EPR data [12] and with studies of pH dependency of catalysis [18]. In this step the CO2 molecule can be released and two electrons transferred to Mo. Alternatively, step II may involve a selenium-carboxylated intermediate, in agreement with an earlier proposal in 1993 by Heider and Böck [19]. These studies, unbiased by any crystal structure, suggested that the free selenol assists formate oxidation through a selenium-carboxylated intermediate stage, before CO2 is released. Indeed, there are several residues close to the new Se position that might catalyze its decarboxylation in the outward position (e.g., Arg333, Arg138, His141).
In step III, electrons from MoIV are transferred via the [4Fe–4S] center to an external electron acceptor and the catalytic cycle is completed.
The crucial role of Se-Cys in the mechanism (namely, in its unbound form) is supported by the fact that Fdh-H is completely inactivated by the presence of iodoacetamide, but only after reduction of the enzyme with formate [18]. The same study showed that this inactivation is pH-dependent and correlates with the pKa value of ionized selenol. In fact, loop 138–146 in the new trace is quite exposed to solvent, in the cavity which leads into the active site. In this conformation the free selenol is clearly more accessible for alkylation with iodoacetamide than when buried and bound to the Mo atom (Fig. 2d).
This new reaction mechanism is supported by EPR studies on D. desulfuricans Fdh (Brondino and Moura, private communication) which show that the OH/SH ligand is present in the MoV state of the enzyme, formed after CO2 release (step III).
This alternative reaction mechanism for Fdhs is the subject of on-going crystallographic studies with the homologous enzyme from D. gigas.
Atomic coordinates have been deposited in the Protein Data Bank under accession code 2iv2.
Abbreviations
- EPR:
-
Electron paramagnetic resonance
- EXAFS:
-
Extented X-ray absorption fine structure
- Fdh:
-
Formate dehydrogenase
- Fdh-H:
-
Formate dehydrogenase component of the formate-hydrogen lyase complex of E. coli
- Fdh-N:
-
Formate dehydrogenase expressed in E. coli when growing on nitrate
- Fdh-O:
-
Formate dehydrogenase expressed in E. coli for oxygen metabolism
- MGD:
-
Molybdopterin guanine dinucleotide
References
Hille R (1996) Chem Rev 96:2757–2816
Romão MJ, Knäblein J, Huber R, Moura JJG (1997) Prog Biophys Mol Biol 68:121–144
Moura JJG, Brondino CD, Trincao J, Romao MJ (2004) J Biol Inorg Chem 9:791–799
Vorholt JA, Thauer RK (2002) In: Sigel A, Sigel H (eds) Metal ions in biological systems, vol 39. Dekker, New York, pp 571–619
Berg BL, Li J, Heider J, Stewart V (1991) J Biol Chem 266:22380–22385
Abaibou H, Pommier J, Giordano G, Mandrand-Berthelot MA (1995) J Bacteriol 177:7141–7149
Boyington JC, Gladyshev VN, Khangulov SV, Stadtman TC, Sun PD (1997) Science 275:1305–1308
Jormakka M, Tornroth S, Byrne B, Iwata S (2002) Science 295:1863–1868
Raaijmakers H, Teixeira S, Dias JM, Almendra MJ, Brondino CD, Moura I, Moura JJ, Romao MJ (2001) J Biol Inorg Chem 64:398–404
Raaijmakers H, Macieira S, Dias JM, Teixeira S, Bursakov S, Huber R, Moura JJ, Moura I, Romao MJ (2002) Structure 10:1261–1272
George GN, Colangelo CM, Dong J, Scott RA, Khangulov SV, Gladyshev VN, Stadtman TC (1998) J Am Chem Soc 120:1267–1273
Gladyshev VN, Boyington JC, Khangulov SV, Grahame DA, Stadtman TC, Sun PD (1996) J Biol Chem 271:8095–8100
Jones TA, Zou JY, Cowan SW, Kjeldgaard M (1991) Acta Crystallogr Sect A 47:110–119
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr Sect D 53:240–245
Perrakis A, Morris RJ, Lamzin VS (1999) Nat Struct Biol 6:458–463
Barber M, May H, Ferry JG (1986) Biochemistry 25:8150–8155
Khangulov SV, Gladyshev VN, Dismukes GC, Stadtman TC (1998) Biochemistry 37:3518–3528
Axley MJ, Bock A, Stadtman TC (1991) Proc Natl Acad Sci USA 88:8450–8454
Heider J, Böck A (1993) Adv Microb Physiol 35:71–109
DeLano WL (2002) The PyMOL molecular graphics system. DeLano Scientific, San Carlos. http://www.pymol.org
Acknowledgements
This work was supported by EC-TMR/FMRX-CT980204 and POCTI/QUI/57641/2004. We thank Jeff Boyington for providing the structure factors corresponding to the deposited coordinates (pdb codes 1aa6 and 1fdo), and A. Boeck for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Raaijmakers, H.C.A., Romão, M.J. Formate-reduced E. coli formate dehydrogenase H: the reinterpretation of the crystal structure suggests a new reaction mechanism. J Biol Inorg Chem 11, 849–854 (2006). https://doi.org/10.1007/s00775-006-0129-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-006-0129-2