Abstract
S100A7 (psoriasin) and S100A15 (koebnerisin) were first identified in inflamed psoriatic skin. They are of major interest because of their putative functional roles in innate immunity, epidermal cell maturation, and epithelial tumorigenesis. Human S100A7 and S100A15 have lately evolved by gene duplications within the epidermal differentiation complex (chromosome 1q21) during primate evolution forming a novel S100 subfamily. Therefore, S100A7 and S100A15 are almost identical in sequence (>90%) and are difficult to discriminate. Despite their high homology, S100A7 and S100A15 are distinct in tissue distribution, regulation, and function, and thus, exemplary for the diversity within the S100 family. Their different properties are compelling reasons to discriminate S100A7 (psoriasin) and S100A15 (koebnerisin) in epithelial homeostasis, inflammation, and cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
S100 proteins are small (9–13 kDa) and constitute the largest, multigenic family of calcium-binding EF-hand proteins (Donato 2003). Their distinct tissue- and cell-type expression patterns indicate local specification for homeostasis and disease.
S100 proteins are produced as monomers and spontaneously assemble to dimers/multimers. To be active, calcium binding then induces the exposure of a binding surface with which the S100-mers interact with their target proteins (Zimmer et al. 2003). Thus, members of this protein family are involved in calcium-dependent and, in some cases, zink- or copper-dependent cellular functions. They modulate cell metabolism, proliferation, and maturation by regulating gene transcription, protein activation, and intracellular trafficking by structural participation in membranes. Some S100 members are released into the extracellular space where they act as chemoattractants for leukocytes or regulate their activity (Heizmann et al. 2002).
Most S100 family members are encoded in the epidermal differentiation complex (EDC) located on human chromosome 1q21 (Hardas et al. 1996). This region is of particular interest as it encodes many genes that have been linked to epidermal differentiation and inflammation (Mischke et al. 1996; South et al. 1999; de Cid et al. 2009; Zhang et al. 2009). Two genes in that locus were cloned, S100A7 (psoriasin) and subsequently S100A15 (koebnerisin), because of their particularly high expression in inflamed psoriatic lesions, which are characterized by disturbed epidermal differentiation and inflammation (Madsen et al. 1991; Wolf et al. 2003). Despite their small size and conserved functional domains, S100 gene duplications throughout vertebrate evolution led to an increase in number and diversity within the S100 family (Kulski et al. 2003). Minor sequence variations can lead to structural differences that subsequently result in different binding to target proteins and functions. The human S100A7/S100A15 subfamily shares over 90% sequence identity. Despite the high homology, S100A7 (psoriasin) and S100A15 (koebnerisin) are distinct and therefore exemplary for the functional and expressional diversity within the S100 family.
Identification, genomic organization, and protein structure
Human S100A7 has been identified almost two decades ago and named ‘psoriasin’ because it is overexpressed by psoriatic keratinocytes (Madsen et al. 1991). Similarly, the human S100A15 has been discovered by analyzing the differential gene expression in psoriasis (Wolf et al. 2003). Due to overexpression in ‘koebnerized’ psoriatic skin, S100A15 proposed name is ‘koebnerisin’. Both genes map to the S100 gene cluster within the EDC (chromosome 1q21), which has been identified as one of the psoriasis candidate loci (PSORS4) (Hardas et al. 1996; Semprini et al. 1999, 2002).
In contrast to other S100 members, S100A15 reveals an unusual genomic organization. Whereas most S100 genes including S100A7 encode for a single transcript, two alternate S100A15 mRNA-isoforms have been discovered. They share the same coding region, but show differences in UTR composition and length (0.5 vs. 4.4 kb). Both S100A15 splice variants are differently expressed in psoriatic skin suggesting regulation through alternate promoters (Fig. 1).
Genomic structure of the human S100A7/S10015 subfamily. Schematic representation of the genomic and exon/intron organization of the S100A7 and the S100A15 genes. Boxes represent exons; hatched regions indicate the coding sequence; intervening lines denote introns. The alternate S100A15 mRNA-splice variants are marked A for S100A15-L (long) and B for S100A15-S (short)
Analysis of the S100A15-deduced amino acid sequence reveals a conserved C-terminal and a variant N-terminal EF-hand typical for S100 proteins (101 amino acids, 11.305 Da, calculated pI of 7.57). The S100A15 protein is highly homologous (93% sequence identity) to S100A7 (psoriasin, 101 amino acids, 11.326 kD, calculated pI of 6.77) (Fig. 2). The closest match to the S100A7/S100A15 subfamily is hS100A11 (Calgizzarin), which shows a skin expression profile similar to S100A15 (Broome and Eckert 2004). Calgizzarin is important for plasma membrane remodeling during terminal differentiation, and the calcium-dependent S100A11-homodimer interaction with annexin I might participate in the annexins calcium-channel activity (Bianchi et al. 2003; Broome et al. 2003; Dempsey et al. 2003).
Protein structure of the human S100A7/S10015 subfamily. Alignment of the predicted amino acid sequences of the human S100A7 (S100A7) with the human S100A15 (S100A15) compared to the human S100A11 (hS100A11) representing the closest human S100 member outside the S100A7/S100A15 subfamily. Identical amino acid residues are indicated as blue boxes, and chemically similar amino acids are marked as gray boxes. Predicted EF-hand motifs for both proteins are marked above the sequences (variant S100-specific motif: amino acids 12–39, canonical EF-hand: amino acids 54–82)
The main differences between S100A15 and S100A7 at the putative N-terminus lead to the prediction of a calcium-binding EF-hand motif for the S100A15, which is not found to be functional in S100A7. Further, zinc binding of S100A15 could be impacted as one out of four sites important for zinc binding in S100A7 is missing in the S100A15 sequence (Asp24Gly) (Brodersen et al. 1999). Because of the predicted differences between highly homologous S100A7 and S100A15, studies were needed to discriminate their expression, regulation, and function in epithelial homeostasis and disease.
Epithelial maturation
The gene locus of the S100A7/S100A15 subfamily is linked to epidermal maturation (EDC; human chromosome 1q21). This chromosomal region encodes additional genes (involucrin, filaggrin, trichoyalin, repetin, etc.) that are sequentially expressed in maturing epidermis (Mischke et al. 1996; South et al. 1999). The epidermis is constantly exposed to a variety of physicochemical and microbial challenges. The upper differentiated layers serve as a physical (cornified envelope) and biological (antimicrobial lipids, antimicrobial proteins) protective hurdle supported by normal microflora on the skin surface (Schroder and Harder 2006). The cornified envelope is a protective structure that is assembled adjacent the inner surface of the cell plasma membrane during the terminal stages of keratinocyte differentiation (Eckert et al. 2004). It is assembled from a pool of precursor proteins that are covalently crosslinked to one another via the action of the membrane-anchored enzyme type I transglutaminase (Steinert et al. 1996). S100A7 (psoriasin) and other S100 proteins are transglutaminase substrates, and several S100 proteins are components of the keratinocyte-cornified envelope (Robinson et al. 1997). Accordingly, the S100A7 transcript shows a calcium- and differentiation-dependent regulation in human keratinocytes (Martinsson et al. 2005). During keratinocyte differentiation in epidermis, S100A7 redistributes to the cell periphery (Broome et al. 2003; Ruse et al. 2003), suggesting that S100A7 is released from differentiated keratinocytes. Studies show that extracellular S100A7 functions as an antibacterial agent reducing survival of E. coli and other strains (Glaser et al. 2005). However, considering the difficulties in distinguishing the highly homologous S100A7 and S100A15, both proteins may have contributed to previously reported features. Moreover, many of the customized and commercial S100A7 antibodies are cross-reactive with both S100 proteins (Wolf et al. 2009). Thus, S100A15 (koebnerisin)-specific antibodies have been generated that did not cross-react with related recombinant hS100 proteins, particularly S100A7 (Wolf et al. 2008).
Using these antibodies, distinct distribution patterns of S100A15 and S100A7 were found in normal skin. S100A7 expression in the epidermis is confined to the granular/cornified layer only. S100A15 co-localized there, but unlike S100A7, it is also expressed by epidermal basal keratinocytes and keratin 5 negative dendritic-shaped cells. These epidermal cells were identified as melanocytes (MART-1 positive), dendritic cells, and Langerhans cells (MHCII positive). Within the pilosebaceous unit, S100A15 is co-localized with S100A7 in the inner root sheath. Unlike S100A7, S100A15 is expressed in K14-positive cells (external root sheath, basal layer of the sebaceous gland). In the dermis, S100A15 is expressed by SMA-positive smooth muscle cells and endothelial cells, whereas S100A7 could not be detected outside epidermal structures (Fig. 3). Similar to skin, a differential staining pattern has been shown in normal breast tissue. There, both proteins are expressed by cellular subsets of alveolar and small duct luminal cells within normal breast (Wolf et al. 2009). That S100A15 is expressed by epithelial-derived myoepithelial cells around acini and by surrounding blood vessels may reflect its biological diversity from S100A7 with additional, distinct functions for S100A15.
As S100A15 is expressed within the differentiated epidermal layers of normal skin, the regulation of S100A15 during calcium-induced keratinocyte differentiation was investigated. While both S100A15 mRNA variants were induced by calcium, the S100A15-L response was more pronounced (Wolf et al. 2007). Induction was more rapid at higher calcium concentrations concordant with expression of late differentiation markers similar to S100A7 (Martinsson et al. 2005).
S100A15 (koebnerisin) presence in differentiated layers at the epidermal surface suggests participation in the antimicrobial defense. Similar to S100A7, S100A15 functions as an antibacterial agent reducing survival of E. coli (Glaser et al. 2005; Buchau et al. 2007; Lee and Eckert 2007). E. coli-induced expression of both S100A7 and S100A15 is dependent on Toll-like-receptor (TLR) 4. In contrast to S100A7, S100A15 is also strongly regulated by several bacterial components, such as P. aeruginosa and S. aureus. As in human skin, both proteins might participate in the microbial homeostasis within the host as well as in the digestive tract of breast-feeding newborns (Wolf et al. 2009). That S100A7 (psoriasin) and S100A15 (koebnerisin) are co-regulated through common inducers of differentiation but by different antimicrobial agents make them co-operate to potentiate both the mechanical and the antimicrobial host defense (Glaser et al. 2005; Buchau et al. 2007; Abtin et al. 2008).
These studies emphasize the importance to discriminate the highly related S100A7 and S100A15 paralogs and open the opportunity to further dissect their differential physiological functions in normal tissues and their use as distinct markers in epithelial pathologies in the skin and beyond.
Epithelial tumorigenesis
Disruption of the calcium signaling pathway has been implicated as a central mechanism in tumorigenesis, specifically tumor invasion and metastasis (Kohn and Liotta 1995).
In normal skin, S100A7 and S100A15 are coregulated and coexpressed with squamous cell differentiation (Moubayed et al. 2007; Wolf et al. 2007, 2008). In epithelial tumors, upregulation of S100A7 is considered a useful marker for recognizing in situ carcinomas and pre-invasive foci. S100A7 expression is often decreased in invading carcinomas; however, its persistent expression in invading tumors is associated with poor prognosis (Emberley et al. 2003, 2004a). While factors related to cellular differentiation clearly comprise an important aspect of the regulation of the S100A7/S100A15 subfamily, the downregulation that is frequently seen with invasion suggests regulation by additional factors that may also be associated with the invasive process in these tumors (Alowami et al. 2003; Emberley et al. 2004b). Since much of the previous work on S100A7 proceeds the discovery of the highly homologous S100A15, both proteins might have detected in association with tumor progression.
The ability to distinguish the closely related S100A7 and S100A15 at the RNA and protein level reveals significant distinctions in their regulation. Breast cancer specimens link high expression levels of both S100A7 and S100A15 transcripts with ER negativity and imply a correlation to clinical outcome as previously indicated for S100A7 only (Emberley et al. 2003, 2004a). While S100A7 protein expression closely follows corresponding RNA levels, the S100A15 protein is ubiquitous in invasive carcinomas and appears to be preferentially modified/cross-linked, and thus, provides a more stable and potentially longer lived protein even when transcript levels are low (Wolf et al. 2009). The coincidental but differential expression and intracellular localization of these almost identical S100 paralogs could have significant biological implications for normal breast and breast cancer. Co-expression of transcripts for both S100A7 and S100A15 proteins in ER/PR negative tumors suggests a joint regulation related to tumor progression. While the secreted proteins have distinct roles as chemoattractants (Wolf et al. 2008), they also act synergistically to enhance inflammation, and thus, could influence breast tumor progression.
Beyond skin, these studies emphasize the S100A7 and S100A15 differential roles and their use as distinct markers in breast cancer pathogenesis. Their cooperate action could be important in other epithelial tumors, such as lung, gastric, and bladder cancer, where S100A7 has been previously described (Yao et al. 2007; Zhang et al. 2007; Liu et al. 2008).
Skin inflammation
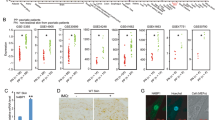
The human calcium-binding protein S100A15 (koebnerisin) was first identified in inflamed hyperplastic psoriatic skin, where the S100A15 gene is transcribed into two mRNA splice variants, S100A15-S (short isoform) and S100A15-L (long isoform) (Wolf et al. 2003). Both isoforms showed elevated levels in lesional psoriatic skin, where S100A15-L was more pronounced than S100A15-S. There, the detection of the S100A15-L transcript was pronounced in the basal and granular layer of non-lesional psoriatic skin and further extended throughout the hyperplastic epidermis of lesional psoriatic skin. Similar to psoriasis, increased S100A15 mRNA expression was observed throughout the epidermis in the skin of chronic atopic eczema (Wolf et al. 2007), whereas S100A7 (psoriasin) transcripts are restricted to the upper epidermis of both atopic and psoriatic skin (Madsen et al. 1991; Glaser et al. 2009). In contrast to S100A7, sporadic staining of S100A15 transcripts in single cells and cell clusters was detected in the dermis of inflammatory psoriatic and eczematous skin. Compared to normal, both S100A7 and S100A15 proteins are upregulated in inflamed lesional psoriatic skin and co-expressed by the epidermal suprabasal compartments (Fig. 4a). In addition, S100A15 is highly expressed by clusters of basal psoriatic keratinocytes at the epidermal–dermal junction. Both S100A7 and S100A15 proteins are expressed/secreted at a similarly high rate but at a different ratio (psoriasin 1/3.8; koebnerisin 1/2.5) by cultured psoriatic keratinocytes, which might be important to further understand their distinct intracellular roles and functions as chemoattractants (Wolf et al. 2008).
S100A7 and S100A15 function through distinct classes of receptors. a Frozen skin sections were stained for S100A7 (green) and S100A15 (red) showing their upregulation and differential distribution in normal skin compared to psoriasis. Nuclei were stained with DAPI (blue). Bar 50 μm. b S100A7 mediates leukocyte chemotaxis through RAGE (receptor of advanced glycated end products). This atypical chemotaxis receptor is pertussis toxin-insensitive, which helps to distinguish ligand activity through classical chemokine receptors. S100A15 chemotactic activity is Gi protein-dependent, but the receptor has yet to be specified. The distinct mechanisms of actions within the S100A7 (psoriasin)/S100A15 (koebnerisin) subfamily contribute to their synergistic effect in inflammation
Psoriasis and chronic atopic eczema are chronic inflammatory skin diseases characterized by skin-infiltrating immune cells. These cells are known to secrete proinflammatory cytokines which are mainly produced by granulocytes, macrophages, and Th1/Th17-differentiated lymphocytes (Numerof and Asadullah 2006). S100A15 expression is induced in cultured human keratinocytes upon treatment with TNF-α and IFN-γ as well as IL-1β, suggesting that the proinflammatory environment in diseased skin contributes to S100A15 expression in the epidermis. Whereas S100A15-S is only weakly induced by Th1 cytokines, IL-1β solely induces S100A15-L, which further indicates specific S100A15 isoform regulation by alternate promoters. The unresponsiveness of keratinocytes to regulate S100A15 by the Th2-derived cytokines IL-4 and IL-13 suggests that S100A15 is preferentially induced by Th1-driven psoriasis and late chronic atopic eczema inflammation rather than in Th2-dominated diseases (Grewe et al. 1998). A similar regulation pattern through Th1 cytokines has been shown for S100A7 concordant with co-regulation of both S100A7 and S100A15 in inflammation (Glaser et al. 2005, 2009). Moreover, the S100A7/S100A15 subfamily is regulated by Th17 and Th22 cytokines important in the pathogenesis of psoriasis and other inflammatory skin diseases (Sabat et al. 2007; Eyerich et al. 2009); however, a specific distinction between S100A7 and S100A15 is needed in future investigations.
That S100A7 and S100A15 are co-upregulated in similar pathophysiological conditions through similar epidermotropic proinflammatory mediators suggests that both proteins functionally cooperate in inflammation. The upregulation and secretion of both human S100A7 and S100A15 in chronic inflammatory diseases suggests that they contribute to the inflammatory phenotype (Fig. 4a). When extracellular, either hS100 protein induced an inflammatory response as shown by intraperitoneal injection into mice (Wolf et al. 2008). When injected together, the inflammatory response was amplified resembling the increased expression and release of both S100A7 and S100A15 by psoriatic keratinocytes with implications for their pathogenetic role in the disease (Fig. 4b). Both proteins are chemoattractants, but differ in their chemotactic activity toward specific leukocyte subtypes. With the discovery of S100A7 (Madsen et al. 1991), structural and functional data had stimulated the quest for mechanistic clarification of its extracellular action. The multiligand receptor of advanced glycated end products (RAGE) is implicated in inflammatory processes including leukocyte migration (Zen et al. 2007; Ramasamy et al. 2008). RAGE is expressed at low levels in normal tissues and becomes upregulated wherever its ligands accumulate. Ligands initiate a sustained cellular activation through MAP kinases culminating in the activation of NFκB. Through recognition of β-sheet fibrillar structures, proinflammatory cytokine-like mediators of the S100/calgranulin family or high mobility group box-1 (HMGB-1) protein, RAGE participates in the phenotype of inflammatory skin diseases, diabetes, and amyloidosis, and promotes tumor progression (Taguchi et al. 2000; Santilli et al. 2009; Sourris and Forbes 2009; Yan et al. 2009). Mechanistically, S100A7 and S100A15 stimulate chemotaxis through activation of different classes of receptors. S100A15-mediated chemotaxis is blocked by pertussis toxin suggesting a signaling through a classical Gi protein-coupled receptor. In contrast, S100A7-mediated chemotaxis is pertussis toxin-insensitive and is mediated through the pattern recognition receptor RAGE. Further, S100A7 but not S100A15 binds and directly mediates chemotaxis through RAGE in both in vitro chemotaxis assays and in vivo in mouse models (Wolf et al. 2008).
S100A7-RAGE binding, signaling, and chemotaxis are zinc dependent, reflecting the zinc-mediated changes in the S100A7 dimer structure. This finding identifies zinc as an important mediator of S100A7 chemotactic activity similar to S100A7 zinc-dependent antimicrobial action (Glaser et al. 2005; Lee and Eckert 2007). That RAGE is not the receptor for both S100 paralogs is likely due to the structural disparity between S100A7 and S100A15. Whereas most S100 proteins including S100A7 and S100A15 bind calcium at their conserved C-terminal EF-hand, the calcium binding at the variant N-terminal EF hand is impaired in S100A7 due to lack of glutamate residues (Donato 2003; Zimmer et al. 2003). In contrast to S100A7, the S100A15 protein features those glutamate residues suggesting that S100A15 binds calcium at its N-terminal EF-hand which may contribute to a quaternary structure distinct from S100A7 (Boeshans et al. 2006).
The RAGE is thought to recognize spatial structures rather than amino acid sequences (Bierhaus et al. 2005). Since sequences and secondary structures of the S100A7 and S100A15 monomers are alike, their distinct quaternary structures may determine if they are either perceptible by RAGE (S100A7) or not (S100A15). Similar results have been reported for hS100A12 binding to RAGE in a multimeric form (Moroz et al. 2009).
The proposed structural differences and distinct functional mechanisms between S100A7 and S100A15 provide evidence for disparity within the S100A7/S100A15 subfamily beyond their differences in expression. Their independent actions through distinct receptors regulate physiological functions and potentiate their action in inflammatory diseases (Fig. 4).
Perspective
S100A7 (psoriasin) and S100A15 (koebnerisin) were cloned because of their particularly high expression in psoriatic lesions. They are encoded within the S100 protein complex on chromosome 1q21 (PSOR4) that has been genetically linked to disturbed differentiation and inflammation. Although both proteins are highly homologous, they are differentially expressed and regulated in normal and diseased tissues and have distinct functions and mechanisms of action. It is therefore important to discriminate S100A7 (psoriasin) and S100A15 (koebnerisin) and to learn more about their distinct functional roles and synergistic action in immunity and tumorigenesis. This understanding is crucial for developing therapeutic interventions for pathological conditions mediated by the S100A7 (psoriasin)/S100A15 (koebnerisin) subfamily.
Abbreviations
- UTR:
-
Untranslated region
- EDC:
-
Epidermal differentiation complex
- S100A15-L:
-
Long human S100A15 transcript
- S100A15-S:
-
Short human S100A15 transcript
- TLR:
-
Toll-like receptor
- RAGE:
-
Receptor of advanced glycated end products
References
Abtin A, Eckhart L, Mildner M, Gruber F, Schroder JM et al (2008) Flagellin is the principal inducer of the antimicrobial peptide S100A7c (psoriasin) in human epidermal keratinocytes exposed to Escherichia coli. FASEB J 22(7):2168–2176
Alowami S, Qing G, Emberley E, Snell L, Watson PH (2003) Psoriasin (S100A7) expression is altered during skin tumorigenesis. BMC Dermatol 3:1
Bianchi R, Giambanco I, Arcuri C, Donato R (2003) Subcellular localization of S100A11 (S100C) in LLC-PK1 renal cells: calcium- and protein kinase c-dependent association of S100A11 with S100B and vimentin intermediate filaments. Microsc Res Tech 60(6):639–651
Bierhaus A, Humpert PM, Morcos M, Wendt T, Chavakis T et al (2005) Understanding RAGE, the receptor for advanced glycation end products. J Mol Med 83(11):876–886
Boeshans KM, Wolf R, Voscopoulos C, Gillette W, Esposito D et al (2006) Purification, crystallization and preliminary X-ray diffraction of human S100A15. Acta Crystallogr Sect F Struct Biol Cryst Commun 62(Pt 5):467–470
Brodersen DE, Nyborg J, Kjeldgaard M (1999) Zinc-binding site of an S100 protein revealed. Two crystal structures of Ca2+-bound human psoriasin (S100A7) in the Zn2+-loaded and Zn2+-free states. Biochemistry 38(6):1695–1704
Broome AM, Eckert RL (2004) Microtubule-dependent redistribution of a cytoplasmic cornified envelope precursor. J Invest Dermatol 122(1):29–38
Broome AM, Ryan D, Eckert RL (2003) S100 protein subcellular localization during epidermal differentiation and psoriasis. J Histochem Cytochem 51(5):675–685
Buchau AS, Hassan M, Kukova G, Lewerenz V, Kellermann S et al (2007) S100A15, an antimicrobial protein of the skin: regulation by E. coli through Toll-like receptor 4. J Invest Dermatol 127(11):2596–2604
de Cid R, Riveira-Munoz E, Zeeuwen PL, Robarge J, Liao W et al (2009) Deletion of the late cornified envelope LCE3B and LCE3C genes as a susceptibility factor for psoriasis. Nat Genet 41(2):211–215
Dempsey AC, Walsh MP, Shaw GS (2003) Unmasking the annexin I interaction from the structure of Apo-S100A11. Structure 11(7):887–897
Donato R (2003) Intracellular and extracellular roles of S100 proteins. Microsc Res Tech 60(6):540–551
Eckert RL, Broome AM, Ruse M, Robinson N, Ryan D et al (2004) S100 proteins in the epidermis. J Invest Dermatol 123(1):23–33
Emberley ED, Niu Y, Njue C, Kliewer EV, Murphy LC et al (2003) Psoriasin (S100A7) expression is associated with poor outcome in estrogen receptor-negative invasive breast cancer. Clin Cancer Res 9(7):2627–2631
Emberley ED, Murphy LC, Watson PH (2004a) S100A7 and the progression of breast cancer. Breast Cancer Res 6(4):153–159
Emberley ED, Alowami S, Snell L, Murphy LC, Watson PH (2004b) S100A7 (psoriasin) expression is associated with aggressive features and alteration of Jab1 in ductal carcinoma in situ of the breast. Breast Cancer Res 6(4):R308–R315
Eyerich K, Pennino D, Scarponi C, Foerster S, Nasorri F et al (2009) IL-17 in atopic eczema: linking allergen-specific adaptive and microbial triggered innate immune response. J Allergy Clin Immunol 123(1):59–66 e54
Glaser R, Harder J, Lange H, Bartels J, Christophers E et al (2005) Antimicrobial psoriasin (S100A7) protects human skin from Escherichia coli infection. Nat Immunol 6(1):57–64
Glaser R, Meyer-Hoffert U, Harder J, Cordes J, Wittersheim M et al (2009) The antimicrobial protein psoriasin (S100A7) is upregulated in atopic dermatitis and after experimental skin barrier disruption. J Invest Dermatol 129(3):641–649
Grewe M, Bruijnzeel-Koomen CA, Schopf E, Thepen T, Langeveld-Wildschut AG et al (1998) A role for Th1 and Th2 cells in the immunopathogenesis of atopic dermatitis. Immunol Today 19(8):359–361
Hardas BD, Zhao X, Zhang J, Longqing X, Stoll S et al (1996) Assignment of psoriasin to human chromosomal band 1q21: coordinate overexpression of clustered genes in psoriasis. J Invest Dermatol 106(4):753–758
Heizmann CW, Fritz G, Schafer BW (2002) S100 proteins: structure, functions and pathology. Front Biosci 7:d1356–d1368
Kohn EC, Liotta LA (1995) Molecular insights into cancer invasion: strategies for prevention and intervention. Cancer Res 55(9):1856–1862
Kulski JK, Lim CP, Dunn DS, Bellgard M (2003) Genomic and phylogenetic analysis of the S100A7 (Psoriasin) gene duplications within the region of the S100 gene cluster on human chromosome 1q21. J Mol Evol 56(4):397–406
Lee KC, Eckert RL (2007) S100A7 (Psoriasin)—mechanism of antibacterial action in wounds. J Invest Dermatol 127(4):945–957
Liu J, Li X, Dong GL, Zhang HW, Chen DL et al (2008) In silico analysis and verification of S100 gene expression in gastric cancer. BMC Cancer 8:261
Madsen P, Rasmussen HH, Leffers H, Honore B, Dejgaard K et al (1991) Molecular cloning, occurrence, and expression of a novel partially secreted protein “psoriasin” that is highly up-regulated in psoriatic skin. J Invest Dermatol 97(4):701–712
Martinsson H, Yhr M, Enerback C (2005) Expression patterns of S100A7 (psoriasin) and S100A9 (calgranulin-B) in keratinocyte differentiation. Exp Dermatol 14(3):161–168
Mischke D, Korge BP, Marenholz I, Volz A, Ziegler A (1996) Genes encoding structural proteins of epidermal cornification and S100 calcium-binding proteins form a gene complex (“epidermal differentiation complex”) on human chromosome 1q21. J Invest Dermatol 106(5):989–992
Moroz OV, Burkitt W, Wittkowski H, He W, Ianoul A et al (2009) Both Ca2+ and Zn2+ are essential for S100A12 protein oligomerization and function. BMC Biochem 10:11
Moubayed N, Weichenthal M, Harder J, Wandel E, Sticherling M et al (2007) Psoriasin (S100A7) is significantly up-regulated in human epithelial skin tumours. J Cancer Res Clin Oncol 133(4):253–261
Numerof RP, Asadullah K (2006) Cytokine and anti-cytokine therapies for psoriasis and atopic dermatitis. BioDrugs 20(2):93–103
Ramasamy R, Yan SF, Herold K, Clynes R, Schmidt AM (2008) Receptor for advanced glycation end products: fundamental roles in the inflammatory response: winding the way to the pathogenesis of endothelial dysfunction and atherosclerosis. Ann N Y Acad Sci 1126:7–13
Robinson NA, Lapic S, Welter JF, Eckert RL (1997) S100A11, S100A10, annexin I, desmosomal proteins, small proline-rich proteins, plasminogen activator inhibitor-2, and involucrin are components of the cornified envelope of cultured human epidermal keratinocytes. J Biol Chem 272(18):12035–12046
Ruse M, Broome AM, Eckert RL (2003) S100A7 (psoriasin) interacts with epidermal fatty acid binding protein and localizes in focal adhesion-like structures in cultured keratinocytes. J Invest Dermatol 121(1):132–141
Sabat R, Philipp S, Hoflich C, Kreutzer S, Wallace E et al (2007) Immunopathogenesis of psoriasis. Exp Dermatol 16(10):779–798
Santilli F, Vazzana N, Bucciarelli LG, Davi G (2009) Soluble forms of RAGE in human diseases: clinical and therapeutical implications. Curr Med Chem 16(8):940–952
Schroder JM, Harder J (2006) Antimicrobial skin peptides and proteins. Cell Mol Life Sci 63(4):469–486
Semprini S, Capon F, Bovolenta S, Bruscia E, Pizzuti A et al (1999) Genomic structure, promoter characterisation and mutational analysis of the S100A7 gene: exclusion of a candidate for familial psoriasis susceptibility. Hum Genet 104(2):130–134
Semprini S, Capon F, Tacconelli A, Giardina E, Orecchia A et al (2002) Evidence for differential S100 gene over-expression in psoriatic patients from genetically heterogeneous pedigrees. Hum Genet 111(4–5):310–313
Sourris KC, Forbes JM (2009) Interactions between advanced glycation end-products (AGE) and their receptors in the development and progression of diabetic nephropathy—are these receptors valid therapeutic targets. Curr Drug Targets 10(1):42–50
South AP, Cabral A, Ives JH, James CH, Mirza G et al (1999) Human epidermal differentiation complex in a single 2.5 Mbp long continuum of overlapping DNA cloned in bacteria integrating physical and transcript maps. J Invest Dermatol 112(6):910–918
Steinert PM, Kim SY, Chung SI, Marekov LN (1996) The transglutaminase 1 enzyme is variably acylated by myristate and palmitate during differentiation in epidermal keratinocytes. J Biol Chem 271(42):26242–26250
Taguchi A, Blood DC, del Toro G, Canet A, Lee DC et al (2000) Blockade of RAGE-amphoterin signalling suppresses tumour growth and metastases. Nature 405(6784):354–360
Wolf R, Mirmohammadsadegh A, Walz M, Lysa B, Tartler U et al (2003) Molecular cloning and characterization of alternatively spliced mRNA isoforms from psoriatic skin encoding a novel member of the S100 family. FASEB J 17(13):1969–1971
Wolf R, Lewerenz V, Buchau AS, Walz M, Ruzicka T (2007) Human S100A15 splice variants are differentially expressed in inflammatory skin diseases and regulated through Th1 cytokines and calcium. Exp Dermatol 16(8):685–691
Wolf R, Howard OM, Dong HF, Voscopoulos C, Boeshans K et al (2008) Chemotactic activity of S100A7 (Psoriasin) is mediated by the receptor for advanced glycation end products and potentiates inflammation with highly homologous but functionally distinct S100A15. J Immunol 181(2):1499–1506
Wolf R, Voscopoulos C, Winston J, Dharamsi A, Goldsmith P et al (2009) Highly homologous hS100A15 and hS100A7 proteins are distinctly expressed in normal breast tissue and breast cancer. Cancer Lett 277(1):101–107
Yan SD, Bierhaus A, Nawroth PP, Stern DM (2009) RAGE and Alzheimer’s disease: a progression factor for amyloid-beta-induced cellular perturbation? J Alzheimers Dis 16(4):833–843
Yao R, Lopez-Beltran A, Maclennan GT, Montironi R, Eble JN et al (2007) Expression of S100 protein family members in the pathogenesis of bladder tumors. Anticancer Res 27(5A):3051–3058
Zen K, Chen CX, Chen YT, Wilton R, Liu Y (2007) Receptor for advanced glycation endproducts mediates neutrophil migration across intestinal epithelium. J Immunol 178(4):2483–2490
Zhang H, Wang Y, Chen Y, Sun S, Li N et al (2007) Identification and validation of S100A7 associated with lung squamous cell carcinoma metastasis to brain. Lung Cancer 57(1):37–45
Zhang XJ, Huang W, Yang S, Sun LD, Zhang FY et al (2009) Psoriasis genome-wide association study identifies susceptibility variants within LCE gene cluster at 1q21. Nat Genet 41(2):205–210
Zimmer DB, Wright Sadosky P, Weber DJ (2003) Molecular mechanisms of S100-target protein interactions. Microsc Res Tech 60(6):552–559
Acknowledgments
This work was supported by grants from the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research, and the German Research Foundation (DFG).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wolf, R., Ruzicka, T. & Yuspa, S.H. Novel S100A7 (psoriasin)/S100A15 (koebnerisin) subfamily: highly homologous but distinct in regulation and function. Amino Acids 41, 789–796 (2011). https://doi.org/10.1007/s00726-010-0666-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-010-0666-4