Abstract
Despite the fact that when subjected to salinity stress most plants accumulate high concentrations of sodium (Na+) and chloride (Cl−) ions in their tissues, major research has however been focused on the toxic effects of Na+. Consequently, Cl− toxicity mechanisms in annual plants, particularly in inducing oxidative stress, are poorly understood. Here, the extent to which Na+ and/or Cl− ions contribute in inducing oxidative stress and regulating the adaptive antioxidant defense is shown in two Indica rice genotypes differing in their salt tolerance. Equimolar (100 mM) concentrations of Na+, Cl−, and NaCl (EC ≈ 10 dS m−1) generated free-radical (O2 •−, •OH) and non-radical (H2O2) forms of reactive oxygen species (ROS) and triggered cell death in leaves of 21-day-old hydroponically grown rice seedlings as evident by spectrophotometric quantifications and histochemical visualizations. The magnitude of ROS-mediated oxidative damage was higher in sensitive cultivar, whereas NaCl proved to be most toxic among the treatments. Salt treatments significantly increased activities of antioxidant enzymes and their isozymes including superoxide dismutase, catalase, peroxidase, ascorbate peroxidase, and glutathione reductase. Na+ and Cl− ions showed additive effects under NaCl in activating the antioxidant enzyme machinery, and responses were more pronounced in tolerant cultivar. The expression levels of SodCc2, CatA, and OsPRX1 genes were largely consistent with the activities of their corresponding enzymes. Salt treatments caused an imbalance in non-enzymatic antioxidants ascorbic acid, α-tocopherol, and polyphenols, with greater impacts under NaCl than Na+ and Cl− separately. Results revealed that though Cl− was relatively less toxic than its counter-cation, its effects cannot be totally ignored. Both the cultivars responded in the same manner, but the tolerant cultivar maintained lower Na+/K+ and ROS levels coupled with better antioxidant defense under all three salt treatments.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Salinity stress greatly limits the growth and yield of major crops, and its high scale of impact coupled with wide distribution makes it one of the most severe global environmental threats for crop production. The problem has been aggravated of late, owing to the climate changes, agricultural practices including irrigation with saline water, faulty water management, excessive use of fertilizers, and rising water tables (Munns et al. 2012; Huang et al. 2013). Global food requirements are estimated to increase by 70–110 %, and with increasing land degradation, urban spread, and sea water intrusion, gains in agricultural productivity are a must for the saline lands (Tilman et al. 2011; Munns et al. 2012). Production of salt-tolerant plants necessitates deciphering various complicated physiological and molecular mechanisms; plants develop in response to and to counteract the deleterious effects of salt stress.
The effects of salinity stress are generally threefold as reduced plant growth and yield takes place through osmotic stress, ion toxicities and imbalances, and oxidative stress (Kumar et al. 2009). An unfortunate consequence of salinity stress in plants is the excessive generation of reactive oxygen species (ROS) due to decreased stomatal conductivity under low water potential, over-reduction of electron transport in cellular organelles, and accumulation of Na+ and/or Cl− ions (Turkan and Demiral 2009). These ROS are major causative factors of oxidative damage to lipid membranes and other essential macromolecules including pigments, proteins, DNA, and RNA and ultimately provokes cell death (Qureshi et al. 2013; Yildiztugay et al. 2014). However, they also have the ability to work as signaling molecule and regulate responses of development as well as various aspects of stress (Ismail et al. 2014).
Plants respond to salinity stress through a set of combating mechanisms. Among them, selective accumulation, exclusion, and/or compartmentalization of ions at the whole plant and cellular levels, synthesis of compatible solutes, and induction of enzymatic and non-enzymatic antioxidants for ROS scavenging are considered crucial biochemical strategies (Qureshi et al. 2013; Yildiztugay et al. 2014). Both enzymatic and non-enzymatic processes participate in the detoxification of ROS (Munns and Tester 2008).
Low molecular weight antioxidants, both hydrophilic, like ascorbic acid (AsA), reduced glutathione (GSH), flavonoids, and phenolic compounds, and lipophilic, including α-tocopherol and carotenoids, can quench all types of ROS. However, in spite of various roles these non-enzymatic antioxidants play, relatively very few are well characterized and have been attributed to enhanced plant salt tolerance. Antioxidant enzymes, on the other hand, like superoxide dismutase (SOD), catalase (CAT), peroxidase (POX), and glutathione reductase (GR), are well documented for detoxifying ROS. These enzymes can be divided into two categories: one that reacts with ROS and keeps them at low levels (POX, SOD, and CAT) and one that regenerates the oxidized antioxidants (ascorbate peroxidase, APX, and glutathione reductase, GR) to maintain redox balance (Kumar et al. 2009). SOD decomposes O2 •− to H2O2, which is further scavenged by POX in extracellular space and cytosol and by CAT in peroxisomes. APX, an Asada–Halliwel enzyme, also decomposes H2O2 in cellular compartments. These enzymatic and non-enzymatic antioxidants contribute significantly in maintaining the proper functioning of metabolic pathways and integrity of cell structures via reacting directly or indirectly with ROS (Gill and Tuteja 2010). The stimulation of antioxidant defense is one of the major components of tolerance mechanism against oxidative damage, and in accordance, salt tolerance of various plants is often correlated with their better antioxidant capacities (Sairam et al. 2005; Qureshi et al. 2013). However, in order to assess the tolerance mechanisms and threshold of a particular plant, the processes underlying the antioxidant responses to oxidative stress need to be clearly understood.
The principal cations of total soluble salts in saline soils comprise sodium (Na+), calcium (Ca2+), and magnesium (Mg2+), whereas major anions are chloride (Cl−), sulfate (SO4 2−), and carbonates (including bicarbonates) (Tavakkoli et al. 2011). However, NaCl is the most prevalent salt (making 50 to 80 % of the total soluble salts) where Na+ dominates the cations and Cl− dominates the anions in majority of saline soils (Munns and Tester 2008). It is well documented that most plants accumulate high Na+ and Cl− concentrations under salinity stress and their high cytoplasmic concentrations are metabolically toxic to plants (Teakle and Tyerman 2010; Tavakkoli et al. 2011). Therefore, both these ions deserve equal consideration; however, most research on salt stress tolerance in cereals has been focused on Na+ (Kumar and Khare 2014). Consequently, the role of high Cl− concentrations in salt stress and the tolerance mechanisms involved therein are poorly understood as compared with that of Na+. Though Cl− is an essential micronutrient which plays important roles in regulating enzyme activities in cytoplasm, stomatal closure, acts as a co-factor in photosynthesis, and is involved in turgor and pH regulation (Xu et al. 2000; White and Broadley 2001), it exerts toxic effects on plants at high concentrations (Moya et al. 2003). There is a recent argument about the importance of Cl− ions under salinity stress and its recognition as a vital cause of damage (Teakle and Tyerman 2010), and in some cases, its concentration in soil has been ascribed as more important to growth and yield reduction than Na+ (Dang et al. 2008; Zhang et al. 2011). However, very few attempts have been made so far to evaluate the extent to which Na+ and Cl− contribute to ion toxicity besides their possible additive effects under NaCl and whether salt-tolerant genotypes have more differential mechanisms than their sensitive counterparts in this regard.
The present investigation was undertaken with the following objectives: first, to assess the extent to which Na+ and Cl− contribute individually in the generation of ROS leading to oxidative stress damages and cell death besides their possible additive effects on two rice cultivars with contrasting salt tolerance; second, to study the detailed non-enzymatic and enzymatic antioxidant defense responses (including expression of genes associated with antioxidant defense) of rice cultivars under Cl−, Na+, and NaCl stress; and third, to investigate whether tolerant genotypes have better abilities to exclude Na+ and Cl− from their shoot tissues and maintain relatively higher K+ concentrations coupled with better enzymatic and non-enzymatic antioxidant capacities under salinity stress.
Materials and methods
Plant material, salt treatments, and growth conditions
Seeds of two rice (Oryza sativa L.) cultivars, Panvel-3 (salt tolerant) and Sahyadri-3 (salt sensitive), were kindly provided by the Saline Land Research Station, Panvel, India, and the Regional Rice Research Station, Karjat, India, respectively. The dehusked seeds were washed thoroughly with water containing few drops of Tween-20 for 10 min and then rinsed five times with distilled water. Seeds were surface-sterilized with 0.1 % HgCl2 for 5 min and rinsed several times before placing them on wet filter paper for 2 days in dark condition for germination. Equally grown seedlings were transferred to a hydroponic culture system. Yoshida’s nutrient solution (YNS) for rice was used as a background to evaluate individual and additive effects of Na+ and Cl− ions on rice cultivars. The experiment compared the effect of 100 mM each of Na+ (applied as a range of sulfate, nitrate, and phosphate salts), Cl− (applied as a range of calcium, magnesium, and potassium salts), and 100 mM NaCl at similar electrical conductivity (EC) (9.6 ± 0.5) and pH (6.0) as described by Tavakkoli et al. (2010). Four treatments were compared which consisted of control (YNS medium, no amendment), Na+-dominant salts (YNS plus 15 mM Na2SO4, 15 mM Na2HPO4, 40 mM NaNO3), Cl−-dominant salts (YNS plus 15 mM CaCl2, 15 mM MgCl2, 40 mM KCl), and NaCl (YNS plus 100 mM NaCl). Milli-Q water was used to dissolve the salts and for preparation of culture medium. Seedlings were kept at 25 ± 2 °C and 80–85 % relative humidity. Samples (first leaves) were collected for different experiments on the 21st day after treatments. Data obtained are from three replicates under identical conditions.
Determination of ion content
Leaves were analyzed for mineral nutrients by modified Anil et al. (2005) method; briefly, tissues were washed with distilled water to remove any surface contaminants followed by drying at 70 °C for 48 h. The dried tissues were ground into powder in liquid nitrogen, which was acid digested overnight in 5 ml of concentrated nitric acid. Five milliliters of a 10:4 diacid mixture of nitrate and perchlorate was added to the partially digested tissue powder and allowed to completely digest for 2 h on a sand bath. Na+ and K+ were measured by atomic absorption spectrophotometer (AA201; Chemito, India), while Cl was estimated by potentiometric method (Chapman and Pratt 1961).
In-leaf estimation and histochemical visualization of ROS
Cellular generation of O2 ˙ − (Chaitanya and Naithani, 1994), H2O2 (Loreto and Velikova 2001), and •OH (Halliwell et al. 1987) were measured using UV-visible spectrophotometer (UV-1800; Shimadzu Corp., Japan) and visualized histochemically in leaf (Romeo-Puertas et al. 2004). For measuring O2˙−, 100 mg fresh tissues was homogenized in cold phosphate buffer (0.2 M, pH 7.2) containing 1 mM sodium diethyl dithiocarbamate and centrifuged at 10,000×g at 4 °C for 10 min. The reaction mixture containing 250 μL extract, 100 μL of 2.25 mM NBT, 50 μL of 1.5 M Na2CO3, 100 μL of 3 mM EDTA, 200 μL 200 mM l-methionine, and 2.3 mL distilled water was allowed to incubate at 30 °C for 30 min. The O2 ˙ − content is expressed as ΔA540 min−1 g−1 fresh weight (FW).
For in-leaf visualization of O2˙−, 21-day-old leaves of equal size were immersed in a 6-mL reaction mixture containing 50 mM Tris–HCl buffer (pH 6.4), 0.2 mM nitro blue tetrazolium (NBT), 0.2 mM NADH, and 250 mM sucrose, vacuum-infiltrated for 15 min and illuminated at 200 μM m−2 s−1 for 24 h to develop color, characteristic of blue monoformazan precipitation. Stained leaves were bleached in 95 % ethanol at 90 °C for 15–20 min to localize cellular O2 ˙ − generation and were macro-photographed against white fluorescent light background.
For measurement of H2O2, 100 mg leaves was homogenized in 1 mL trichloroacetic acid (TCA, 0.2 %) at 4 °C. The homogenate was centrifuged at 12,000×g for 15 min and 0.5 mL of supernatant was mixed with 1 mL of 10 mM sodium phosphate buffer (pH 7) and 2 mL of 1 M KI. The H2O2 content was measured by comparing its absorbance at 390 nm with a standard calibration curve.
For histochemical visualization of H2O2, leaves of equal size were immersed in 1 % solution of 3,3′-diaminobenzidine (DAB) (pH 3.8), vacuum infiltrated for 10–15 min, and then incubated at room temperature for 8 h in absence of direct light. Leaves were then illuminated until the appearance of brown color, characteristic of polymerization reaction between DAB and H2O2. Following the bleaching, the leaves were macro-photographed against white fluorescent light.
For estimating •OH content, 100 mg leaves was homogenized in 1 mL of 10 mM phosphate buffer (pH 7.4) containing 15 mM 2-deoxy-d-ribose and centrifuged at 12,000×g for 15 min. The supernatants were incubated at 37 °C for 2 h. Aliquots of 0.2 mL of the supernatant were added to the reaction mixture containing 3 mL of 0.5 % (w/v) thiobarbituric acid (TBA, 1 % stock solution made in 5 mM NaOH) and 1 mL glacial acetic acid, heated at 100 °C in water bath for 30 min. The mixture was cooled to 4 °C. The absorbance of malondialdehyde (MDA) was measured at 532 nm and concentration was calculated using extinction coefficient (ε = 155 mM−1 cm−1) and expressed in nanomoles per gram FW.
In-leaf estimation and histochemical visualization of cell death
Cell death was visualized in leaf tissues histochemically as well as quantified by Evan’s blue staining method (Romeo-Puertas et al. 2004). Freshly excised leaves were immersed and vacuum infiltrated in 0.25 % (w/v) aqueous solution of Evan’s blue for 6 h and left overnight in the same solution. The leaves were then bleached in boiling 95 % ethanol to remove chlorophyll. Cell death was visualized as blue patches on the leaf surface and were macro-photographed against white fluorescent light. For quantification of cell death, five stained leaves weighing about 100 mg from each treatment were immersed in dye-extraction solution consisting of 50 % ethanol in 1 % SDS (sodium dodecyl sulfate) for 1 h at 50 °C. The absorbance of Evan’s blue released into solution was recorded at 600 nm.
Antioxidant enzyme extraction and assays
For enzyme extraction, freshly harvested leaves (500 mg) were ground to fine powder in liquid nitrogen using mortar and pestle, and then homogenized with 100 mM phosphate buffer (pH 7.5) containing 0.5 mM EDTA. The homogenate was centrifuged at 15,000×g for 20 min at 4 °C. Supernatant was collected as a crude enzyme extract for further enzyme measurements. All the steps were carried out at 0–4 °C. The absorbance for all the enzymes was taken on a UV-VIS spectrophotometer (UV-1800; Shimadzu Corp., Tokyo, Japan).
SOD (EC 1.15.1.1) activity was estimated by recording the decrease in absorbance of formazan produced by superoxide–nitro blue tetrazolium (NBT) complex by the enzyme as described by Dhindsa et al. (1981). The three-milliliter reaction mixture contained 0.1 mL of 1.5 M sodium carbonate, 0.2 mL of 200 mM methionine, 0.1 mL of 3 mM EDTA, 0.1 mL of 2.25 mM NBT, 1.5 mL of 100 mM phosphate buffer, 0.9 mL distilled water, and 0.1 mL enzyme taken in the test tubes in duplicate from each enzyme sample. Reaction was started by adding 0.1 mL riboflavin (60 mM) and placing the tubes under two 15-W florescent lamps for 15 min. The reaction was stopped by switching off the light and covering the tubes with black cloth. The tube without enzyme developed maximal color (control) while the non-irradiated tube did not develop any color and served as blank. Absorbance was recorded at 560 nm and 1 U of enzyme activity was taken as the quantity of enzyme which reduced the absorbance reading of samples by 50 % in comparison with tubes lacking enzymes.
CAT (EC 1.11.1.6) activity was determined according to Aebi (1984) by measuring the rate of decrease in absorbance of H2O2 at 240 nm. The reaction mixture consists of 50 mM phosphate buffer, 12.5 mM H2O2, and 50 μL enzyme.
POX (EC 1.11.1.7) activity was measured in terms of increase in absorbance at 470 nm due to the oxidation of guaiacol to tetra-guaiacol (Castillo et al. 1984). The reaction mixture contained 50 mM phosphate buffer (pH 6.1), 16 mM guaiacol, 2 mM H2O2, and 0.1 mL enzyme extract.
GR (EC 1.6.4.2) activity was assayed by following Smith et al. (1988) by recording the increase in the presence of oxidized glutathione and DTNB (5,5′-dithiobis-2-nitrobenzoic acid). The three-milliliter reaction mixture contained 1 mL of 0.2 M potassium phosphate buffer (pH 7.5) containing 0.1 mM EDTA, 0.5 mL of 3 mM DTNB in 10 mM potassium phosphate buffer (pH 7.5), 0.1 mL of 2 mM NADPH, 0.1 mL enzyme extract, and 0.3 mL distilled water. The increase in absorbance at 412 nm was recorded over a period of 5 min.
APX (EC 1.11.1.11) was assayed by recording the decrease in absorbance due to ascorbic acid at 290 nm (Nakano and Asada 1981) in a 3-mL reaction mixture which contained phosphate buffer (50 mM, pH 7.0), ascorbic acid (0.5 mM), 0.1 mM EDTA, 0.1 mM H2O2, and 100 μL enzyme extract. Reaction was initiated by addition of H2O2 and decrease in absorbance was recorded at 30-s time interval.
Gel electrophoresis and identification of isozymes
Isozymes were separated on non-denaturing (native) polyacrylamide gels at 4 °C using a PROTEAN-II xi-Cell Electrophoresis Unit (BIO-RAD, USA). Estimation of proteins was carried out prior to loading by Bradford (1976) method.
A protein concentration of 50 µg from leaves under non-saline and saline conditions was applied into each well and then electrophoresed at 30 mA for 2 h. SOD activity staining was performed using modified Beauchamp and Fridovich (1971) method and inhibitor studies to distinguish various SOD isoforms as standardized in the authors’ laboratory (Kumar et al. 2009). The gel was soaked in 50 mL of 100 mM potassium phosphate buffer, pH 7.5 containing 1.23 mM NBT, 0.02 mM riboflavin, and 28 mM TEMED for 20 min in the dark and then illuminated under light source to initiate the photochemical reaction.
CAT isoenzymes were separated on 10 % non-denaturing acrylamide gel run at 30 mA for 24 h at 4 °C and were visualized by following Woodbury et al. (1971). Gels were rinsed in double distilled water (DDW) followed by incubation in 0.01 % H2O2 for 5 min, washed twice with DDW, and then stained in a solution of 0.5 % potassium ferricyanide and 0.5 % ferric chloride. The reaction was terminated with 1 % HCl after development of bands.
POX isozymes separation and staining was carried out as per Anderson et al. (1995). POX isozymes were separated on 8 % non-denaturing polyacrylamide gel and the electrophoresis was done at 30 mA for 150 min at 4 °C. Isoforms of POX were visualized by staining the gel in a 50-mL solution containing 100 mM potassium phosphate buffer (pH 6.4), 20 mM guaiacol, and 5.55 mM H2O2.
GR isoenzymes were separated on non-denaturing 8 % gel at 50 mA for 3 h at 4 °C and gels were stained in 50 mL of 250 mM Tris–HCl buffer (pH 7.5) containing 10 mg MTT, 3 mL of 3.4 mM DCPIP, 18.6 mg NADPH, and 52 mg oxidized glutathione (Anderson et al. 1990).
APX isozymes were visualized in-gel following Mittler and Zilinskas (1993). Resolving gel (8 %) containing 10 % glycerol was used for separation of isozymes. Electrophoresis was carried out at 50 mA for 2 h at 4 °C. Gels were stained in phosphate buffer (100 mM, pH 7.0) with 4 mM ascorbic acid and 2 mM H2O2 until desired contrast was achieved.
RNA extraction and gene expression of antioxidant enzymes
Freshly harvested leaf samples from hydroponically cultivated plants were ground in mortar and pestle containing liquid nitrogen, and total RNA was isolated using silica-based spin columns following the manufacturer’s instructions (GeneiPureTM Total RNA isolation Kit—Plants; GeneiTM, Bangalore, India). The purified total RNAs were quantified spectrophotometrically at 260 nm and the purity was checked by means of the A260/A280 ratio. The integrity and quality of the total RNA were verified by running the samples on 1 % denaturing agarose gel containing 2 M formaldehyde. Semi-quantitative RT-PCR was performed using One Step M-MuLV RT-PCR Kit (GeneiTM, Bangalore, India) according to the manufacturer’s instructions. First strand of cDNA was synthesized using 2 μg of total RNA by incubating the reaction mixture at 50 °C for 30 min. PCR reaction was performed using a thermal cycler (Mastercycler® personal, Eppendorf). Twenty-five microliters of reaction mixture contained 10 pmol of specific primers for the genes SodCc2, CatA, OsPRX1, and GAPDH (rice glyceraldehyde 3-phosphate dehydrogenase) as internal control (Table 1). PCR amplification reaction were initially incubated at 95 °C for 5 min, followed by 35 cycles at 94 °C for 30 s, 57–59 °C (depending upon the melting temperature of each gene-specific primer) for 30 s, and 72 °C for 45 s. Final extension was performed at 72 °C for 10 min. Amplified products were loaded and separated on 1 % agarose gel spiked with ethidium bromide. The gel was visualized under UV light and photographed using Gel Doc XR documentation system (BIO-RAD, USA).
Estimation of non-enzymatic antioxidants
Ascorbic acid was determined according to Mukharjee and Choudhari (1983). Fresh leaf sample (500 mg) was extracted in 10 mL of 6 % TCA. The homogenate was centrifuged at 10,000×g at 4 °C for 10 min and 4 mL of supernatant was mixed with 2 mL of 2 % dinitrophenylhydrazine (in acidic medium) followed by addition of one drop of 10 % thiourea prepared in 70 % ethanol. Reaction mixture is heated in boiling water for 15 min, and after cooling to room temperature, 5 mL of 80 % sulfuric acid was added to the mixture at 0 °C. The absorbance was recorded at 530 nm and concentration was calculated from standard curve plotted with known concentrations of ascorbic acid.
The α-tocopherol levels in the plant samples were estimated spectrophotometrically (Rosenberg 1992). Plant samples (2.5 g) were homogenized in 5 mL of 0.1 N sulfuric acid and volume was finally made up to 50 mL by adding 0.1 N sulfuric acid without shaking, and the content was allowed to stand overnight. Next day, the contents in flask were shaken vigorously and filtered through Whatman No. 1 filter paper. Aliquots of filtrate were used for estimation of α-tocopherol. To 1.5 mL of aliquot, 1.5 mL each of absolute ethanol and xylene were added, stoppered, mixed well, and centrifuged. After centrifugation, 1 mL of xylene layer was transferred into another tube carefully followed by addition of 1 mL of 2,2′-dipyridyl reagent (1.2 % in 1-propanol). Absorbance of mixture was read against blank at 460 nm. Then, in turn, beginning with the blank, 0.33 mL of ferric chloride solution (1.2 % in ethanol) was added, mixed well, and absorbance read at 520 nm after 15 min. Tocopherol levels were calculated using d, l-α-tocopherol acetate as standard.
The levels of reduced glutathione (GSH) were estimated by the method of Moron et al. (1979). Plant tissue (0.5 g) was homogenized in 2.5 mL of 5 % TCA. The homogenate was immediately acidified by adding 125 μL of 25 % TCA to prevent aerial oxidation of glutathione. Precipitated protein was centrifuged at 10,000×g for 10 min. Homogenate was cooled and 0.1 mL of supernatant was used for estimation by diluting it up to 1 mL with 0.2 M sodium phosphate buffer (pH 8.0). Two milliliters of freshly prepared DTNB solution was added to the tubes and intensity of the yellow color was read at 412 nm after 10 min. Standard curve was plotted using GSH and values were expressed as nanomoles of GSH per gram of tissue.
Total phenols were estimated according to Malick and Singh (1980). The tissue homogenate was prepared in 80 % ethanol and centrifuged at 10,000×g for 20 min. The residue was re-extracted with 80 % ethanol, and supernatants were pooled and evaporated to dryness. The residue was dissolved in known volume of distilled water. Aliquots were diluted to 3 mL followed by addition of Folin–Ciocalteu reagent. After 5 min, 2 mL of 20 % Na2CO3 was added to each tube. After thorough mixing, tubes were kept in boiling water bath for 1 min and allowed to cool. The absorbance was recorded at 650 nm. Total phenols were calculated using catechol as standard.
Flavonoids were estimated by the method of Cameron et al. (1943). Plant samples were extracted first with methanol/water mixture (2:1) and secondly with the same mixture in a ratio of 1:1. Extracts were mixed well and allowed to stand overnight; supernatants were pooled and volume was noted. An aliquot of the above extract was pipetted out and evaporated to dryness. Four milliliters of vanillin reagent was added and tubes were heated for 15 min in boiling water. Absorbance was read at 340 nm and total flavonoids were calculated using standard catechin.
Statistical analyses
The experiment was designed as factorial to study the individual and/or additive effects of Cl− and Na+ ions on two rice cultivars. The results are presented as means ± standard error, and means were compared using Duncan’s multiple range test (DMRT) at P ≤0.05 using MSTAT-C statistical software. The graphs were plotted using Microcal Origin 6.0 software.
Results and discussion
Effects of Cl−, Na+, and NaCl on dry matter and ion accumulation
Since NaCl is the most abundant soluble salt of saline soils, its constituent ions Na+ and Cl− have understandably become two most important toxic components of ionic stress. However, plant’s integral tolerance to these ions differs, and the relative value often varies among the plant species (Zhang et al. 2011). In order to assess the relative impact of NaCl and its individual constituents (Cl− and Na+), two rice cultivars with contrasting salt tolerance abilities were subjected to 100 mM solutions of each salt with similar EC. Four treatments consisting of control (YNS, no amendment), Na+-dominant salts (YNS plus 15 mM Na2SO4, 15 mM Na2HPO4, 40 mM NaNO3), Cl−-dominant salts (YNS plus 15 mM CaCl2, 15 mM MgCl2, 40 mM KCl), and NaCl (YNS plus 100 mM NaCl) were used to initiate the desired conditions. Since Na+ and Cl− are combined with their counter-anions/counter-cations in natural soils, getting them solely is difficult (Luo et al. 2005). Therefore, to stimulate the ion states of Na+ and Cl−, their different salt combinations were applied for maintaining their equimolar concentrations and to avoid the increase of particular counter-anions/counter-cations, besides maintaining the EC as suggested by Tavakkoli et al. (2011). However, despite the usefulness of this approach, unlike control and Na+- and NaCl-stressed plants, the possibility of enrichment of Cl−-stressed plants with some nutrient ions such as potassium, magnesium, and calcium cannot be ruled out, owing to the use of their respective salts. However, Yoshida’s nutrient solution (YNS) was used instead of water as a background for all the treatments to keep this disparity under check as well as to avoid other nutritional deficiencies or toxicities in plants other than those targeted by the salts.
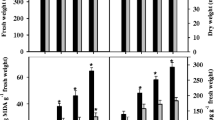
Though both Cl− and Na+ caused significant reductions in dry matter production (Table 2), the degree of DW reduction did not vary significantly (at P ≤ 0.05) between the individual ions. In contrast, NaCl induced a sharp decrease in DW. Genotypic differences were observed in salinity-induced growth reductions, with 28 % DW reduction in Panvel-3 against 48 % in Sahyadri-3 at 100 mM NaCl stress than their respective controls. Decrease in dry matter under the influence of ionic stress is a well-documented phenomenon (Kamyab-Talesh et al. 2014). The results revealed that ionic stresses significantly diminished the biomass production, indicated by reduced DW in the following manner: NaCl > Na+ > Cl−, suggesting possible additive effects of individual Cl− and Na+ ions on rice seedlings. The use of nutrient medium with similar EC and molar concentrations but different combinations of Na+ and Cl− dominant salts may be attributed to the comparative effects of these ions on dry matter accumulation. These trends are in confirmation of earlier findings where greater reduction in biomass occurred in rice (Lin and Kao 2001), faba bean (Tavakkoli et al. 2010), and barley (Tavakkoli et al. 2011) treated with NaCl than its constituent individual ions.
The endogenous levels of Cl− and Na+ ions increased under individual and combined ionic stresses, but rice cultivars showed considerable genotypic variations (Table 2). Application of Na+-dominant salts resulted into a 4.75-fold more Na+ content in Panvel-3 against 7.17 times higher Na+ in Shaydri-3 seedlings as compared to their respective controls. Similar trends were evidenced with Cl− treatments where Panvel-3 and Sahyadri-3 seedlings accumulated 2.69 and 3.25 times higher Cl− content than their non-saline counterparts, respectively. This indicates the inability of sensitive cultivar to restrict the entry of ions into its leaf tissues under salinity stress. Among three salt treatments, the highest magnitude of ion accumulation occurred upon exposure of each cultivar to NaCl, suggesting combined effects of the constituent ions. Panvel-3 showed its ability to maintain Na/K ratio below 1.0 (under Na as well as NaCl stress), which is considered as critical to maintain metabolic activities (Tavakkoli et al. 2011). In contrast, Sahyadri-3 failed to maintain higher K+ levels under applied stresses and showed higher Na+/K+ ratio both under Na+ (1.62) and NaCl (2.20) stresses. Better growth performance by tolerant plants usually gets correlated with their ability to maintain low Na+ concentrations in their tissues (Munns et al. 2012). Similarly, acquisition of K+ ions in order to maintain a low Na/K ratio has always been a key feature of salt-tolerant crop plants (Khan et al. 2009). Lower K+ concentrations under Na+ and NaCl stresses in both the cultivars might be attributed to the leakage of K+ by the outward rectifying channels.
Effect of Cl−, Na+, and NaCl stress on ROS generation and cell death
Though there are numerous reports on NaCl-induced ROS generation and concurrent antioxidative defense machinery employed by the plants to cope with the deleterious effects of NaCl, however, to our knowledge, this is the first report assessing the individual and combined effects of Na+ and Cl− ions on the relative induction of oxidative stress, cell death, and responsive antioxidant defense in the leaf tissues of two rice cultivars with contrasting salinity tolerance abilities.
It is a well-established fact now that plant cells maintain a delicate balance between ROS generation and their scavenging, largely via enzymatic and non-enzymatic antioxidant machinery. At lower levels, they do not exert toxic effects on plant cells and tissues and even work as signaling molecules. However, this balance gets perturbed under stressful conditions including salinity, and this disturbed equilibrium leads to sudden upsurge in intracellular ROS levels, which ultimately causes significant damages to cell structures besides compromising their functional abilities (Gill and Tuteja 2010). One of the major objectives of this work was to establish a missing link between the roles played by Na+ and Cl− ions (individually and in combination) in ROS generation (and resultant oxidative damage), besides elucidating whether tolerant cultivars behave differently than their sensitive counterparts in this regard.
Levels of ROS generation, a measure of oxidative eruption under adverse conditions, were monitored by spectrophotometric analyses along with their histochemical visualization in the first leaf tissues of 21-day-old rice seedlings grown under salinity treatments. Histochemical localization of O2 •− as indicated by the formation of blue-colored formazan complex (reduced NBT), well supported by the data obtained via spectrophotometric analysis, showed that all three types of equimolar salt treatments stimulated superoxide anion production in rice leaves. However, much higher O2 •− accumulation occurred in leaves under NaCl stress as compared to its constituent Cl− and Na+ ions in both cultivars (Fig. 1).
Cl−, Na+, and NaCl induced O2 •− generation in the leaves of 21-day-old hydroponically grown two rice cultivars: spectrophotometric estimation (a) and histochemical visualization (b). P Panvel-3, S Sahyadri-3. The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
H2O2 is considered as the major ROS involved in inducing oxidative burst in plants, owing to its longevity and ability to cross plant cell membranes and thereby acts as a diffusible and relatively lasting signal (Huseynova et al. 2014). All three ionic stresses induced a significant increase in leaf H2O2 content over their non-saline counterparts, however, with much higher magnitude in Sahyadri-3 than in Panvel-3 (Fig. 2). H2O2 content did not vary significantly among the Cl−- and Na+-stressed leaves; however, NaCl treatment caused a sharp increase in leaf H2O2 concentration, particularly in Sahyadri-3, as revealed by spectrophotometric estimation and histological visualization. The univalent reduction of O2 •− produces H2O2. Therefore, elevated H2O2 production is considered to be driven by an increase in the concentrations of O2 •− in plant tissues as evidenced in the present work. H2O2 was histochemically localized in the leaf tissues of rice cultivars subjected to Cl−, Na+, and NaCl stress using DAB staining. DAB polymerizes to produce a brown precipitate due to contact with H2O2 in the presence of peroxidase and thus provides a useful marker of peroxide accumulation.
Cl−, Na+, and NaCl induced H2O2 generation in the leaves of 21-day-old hydroponically grown two rice cultivars: spectrophotometric estimation (a) and histochemical visualization (b). P Panvel-3, S Sahyadri-3. The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
No significant difference was observed between two cultivars in terms of •OH generation in their leaves under non-stressed conditions; however, all salt treatments caused a marked increase in leaf •OH concentration (estimated in terms of MDA content), with highest magnitude under combined stress of Na+ and Cl−, the NaCl (Fig. 3). Once again, Panvel-3 displayed its ability to keep a check on the ionic stress-induced •OH generation as its leaves accumulated significantly lesser amount of •OH than Sahyadri-3.
Generation of •OH by Cl−, Na+, and NaCl stress in the leaves of 21-day-old hydroponically grown two rice cultivars. The DMRT at P ≤ 0.05 was applied. Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
Hydroxyl radicals are considered as one of the highest reactive ROS, which can be produced from O2 •− and H2O2 in the presence of suitable transitional metals via O2 •−-driven Fenton reaction. These radicals are thought to be largely responsible for mediating oxygen toxicity (Gill and Tuteja 2010) and can potentially react with all biological molecules and cellular constituents. Excess production of •OH may ultimately lead to cell death (Vranova et al. 2002) as evidenced in the present investigation. The frequency of cell death varied considerably between salt types, and NaCl provoked maximum cell death events followed by Na+ and Cl−, respectively, with higher deleterious effects on Sahyadri-3, as revealed by spectrophotometric and histochemical visualization (Fig. 4). Evans blue staining was used to histochemically visualize the cell death, which is based on the disintegration of plasma membrane and is considered as a reliable parameter to assess cell death (Achary et al. 2012). A strong correlation may be established between the amounts of ROS generated under the influence of Na+ and Cl− ions (singly and as their combined salt) and ROS-induced cell death in the leaf tissues subjected to salt stress.
Induction of cell death by Cl−, Na+, and NaCl stress in the leaves of 21-day-old hydroponically grown two rice cultivars: spectrophotometric estimation (a) and histochemical visualization (b). P Panvel-3, S Sahyadri-3. The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
Effect of Cl−, Na+, and NaCl stress on antioxidant enzyme/isozyme activities
In order to protect themselves against toxic oxygen intermediates, plant cells and organelles deploy antioxidant defense system consisting of non-enzymatic and enzymatic components. A considerable amount of research has established that timely induction of cellular antioxidant machinery is vital for plant protection against various abiotic stresses via scavenging or detoxifying the ROS generated therein. The enzymatic antioxidants include one that reacts with ROS and keep them at low levels (SOD, CAT, and POX) and another that regenerates the oxidized antioxidants (APX and GR) (Chawla et al. 2013). Very often, increased salinity tolerance of tolerant glycophytic crop plants is correlated with better antioxidant capacities than their sensitive counterparts (Kumar et al. 2009; Chawla et al. 2013).
Figure 5a reveals that Panvel-3 seedlings showed significantly higher SOD activities than Sahyadri-3, both constitutive as well as salinity-induced. No genotypic difference was evidenced under Na+ and Cl− stress; however, NaCl-stressed seedlings of Panvel-3 showed 1.56 times more activity of SOD than Sahyadri-3. Spectrophotometric results were well supported by isozymatic patterns (Fig. 5b). Four isoforms were identified, one each of Mn-SOD and Cu/Zn-SOD besides two Fe-SOD in Panvel-3, whereas Sahyadri-3 leaves lacked one Fe-SOD isoform under stressed as well as non-stressed conditions. De novo synthesis of Fe-SOD1 isoform and an increase in the intensity of Fe-SOD2 in Panvel-3 suggested that specific activity of Fe-SOD seems to contribute to increased activities under salt stress, although Cu/Zn-SOD was the predominant form in both cultivars. SODs are metallo-enzymes and form the foremost line of defense against ROS by rapidly catalyzing the dismutation of O2 •− in the cell, where one O2 •− gets reduced to H2O2 and another oxidized to O2 (Mahanty et al. 2012). This in turn decreases the risk of •OH formation via the metal catalyzed Haber–Weiss type reaction. Thus, SOD seems to be playing a crucial role in conferring the salt tolerance abilities of Panvel-3.
Effects of Cl−, Na+, and NaCl stress separately on induction of SOD activities in the leaves of 21-day-old hydroponically grown two rice cultivars assessed by spectrophotometric estimation (a) and in-gel visualization (b). The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
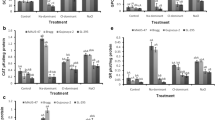
Unlike SOD, the constitutive levels of all other antioxidant enzymes did not vary significantly (at P ≤ 0.05) among the two genotypes examined in non-stressed tissues. However, all three types of salinity stresses induced a marked increase in antioxidant enzymes activities, though in a differential salt- and genotype-dependent manner. CAT activities gradually increased in rice cultivars in an order control < Cl < Na < NaCl, showing additive effects of the constituent ions (Fig. 6a). However, the increase was relatively higher in Panvel-3 than Sahyadri-3. Two isoforms of CAT were seen in the gel, both likely contributing significantly to its activity (Fig. 6b). The CAT activity obtained via spectrophotometric analysis coincided with the in-gel assay. CATs are tetrameric heme-containing enzymes, which effectively convert H2O2 into H2O and O2 and are considered indispensable for detoxification of elevated amounts of ROS generated under stressed conditions. The present results are in accordance with the previous reports indicating higher CAT activities in salt-tolerant rice cultivars (El-Shabrawi et al. 2010; Turan and Tripathy 2013).
Effects of Cl−, Na+, and NaCl stress separately on induction of CAT activities in the leaves of 21-day-old hydroponically grown two rice cultivars assessed by spectrophotometric estimation (a) and in-gel visualization (b). The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
Cl− stress failed to induce any significant change in POX activities of both the cultivars over their non-stressed counterparts, indicating that this ion is less toxic compared to Na+. Na+ and NaCl, on the other hand, increased the POX activities sharply with more pronounced effects in Panvel-3 (Fig. 7a), which was further confirmed by the band intensities of three isoforms obtained via in-gel assay (Fig. 7b). Overall, NaCl showed synergistic effects on modulation of POX activities against its constituent ions.
Effects of Cl−, Na+, and NaCl stress separately on induction of POX activities in the leaves of 21-day-old hydroponically grown two rice cultivars assessed by spectrophotometric estimation (a) and in-gel visualization (b). The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
APX utilizes ascorbate as an electron donor to scavenge the toxic H2O2 in water-water and ascorbate-GSH cycles. It is thought to be the most essential antioxidant enzyme, owing to its higher affinity for H2O2 than CAT and POX and a presumably more crucial role in the management of ROS during environmental stresses. APX activities constantly increased in two rice cultivars under the influence of individual ions, and NaCl showed high stimulus and induced 2-fold increase in Sahyadri-3 against 2.5-fold in Panvel-3 (Fig. 8a). APX exists in several isoforms, as seen in the present investigation with a total of five APX isoforms, where APX-IV seems not only the major contributor to overall enzyme activity in both the cultivars but also showed visible differential expression among the tolerant and sensitive genotype (Fig. 8b). The present results reaffirm the hypotheses that tolerant cultivars activate APX enzyme to detoxify the ROS generated under stress conditions (Turan and Tripathy 2013).
Effects of Cl−, Na+, and NaCl stress separately on induction of APX activities in the leaves of 21-day-old hydroponically grown two rice cultivars assessed by spectrophotometric estimation (a) and in-gel visualization (b). The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
GR levels did not vary significantly among the non-stressed leaves of rice cultivars; however, the genotypes exhibited contrasting behavior under salinity stress. Individual ions did not show momentous effects on GR activities in Sahyadri-3; on the contrary though, both Na+ and Cl− ions increased the enzyme activities markedly over their corresponding controls (Fig. 9a). The two genotypes showed no significant (at P ≤ 0.05) differences in GR activities under Na+ or Cl− ions but recorded noticeable changes among them under NaCl (Fig. 9a), and Sahyadri-3 proved relatively inferior on this count. The isozymatic analysis of GR in leaves as given in Fig. 9b clearly supports these findings. GR belongs to the family of NADPH-dependent oxidoreductase and comprises the last enzyme of AsA-GSH cycle. It holds great significance in the regulation of cytosolic redox environment, which is considered vital for cell endurance. GR plays a central role in cellular defense against the reactive oxygen metabolites by catalyzing the reduction of GSSG to GSH with the accompanying oxidation of NADPH and thus by maintaining the reduced-GSH pool under adverse conditions, which is considered crucial for active functioning of cells (Anjum et al. 2012; Trivedi et al. 2013). Therefore, salt tolerance character of genotypes is often linked to stimulated GR activity (Sairam et al. 2005).
Effects of Cl−, Na+, and NaCl stress separately on induction of GR activities in the leaves of 21-day-old hydroponically grown two rice cultivars assessed by spectrophotometric estimation (a) and in-gel visualization (b). The DMRT at P ≤ 0.05 was applied (a). Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
Differential antioxidant gene expression under Cl−, Na+, and NaCl stress
Though there are few reports on physiological disturbances produced by Cl− and Na+, very little is known about the relation of this ion accumulation to induction of antioxidant system. To probe the effects of individual Na+ and Cl− ions and their combined salt on antioxidant enzymes at the transcriptional level, the expression levels of certain genes involved in antioxidant activities were investigated. All three selected genes corresponding to SOD, CAT, and POX besides one housekeeping gene (rice-GAPDH) were analyzed by semi-quantitative RT-PCR using gene-specific primers (Table 2). The transcriptional expression level of GAPDH was investigated as housekeeping mRNA, which has been well established as normalization for relative gene(s) expression patterns in rice. As shown in Fig. 10, in consonance with the enzyme activities, the SODCc2 transcripts were up-regulated in plants of both cultivars, subjected to all three types of stress, though with higher expression levels in Panvel-3 over its sensitive counterpart. Among the multiple forms of SODs, the Cu/Zn-SODs are most abundantly prevalent isozymes in plants (Mahanty et al. 2012). The expression level of SodCc2 gene encoding Cu/Zn-SOD was monitored under NaCl and its constituent ions, which was higher in Panvel-3 under controlled as well as all three salt treatments. In the same vein, Turan and Tripathy (2013) reported an increase in rice Cu/Zn-SOD transcripts in response to salinity stress and suggested that increase in the antioxidative enzyme activity may include its respective isoform whose transcript abundance increases under stress conditions. Along with SODs, CATs constitute a front-line defense against ROS, converting H2O2 into water. In Panvel-3 seedlings, a slight and gradual up-regulation of CatA was noticed in Cl−-, Na+-, and NaCl-treated plants, respectively. On the contrary, CatA levels decreased in Sahyadri-3 submitted to Cl− and NaCl stresses, while a sharp increase in transcript levels was evidenced under Na+ stress. Identical induction of CAT gene has been reported by Menezes-Benavente et al. (2004) in rice and halotolerant Chenopodium album (Yao et al. 2010). However, any of the salts failed to induce significant changes in the OsPRX1 expression levels in Panvel-3 leaves, whereas its expression was significantly reduced in Sahyadr-3 leaves under Na+ and Cl− stress but increased again under NaCl stress. This disagreement between enzyme activities and expression levels of their pathway genes may be explained by the fact that the gene expression was analyzed only for one of the isoforms of selected antioxidants, while enzyme activity of an enzyme is a cumulative effect of the expression of different isoforms in cellular compartments (Turan and Tripathy 2013). On the other hand, constitutive as well as salinity-induced expression of GAPDH did not vary substantially in any of the cultivars. To sum up, NaCl treatment resulted in the up-regulation of CAT transcript levels followed by Na+ and Cl−, respectively, which is falling in line of total enzyme activities obtained, with more pronounced effects on Panvel-3. It appears, therefore, that the tolerant lines have the ability to up-regulate the genes that encode antioxidant enzymes and failure of sensitive plants in doing so under salinity stress conditions. This is probably the first comprehensive attempt to investigate the effects of equimolar concentrations of NaCl and its constituent ions on induction of antioxidant enzymes including the expression levels of their pathway genes.
Effects of Cl−, Na+, and NaCl stress separately on gene expressions as visualized on agarose gel electrophoresis. Expression levels of SOD, CAT, POX, and control (GAPDH) are observed after RT-PCR amplification using respective specific primers in the leaves of 21-day-old hydroponically grown two rice cultivars
Effect of Cl−, Na+, and NaCl stress on non-enzymatic antioxidants
To detoxify and eliminate the excessive levels of ROS, plants deploy an antioxidant defense armory consisting of two components, enzymatic and non-enzymatic ROS detoxifiers such as AsA, GSH, flavonoids, and tocopherols. Both these components work in tandem in providing metabolic acclimation and consequential improved salinity tolerance to plants. The SOD catalyzes the dismutation of O2 •− to 3O2 and H2O2, CAT, and POX then remove H2O2 (Yao et al., 2010), and in parallel, the non-enzymatic antioxidants such as AsA, GSH, and carotenoids can effectively prevent the 1O2 formation and scavenge the ROS (Mittler 2002). Despite a credible number of reports on NaCl-induced ROS generation and concomitant modulation of enzymatic antioxidants, relatively much lesser efforts have been made to decipher the roles of non-enzymatic antioxidants. Moreover, there is a serious dearth of investigations focused at dissecting the individual roles and relative importance of Na+ and Cl− ions in ROS generation and in inducing responsive antioxidant machinery, and to our knowledge, this the first such report.
Panvel-3 leaves with or without salinity exposure accumulated relatively higher amounts of total phenols, AsA, α-tocopherols, and GSH than Sahyadri-3 (Fig. 11). Each of the three treatments induced different non-enzymatic antioxidant responses in rice, which were largely in accord with the trends obtained with their enzymatic counterparts. Leaf non-enzymatic antioxidant levels were largely unaffected by Cl− treatment. Na+ and NaCl, on the other hand, exerted sharp changes in the non-enzymatic antioxidant levels. Total phenols did not vary significantly among the cultivars grown under Na and Cl treatments; however, Panvel-3 accumulated 189 % higher phenol levels in its leaves as compared with 143 % in Sahyadri-3 under NaCl stress over their respective controls (Fig. 11).
Effects of Cl−, Na+, and NaCl stress separately on ascorbic acid (AsA), reduced glutathione (GSH), α-tocopherol, flavonoids, and total phenols content in the leaves of 21-day-old hydroponically grown two rice cultivars. The DMRT at P ≤ 0.05 was applied. Treatments followed by the same lowercase letter for a particular genotype do not differ statistically. Genotypes followed by the same uppercase letter in a particular treatment do not differ statistically. Bar represents standard error
Similarly, concentrations of AsA, GSH, and total flavonoids increased under Na+ and NaCl but not in Cl− stress (Fig. 11). There was a sharp change in total flavonoid content in two rice cultivars when subjected to Na+ or NaCl stress, and once again NaCl exerted sharper increase in flavonoids concentration, while Cl− failed to induce any notable amendment as compared with controls. Flavonoids are considered as important constituents of plant phenolics, and their accumulation is crucial for their role in scavenging ROS. Total AsA content was markedly increased after exposure to Na+ (1.73-fold) and NaCl (1.93-fold) in Panvel-3, whereas the increase was relatively lesser (1.59-fold under Na+ and 1.63-fold under NaCl) in Sahyadri-3. Though out of many functions ascribed to AsA only a few are well characterized, however, it is quite clear that it is a major antioxidant which directly reacts with hydroxyl radicals, superoxide and singlet oxygen, and have a role in preserving the enzymes that contain prosthetic transition metal ions.
A shift from non-saline to saline environment caused a steady decrease in α-tocopherol concentration, and again the constituent ions showed additive effects with the addition of NaCl. Significantly lesser antagonistic effects were recorded in Panvel-3 than Sahyadri-3, with 32 and 65 % reductions over their controls, respectively. The increased GSH content in NaCl-stressed leaves of Panvel-3 (Fig. 11) might probably be reflecting its increased demand as a substrate for the enzymes participating in the detoxification of ROS and their end-products. GSH is known to facilitate AsA regeneration, as evident in this study. It is a major regulator of protein thiol-disulfide exchange reaction and seemingly works as an important signal molecule by forming a link between environmental stress and the key adaptive responses (Qureshi et al. 2013). The GSH content and GR activity are considered to be intricately linked and are differentially modulated by the stressors. Even though there are a number of reports signifying the roles of GR and GSH in plant stress protection, there are a few investigations on the cross-talking of these two enigmatic components of plant defense machinery (Huseynova et al. 2014). The present results indicated that AsA, GR, and GSH perform together the scavenging of ROS and their reaction products, and play important roles in conferring tolerance to stress-exposed rice seedlings.
Conclusion
The results comprehensively suggested that equimolar treatments of Cl−, Na+, and NaCl applied using hydroponic systems induced ionic imbalance, excess ROS generation, and oxidative damage in leaves of 21-day-old rice seedlings, though with a disparity of magnitude between two rice cultivars. Of the three salts, NaCl proved most detrimental than its constituent ions, followed by Na+ and Cl−, respectively. Na+ and Cl− ions showed additive effects under NaCl in terms of generation of ROS, resultant oxidative damage, and cell death events with a higher magnitude in the sensitive cultivar. Though Cl− proved relatively less toxic than Na+ or NaCl, its effect, however, cannot be totally ignored on the induction of oxidative stress and responsive antioxidant defense in rice. The salt-tolerant nature of Panvel-3 against individual as well as combined salt of Na+ and Cl− ions may be attributed to its stronger ability to maintain significantly lower Na+/K+ ratio. Also, the tolerant line was able to combat the oxidative stress by up-regulating the gene expressions and their encoded enzyme activities coupled with the accumulation of important non-enzymatic antioxidants in its leaves.
Abbreviations
- APX:
-
Ascorbate peroxidase
- CAT:
-
Catalase
- DAB:
-
Diaminobenzidine
- EC:
-
Electrical conductivity
- GAPDH :
-
Rice glyceraldehyde 3-phosphate dehydrogenase
- GR:
-
Glutathione reductase
- GSH:
-
Reduced glutathione
- GSSG:
-
Oxidized glutathione
- NBT:
-
Nitro blue tetrazolium
- POX:
-
Peroxidase
- ROS:
-
Reactive oxygen species
- RT-PCR:
-
Reverse transcriptase polymerase chain reaction
- SOD:
-
Superoxide dismutase
References
Achary VMM, Patnaik AR, Panda BB (2012) Oxidative biomarkers in leaf tissue of barley seedlings in response to aluminum stress. Ecotoxicol Environ Saf 75:16–26. doi:10.1016/j.ecoenv.2011.08.015
Aebi H (1984) Catalase in vitro. Meth Enzymol 105:121–126. doi:10.1016/S0076-6879(84)05016-3
Anderson JV, Hess JL, Chevone BI (1990) Purification, characterization, and immunological properties for two isoforms of glutathione reductase from eastern white pine needles. Plant Physiol 94:1402–1409. doi:10.1104/pp.94.3.1402
Anderson MD, Prasad TK, Stewart CR (1995) Changes in isozyme profiles of catalase, peroxidase, and glutathione reductase during acclimation to chilling in mesocotyls of maize seedlings. Plant Physiol 109:1247–1257. doi:10.1104/pp.109.4.1247
Anil VS, Krishnamurthy P, Kuruvilla S, Sucharitha K, Thomas G, Mathew MK (2005) Regulation of the uptake and distribution of Na+ in shoots of rice (Oryza sativa) variety Pokkali: role of Ca2+ in salt tolerance response. Physiol Plant 124:451–464. doi:10.1111/j.1399-3054.2005.00529.x
Anjum NA, Ahmad I, Mohmood I, Pacheco M, Duarte AC, Pereira E, Umar S, Ahmad A, Khan NA, Iqbal M, Prasad NMV (2012) Modulation of glutathione and its related enzymes in plants’ responses to toxic metals and metalloids—a review. Environ Exp Bot 75:307–324. doi:10.1016/j.envexpbot.2011.07.002
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–287. doi:10.1016/0003-2697(71)90370-8
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254. doi:10.1016/0003-2697(76)90527-3
Cameron GR, Milton RF, Allan JW (1943) Toxicity of tannic acid: an experimental investigation. Lancet 242:179–186. doi:10.1016/S0140-6736(00)87163-9
Castillo FI, Penel I, Greppin H (1984) Peroxidase release induced by ozone in Sedum album leaves. Plant Physiol 74:846–851. doi:10.1104/pp.74.4.846
Chaitanya KSK, Naithani SC (1994) Role of superoxide, lipid peroxidation and superoxide dismutase in membrane perturbation during loss of viability in seeds of Shorea robusta Gaertn.f. New Phytol 126:623–627. doi:10.1111/j.1469-8137.1994.tb02957.x
Chapman HD, Pratt PF (1961) Method for analysis of soil, plants and waters. University of California, Berkeley, CA
Chawla S, Jain S, Jain V (2013) Salinity induced oxidative stress and antioxidant system in salt-tolerant and salt-sensitive cultivars of rice (Oryza sativa L.). J Plant Biochem Biotechnol 22:27–34. doi:10.1007/s13562-012-0107-4
Dang YP, Dalal RC, Mayer DG, McDonald M, Routley R, Schwenke GD, Buck SR, Daniells IG, Singh DK, Manning W, Ferguson N (2008) High subsoil chloride concentrations reduce soil water extraction and crop yield on Vertisols in north-eastern Australia. Aust J Agr Res 59:321–330. doi:10.1071/AR07192
Dhindsa RA, Plumb-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased permeability and lipid peroxidation and decreased levels of superoxide dismutase and catalase. J Exp Bot 126:93–101. doi:10.1093/jxb/32.1.93
El-Shabrawi H, Kumar B, Kaul T, Reddy MK, Singla-Pareek SL, Sopory SK (2010) Redox homeostasis, antioxidant defense and methylglyoxal detoxification as markers for salt tolerance in Pokkali rice. Protoplasma 245:85–96. doi:10.1007/s00709-010-0144-6
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930. doi:10.1016/j.plaphy.2010.08.016
Halliwell B, Gutteridge JMC, Aruma O (1987) The deoxyribose method: a simple ‘test tube’ assay for determination of rate constant for reaction of hydroxyl radicals. Anal Biochem 165:215–219. doi:10.1016/0003-2697(87)90222-3
Huang Z, Zhao L, Chen D, Liang M, Liu Z, Hongbo S, Xiaohua L (2013) Salt stress encourages proline accumulation by regulating proline biosynthesis and degradation in Jerusalem artichoke plantlets. PLoS ONE 8(4):e62085. doi:10.1371/journal.pone.0062085
Huseynova IM, Sultanova NF, Aliyev JA (2014) Histochemical visualization of ROS and antioxidant response to viral infections of vegetable crops grown in Azerbaijan. Plant Physiol Biochem 81:26–35. doi:10.1016/j.plaphy.2014.03.002
Ismail A, Takeda S, Nick PR (2014) Life and death under salt stress: same players, different timing? J Exp Bot. doi:10.1093/jxb/eru159
Kamyab-Talesh F, Mousavi SF, Asadi R, Rezaei M, Khaledian MR (2014) Evaluation of some rice cultivars response to salinity stress using resistance indices. Arch Agron Soil Sci 60:1303–1314. doi:10.1080/03650340.2014.891730
Khan MA, Shirazi MU, Khan MA, Mujtaba SM, Islam E, Mumtaz S (2009) Role of proline, K/Na ratio and chlorophyll content in salt tolerance of wheat (Triticum aestivum L.). Pak J Bot 41:633–638
Kumar V, Khare T (2014) Individual and additive effects of Na+ and Cl− ions on rice under salinity stress. Arch Agron Soil Sci. doi:10.1080/03650340.2014.936400
Kumar V, Shriram V, Nikam TD, Jawali N, Shitole MG (2009) Antioxidant enzyme activities and protein profiling under salt stress in indica rice genotypes differing in salt tolerance. Arch Agron Soil Sci 55:379–394. doi:10.1080/03650340802595543
Lin CC, Kao CH (2001) Relative importance of Na+, Cl−, and abscisic acid in NaCl induced inhibition of root growth of rice seedlings. Plant Soil 237:165–171. doi:10.1023/A:1013321813454
Loreto F, Velikova V (2001) Isoprene produced by leaves protects the photosynthetic apparatus against ozone damage, quenches ozone products, and reduces lipid peroxidation of cellular membranes. Plant Physiol 127:1781–1787. doi:10.1104/pp.010497
Luo Q, Yu B, Liu Y (2005) Differential sensitivity to chloride and sodium ions in seedlings of Glycine max and G. soja under NaCl stress. J Plant Physiol 162:1003–1012. doi:10.1016/j.jplph.2004.11.008
Mahanty S, Kaul T, Pandey P, Reddy RA, Mallikarjuna G, Reddy CS et al (2012) Biochemical and molecular analyses of copper–zinc superoxide dismutase from a C4 plant Pennisetum glaucum reveals an adaptive role in response to oxidative stress. Gene 505:309–317. doi:10.1016/j.gene.2012.06.001
Malick CP, Singh MB (1980) Plant enzymology and histo-enzymology. Kalyani, New Delhi
Menezes-Benavente L, Teixeira FK, Alvim Kamei CL, Margis-Pinheiro M (2004) Salt stress induces altered expression of genes encoding antioxidant enzymes in seedlings of a Brazilian indica rice (Oryza sativa L.). Plant Sci 166:323–331. doi:10.1016/j.plantsci.2003.10.001
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7:405–410. doi:10.1016/S1360-1385(02)02312-9
Mittler R, Zilinskas BA (1993) Detection of ascorbate peroxidase activity in native gels by inhibition of the ascorbate-dependent reduction of nitroblue tetrazolium. Anal Biochem 212:540–546. doi:10.1006/abio.1993.1366
Moron MS, Depierre JW, Mannervik B (1979) Levels of glutathione, glutathione reductase and glutathione S-transferase activities in rat lung and liver. Biochim et Biophys Acta 582:67–78. doi:10.1016/0304-4165(79)90289-7
Moya JL, Gomez-Cadenas A, Primo-Millo E, Talon M (2003) Chloride absorption in salt-sensitive Carrizo citrange and salt-tolerant Cleopatra mandarin citrus rootstocks in linked to water use. J Exp Bot 54:825–833. doi:10.1093/jxb/erg064
Mukharjee SP, Choudhari MA (1983) Implication of water stress-induced changes in the levels of endogenous ascorbic acid and hydrogen peroxide in Vigna seedlings. Physiol Plant 58:166–170. doi:10.1111/j.1399-3054.1983.tb04162.x
Munns R, Tester S (2008) Mechanism of salinity tolerance. Annu Rev Plant Biol 59:651–681. doi:10.1146/annurev.arplant.59.032607.092911
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C et al (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364. doi:10.1038/nbt.2120
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Qureshi MI, Abdin MZ, Ahmad J, Iqbal M (2013) Effect of long-term salinity on cellular antioxidants, compatible solute and fatty acid profile of Sweet Annie (Artemisia annua L.). Phytochemistry 95:215–223. doi:10.1016/j.phytochem.2013.06.026
Romeo-Puertas MC, Rodriguez-serrano M, Corpas FJ, Gomez M, del-Rio LA, Sandalio LM (2004) Cadmium-induced subcellular accumulation of O2 •− and H2O2 in pea leaves. Plant Cell Environ 27:1122–1134. doi:10.1111/j.1365-3040.2004.01217.x
Rosenberg HR (1992) Chemistry and physiology of vitamins. Inter Science, New York, pp 452–453
Sairam RK, Srivastava GC, Agarwal S, Meena RC (2005) Differences in antioxidant activity in response to salinity stress in tolerant and susceptible wheat genotypes. Biol Plant 49:85–91. doi:10.1007/s10535-005-5091-2
Smith IK, Vierheller TL, Thorne CA (1988) Assay of glutathione reductase in crude tissue homogenate using 5,5′-dithiobis (2-nitrobenzoic acid). Anal Biochem 175:408–413. doi:10.1016/0003-2697(88)90564-7
Tavakkoli E, Rengasamy P, McDonald GK (2010) High concentrations of Na+ and Cl− ions in soil solution have simultaneous detrimental effects on growth of faba bean under salinity stress. J Exp Bot 61:4449–4459. doi:10.1093/jxb/erq251
Tavakkoli E, Fatehi F, Coventry S, Rengasamy P, McDonald GK (2011) Additive effects of Na+ and Cl− ions on barley growth under salinity stress. J Exp Bot 62:2189–2203. doi:10.1093/jxb/erq422
Teakle NL, Tyerman SD (2010) Mechanisms of Cl− transport contributing to salt tolerance. Plant Cell Environ 33:566–589. doi:10.1111/j.1365-3040.2009.02060.x
Tilman D, Balzer C, Hill J, Belfort BL (2011) Global food demand and the sustainable intensification of agriculture. Proc Natl Acad Sci USA 108:20260–20264. doi:10.1073/pnas.1116437108
Trivedi D, Gill SS, Yadav S, Tuteja N (2013) Genome-wide analysis of glutathione reductase (GR) genes from rice and Arabidopsis. Plant Signal Behav 8:e2301–e2307. doi:10.4161/psb.23021
Turan S, Tripathy BC (2013) Salt and genotype impact on antioxidative enzymes and lipid peroxidation in two rice cultivars during de-etiolation. Protoplasma 250:209–222. doi:10.1007/s00709-012-0395-5
Turkan I, Demiral T (2009) Recent development in understanding salinity tolerance. Environ Exp Bot 67:2–9. doi:10.1016/j.envexpbot.2009.05.008
Vranova E, Inze D, Van Breusegem F (2002) Signal transduction during oxidative stress. J Exp Bot 53:1227–1236. doi:10.1093/jexbot/53.372.1227
White PJ, Broadley MR (2001) Chloride in soils and its uptake and movement within the plant: a review. Ann Bot 88:967–988. doi:10.1006/anbo.2001.1540
Woodbury W, Spencer AK, Stahmann MA (1971) An improved procedure using ferricyanide for detecting catalase isozymes. Anal Biochem 44:301–305. doi:10.1016/0003-2697(71)90375-7
Xu G, Magen H, Tarchitzky J, Kafkafi U (2000) Advances in chloride nutrition of plants. Adv Agron 68:97–150. doi:10.1016/S0065-2113(08)60844-5
Yao S, Chen S, Xu D, Lan H (2010) Plant growth and responses of antioxidants of Chenopodium album to long-term NaCl and KCl stress. Plant Growth Regul 60:115–125. doi:10.1007/s10725-009-9426-4
Yildiztugay E, Ozfidan-Konakci C, Kucukoduk M (2014) Modulation of osmotic adjustment and antioxidant status in salt-stressed leaves of Thermopsis turcica. Acta Physiol Plant 36:125–138. doi:10.1007/s11738-013-1393-8
Zhang XK, Zhou QH, Cao JH, Yu BJ (2011) Differential Cl− salt tolerance and NaCl-induced alternations of tissue and cellular ion fluxes in Glycine max, Glycine soja and their hybrid seedlings. J Agron Crop Sci 197:329–339. doi:10.1111/j.1439-037x.2011.00467.x
Acknowledgments
Financial support from the Science and Engineering Research Board (SERB), Government of India (grant number SR/FT/LS-93/2011) for carrying out this work is gratefully acknowledged. The authors also like to acknowledge the use of facilities created under DST-FIST and DBT Star College Schemes implemented at Modern College, Ganeshkhind, Pune, and thank the college authorities for permitting to utilize these facilities.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Peter Nick
Rights and permissions
About this article
Cite this article
Khare, T., Kumar, V. & Kishor, P.B.K. Na+ and Cl− ions show additive effects under NaCl stress on induction of oxidative stress and the responsive antioxidative defense in rice. Protoplasma 252, 1149–1165 (2015). https://doi.org/10.1007/s00709-014-0749-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-014-0749-2