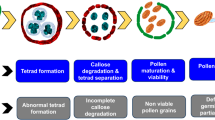
Abstract
The endosporic male gametophyte of the water fern, Marsilea vestita, provides a unique opportunity to study the mechanisms that control cell fate determination during a burst of rapid development. In this review, we show how the spatial and temporal control of development in this simple gametophyte involves several distinct modes of RNA processing that allow the translation of specific mRNAs at distinct stages during gametogenesis. During the early part of development, nine successive cell division cycles occur in precise planes within a closed volume to produce seven sterile cells and 32 spermatids. There is no cell movement in the gametophyte; so, cell position and size within the spore wall define cell fate. After the division cycles have been completed, the spermatids become sites for the de novo formation of basal bodies, for the assembly of a complex cytoskeleton, for nuclear and cell elongation, and for ciliogenesis. In contrast, the adjacent sterile cells exhibit none of these changes. The spermatids differentiate into multiciliated, corkscrew-shaped gametes that resemble no other cells in the entire plant. Development is controlled post-transcriptionally. The transcripts stored in the microspore are released (unmasked) in the gametophyte at different times during development. At the start of these studies, we identified several key mRNAs that undergo translation at specific stages of gametophyte development. We developed RNA silencing protocols that enabled us to block the translation of these proteins and thereby establish their necessity and sufficiency for the completion of specific stages of gametogenesis. In addition, RNAi enabled us to identify additional proteins that are essential for other phases of development. Since the distributions of mRNAs and the proteins they encode are not identical in the gametophyte, transcript processing is apparently important in allowing translation to occur under strict temporal and spatial control. Transcript polyadenylation occurs in the spermatogenous cells in ways that match the translation of specific mRNAs. We have found that the exon junction complex plays key roles in transcript regulation and modifications that underlie cell specification in the gametophyte. We have recently become interested in the mechanisms that control the unmasking of the stored transcripts and have linked the synthesis and redistribution of spermidine in the gametophyte to the control of mRNA release from storage during early development and later to basal body formation, cytoskeletal assembly, and nuclear and cell elongation in the differentiating spermatids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction—post-transcriptional control of rapid development
Across the living realm, there are numerous examples of organisms that exhibit rapid bursts of development. These changes usually occur in cells that have special characteristics and the shift to rapid developmental activity is usually preceded by a period of quiescence (e.g., oocytes before fertilization, seeds and spores after desiccation, certain hibernating animals, etc.). Rapid development often involves a change in physiology, proliferation or differentiation without cell growth, and new proteins needed for the development process are made from pre-existing mRNAs. Thus, the regulation of the developmental burst is often controlled post-transcriptionally. The triggers responsible for entry into quiescence can involve changes in the environment (e.g., desiccation, nutrition) or developmental alterations that involve neighboring cells (e.g., oocyte formation in a number of animals). The mechanisms responsible for mRNA storage and ribosome accumulation are not well defined, though they are apparently accompanied or followed by shifts in cytoplasmic and nuclear composition and organization that allow the cell to activate a rapid developmental program when the appropriate stimulus releases the cell from its quiescent state.
In this review, we focus on the rapid development of the male gametophyte of the water fern, Marsilea vestita. This organism produces spores that undergo desiccation as part of a normal maturation process. During desiccation, the spores accumulate and store transcripts that will be utilized during a rapid burst of growth and development that is initiated when the dry spores are placed into water. The rapid burst of development of these gametophytes results in the production of multiciliated spermatozoids, which look nothing like the cells that gave rise to them. Development in this gametophyte is controlled post-transctiptionally; herein, we describe mechanisms that provide spatial and temporal control during the rapid developmental process that results in the formation and maturation of motile spermatozoids.
Stored RNA and the regulation of its translation
Initial indications that some mRNA species undergo a prolonged period of inactivity before translation associated with rapid development came from studies of the unfertilized sea urchin egg when non-nucleated fragments of the eggs were parthenogenetically stimulated to undergo cleavage and form blastulas (Harvey 1936, 1940). Latter studies carried out in the presence of the transcriptional inhibitor actinomycin D demonstrated that protein synthesis is necessary for cell division (Gross and Cousineau 1963, 1964). These experiments provided some of the earliest hints that the “masking” or storage of mRNA combines with regulated translation to mediate the bursts of development. Today, overwhelming evidence indicating that the regulated translation of stored mRNA is an important mechanism underlying the rapid development in a variety of organisms and cell types exists.
Stored transcripts in angiosperm seeds were first identified in 1965 and are now viewed as ubiquitous in the seeds of all flowering plants. In Arabidopsis thalania seeds, there can be as many as 10,000 different species of stored mRNAs (Dure and Waters 1965; Nakabayashi et al. 2005). Stored mRNA is utilized during the early stages of embryo development after the seed is hydrated during a process known as imbibition. Hydration triggers the activation of translation for these stored transcripts (Hughes and Galau 1989, 1991; Comai et al. 1989). Kimura and Nambara (2010) completed a microarray analysis of stored mRNA from different Arabidopsis ecotypes. Their findings suggest that the classes of mRNA stored in the seeds of different Arabidopsis accessions are similar and that there is an abundant accumulation of transcripts from the late embryogenesis abundant, seed storage protein, stress protein, lipid storage, and metabolism classes (Kimura and Nambara 2010).
Arousal from hibernation has been used as a model to study the maturation of stored pre-mRNA in a number of different kinds of animals. In hibernating dormice (Malatesta et al. 1994, 1999), pre-mRNAs are stored within nuclei at different stages of maturation in a tissue-specific manner. For example, in the liver of hibernating dormice, pre-mRNA is accumulated predominantly at the splicing stage, whereas in brown adipose tissue pre-mRNA is found to be mostly stored in its cleavage stage of maturation. Interestingly, the dramatic redistributions of pre-mRNA maturation machinery are observed within these tissues during hibernation, and the accumulation of this machinery mirrors the pre-mRNA storage state. This pattern of regulation assures that, upon arousal, dormouse tissues high in pre-mRNA at the cleavage state also have abundant stores of cleavage and polyadenylation machinery within their subnuclear processing bodies. Similarly, tissues with high levels of pre-mRNA stored at the splicing stage of maturation have a large distribution of splicing machinery localized to the subnuclear processing bodies. In this way, stored pre-mRNAs are regulated such that, upon arousal, certain tissues are metabolically activated in a prioritized fashion, with brown adipose tissue (cleavage pre-mRNA) activating first, followed by the liver (splicing pre-mRNA), and then finally the de novo transcription in all tissues (Malatesta et al. 1994, 1999).
Brine shrimp larvae undergo desiccation. Upon hydration, these organisms utilize and translate stored mRNAs (Muthukrishnan et al. 1975). The analyses of transcripts isolated from imbibed larvae indicate that very few (if any) mRNAs are associated with polysomes at the resumption of development, but by 22 h a large portion of these transcripts become associated with the translational complexes (Amaldi et al. 1977; Grosfeld and Littauer 1975). Additionally, a subset of messages has been identified as being pre-loaded with the 40S subunit, a condition that appears to destine transcripts for early translation.
The role of mRNA modification in the post-transcriptional regulation of translation
RNA modifications can underlie the regulation of translational activation. Both splicing and cytoplasmic polyadenylation have been shown to affect the translational activity of RNA in a variety of organisms. The translation of a specific transcript can be activated or repressed by the lengthening or shortening of its poly(A) tail (Gorgoni and Gray 2004). This event is mediated by specific sequences within the 3′ untranslated region of mRNA, which is recognized by cytoplasmic polyadenylation factors (Fox et al. 1989). The polyadenylation of transcripts begins in the nucleus, where adenines are added to the 3′ end of pre-mRNA before getting exported to the cytoplasm. Once exported from the nucleus, transcripts with the correct cytoplasmic polyadenylation sequences may be modified further. Lengthening or stabilizing a long poly(A) tail, ~80–250 bases, has been shown to increase transcript stability and translational activity. Shortening of the tail to ~20–40 bases has been shown to promote the storage and diminish the translational rates of mRNA (Paris and Phillippe 1988; Rosenthal et al. 1983; Vassalli et al. 1989; Fox et al. 1989; McGrew et al. 1989). The extent of polyadenylation apparently has a direct effect on the ability of a transcript to form into complexes with translational machinery in a competitive manner (Proweller and Butler 1994).
The splicing of transcripts has been shown to increase translation significantly, a phenomenon that has been attributed to the deposition of exon junction complex (EJC) proteins on the spliced mRNA, which are believed to mediate polysome association with the mRNA (Nott et al. 2004). Splicing has also been shown to have an effect on the rate or timing at which a particular mRNA is exported from the nucleus to the cytoplasm or, in a few cases, to alter the subcellular distribution of specific transcripts (Luo and Reed 1999; Zhou et al. 2000; LeHir et al. 2001a; Ryu and Mertz 1989; Rafiq et al. 1997). Unspliced transcripts with premature stop codons are known to be targets of nonsense-mediated decay (NMD) and therefore undergo accelerated degradation. Splicing of premature stop codons allows for the passage of a transcript through the NMD pathway, allowing for a downstream translation of the message (Maquat and Carmichael 2001; Wilusz et al. 2001; Wilkinson and Shyu 2002).
M. vestita as a model system for studies on cell fate determination during rapid development
In certain algae and in most of the nonflowering, vascular plants (a group of basal plants including the bryophytes, the pteridophytes, and some gynmosperms), spermatogenesis is a complex process that results in the production of multiciliated gametes that look nothing like the cells that gave rise to them (Renzaglia and Garbary 2001). The process involves the formation of basal bodies in a cytoplasmic particle known as a blepharoplast (Webber 1897) and is a de novo event since it occurs in the absence of any preexisting centrioles in any cells in the organism. Basal body assembly itself occurs only in the spermatids, where as few as two basal bodies form in Charophycean algae (Chapman and Henk 1983), bryophytes (Carothers and Kreitner 1967; Carothers 1973; Kreitner and Carothers 1976; Renzaglia and Carothers 1986; Brown and Carothers 1986; Carothers and Rushing 1990), and lycopods (Carothers et al. 1975; Robert 1977) to ~32–150 in ferns and fern allies (Mizukami and Gall 1966; Duckett 1973; Hepler 1976; Myles and Hepler 1977, 1982; Marc and Gunning 1986; Hoffman and Vaughn 1995) or as many as several thousands in the spermatids of Ginkgo (Webber 1897; Gifford and Lin 1975; Li et al. 1989; Vaughn and Renzaglia 2006) and the cycads (Chamberlain 1909; Mizukami and Gall 1966; Norstog 1974). Spermatid maturation also involves the formation of a remarkably complicated cytoskeletal apparatus known as the multilayered structure (MLS; Paolillo et al. 1968; Myles and Hepler 1977; Vaughn et al. 1993), the extensive remodeling and reshaping of the gamete nucleus (Kreitner 1977; Myles and Hepler 1977, 1982), and ciliogenesis (Myles et al. 1978). These events combine to produce a more or less spirally shaped gamete that is ciliated on the dorsal face of the elongated and coiled cell body. The gametes formed lack walls and they can swim for long distances, changing their swimming patterns in response to the chemical cues that they receive from the environment. Thus, these gametes are chemotactic and the word chemotaxis was originally coined to describe the changes in the swimming patterns as exhibited by the spermatozoids of bracken fern (Pteridium aquilinum) by Pfeffer (1884).
Lester Sharp (1912, 1914) characterized the development of male gametophytes of the water fern, M. vestita, and the scouring rush, Equisetum arvense, and in these studies provided a descriptive roadmap for multiciliate spermatozoid formation within the antheridium. He showed that M. vestita provides an unusually simple system for studies on the mechanisms that underlie cell fate formation in the endosporic gametophyte and morphogenesis of the spermatids during its short developmental cycle. Development leading to spermatozoid formation proceeds rapidly following the hydration of dry microspores in water (Sharp 1914; Mizukami and Gall 1966; Hepler 1976; Myles and Hepler 1977, 1982; Hyams et al. 1983; Pennell et al. 1986, 1988; Hart and Wolniak 1998, 1999; Wolniak et al. 2000; Klink and Wolniak 2001, 2003; van der Weele et al. 2007).
The male gametophyte of M. vestita begins as a single progenitor cell contained within the microspore wall. The microspore is a meiotic product, and after desiccation within a leaf-based structure known as a sporocarp, the spore can remain quiescent for decades. Spore hydration initiates a complex series of developmental events, starting with cytoplasmic reorganization (Fig. 1a, b), followed by nine rapid mitotic division cycles that produce a total of 39 cells; when the division cycles are complete, there are seven sterile cells and 32 spermatids (Sharp 1914). The positions, sizes, and fates of the cells produced by these divisions are constant among the gametophytes (Fig. 1c–k), making a fate map simple to construct (Klink and Wolniak 2001). The division cycles are synchronous in populations of developing gametophytes, and all divisions occur in predictable planes within the microspore wall (Fig. 1; Sharp 1914; Mizukami and Gall 1966; Hepler 1976; Pennell et al. 1986, 1988; Klink and Wolniak 2001). This spatial and temporal precision among identically treated microspores makes it easy to determine the extent of progress in a population of gametophytes and straightforward to discern anomalies in the spatial patterns of cell division and differentiation (Klink and Wolniak 2001, 2003; Tsai and Wolniak 2001; van der Weele et al. 2007; Deeb et al. 2010).
Normal development in the male gametophyte of M. vestita involves nine successive mitotic division cycles, shown here in sections of spores fixed at particular time intervals and stained with toluidine blue-O. a At the time the dry spore is placed into water, the gametophyte consists of a single cell with a large, centrally positioned nucleus (n) and numerous starch-containing plastids. b Within a few minutes after being placed into water, the nucleus becomes acentrically positioned, and the starch-containing plastids (st) migrate to a site opposite the nucleus. c The first mitotic division occurs at 90 min, is asymmetric, and produces a small prothallial cell (p) and a large cell that will continue to divide. d The second division occurs about 30 min, is symmetric, and produces two antheridial initials. e–g The third, fourth, and fifth divisions are asymmetric and produce three sterile jacket cells from each antheridial initial (spi). Each of the jacket cells (jc) is significantly smaller than the spermatogenous cell that gave rise to it. The jacket cells completely surround the larger, centrally placed initial, and the jacket cells contain all of the large plastids with the large starch grains. The jacket cell cytoplasm does not exhibit the dark purple staining of the spermatogenous initial. The jacket cells lose the capacity to proliferate further. h–j Each spermatogenous initial undergoes four symmetric divisions, producing a total of 16 spermatids. k, l After the divisions are completed, the spermatids (sp) begin to differentiate; the cells become round, small starch-bearing plastids appear in the cytoplasm, and each spermatid assembles a ciliary apparatus. The processes of cytoskeletal development and nuclear elongation are described in detail elsewhere (Myles and Hepler 1977). The jacket cells become less conspicuous and the large starch grains in the plastids become considerably smaller over time. Brightfield images obtained with transmitted light. Bar = 25 μm
Once formed, the spermatids undergo drastic morphogenetic changes to become elongated, coiled, motile gametes that, when released, are known as spermatozoids. Spermatid differentiation involves the de novo synthesis of basal bodies in a cytoplasmic precursor particle known as a blepharoplast (Webber 1897; Chamberlain 1909; Sharp 1914; Mizukami and Gall 1966; Hepler 1976). During its ontogeny, the blepharoplast first serves as a functional centrosome (Chamberlain 1898) for the last mitotic division and then differentiates as a basal body factory (Hepler 1976; Wolniak et al. 2000). This process is followed by the assembly of the MLS by nuclear and cell reshaping (Myles and Hepler 1977, 1982) and then by ciliogenesis (Myles and Hepler 1977; Myles et al. 1978). The spermatid undergoes these extensive changes to become a spermatozoid in only 4–5 h. Each spermatozoid is a freely swimming cell that possesses ~140 cilia (Sharp 1914; Myles and Hepler 1977; Myles et al. 1978).
In a series of early experiments aimed at understanding the mechanisms that underlie rapid development in the gametophyte, we analyzed protein abundance changes over time (Hart and Wolniak 1998). Coomassie-stained electrophoretic gels revealed a few discernable differences in total extractable protein in the gametophytes over time, while autoradiograms of pulse-labeled spores at the time of germination showed that certain groups of polypeptides became intensely labeled as the gametophytes developed (Hart and Wolniak 1998). The patterns of new translation were identical in gametophytes treated with the transcriptional inhibitor α-amanitin, but translation was abolished by treatments with the translational inhibitor cycloheximide (Hart and Wolniak 1998). Direct microscopic observations of gametophytes showed that treatments with α-amanitin (Fig. 2) allowed development to reach completion, while treatments with cycloheximide arrested development prior to the first mitotic division (Klink and Wolniak 2001). These experiments provided the initial hints that the development in the gametophyte is regulated at a post-transcriptional level. In unpublished extensions of these studies (Fig. 2), Vincent Klink showed that treatments of gametophytes with the transcriptional activators trichostatin A or 5-Azo-cytosine resulted in developmental arrest. Collectively, these results suggested that gametophyte growth and development are dependent on the translation of existing mRNAs and not upon the transcription of new mRNAs.
Spermatogenesis in M. vestita occurs in the absence of new transcription. Gametophytes were grown for 1, 2, 4, 8, and 11 h at 20°C under various experimental conditions as indicated in the figure. Untreated gametophytes develop fully, and by 11 h many have released their mature spermatozoids, so the spore appears to be empty. α-Amanitin treatments develop normally and almost completely. At 8 hours of development in the presence of α-amanitin, the ciliary apparatus is being assembled, but at 11 h the spermatozoids have not been released from the antheridia. Gametophytes grown in the transcriptional activators, trichostatin A (TSA), and 5-aza-cytidine (5-aza-c), fail to develop for the first 8 h, but by 11 h some abnormal divisions are apparent. Bar = 25 mm
Formation of the blepharoplast and its role in the de novo assembly of basal bodies
The blepharoplast in M. vestita is a discrete, spherical particle that appears in the cytosol of the spermatid mother cell or the spermatid itself. It cannot be found in any other cells of the organism, so it represents a distinguishing morphological feature of spermatid mother cells and spermatids. When it appears in the spermatid mother cell, approximately 4 h after the spores are hydrated, the blepharoplast functions as a centrosome for the last mitotic division cycle in the developmental program, which results in the production of the spermatids. In this role, it splits in half and resides at the poles of a biconical mitotic spindle (Hepler 1976). After the division reaches completion, it disappears from the cells. It then reappears in the cytosol of each spermatid and begins its own maturation process, which has been documented ultrastructurally. These studies show that, initially, the centrosomal particle exhibits no substructure beyond electron dense flocculence, but later it forms electron-lucent channels that appear to be basal body cores (Mizukami and Gall 1966; Hepler 1976). The de novo formation of basal bodies first involves the expansion of the blepharoplast particle and the rapid assembly of A-subfibers around the core regions. This is followed by the sequential addition of B- and then C-subfibers to the nascent basal bodies. The basal bodies remain clustered for a short period as the spermatid assembles its complex cytoskeleton. Since de novo basal body formation occurs only in the spermatids of the gametophyte, the presence of basal bodies also serves as a diagnostic marker for spermatid cell specification. It is reasonable to conclude that processes responsible for blepharoplast formation are linked tightly to the mechanism of cell fate determination in the male gametophyte.
Using immunolabeling, we showed that the blepharoplast contains at least one centrin isoform (Klink and Wolniak 2001), which we have named Mv-Cen1 (Hart and Wolniak 1999). We next asked if Mv-Cen1 translation was essential for blepharoplast (and basal body) formation. We developed RNAi strategies (Fire et al. 1998), where the direct addition of Mv-Cen1 dsRNA to the spores at the time of hydration resulted in the inhibition of centrin translation in the developing gametophytes (Klink and Wolniak 2001). We found that dsRNA was 50–200 times more effective in silencing centrin than either sense- or anti-sense RNA strands added independently to identical spores. Immunolabeling and immunoblotting experiments showed that centrin protein was undetectable after treatments with dsRNA derived from the centrin cDNA (Klink and Wolniak 2001) isolated from our M. vestita male gametophyte library (Hart and Wolniak 1999). In the absence of centrin, blepharoplasts failed to form. In the absence of blepharoplasts, basal bodies did not assemble, and further development of the spermatids was arrested.
In performing these RNAi experiments, we found that the microspores are unusually amenable to the introduction of normally impermeant molecules into the cytoplasmic space of the single cell within the spore wall (Klink and Wolniak 2001). The dry spores allow the entry of double-stranded RNA (dsRNA) and other molecules (e.g., drugs) at the time of hydration. We have found that dsRNAs in excess of 500 bp readily enter the cells during the first few minutes after the spores are placed into water. Small molecules (e.g., hydroxyurea) can be taken into the cytosol of gametophytes up to ~3 h after the spores were hydrated (Tsai and Wolniak 2001). We treated populations of spores with dsRNA and elicited an RNAi effect, namely, the production of a phenocopy of a null mutation by destroying a specific mRNA (Fire et al. 1998; Montgomery and Fire 1998; Ngo et al. 1998; Fire 1999; Zamore et al. 2000; Boscher and Labouesse 2000) simultaneously in thousands of synchronously developing gametophytes. We found that the direct addition of dsRNA to our gametophytes was far more potent at silencing than either sense- or anti-sense RNA added separately to identical populations of spores, consistent with observations from Fire et al. (1998) in their original characterizations of RNAi mechanisms.
We recognize that the success of our RNAi experiments was enhanced because there is little, if any, new transcription required for spermiogenesis to reach completion in M. vestita (Hart and Wolniak 1998, 1999; Wolniak et al. 2000; Klink and Wolniak 2001). These early RNAi experiments with centrin (Mv-Cen1) dsRNA revealed that centrin translation does not occur and blepharoplasts failed to form (Klink and Wolniak 2001), demonstrating an essential function for this protein in blepharoplast formation and basal body assembly, the first time anyone had shown an essential function for centrin in any eukaryote. In gametophytes treated with Mv-Cen1 dsRNA, blepharoplasts failed to form and the last mitotic division typically did not occur. Under normal conditions of development, centrin translation increases dramatically at 4 h of development (Hart and Wolniak 1998) and is restricted to the spermatogenous cells. In later experiments, we found that centrin translation can take place throughout the gametophyte after 4 h of development when the cell division cycles that usually precede blepharoplast formation in the male gametophyte are blocked (Tsai and Wolniak 2001) by cyclin RNAi treatments or by treatments with olomoucine (Glab et al. 1994; Meier 1996), an inhibitor of cyclin-dependent kinase activity. Thus, although centrin translation is necessary for blepharoplast formation, it is not sufficient for the de novo formation of basal bodies. We suspect that there are additional factors or morphogenetic determinants involved in centrin aggregation that are affected by the division cycles that separate sterile and spermatogenous cells.
Formation and function of the multilayered structure, a cytoskeletal array unique to spermatozoids
The multilayered structure (MLS; Paolillo et al. 1968) is a signature organelle in the motile spermatozoids produced by certain algae, bryophytes, pteridophytes, and certain gymnosperms (Paolillo 1965; Carothers and Kreitner 1967; Duckett 1973; Carothers et al. 1975; Myles and Hepler 1977; Norstog 1974; Hoffman and Vaughn 1995). It is found at the anterior end of the gamete (Fig. 3), and it consists of a lamellar stack of fins and vanes that are attached ventrally to an anterior mitochondrion (Carothers 1975; Myles and Hepler 1977; Vaughn et al. 1993; Hoffman and Vaughn 1995). The uppermost portion of the MLS is a planar ribbon of crosslinked microtubules (Fig. 3) that extends in a posterior direction along the dorsal face of the coiled cell body. The microtubule ribbon is attached to the nuclear envelope. In M. vestita and most other organisms studied, the MLS forms after the basal bodies have been assembled. The clustered basal bodies become situated on the dorsal face of the ribbon (Myles and Hepler 1977; Marc and Gunning 1986; Hoffman and Vaughn 1995), and they then become dispersed and attached at regularly spaced intervals along the microtubule ribbon (Myles et al. 1978).
Electron micrographs of spermatids of Riella americana, an aquatic liverwort. Cells were fixed in glutaraldehyde and OsO4 using standard procedures. a The anterior, apical portion of a spermatid, showing the MLS, with the section plane essentially orthogonal to the longitudinal axis of the microtubule ribbon (spline). The anterior mitocondrion is clearly visible below the lower strata of the MLS. On the dorsal side of the microtubules, portions of basal bodies are visible. b An enlarged view of the micrograph in a showing the MLS. The sections through the ciliary axonemes show no outer dynein arms; they are lacking in these spermatozoids (Wolniak and Cande 1980). c A section of a spermatid cut at about 90° relative to the cell depicted in a and b. Here, the substrata of the MLS are clearly visible as a series of fins and vanes. The dorsal face of the MLS shows parts of a basal body and the ventral side of the MLS is attached to the anterior mitochondrion. Bars = 200 nm
The function of the MLS is not known, though it clearly plays a role in guiding the pattern of cell and nuclear elongation during spermatid maturation (Myles and Hepler 1977, 1982; Hoffman and Vaughn 1995; Lopez-Smith and Renzaglia 2008; Deeb et al. 2010). The fins and vanes of the structure at its anterior end form an obvious attachment site for a mitochondrion (Fig. 3), and the orientation of each stratum is precise, with regular spacing between the repeating fins and vanes. At a minimum, the MLS controls the placement of the basal bodies and their ciliary axonemes along the length of the coiled cell body of the gamete (Marc and Gunning 1986). Since the basal bodies are attached along the dorsal face of the microtubule ribbon (Carothers 1975; Myles and Hepler 1977; Myles et al. 1978; Vaughn et al. 1993), it is possible that the ribbon plays a role in coordinating ciliary metachrony in the swimming gamete (Wolniak and Cande 1980) and in cell body shape as the spermatozoid moves down the archegonial tube just prior to fertilization (Lopez-Smith and Renzaglia 2008). Algal flagellar rootlets are involved in positioning axonemes and they probably function in the coordination of flagellar or ciliary beat (Hyams and Borisy 1978). It is reasonable to suspect that the MLS serves a similar function. Is the MLS therefore related to algal rootlets? Both contain centrin (Salisbury 1983; Melkonian et al. 1988; Vaughn et al. 1993; Vaughn and Renzaglia 2006), but differences in rootlet striation patterns and MLS vane organization hint that the two arrays may have distinctly different compositions.
In some algae, bryophytes, and lycopods, the spermatozoids possess a pair of basal bodies, with their attached ciliary axonemes (Carothers 1975). Since the beat pattern is three-dimensional with a power stroke propagated along the entire length of the axoneme (Wolniak and Cande 1980), we refer to these structures in all of these spermatozoids as cilia. In the ferns and fern allies (e.g., Equisetum), the cell body is coiled and each gamete possesses ~30–150 cilia (Duckett 1973). The basal bodies in these gametes are arranged in single or double rows (Myles et al. 1978), and in some organisms there may be a staggered arrangement of basal bodies (Marc and Gunning 1986). In the spermatozoid-producing gymnosperms (Ginkgo and the cycads: Norstog 1974; Gifford and Lin 1975; Vaughn and Renzaglia 2006), the gametes are large and spherical, and the top layer of the MLS consists of a helical band of microtubules with over 1,000 basal bodies attached distally. The width and the length of the microtubule ribbon vary among organisms (Carothers 1975). For male gametophytes of this plant group, there is a loose, inverse relationship between the numbers of gametes produced and the numbers of cilia present on each spermatozoid. There is also an ironic inverse relationship between ciliary number and distance traveled by the spermatozoids: the gymnosperms, with their thousands of cilia, may have to swim only a few millimeters as they travel from the pollen tube tip to the archegonium of the female gametophyte. In contrast, fern spermatozoids, with (only) dozens of cilia, may swim several to many meters in thin films of water before reaching the archegonium of a suitable female gametophyte.
As a component in the blepharoplast, in basal bodies, and in the MLS, centrin serves as a specific marker for spermatogenous cells
Centrin is a calcium-binding protein (Salisbury 1995, 2007) that was originally described as a component in contractile flagellar rootlet complexes found in algal cells (Salisbury 1983; Melkonian et al. 1988). The protein is now known to be associated with centrosomes (Middendorp et al. 2000), centrioles (Paoletti et al. 1996; Geimer and Melkonian 2005), and basal bodies (Baron et al. 1992; Taillon et al.et al. 1992; Weich et al.et al. 1996; Levy et al. 1996, 1998; Geimer and Melkonian 2005) in a wide variety of organisms. Of particular importance for the development of plant spermatids is the presence of centrin in the blepharoplast (Klink and Wolniak 2001), in basal bodies (van der Weele et al. 2007), and in the MLS (Vaughn et al. 1993; Vaughn and Renzaglia 2006), where it apparently plays essential roles in the formation and function of these macromolecular arrays.
In the course of asking which mRNAs were essential for development of the gametophyte, we found that centrin protein demonstrated a pronounced increase in abundance in the spermatogenous cells approximately 4 h after the spores were hydrated (Hart and Wolniak 1998; Klink and Wolniak 2001). By contrast, centrin protein was undetectable in the jacket cells, which surround the spermatogenous cells of the gametophyte (Klink and Wolniak 2001; Tsai et al. 2004). The levels of β- and γ-tubulins were abundant in the newly hydrated spore and present at high levels throughout development (Klink and Wolniak 2003; Tsai et al. 2004), though these proteins were also predominantly localized in the spermatogenous cells of the gametophyte (Klink and Wolniak 2003; Tsai et al. 2004). The appearance and abundance of centrin protein was unaffected by α-amanitin, though in marked contrast centrin protein was undetectable in gametophytes treated with cycloheximide (Hart and Wolniak 1998). The translation of new centrin protein was dependent on transcripts that were pre-existing in the microspore before desiccation (Hart and Wolniak 1999). Thus, key features in this rapid developmental process rest in how the transcripts are stored in the spore, how they are distributed among the cells of the gametophyte, and how (and where) they are translated at the appropriate stage of development.
The rise in centrin protein abundance occurs at 4 h of development whether or not the division cycles have been suppressed (Tsai and Wolniak 2001), and obviously, under conditions of mitotic inhibition, centrin translation occurs in all cells present inside the spore wall. In spite of the fact that centrin is translated exclusively in spermatogenous cells of normal gametophytes, centrin mRNAs are equally abundant in spermatogenous and sterile cells in the gametophyte (Tsai et al. 2004).
Centrin interactions that may regulate patterns of spermatid development
As they function in association with centrioles or basal bodies (Marshall and Rosenbaum 2000), centrins have been found to interact with a variety of other proteins (Baron et al. 1992; Klotz et al. 1997), such as pericentrin (Doxsey et al. 1994), γ-tubulin, and cytoplasmic dynein (Young et al. 2000). We have not yet found bone fide cytoplasmic dynein or pericentrin cDNAs in our library, but we possess one cDNA that encodes γ-tubulin. On first principles, it made sense for us to look at γ-tubulin, a protein that resides at the minus ends of microtubules and participates in microtubule nucleation and/or assembly (Hoffman et al. 1994; Zheng et al. 1995; Erickson 2000; Murata et al. 2005). γ-Tubulin also associates with centrioles and basal bodies (Dibbayawan et al. 1995) and plays an important role in basal body duplication (Ruiz et al. 1999). However, an inherent problem with our gene silencing approach and γ-tubulin in the gametophyte is that the existing protein in the cell may obscure the effects of silencing, so it was essential for us to focus RNAi treatments on proteins that are either absent or limiting in a cell for a specific process. γ-Tubulin protein does not appear to be limiting in the gametophyte (Hart and Wolniak 1998; Klink and Wolniak 2003). In contrast, centrin is limiting for blepharoplast and basal body formation, while α-tubulin is limiting for basal body assembly, and the later formation of the MLS (van der Weele and Wolniak, unpublished observations).
Beyond γ-tubulin itself, the γ-tubulin nucleation complex is composed of several proteins, which include Xgrip109 (Martin et al. 1998), RanBPM, and other minor components (Schumacher et al. 1998a, b). Xgrip109 and RanBPM interact with γ-tubulin to form a ring complex (γTuRC) (Martin et al. 1998; Moritz et al. 2000; Wiese and Zheng 2000) that functions in a variety of cells to facilitate an orderly microtubule assembly. The γTuRC resides at the basal (minus) end of a microtubule and forms a cap-like structure. Dutcher and Trabuco (1998) found δ-tubulin as a minor, but significant, component of the basal body that is essential for triplet formation and stability. Recently, additional components (ZYG-1, O'Connell et al. 2001; SAS-4, SAS-5, SAS6, Dammermann et al. 2004; Leidel et al. 2005; NEDD1/GCP-WD, Luders et al. 2006) have been found in and around centrioles and basal bodies, and their orthologs could play similar assembly and stability roles in M. vestita. Some of these proteins have been identified in immunoblots of gametophyte isolates from specific stages of development (Klink and Wolniak 2003) and represent likely components under strict regulation during gametogenesis. As one might expect from a complex developmental program that follows a precise time table, the appearance of these proteins is under strict temporal control, and they are translated almost exclusively in the spermatogenous cells of the gametophyte (Klink and Wolniak 2003).
Cell fate determination in the male gametophyte of M. vestita
Histological staining with toluidine blue-O (Fig. 1) or immunolabeling of gametophytes fixed at different stages of development with anti-centrin (Fig. 4) and anti-β-tubulin antibodies reveals that the spermatogenous cells are quite different from the sterile jacket cells, not only in size but also in composition (Klink and Wolniak 2003). The spermatogenous cells are the only cells in the gametophyte that label with anti-β-tubulin antibody, and they appear to be the only sites where centrin is translated (Klink and Wolniak 2001). This nonrandom distribution of centrin and β-tubulin antibody labeling suggests that protein and perhaps mRNA segregation occurs within the gametophyte prior to its partitioning by cytokinesis (Tsai and Wolniak 2001). Since the mechanisms that underlie the compositional asymmetries between adjacent cells remain obscure, we began a series of localization studies as a means to document the distribution of specific mRNAs and proteins over time in the developing gametophytes (Klink and Wolniak 2003; Tsai et al. 2004).
A male gametophyte late in development, fixed 8 h after spores were placed into water. The fixed gametophytes were fixed in paraformaldehyde and embedded in methacylate as described elsewhere (van der Weele et al. 2007). After sectioning, the plastic was dissolved in acetone. After rehydration, the sections were stained with DAPI and then labeled with anti-centrin antibody (a kind gift of J. Salisbury, Rochester, MN, USA); after buffer rinsing, these were labeled with an Alexa 595-conjugated secondary antibody. This image is a set of merged phase contrast (gray) and fluorescence images. The spermatids show elongated nuclei (blue) and basal bodies stained red with anti-centrin antibody. The basal bodies are regularly spaced along the (invisible) microtubule ribbon of the MLS. The spermatid contains small plastids that are clustered in the ventral part of the cell. The jacket cells are mostly inconspicuous at this stage of development. The exine and the intine of the microspore wall often exhibit autofluorescence; note that the colors of the autofluorescence do not match those of the DAPI and Alexa fluorescence emissions (see Deeb et al. 2010). Bar = 25 μm
We extended our analyses of new translation beyond centrin and tubulin (Hart and Wolniak 1998, 1999; Klink and Wolniak 2001) by assaying for the abundance of proteins likely to be present in ciliated cells during gametophyte development. We tested 87 antibodies that were directed against cytoskeletal, centrosomal, and axonemal antigens for binding to gametophyte protein isolates. The antibodies were kindly provided to us by members of the research community, and among the antibodies provided nearly 20 provided single-band binding on western blots at the appropriate, apparent molecular weights. Our aim was to identify the conserved components of the cytoskeleton and ciliary apparatus that could serve as limiting factors for later phases of development, when the spermatid was engaged in forming its massive cytoskeleton or its ciliary array. We found varying patterns of protein abundance during development. Proteins like the tubulins were abundant at the onset of spore hydration and remained among the dominant proteins in the cells during development (Klink and Wolniak 2003). γ-Tubulin amounts appeared not to change throughout the process of development, while gradual increases in α-tubulin during late development have recently become evident. Other proteins increased dramatically in abundance at specific points during development in a fashion similar to centrin. We saw no consistent or striking declines in immunoblot binding during development with these antibodies, though conceivably there could be declines in the abundance of some proteins during spermatozoid maturation.
In parallel with protein abundance assays, we (Klink and Wolniak 2003) performed a series of RNAi silencing treatments to determine whether the silencing of different transcripts affected development at specific stages or, alternatively, that silencing of any single transcript shut down the entire developmental program in a general fashion. We found that, of the 39 cDNAs used to make dsRNA in this study (one third known to encode existing proteins, one third hypothetically linked to known proteins by sequence similarities, and the last third encoding nothing characterized in the databases), the vast majority of these dsRNAs arrested development of the gametophyte at specific time points during gametogenesis. A few of these dsRNAs either allowed spermiogenesis to reach completion or they exerted arrest late enough to make it difficult to discern the timing of their effects. The fact that silencing with these dsRNAs was so effective indicates that (a) there are many new proteins that must be synthesized for gametophyte development to reach completion and (b) the absence of any of a number of proteins creates a situation where development cannot continue; so, each of these proteins can effectively act as a limiting factor to the completion of gamete formation. Thus, even though there is a large pool of stored protein present in the microspore when it is newly hydrated (Hart and Wolniak 1998), this pool does not represent a complete complement of proteins necessary to make spermatozoids.
As mentioned above, the disruption or arrest of division cycles in the gametophyte, either through chemical inhibition or RNAi-induced gene silencing (Tsai and Wolniak 2001), resulted in changes in the distributions of proteins normally made only in spermatogenous cells. These products of translation were present in the cytoplasm of all cells of the gametophyte. Thus, the mediators of translational control may be restricting protein synthesis largely or entirely to the spermatogenous cells. Since the spermatogenous and jacket cells all originate from the same progenitor cell and since the spatial pattern of division planes is highly predictable for the formation of spermatogenous and sterile cells in the gametophyte, the process of defining cell fate may be regulated by positioning particular components in subcellular domains that would later become spermatogenous or jacket cells after the division cycles took place.
Translational capacity in spermatogenous and sterile cells of the male gametophyte of M. vestita
We found that a relatively large number of transcripts can be detected in all cells of the gametophyte during the first 4 h of its development (Tsai et al. 2004). It was surprising to find mRNAs encoding cyclins in the jacket cells since the jacket cells no longer divide once they are formed. It was equally surprising to find centrin transcripts to be present in jacket cells where centrin protein is not normally made. Thus, a critical factor in jacket cell and spermatogenous cell specification is not in the complement of transcripts present in each type of cell but rather on whether the cell can translate the transcripts that are present in its cytoplasm. This surprising discrepancy led us to surmise that the spermatogenous cells exhibit the capacity to translate mRNAs while the sterile jacket cells lack this ability. Could the transcripts present in spermatogenous cells undergo modifications that were not occurring in the adjacent sterile cells? We used in situ hybridization assays to assess the levels of transcript polyadenylation during development, reasoning that polyadenylation of the transcripts could underlie the differences in translation observed in sterile and spermatogenous cells of the gametophyte. Cyclin translation patterns have been found to correlate closely with polyadenylation (De Moor and Richter 1999). Moreover, differences in polyadenylation of related transcripts have been attributed to sequence variations in 3′-UTRs (Tremblay et al. 2005; Slevin et al. 2007). We found that the spermatogenous cells contained large numbers of polyadenylated transcripts, while they were largely absent from the adjacent jacket cells. We have not yet found 3′-UTR variants among cyclin mRNAs or other transcripts in our gametophytes, but we raised the obvious question of how differences in polyadenylation could arise in adjacent cells. We used an immunological assay to detect cytoplasmic polyA polymerase (PAP) and found that the spermatogenous cells exhibited intense labeling while the jacket cells were almost devoid of antibody binding (Tsai et al. 2004). We found it intriguing that the anti-PAP antibody also heavily labeled the cytoplasmic vesicle of the mature spermatozoids, perhaps indicating that this enzyme is carried by the male gamete to the egg, where it could polyadenylate maternal mRNAs during the early division cycles in the embryo. In M. vestita embryos, new transcription does not commence until the eight cell division cycle, and polyadenylated transcripts can be detected in the nuclei only after the 16-cell stage (Kuligowski et al. 1991).
Regulatory factors that mediate fate determination and morphogenesis in the gametophyte
In an effort to understand the mechanism of cell fate determination during the formation of spermatogenous initials, we focused on proteins and genes that are linked to the control of cell fate in other organisms. In a screen of our male gametophyte cDNA library, we isolated a cDNA that encodes a protein with a high level of homology to a polarity regulator from Drosophila melanogaster, which is known as mago nashi (van der Weele et al. 2007). Mago nashi was originally found as a maternal effect mutation in D. melanogaster that disrupts the anterior–posterior axis and embryonic germ cell formation (Boswell et al. 1991). Mago nashi was later characterized as a posterior group gene that functions in the localization of oskar mRNA (Mohr et al. 2001) and Staufen protein. It is involved in the polarization of the microtubule cytoskeleton and in nuclear migration (Newmark and Boswell 1994; Newmark et al. 1997; Micklem et al. 1997). Our Mv-mago RNAi experiments (van der Weele et al. 2007) demonstrate that Mv-mago protein performs multiple roles in the male gametophyte. Mv-mago protein is required for the proper control of the plane of cell division, especially during the normally asymmetric divisions that give rise to the sterile jacket cells. It also appears to be involved in the movement of some mRNAs. Surprisingly, most of the 30 stored mRNAs that we surveyed (e.g., Mv-cen-1, Mv-mago, cyclin A, cyclin B, β-tubulin, etc.) by in situ hybridization were equally abundant in sterile and spermatogenous cells of the gametophyte, though the proteins that they encode were predominantly (or exclusively) made in spermatogenous cells (Klink and Wolniak 2001; Tsai and Wolniak 2001; Tsai et al. 2004). A few mRNAs, such as the one encoding PRP19, a spliceosome factor (Cheng et al. 1993), exhibit a different distribution pattern and are localized in the spermatogenous cells during normal development (Tsai et al. 2004). After the treatment of spores with Mv-mago dsRNA, Mv-PRP19 mRNAs became dispersed throughout the gametophyte. These findings led to the hypothesis that the Mv-mago protein regulates the translation of specific mRNAs in particular cytoplasmic domains in the gametophyte and this activity underlies a central aspect of cell specification since these domains presage the formation of sterile and spermatogenous cells early in gametophyte development.
Mago nashi is part of the exon junction complex, which is deposited onto pre-mRNA during splicing, 20–24 nucleotides upstream of exon–exon junctions, in a sequence-independent fashion (LeHir et al. 2000, 2001a, b; Kataoka et al. 2001; Kim et al. 2001; Shibuya et al. 2004; Tange et al. 2004). The EJC is attached to the mRNA by an RNA helicase, eIF4AIII (Andersen et al. 2006). Mago nashi, with its binding partner Y14 (Lau et al. 2003), inhibits the ATPase activity of eIF4AIII, which keeps it locked onto the mRNA (Ballut et al. 2005). Mago nashi, Y14, and eIF4AIII are three proteins of the heterotetramer that make up the EJC core while the other components of the EJC are transiently associated with it (Palacios et al. 2004; Shibuya et al. 2004; Bono et al. 2004; Tange et al. 2005). We have isolated cDNAs that encode Mv-Y14 and Mv-eIF4AIII from our cDNA library, and we have found that they play important roles in basal body formation and nuclear remodeling in M. vestita spermatids.
As part of the EJC, mago nashi protein is found in the nucleus (Micklem et al. 1997; Newmark et al. 1997), where it can be localized in nuclear speckles (Degot et al. 2004), which are interchromatin regions enriched in pre-mRNA and splicing proteins (Lamond and Spector 2003; Spector and Lamond 2011). Depending on the proteins that associate with the core complex, the EJC can play a role in nuclear mRNA export (Luo and Reed 1999; Zhou et al. 2000; LeHir et al. 2001b), transcript quality control via the nonsense-mediated degradation pathway (Gehring et al. 2003; Tange et al. 2004), and translational enhancement (Wiegand et al. 2003; Nott et al. 2004). The axis formation and patterned movements of transcripts and proteins have been linked to mago function, but the precise signals that trigger the movement of mRNA or RNA binding proteins are unknown. Mago nashi mutant oocytes in Drosophila develop a symmetric microtubule cytoskeleton (Micklem et al. 1997), and it is likely that interactions between mago protein and cytoskeletal elements underlie many spatial determination events linked to mago function (Palacios and St. Johnston 2002).
We have found that the RNAi-induced silencing of Mv-mago affects the division patterns in the developing gametophyte and subsequently allows centrin translation in both sterile and spermatogenous cells of the gametophyte. Remarkably, the newly translated centrin protein aggregated into blepharoplast-like particles in both spermatogenous and jacket cells (van der Weele et al.et al. 2007). In these gametophytes, the normally asymmetric divisions that produce sterile jacket cells in the gametophyte became more symmetric, and the specification of segregated sterile and spermatogenous cells was lost. Since similar shifts of division planes are observed and similar patterns of centrin translation occurred with the silencing of the other core components of the EJC, Y14, or EiF4AIII, it seems clear that the EJC plays a key regulatory role in cell fate determination in the gametophyte.
Immunolabeling showed the distribution of Mv-Mago protein as punctae in the cytoplasm of spermatogenous and sterile cells of the developing gametophytes (van der Weele et al. 2007). We called these cytoplasmic punctae “mago-dots” to distinguish them from nuclear speckles. The silencing of Mago nashi, Y14, or Eif4AIII all resulted in the disappearance of mago dots, thereby suggesting that these dots are sites of EJC activity or are affected by EJC activity. In the absence of Mv-mago, cell specification fails to occur in the gametophyte, which led us to conclude that it may possibly act as a negative regulator for blepharoplast formation or, alternatively, that it affects other components that are essential for the aggregation of centrin protein into the blepharoplast particle. We do not know yet if Mv-mago and the other EJC components are functioning in the cytoplasmic splicing of transcripts.
Spermidine and cell fate determination in the gametophyte
It next became obvious to focus on how the transcripts are stored in the spore and then on how they are released from storage so that they can be translated. Earlier, we found that transcripts encoding PRP-19 and eIF4AIII appeared in in situ hybridization assays as intense granules of labeling in the cytoplasm of spermatogenous cells of the gametophyte, as the last of the division cycles was completed (Tsai et al. 2004). In seemingly unrelated experiments, a random screen of our cDNA library revealed the presence of a cDNA encoding spermidine synthase (SPDS), a key enzyme involved in the production of the polyamine spermidine. We suspected that spermidine might play an important role in mediating the developmental progression in the gametophyte since its activating properties are so widespread throughout the living realm (Tabor and Tabor 1984; Yatin 2002; Kaur-Sawhney et al. 2003); so, we assessed the distribution patterns of SPDS and spermidine in the gametophyte (Deeb et al. 2010). In situ hybridizations revealed that transcripts encoding SPDS became distributed in the peripheral cytoplasm of the spore shortly after hydration and that the abundance of spermidine in the jacket cells rose dramatically as the spermatogenous cells continued to undergo their last four division cycles. In situ hybridizations revealed the sudden appearance of SPDS transcripts in the spermatids after about 4.5 h of development, which also became detectable even if the cells had been treated with α-amanitin. Thus, the SPDS transcripts were already present in the cells, and when they were not detectable by in situ hybridization they were most likely sequestered as masked mRNAs. The appearance of SPDS transcripts was strikingly similar to the appearance of mRNAs encoding PRP-19 and eIF4AIII. Following the rise in transcript abundance in the spermatids, we saw an increase in spermidine levels in these newly formed spermatids.
Changes in the abundance and distribution of spermidine appear to be linked to the temporal and spatial patterns of translation of centrin, α-tubulin, and other components necessary for basal body formation and cytoskeletal assembly (Deeb et al. 2010). Spermidine is also clearly involved in nuclear remodeling later in gametogenesis (Bode et al. 1977; Shin et al. 2007). Soon after we can detect SPDS mRNA in sites where jacket cells will form, we can detect spermidine in the newly formed jacket cells; so, the enzyme is active. At this point in development, we can detect no spermidine in the spermatogenous cells. When the division cycles are completed and the spermatids have formed, immunolocalizations reveal that spermidine begins to accumulate in the spermatids, and it is apparently transported there from the jacket cells. We then see the emergence of a strong in situ hybridization signal for SPDS mRNA in the spermatids. SPDS is translated in the spermatids later than centrin and its increase in abundance is followed by a substantial increase in spermidine in the developing spermatids. If SPDS is silenced, basal body assembly is anomalous and the MLS fails to form properly. The inhibition of SPDS activity by treatment of gametophytes with cyclohexylamine results in a block to nuclear elongation and nuclei remain ovoid. Neither blepharoplasts nor basal bodies form in the gametophyte. Clearly, spermidine plays several important roles in gametogenesis in M. vestita and some of these roles involve the assembly of macromolecular complexes.
We think that it is important that the early production of spermidine occurs in the sterile jacket cells, about the time they are separated from the spermatogenous cells. This creates two distinctly different environments: one relatively rich in spermidine and one poor in spermidine. Normally, centrin translation occurs only in a spermidine-poor environment. We observe centrin translation in all cells if we block cell divisions or if we disrupt the cytoskeleton. Under these conditions, centrin does not aggregate into blepharoplasts. Is centrin aggregation into the blepharoplast controlled by spermidine? Spermidine enters the spermatids as the blepharoplast forms and spermidine levels continue to rise as the basal bodies are assembled and as they separate from each other. Does spermidine induce blepharoplast maturation and basal body assembly? We know that spermidine is essential for chromatin condensation and remodeling in the elongating spermatid nucleus (Deeb et al. 2010). Does spermidine affect MLS assembly and the elongation of the microtubule ribbon, which in turn affects nuclear elongation?
The EJC controls the patterns of centrin translation: in normal gametophytes, centrin is made in the spermatogenous cells, but not in the jacket cells. The silencing of Mv-mago results in centrin translation in all cells, with aggregation of centrin into blepharoplast-like particles. Under normal growth conditions, does spermidine inhibit centrin translation in the jacket cells of normal gametophytes by affecting the EJC? Later in development, the presence of spermidine in the spermatids is followed by the rapid appearance of SPDS mRNAs, which were already present in the cells but apparently packaged in a complex inaccessible to our in situ probes or to the translational machinery. What is the relationship between spermidine and the formation of mago-dots?
Is spermidine essential for the unpacking of stored transcripts in spermatids or is the action of the polyamine restricted to the unmasking of SPDS mRNAs? We tested this idea experimentally by adding spermidine and other polyamines to populations of spores, and we found that a variety of masked transcripts rapidly become detectable by in situ hybridization (Deeb et al. 2010). We think that it is important to point out that, under conditions of polyamine addition where these transcripts become (precociously) detectable, development is arrested. This is exactly what would be expected in a complex developmental program where there is precise temporal and spatial control over the release of masked mRNAs, and their availability for processing and subsequent translation controls the rates of cellular morphogenesis.
Conclusions and future perspectives
The endosporic gametophyte of the water fern, M. vestita, provides a unique opportunity to study the mechanisms that control cell fate determination in a rapidly developing system. This gametophyte utilizes nine successive cell division cycles in precise planes within a closed volume to produce seven sterile cells and 32 spermatids. There is no cell movement in the gametophyte, so cell position, cell size, and cell composition act collectively within the spore wall to define cell fate. After the division cycles are completed, the spermatids are sites for the de novo formation of basal bodies, for the assembly of a complex cytoskeleton, for nuclear and cell elongation, and for ciliogenesis. The spermatids differentiate into multiciliated, corkscrew-shaped gametes that resemble no other cells in the entire plant. The process reaches completion in less than 11 h and is controlled post-transcriptionally; the stored transcripts are released in the cytoplasm and become detectable at different times by in situ hybridization analysis. The silencing of specific mRNAs in the gametophyte blocks development at specific stages, revealing that the translation of these proteins at particular times is essential for development to proceed. Since the distributions of mRNAs and the proteins they encode are not identical in the gametophyte, it is clear that transcript processing is important in allowing translation to occur under strict temporal and spatial control. At least some of the transcripts appear to be modified before they are translated. Transcript polyadenylation occurs in the spermatogenous cells, and it appears to be under the control of a cytoplasmic PAP. It is clear that the EJC plays key roles in transcript processing, which is essential for the translation of proteins necessary for spermatozoid maturation, and it may be involved in the regulation of splicing of these masked mRNAs. The rapid burst of development in the endosporic gametophyte of M. vestita is controlled post-transcriptionally, but even under conditions where there is no cell movement the mechanisms that mediate developmental progression are multifaceted and functioning under strict spatial and temporal controls.
References
Amaldi PP, Felicetti L, Campioni N (1977) Flow of informational RNA from cytoplasmic poly(A)-containing particles to polyribosomes in Artemia salina cysts at early stages of development. Dev Biol 59:49–61
Andersen CBF, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Olivera CLP, Pedersen JS, Seraphin B, LeHir H, Andersen GR (2006) Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science 313:1968–1972
Ballut L, Marchadier B, Baguet A, Tomasetto C, Seraphin B, LeHir H (2005) The exon junction core complex is locked onto RNA by inhibition of eIF4AIII ATPase activity. Nat Struct Mol Biol 12:861–869
Baron AT, Greenwood T, Bazinet C, Salisbury JL (1992) Centrin is a component of the pericentriolar lattice. Biol Cell 76:383–388
Bode J, Willmitzer L, Opatz K (1977) On the competition between protamines and histones: studies directed towards the understanding of spermiogenesis. Eur J Biochem 72:393–403
Bono F, Ebert J, Unterholzner L, Guttler T, Izaurralde E, Conti E (2004) Molecular insights into the interaction of PYM with the Mago-Y14 core of the exon junction complex. EMBO Rep 5:304–310
Boscher JM, Labouesse M (2000) RNA interference: genetic wand and genetic watchdog. Nat Cell Biol 2:E31–E36
Boswell RE, Prout ME, Steichen JC (1991) Mutations in a newly identified Drosophila melanogaster gene, mago nashi, disrupt germ cell formation and result in the formation of mirror-image symmetrical double abdomen embryos. Development 113:373–384
Brown RC, Carothers ZB (1986) Comparative studies of spermatogenesis in the Bryopsida. II. Blepharoplast morphology in Archidium tenerrinum Mitt. The Bryrologist 89:42–48
Carothers ZB (1973) Studies of spermatogenesis in the Hepaticae. IV. On the blepharoplast of Blasia. Am J Bot 60:819–828
Carothers ZB (1975) Comparative studies on spermatogenesis in bryophytes. In: Duckett JG, Racey PA (eds). The biology of the male gamete. Biol J Linn Soc 7(Suppl 1):71–84
Carothers ZB, Kreitner GL (1967) Studies of spermatogenesis in the Hepaticae: I. Ultrastructure of the Vierergruppe in Marchantia. J Cell Biol 33:43–51
Carothers ZB, Rushing AE (1990) Blepharoplast morphology in Treubia tasmanica (Hepiticae: Treubiales). Bryologist 93:409–416
Carothers ZB, Robbins RR, Haas DL (1975) Some ultrastructural aspects of spermatogenesis in Lycopodium complanatum. Protoplasma 86:339–350
Chamberlain CJ (1898) The homology of the blepharoplast. Bot Gaz 26:431–435
Chamberlain CJ (1909) Spermatogenesis in Dioon edule. Bot Gaz 47:215–236
Chapman RL, Henk MC (1983) Ultrastructure of Cephaleuros virescens (Chroolepidaceae; Chlorophyta). IV. Absolute configuration analysis of the cruciate flagellar apparatus and multilayered structures in the pre- and post-release gametes. Amer J Bot 70:1340–1355
Cheng SC, Tarn WY, Tsao TY, Abelson J (1993) PRP19: a novel spliceosomal component. Mol Cell Biol 13:1876–1882
Comai L, Dietrich RA, Maslyar DJ, Baden CS, Harada JJ (1989) Coordinate expression of transcriptionally regulated isocitrase lyase and malate synthase genes in Brassica napus L. Plant Cell 1:293–300
Dammermann A, Muller-Reichert T, Pelletier L, Habermann B, Desai A, Oegema K (2004) Centriole assembly requires both centriolar and pericentriolar material proteins. Dev Cell 7:815–829
De Moor CH, Richter JD (1999) Cytoplasmic polyadenylation elements mediate masking and unmasking of cyclin B1 mRNA. EMBO J 18:2294–2303
Deeb F, van der Weele CM, Wolniak SM (2010) Spermidine is a morphogenetic determinant for cell fate specification in the male gametophyte of the water fern Marsilea vestita. Plant Cell 22:3678–3691
Degot S, Le Hir H, Alpy F, Kedinger V, Stoll I, Wendling C, Seraphin B, Rio M-C, Tomasetto C (2004) Association of the breast cancer protein MLN51 with the exon junction complex via its speckle localizer and RNA binding module. J Biol Chem 279:33702–33715
Dibbayawan T, Harper JDI, Elliot J, Gunning BES, Marc J (1995) A γ-tubulin that associates specifically with centrioles in HeLa cells and the basal body complex in Chlamydomonas. Cell Biol Int 19:559–567
Doxsey SJ, Stein P, Evans L, Calarco P, Kirschner M (1994) Pericentrin, a highly conserved protein of centrosomes involved in microtubule organization. Cell 76:639–650
Duckett JG (1973) An ultrastructural study of the differentiation of the spermatozoid of Equisetum. J Cell Sci 12:95–129
Dure L, Waters L (1965) Long-lived messenger RNA: evidence from cotton seed germination. Science 147:410–412
Dutcher SK, Trabuco EC (1998) The UNI3 gene is required for assembly of basal bodies of Chlamydomonas and encodes delta-tubulin, a new member of the tubulin superfamily. Mol Biol Cell 9:1293–1308
Erickson HP (2000) γ-Tubulin nucleation: template or protofilament? Nat Cell Biol 2:E93–E96
Fire A (1999) RNA-triggered gene silencing. Trends Genet 15:358–363
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Fox CA, Sheets MD, Wickens MP (1989) Poly(A) addition during maturation of frog oocytes: distinct nuclear and cytoplasmic activities and regulation by the sequence UUUUUAU. Genes Dev 3:2151–2162
Gehring NH, Neu-Yilik G, Schell T, Hentze MW, Kulozik AE (2003) Y14 and hUpf3b form an NMD-activating complex. Mol Cell 11:939–949
Geimer S, Melkonian M (2005) Centrin scaffold in Chlamydomonas reinhardtii revealed by immunoelectron microscopy. Eukaryot Cell 4:1253–1263
Gifford EM Jr, Lin J (1975) Light microscope and ultrastructural studies of the male gametophyte in Ginkgo biloba: the spermatogenous cell. Am J Bot 62:974–981
Glab N, Lavidl B, Quin LX, Trehin C, Bergounioux C, Meijer L (1994) Olomoucine, an inhibitor of the cdc2/cdk2 kinases activity, blocks plant cells at the G1 to S and G2 to M cell cycle transitions. FEBS Lett 353:207–211
Gorgoni B, Gray NK (2004) The roles of cytoplasmic poly(A)-binding proteins in regulating gene expression: a developmental perspective. Brief Funct Genomic Proteomic 3:125–141
Grosfeld H, Littauer UZ (1975) Cryptic form of mRNA in dormant Artemia salina cysts. Biochem Biophys Res Commun 67:176–181
Gross PR, Cousineau GH (1963) Effects of actinomycin D on macromolecule synthesis and early development in sea urchin eggs. Biochem Biophys Res Commun 18:321–326
Gross PR, Cousineau GH (1964) Macromolecule synthesis and the influence of actinomycin on early development. Exp Cell Res 33:368–395
Hart PE, Wolniak SM (1998) Spermiogenesis in Marsilea vestita: a temporal correlation between centrin expression and blepharoplast differentiation. Cell Motil Cytoskelet 41:39–48
Hart PE, Wolniak SM (1999) Molecular cloning of a centrin homolog from Marsilea vestita and evidence for its translational control during spermiogenesis. Biochem Cell Biol 77:101–108
Harvey EB (1936) Parthenogenetic merogony or cleavage, without nuclei in Arbacia punctulata. Biol Bull 71:101–121
Harvey EB (1940) A comparison of the development of nucleate and non-nucleate eggs of Arbacia punctulata. Biol Bull 79:166–187
Hepler PK (1976) The blepharoplast of Marsilea: its de novo formation and spindle association. J Cell Sci 21:361–390
Hoffman JC, Vaughn KC (1995) Using the developing spermatogenous cells of Ceratopteris to unlock the mysteries of the plant cytoskeleton. Int J Plant Sci 156:346–358
Hoffman JC, Vaughn KC, Joshi HC (1994) Structural and immunocytochemical characterization of microtubule organizing centers in pteridophyte spermatogenous cells. Protoplasma 179:46–60
Hughes DW, Galau GA (1989) Temporally modular gene expression during cotyledon development. Genes Dev 3:358–369
Hughes DW, Galau GA (1991) Developmental and environmental induction of Lea and LeaA mRNAs and the postabscission program during embryo culture. Plant Cell 3:605–618
Hyams JS, Borisy GG (1978) Isolated flagellar apparatus of Chlamydomonas: characterization of forward swimming and alteration of waveform and reversal of motion by calcium ions in vitro. J Cell Sci 33:235–253
Hyams JS, Vondy KP, Luba A, Bell PR (1983) Structural and macromolecular events associated with basal body morphogenesis in Marsilea. J Submicros Cytol 15:133–138
Kataoka N, Diem MD, Kim VK, Yong J, Dreyfuss G (2001) Magoh, a human homolog of Drosophila mago nashi protein, is a component of the splicing-dependent exon–exon junction complex. EMBO J 20:6424–6433
Kaur-Sawhney R, Tiburcio AF, Atlabells T, Galston AW (2003) Polyamines in plants: an overview. J Cell Mol Biol 2:1–12
Kim VN, Yong J, Kataoka N, Abel L, Diem MD, Dreyfuss G (2001) The Y14 protein communicates to the cytoplasm the position of exon–exon junctions. EMBO J 20:2062–2068
Kimura M, Nambara E (2010) Stored and neosynthesized mRNA in Arabidopsis seeds: effects of cycloheximide and controlled deterioration treatment on the resumption of transcription during imbibition. Plant Mol Biol 73:119–129
Klink VP, Wolniak SM (2001) Centrin is necessary for the formation of the motile apparatus in spermatids of Marsilea. Mol Biol Cell 12:761–776
Klink VP, Wolniak SM (2003) Changes in the abundance and distribution of conserved centrosomal, cytoskeletal and ciliary proteins during spermiogenesis in Marsilea vestita. Cell Motil Cytoskelet 56:57–73
Klotz C, deLoubresse NG, Ruiz F, Beisson J (1997) Genetic evidence for a role of centin-associated proteins in the organization and dynamics of the infraciliary lattice in Paramecium. Cell Motil Cytoskelet 38:172–186
Kreitner GL (1977) Transformation of the nucleus in Marchantia spermatids: morphogenesis. Am J Bot 64:464–475
Kreitner GL, Carothers ZB (1976) Studies of spermatogenesis in the Hepaticae. V. Blepharoplast development in Marchantia polymorpha. Am J Bot 63:545–557
Kuligowski J, Ferrand M, Chenou E (1991) Stored mRNA in early embryos of a fern Marsilea vestita: a paternal and maternal origin. Mol Reprod Dev 30:27–33. doi:10.1002/mrd.1080300104
Lamond AI, Spector DL (2003) Nuclear speckles: a model for nuclear organelles. Nat Rev Mol Cell Biol 4:605–612
Lau C-K, Dlem MD, Dreyfuss G, Van Duyne GD (2003) Structure of the Y14-Magoh core of the exon junction complex. Curr Biol 13:933–941
LeHir H, Izaurralde E, Maquat LE, Moore MJ (2000) The spliceosome deposits multiple proteins 20–24 nucleotides upstream of mRNA exon–exon junctions. EMBO J 19:6860–6869
LeHir H, Gatfield D, Braun IC, Forler D, Izaurralde E (2001a) The protein Mago provides a link between splicing and mRNA localization. EMBO Rep 2:1119–1124
LeHir H, Gatfield D, Izaurralde E, Moore MJ (2001b) The exon–exon junction complex provides a binding platform for factors involved in mRNA export and nonsense-mediated mRNA decay. EMBO J 20:4987–4997
Leidel S, Delattre M, Cerutti L, Baumer K, Gonczy P (2005) SAS-6 defines a protein family required for centrosome duplication in C. elegans and in human cells. Nat Cell Biol 7:115–125
Levy YY, Lai EY, Remillard SP, Heintzelman MB, Fulton C (1996) Centrin is a conserved protein that forms diverse associations with centrioles and MTOCs in Naegleria and other organisms. Cell Motil Cytoskelet 33:298–323
Levy YY, Lai EY, Remillard SP, Fulton C (1998) Centrin is synthesized and assembled into basal bodies during Naegleria differentiation. Cell Motil Cytoskelet 40:249–260
Li Y, Wang FH, Knox RB (1989) Ultrastructural analysis of the flagellar apparatus in sperm cells of Ginkgo biloba. Protoplasma 14:57–63
Lopez-Smith R, Renzaglia KS (2008) Sperm cell architecture, insemination, and fertilization in the model fern, Ceratopteris richardii. Sex Plant Reprod 21:153–167
Luders J, Patel UK, Stearns T (2006) GCP-WD is a γ-tubulin targeting factor required for centrosomal and chromatin-mediated microtubule nucleation. Nat Cell Biol 8:137–147
Luo M, Reed R (1999) Splicing is required for rapid and efficient mRNA export in metazoans. Proc Nat Acad Sci USA 96:14937–14942
Malatesta M, Zancanaro C, Martin TE, Chan EK, Amalric F, Luhrmann R, Vogel P, Fakan S (1994) Cytochemical and immunocytochemical characterization of nuclear bodies during hibernation. Eur J Cell Biol 65:82–93
Malatesta M, Cardinali A, Battistelli S, Zancanaro C, Martin TE, Fakan S, Gazzanelli G (1999) Nuclear bodies are usual constituents in tissue of hibernating dormice. Anat Rec 254:389–395
Maquat LE, Carmichael GG (2001) Quality control of mRNA function. Cell 104:173–176
Marc J, Gunning BES (1986) Immunofluorescent localization of cytoskeletal tubulin and actin during spermatogenesis in Pteridium aquilinum (L.) Kuhn. Protoplasma 134:163–177
Marshall WF, Rosenbaum JL (2000) How centrioles work: lessons from green yeast. Curr Opin Cell Biol 12:119–125
Martin OC, Gunawardane RN, Iwamatsu A, Zheng Y (1998) Xgrip109: a γ-tubulin-associated protein with an essential role in γ-tubulin ring complex (γTuRC) assembly and centrosome function. J Cell Biol 141:675–687
McGrew LL, Dworkin-Rastl E, Dworkin MB, Richter JD (1989) Poly(A) elongation during Xenopus oocyte maturation is required for translational recruitment and is mediated by a short sequence element. Genes Dev 3:803–815
Meier L (1996) Chemical inhibitors of cyclin-dependent kinases. Trends Cell Biol 6:393–397
Melkonian M, Schulze D, McFadden GI, Robenek H (1988) A polyclonal antibody (anticentrin) distinguishes between two types of fibrous flagellar roots in green algae. Protoplasma 144:56–61
Micklem DR, Dasgurpta R, Elliott H, Gergely F, Davidson C, Brand A, Gonzalez-Reyes A, St Johnston D (1997) The mago nashi gene is required for the polarisation of the oocyte and the formation of perpendicular axes in Drosophila. Curr Biol 7:468–478
Middendorp S, Kuntziger T, Abraham Y, Holmes S, Bordes N, Paintrand M, Paoletti A, Bornens M (2000) A role for centrin 3 in centrosome reproduction. J Cell Biol 148:405–415
Mizukami I, Gall J (1966) Centriole replication. II. Sperm formation in the fern, Marsilea and the cycad, Zamia. J Cell Biol 29:97–111
Mohr SE, Dillon ST, Boswell RE (2001) The RNA-binding protein Tsunagi interacts with Mago Nashi to establish polarity and localize oskar mRNA during Drosophila oogenesis. Genes Dev 15:2886–2899
Montgomery MK, Fire A (1998) Double-stranded RNA as a mediator in sequence-specific genetic silencing and co-suppression. Trends Genet 14:255–258
Moritz M, Braunfeld MB, Guenebaut V, Heuser J, Agard DA (2000) Structure of the γ-tubulin ring complex: a template for microtubule nucleation. Nat Cell Biol 2:365–370
Murata T, Sonobe S, Baskin TI, Hyodo S, Hasezawa S, Nagata T, Horio T, Hasebe M (2005) Microtubule-dependent microtubule nucleation based on recruitment of γ-tubulin in higher plants. Nat Cell Biol 7:961–968
Muthukrishnan S, Filipowicz W, Sierra JM, Both GW, Shatkin AJ, Ochoa S (1975) mRNA methylation and protein synthesis in extracts from embryos of brine shrimp, Artemia salina. J Biol Chem 250:9336–9341
Myles DG, Hepler PK (1977) Spermiogenesis in the fern Marsilea: microtubules, nuclear shaping and cytomorphogenesis. J Cell Sci 23:57–83
Myles DG, Hepler PK (1982) Shaping of the sperm nucleus in Marsilea: a distinction between factors responsible for shape generation and shape determination. Dev Biol 90:238–252
Myles DG, Southworth D, Hepler PK (1978) Cell surface topography during Marsilea spermiogenesis: flagellar reorientation and membrane particle arrays. Protoplasma 93:405–417
Nakabayashi K, Okamoto M, Koshiba T, Kamiya Y, Nambara E (2005) Genome-wide profiling of stored mRNA in Arabidopsis thaliana seed germination: epigenetic and genetic regulation of transcription in seed. Plant J 41:697–709
Newmark PA, Boswell RE (1994) The mago nashi locus encodes an essential product required for germ plasm assembly in Drosophila. Development 120:1303–1313
Newmark PA, Mohr SE, Gong L, Boswell RE (1997) Mago nashi mediates the posterior follicle cell-to-oocyte signal to organize axis formation in Drosophila. Development 124:3197–3207
Ngo J, Tshudi C, Gull K, Ullu E (1998) Double-stranded RNA induces mRNA degradation in Trypanosoma brucei. Proc Nat Acad Sci USA 95:14687–14692
Norstog K (1974) Fine structure of the spermatozoid of Zamia: the vieregruppe. Am J Bot 61:449–456
Nott A, Le Hir H, Moore MJ (2004) Splicing enhances translation in mammalian cells: an additional function of the exon junction complex. Genes Dev 18:210–222
O'Connell KF, Caron C, Kopish KR, Hurd DD, Kemphues KJ, Li Y, White J (2001) The C. elegans zyg-1 gene encodes a regulator of centrosome duplication with distinct maternal and paternal roles in the embryo. Cell 105:547–558
Palacios IM, St. Johnston D (2002) Kinesin light chain-independent function of the Kinesin heavy chain in cytoplasmic streaming and posterior localisation in the Drosophila oocyte. Development 129:5473–5485
Palacios IM, Gatfield D St, Johnston D, Izaurralde E (2004) An eIF4AIII-containing complex required for mRNA localization and nonsense-mediated mRNA decay. Nature 427:753–757
Paoletti A, Moudjou M, Paintrand M, Salisbury J, Bornens M (1996) Most of the centrin in animal cells is not centrosome-associated and centrosomal centrin is confined to the distal lumen of centrioles. J Cell Sci 109:3089–3102
Paolillo DJ Jr (1965) On the androcyte of Polytrichum, with special reference to the Dreiergruppe and the limosphere (Nebenkern). Can J Bot 43:669–676
Paolillo DJ Jr, Kreitner GL, Reighard J (1968) Spermatogenesis in Polytrichum junperinum. I. The origin of the apical body and elongation of the nucleus. Planta 78:226–247 (Berl)
Paris J, Phillippe M (1988) Poly(A) metabolism and polysomal recruitment of maternal mRNAs during early Xenopus development. Dev Biol 140:221–224
Pennell RI, Hyams JS, Bell PR (1986) The blepharoplast of Marsilea: a structure concerned with basal body assembly lacking tubulin. Eur J Cell Biol 40:238–241
Pennell RI, Vondy K, Bell PR, Hyams JS (1988) Composition and function of the blepharoplast of Marsilea vestita. Eur J Cell Biol 46:51–60
Pfeffer W (1884) Locomotorische Richtungsbewegungen durch chemische Reize. Unters Bot Inst Tubingen 1:363–482
Proweller A, Butler S (1994) Efficient translation of poly(A)-deficient mRNAs in Saccharomyces cerevisiae. Genes Dev 8:2629–2640
Rafiq M, Suen CKM, Choudhury N, Joannou CL, White KN, Evans RW (1997) Expression of recombinant human ceruloplasmin—an absolute requirement for splicing signals in the expression cassette. FEBS 407:132–136
Renzaglia KS, Carothers ZB (1986) Ultrastructural studies of spermatogenesis in the Anthocerotales. IV. The blepharoplast and mid-stage spermatid of Notothylas. J Hattori Bot Lab 60:97–104
Renzaglia KS, Garbary DJ (2001) Motile gametes of land plants: diversity, development and evolution. Crit Rev Plant Sci 20:107–213
Robert D (1977) Le noyau du spermnatozoide du Selaginella kraussiana: etude cytochemique en microscopie electronique. J Ultrastruct Res 58:178–195
Rosenthal ET, Tansey TR, Ruderman JV, Gottesman M (1983) Sequence-specific adenylation and deadenylations accompany changes in the translation of maternal messenger RNA after fertilization of Spisula oocytes. J Mol Biol 166:309–327
Ruiz F, Beisson J, Rossier J, Dupuis-Williams P (1999) Basal body duplication in Paramecium requires γ-tubulin. Curr Biol 9:43–46
Ryu WS, Mertz JE (1989) Simian virus 40 late transcripts lacking excisable intervening sequences are defective in both stability in the nucleus and transport to the cytoplasm. J Virol 63:4386–4394
Salisbury JL (1983) Contractile flagellar roots: the role of calcium. J Submicros Cytol 15:105–110
Salisbury JL (1995) Centrin, centrosomes, and mitotic spindle poles. Curr Opin Cell Biol 7:39–45
Salisbury JL (2007) A mechanistic view on the evolutionary origin for centrin-based control of centriole duplication. J Cell Physiol 213:420–428
Schumacher J, Ashcroft N, Donovan PJ, Golden A (1998a) A highly conserved centrosomal kinase, AIR-1, is required for accurate cell cycle progression and segregation of developmental factors in Caenorhabditis elegans embryos. Development 125:4391–4402
Schumacher J, Golden A, Donovan PJ (1998b) AIR-2: an Aurora/Ipl1-related protein kinase associated with chromosomes and midbody microtubules is required for polar body extrusion and cytokinesis in C. elegans embryos. J Cell Biol 143:1635–1646
Sharp LW (1912) Spermatogenesis in Equisetum. Bot Gaz 54:89–119
Sharp LW (1914) Spermatogenesis in Marsilia. Bot Gaz 58:419–431
Shibuya T, Tange TØ, Sonenberg N, Moore MJ (2004) eIF4AIII binds spliced mRNA in the exon junction complex and is essential for nonsense-mediated decay. Nat Struct Mol Biol 11:346–351
Shin M, Larsson L-I, Fujiwara K (2007) Polyamines in spermatocytes and residual bodies of rat testis. Biochem Cell Biol 127:649–655
Slevin MK, Gourronc F, Hartley RS (2007) ElrA binding to the 3′UTR of cyclin E1 mRNA requires polyadenylation elements. Nucleic Acids Res 35:2167–2176. doi:10.1093/nar/gkm084
Spector DL, Lamond AI (2011) Nuclear speckles. Cold Spring Harb Perspect Biol. doi:10.1101/cshperspect.a000646
Tabor CW, Tabor H (1984) Polyamines. Annu Rev Biochem 53:749–790
Taillon BE, Adler SA, Suhan JP, Jarvik JW (1992) Mutational analysis of centrin: an EF-hand protein associated with three distinct contractile fibers in the basal body apparatus of Chlamydomonas. J Cell Biol 119:1613–1624
Tange TØ, Nott A, Moore MJ (2004) The ever-increasing complexities of the exon junction complex. Curr Opin Cell Biol 16:279–284
Tange TØ, Shibuya T, Jurica MS, Moore MJ (2005) Biochemical analysis of the EJC reveals two new factors and a stable tetrameric protein core. RNA 11:1869–1883
Tremblay K, Vigneault C, McGraw S, Sirard M-A (2005) Expression of cyclin B1 messenger RNA isoforms and initiation of cytoplasmic polyadenylation in the bovine oocyte. Biol Rep 72:1037–1044. doi:10.1095/biolreprod.104.034793
Tsai CW, Wolniak SM (2001) Cell cycle arrest allows centrin translation but not basal body formation during spermiogenesis in Marsilea. J Cell Sci 114:4265–4272
Tsai CW, van der Weele CM, Wolniak SM (2004) Differential segregation and modification of mRNA during spermiogenesis in Marsilea vestita. Dev Biol 269:319–330
van der Weele CM, Tsai CW, Wolniak SM (2007) Mago nashi is essential for spermatogenesis in Marsilea. Mol Biol Cell 18:3711–3722
Vassalli J, Huarte J, Belin D, Gubler P, Vassalli A, O’Connell ML, Parton LA, Rickles RJ, Strickland S (1989) Regulated polyadenylation controls mRNA translation during meiotic maturation of mouse oocytes. Genes Dev 3:2163–2171
Vaughn KC, Renzaglia KS (2006) Structural and immunocytochemical characterization of the Ginkgo biloba L. sperm motility apparatus. Protoplasma 227:165–183
Vaughn KC, Sherman TD, Renzaglia KS (1993) A centrin homologue is a component of the multilayered structure in bryophytes and pteridophytes. Protoplasma 175:58–66
Webber HJ (1897) Notes on the fecundation of Zamia and the pollen tube apparatus of Ginkgo. Bot Gaz 24:225–235
Weich H, Grier B, Paschke T, Spang A, Grein K, Steinkotter J, Melkonian M, Scheibel E (1996) Characterization of green alga, yeast, and human centrins. J Biol Chem 271:22453–22461
Wiegand HL, Lu S, Cullen BR (2003) Exon junction complexes mediate the enhancing effect of splicing on mRNA expression. Proc Natl Acad Sci USA 100:11327–11332
Wiese C, Zheng Y (2000) A new function for the g-tubulin ring complex as a microtubule minus-end cap. Nat Cell Biol 2:358–364
Wilkinson MF, Shyu A (2002) RNA surveillance by nuclear scanning? Nat Cell Biol 4:144–147
Wilusz CJ, Wormington M, Peltz SW (2001) The cap-to-tail guide to mRNA turnover. Nat Rev Mol Cell Biol 2:237–246
Wolniak SM, Cande WZ (1980) Physiological requirements for ciliary reactivation of bracken fern spermatozoids. J Cell Sci 43:195–207
Wolniak SM, Klink VP, Hart PE, Tsai CW (2000) Control of development and motility in the spermatozoids of lower plants. Grav Sp Biol Bull 13:85–93
Yatin M (2002) Polyamines in living organisms. J Cell Mol Biol 1:57–67
Young A, Dictenberg JB, Purohit A, Tuft R, Doxsey SJ (2000) Cytoplasmic dynein-mediated assembly of pericentrin and γ-tubulin onto centrosomes. Mol Biol Cell 11:2047–2056
Zamore PD, Tuschl T, Sharp PA, Bartel DP (2000) RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101:25–33
Zheng Y, Wong ML, Alberts B, Mitchison T (1995) Nucleation of microtubule assembly by a γ-tubulin-containing ring structure. Nature 378:578–583
Zhou Z, Luo M, Straesser K, Katahira J, Hurt E, Reed R (2000) The Protein Aly links pre-messenger-RNA splicing to nuclear export in metazoans. Nature 407:401–405
Acknowledgements
We gratefully acknowledge the support for this project in the form of grants from NSF (MCB 0720486 and DBI 0842525). We are also grateful for the support from Maryland Agriculture Experiment Station for this project. Faten Deeb was partially supported by a USDA Food and Agricultural Sciences National Needs Graduate and Postdoctoral Fellowship Grant (#20053842015761) while this work was being performed.
Conflicts of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wolniak, S.M., van der Weele, C.M., Deeb, F. et al. Extremes in rapid cellular morphogenesis: post-transcriptional regulation of spermatogenesis in Marsilea vestita . Protoplasma 248, 457–473 (2011). https://doi.org/10.1007/s00709-011-0276-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-011-0276-3