Abstract
In the light of our previous work, we know that there is a relationship between bound polyamines and the chloroplast differentiation process. This relationship may represent an important component of the process and be part of the mechanism of kinetin action, which stimulates chloroplast differentiation. To clarify the nature of the binding of polyamines to chloroplast structures, the possible involvement of transglutaminases in kinetin-stimulated chloroplast photodevelopment was investigated. Immunodetection of transglutaminases revealed bands at 77, 50 and 30 kDa both in etioplasts and chloroplasts. The data indicated a positive correlation between enzyme level and activity. It also demonstrated the regulation of transglutaminase protein expression by kinetin. The suborganellar location of transglutaminases by electron microscopy showed that the enzyme is peculiarly localised, mainly in pro-thylakoids and appressed grana thylakoids. The data corroborated that spermidine post-translational modification of certain plastid proteins of 58, 29, 26 and 12 kDa occurred. The results we obtained suggest that transglutaminases take part in the formation of the chloroplast structure via a mechanism whereby polyamines bind to their protein substrates. These findings about the effect of kinetin on conjugation provide a new contribution to the understanding of the mechanism of kinetin action on etioplast-to chloroplast transformation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The light-dependent development of etioplasts into chloroplasts is one of the most interesting differentiation processes that occurs in plants. The chloroplast is an organelle of great metabolic complexity and its biogenesis is critical in determining the photosynthetic capacity of plants. One of the metabolic activities is the biosynthesis and catabolism of polyamines (PAs) and their conjugation to chloroplast structures (Kotzabasis et al. 1993, 1999; Del Duca et al. 1994, 2000; Dondini et al. 2003).
PAs are synthesised and oxidised in chloroplasts (Kotzabasis 1995; Andreadakis and Kotzabasis 1996; Bernet et al. 1999), and a correlation has been observed between PA levels, chlorophyll biosynthesis and the photosynthetic rate (Beigbeder et al. 1995). Kotzabasis et al. (1993) reported that the main polyamines, putrescine (Put), spermidine (Spd) and spermine (Spm), are associated with the light-harvesting complex (LHC) and the photosystem II (PSII) of spinach. PAs can be associated with many molecules by different types of binding (Del Duca and Serafini-Fracassini 1993). One of these is conjugation to protein via transglutaminases (TGases; R-glutaminyl-peptide: amine(γ-glutamyl)transferase, EC 2.3.2.13), a family of enzymes that catalyse the covalent binding of substrates with primary amine groups, like PAs, to the γ-carboxyamide group of protein endo-glutamine residues (Lorand and Graham 2003; Del Duca and Serafini-Fracassini 2005).
It has been suggested that the covalent binding of PAs to proteins might play a significant role in the post-translational modifications of structural proteins or enzymes in chloroplasts (Del Duca et al. 1994, 2000; Della Mea et al. 2004b). Chloroplast TGases have been reported to catalyse the conjugation of PAs to thylakoid membranes and stromal proteins (Del Duca et al. 1994; Dondini et al. 2003; Della Mea et al. 2004a). The first chloroplast TGase has recently been cloned and characterised (Villalobos et al. 2004; Carvajal-Vallejos et al. 2007).
There is limited data available concerning the potential role of PAs during the hormone-modulated differentiation processes and only a small amount of evidence suggests that changes in PA levels and their biosynthesis are linked to hormone action (Legocka and Żarnowska 1999, 2000, 2002).
Our previous results indicated that changes in the level of free PAs and their metabolism during cytokinin-stimulated greening are not essential for plant hormone action (Sobieszczuk-Nowicka et al. 2007b), whereas the bound form of PAs appeared to be important for this developmental process (Sobieszczuk-Nowicka et al. 2007a). These studies showed that PAs are an important component of etioplast membrane structure and the mature chloroplast complex. The same data suggested that the mechanism by which PAs could affect this membrane assembly might be, at least in part, mediated by their covalent linkage to these membranes (Sobieszczuk-Nowicka et al. 2007a).
With this in mind, we selected the cucumber cotyledon greening test as a model (Parthier 1979) for the present work, with the aim of improving the understanding of cytokinin-stimulated chloroplast development through covalent binding of PAs to chloroplast structures via TGases. We did this in order to determine whether the relationship between this enzyme and the chloroplast differentiation process may represent an important component of the process itself and/or of the mechanism of kinetin action. We examined the localisation of TGase by immunogold staining and electron microscopy, and determined its level and activity. The resulting data are complemented by the identification of plastid proteins acting as substrates for TGases.
Materials and methods
Plant material
Cucumber seeds (Cucumis sativus L. cv. Racibór) were germinated on several layers of moistened tissue paper in the dark at 25°C for 6 days. The etiolated cotyledons were then excised under dim green light and subjected to pre-treatment: placed onto Petri dishes on filter paper wetted with water (control) or 100 μM aqueous kinetin (Sigma, St. Louis, MO, USA) solution at 25°C in the dark for 24 h. After the pre-treatment, the Petri dishes with cotyledons were exposed to light (146 μmol m–2 s–1) for 6 or 24 h at 25°C, and then used for analyses.
Immunocytochemistry
Cotyledons were fixed in 4% paraformaldehyde and 0.5% glutaraldehyde (v/v) in phosphate buffered saline (PBS) for 3 h at room temperature. Following this, the material was washed in PBS and dehydrated in a graded aqueous ethanol series.
After dehydration, the material was progressively infiltrated with LR Gold acrylic resin (Sigma, St. Louis, MO, USA) and then embedded in gelatine capsules. Polymerisation using 1% benzyl peroxide as the accelerator was allowed to proceed for 2 weeks at room temperature.
Ultrathin sections were cut using a diamond knife on an Ultracut S ultramicrotome (Leica-Reichert, Bensheim, Germany) and collected on carbon-coated nickel grids (200 meshes).
For immunolabelling, the sections were first treated with blocking solution (2% bovine serum albumin in PBS) for 15 min and were then incubated for 2 h at room temperature with TGase antibody (RB-060, Ab-4, NeoMarkers™, Fremont, CA, USA) diluted 1:100. After washing in blocking solution, sections were incubated for 1 h at room temperature in a solution of 10 nm diameter colloidal gold-affinipure (Sigma, St. Louis, MO, USA) diluted at a ratio of 1:20. The sections were then washed with saline buffer and distilled water and subsequently stained with 3% uranyl acetate solution. As a control, some samples were only treated with blocking solution instead of the antibody, following the same protocol described above.
Microscopy
Immunogold observations were carried out using a JEM-1200 EXII JEOL transmission electron microscope (Tokyo, Japan).
To quantify the observed differences, the number of gold particles per squared micrometer of thylakoids in samples from the different stages of chloroplast development was calculated.
Isolation of plastids
Cucumber cotyledons were homogenised with three volumes of 0.05 M Tris-HCl, pH 7.8, containing 0.4 M saccharose, 0.01 M KCl, 0.01 M MgCl2 and 6 mM mercaptoethanol. The extract was filtered through two layers of 20-μM nylon mesh and centrifuged at 1,000×g at 4°C for 5 min. Plastids were recovered by means of a pointed brush from a soft layer above the pellet, which mainly contained starch and nuclei, and were resuspended in the buffer. This was then diluted up to one-eighth of the initial volume with the same extraction buffer and again centrifuged. This protocol allows plastids to be isolated with a high degree of intactness.
Colorimetric assay of TGase activity
To determine the putative TGase activity, we carried out colorimetric enzyme assays. The biotin cadaverine incorporation assay was carried out as described by Slaughter et al. (1992) following the modifications of Lilley et al. (1998). The presence of endogenous TGase activity was measured as covalent binding of biotinylated cadaverine to endo-glutamyl residues of the N’, N’-dimethylcasein (DMC; 10 mg/ml) used to precoat the assay microplate. The 5 mM Ca2+ was replaced by 1 mM EDTA in the negative control.
Identification of plastid proteins as TGase substrates
Measurement of TGase enzyme activity was preceded by overnight dialysis (10 mM Tris, pH 8.5, and 1 mM 2-mercaptoethanol). Isolated plastids were assayed for the presence of both the TGase and its endogenous substrates in the presence of 56 kBq [1,4–14C]Spd (specific activity 4.4 GBq mmol–1, Amersham, Piscataway, NY, USA), which provides primary amino groups used as acyl acceptors of glutamyl residues of plastid proteins. The assay was carried out as described by Dondini et al. (2003). After 1 h, the reaction was stopped and samples were centrifuged at 15,000×g for 20 min. The pellet was re-suspended in sample buffer (50 mM Tris, pH 6.8, 100 mM DTT, 1% bromophenol blue, 10% glycerol and 2% sodium dodecyl sulfate (SDS)). Proteins (20 μg per sample) were separated using 15% polyacrylamide slab gel electrophoresis and then transferred from the gel onto a polyvinylidene fluoride (PVDF) membrane (Immobilon P transfer membrane™, Millipore, Bilerica, MA, USA) using a semi-dry electrophoretic blotting system (Sigma, St. Louis, MO, USA). The SDS-polyacrylamide gel electrophoresis (PAGE) gel was made according to Laemmli (1970).
Autoradiography
PVDF membranes were exposed at room temperature to an imaging plate sensitive to [14C] (3543 MS IP) for 1 week, which was then scanned by an FLA-5100 laser system (Radioisotope image analyser, Fuji Photo Film. Co., Tokyo, Japan).
Western-blot analysis
An SDS-PAGE gel was made according to Laemmli (1970). Proteins (5 μg) from isolated plastids were dissolved in sample buffer and separated using 12% polyacrylamide slab gel electrophoresis. Prestained Protein Molecular Weight Marker and Page Ruler™ Protein Ladder (Fermentas, Ontario, Canada) were used as the protein standards according to the manufacturer’s instructions.
Proteins were transferred from polyacrylamide gels onto PVDF membranes using a semi-dry blotting system and immunoblotting was performed with the use of the ExtrAvidin Peroxidase Staining Kit (Sigma, St. Louis, MO, USA) according to the manufacturer’s instructions. The optimal dilution of the anti-TGase sera (RB-060, Ab-4, NeoMarkers™, Fremont, CA, USA), as estimated by dot-blot, was 1:5000.
Densitometry
To perform a quantification of immunostained TGase bands, PVDF membranes were scanned using an UMAX Alpha Vista II scanner and the staining intensity of the bands was measured and calculated with the Multi Gauge programme (Fuji Photo Film Co., Ltd., Tokyo, Japan). As the measurement system was not calibrated, the parameters are expressed in arbitrary units.
Protein assay
The protein content of the plastid fraction was determined using the bicinchoninic acid method (Brown et al. 1989). Bovine serum albumin was used as the standard protein.
Statistical analyses
Each data point represents a mean of three replicates obtained from three independent experiments. The statistical analysis was performed using Standard Method STATISTICA (Stat Soft Inc., Tulsa, OK, USA).
Results
Immunodetection of plastid TGases
We detected the plastid TGase protein in each sample group—0 h control, 0 h kinetin, 6 h control, 6 h kinetin, 24 h control and 24 h kinetin. In etioplasts (0 h), antibodies recognised bands at 77 and 30 kDa, and an extra 50-kDa band was detected in chloroplasts (6 and 24 h; Fig. 1). Generally, during the early stage of chloroplast development (0–6 h), all band intensities increased. In etioplasts, kinetin stimulated the accumulation of the 30-kDa band by about 50%, and at 6 h, the intensity of each band was greater in kinetin-treated samples compared to those of the control. After 24 h of light exposure, this difference in band intensity between the control and kinetin samples was lost and the intensity itself decreased.
Immunodetection of TGases by Western blot analysis. Proteins were isolated from plastids of cucumber cotyledons. Six-day-old etiolated cucumber cotyledons were excised from the seedling, pre-incubated in the dark for 24 h in water as a control (H2O) or 100 μM kinetin (KIN, 0 h) and exposed to light for 6 or 24 h. Isolated proteins were electrophoresed by 12% (w/v) SDS-PAGE. Densitometry analysis of the immunostained band was carried out, and the results from the 30 kDa band are plotted on the histogram. The water control at 0 h was taken as 100%. Data points represent means ±SE of three replicates
Immunocytochemical localisation of plastid TGases
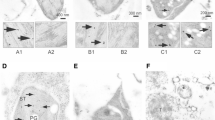
We could observe specific suborganellar enzyme localisation during etioplast-to-chloroplast transformation, when an alteration of etioplast ultrastructure took place (Fig. 2).
Immunocytochemical localisation of plastid TGases. Cotyledons were treated as described for Fig. 1. A and A1. Control (0 h) etioplast after pre-incubation in the dark. Gold particles (arrowheads) indicate TGases localised in the PLB and PTs. B Hormone-treated (KIN 0 h) etioplast after pre-incubation in the dark. Gold particles (arrowheads) indicate TGases localised in the PLB and PTs. C and C1. Control etiochloroplast after 6-h light exposure. Gold particles (arrowheads) indicate TGases localised in Ts. D and D1. Hormone-treated (KIN) chloroplast after 6-h light exposure. No PLB is visible. Gold particles (arrowheads) indicate TGases localised in Ts. E. Control chloroplast after 24-h light exposure. Gold particles (arrowheads) indicate TGases localised in Ts. F. Hormone-treated (KIN) chloroplast after 24-h light exposure. Gold particles (arrowheads) indicate TGases localised in Ts. Bars = 100 nm Legend: TGase; transglutaminase, PLB; prolamellar body, PT; pro-thylakoid, T; thylakoid
This included the conversion of the distinctive prolamellar body into stromal and stacked thylakoid structures, which are characteristic of mature chloroplasts. Exposure of etiolated cotyledons (0 h) to light caused an almost complete thylakoid differentiation by 24 h (Fig. 2). The process of etioplast-to-chloroplast transformation was markedly accelerated by kinetin (Fig. 2B, D, F). Gold labelling, indicating the presence of TGases, was present in all types of plastid—in etioplasts (Fig. 2A, B), etiochloroplasts (Fig. 2C) and mature chloroplasts (Fig. 2D–F). The location of cucumber cotyledon TGase by electron microscopy indicated that the plastid TGase was peculiarly localised (Fig. 2A1, C1, D1). In etioplasts, TGases were detected mainly in pro-thylakoids (Fig. 2A); however, there was some gold labelling in the prolamellar body (Fig. 2A, A1) and stroma. In etiochloroplasts and chloroplast, mainly thylakoid grana were labelled, but there were also gold particles present in the stroma (Fig. 2C–F). The number of gold particles in plastids increased at the early stages of etioplast-to-chloroplast transformation (Fig. 2A, C) and was further increased by the hormone (Fig. 2B, D). Conversely, as the transformation advanced in later stages, the number of gold particles decreased (Fig. 2E, F).
TGase activity
We found that the TGase activity of isolated plastids varied at different times of light exposure (Fig. 3). If the cotyledons were pre-incubated with water, we observed an increase in activity from 0 to 6 h and an insignificant decrease between 6 and 24 h of light exposure. However, following pre-incubation with kinetin, the endogenous TGase activity showed significant differences, mainly at 6 h. The enzyme activity of the kinetin-treated samples increased very strongly at 6 h, to about 128% and 60% of that of the kinetin sample at 0 h and the control at 6 h, respectively (Fig. 3). At 24 h, the high TGase activity decreased.
TGase activity in cucumber cotyledon plastids, treated as described for Fig. 1. The TGase activity was evaluated as biotin cadaverine binding to DMC. Data represent means ± SE of three replicates. Differences between control and kinetin samples are most significant at 6 h at P < 0.5
Plastid proteins as TGase substrates
Spd is the PA most efficiently conjugated to the thylakoid membrane during the etioplast-to-chloroplast transformation (Sobieszczuk-Nowicka et al. 2007a), whereas the conjugation of Put and Spm via TGase is very low. Therefore, we used [1,4–14C]Spd to identify plastid proteins acting as TGase substrates throughout the chloroplast development process. Autoradiography of plastid proteins separated by SDS gel electrophoresis revealed four bands at 58, 29, 26 and 12 kDa. The 58-kDa band was present in the etioplast fraction in both the control and kinetin-treated cotyledons at 0 h and only in the control at 6 h (Fig. 4). The band was no longer labelled by [14C]Spd at 6 h in kinetin-treated cotyledons or at 24 h in both the control and kinetin samples. Conversely, the 29-, 26- and 12-kDa bands did not appear in plastids before the cotyledons had been exposed to light for 6 h. The intensity of the these bands increased with the time of light exposure. Kinetin accelerated the covalent binding of [14C]Spd to the 29-, 26- and 12-kDa protein TGase substrates (Fig. 4).
Autoradiography of SDS gel electrophoresis of plastid proteins isolated from cucumber cotyledons, treated as described for Fig. 1 and incubated in the presence of [14C]Spd
Discussion
TGases have been found in green plant cells, in isolated chloroplasts and in unicellular green algae (Signorini et al. 1991; Falcone et al. 1993; Del Duca et al. 1994, 2000; Dondini et al. 2000, 2003; Villalobos et al. 2001, 2004; Carvajal-Vallejos et al. 2007). Most of this data was obtained from fully differentiated leaves, whereas scarce information is available on the presence of TGases and their activity during the chloroplast photodevelopment process (Del Duca et al. 1993; Villalobos et al. 2001, 2004). Preliminary reports have suggested that the enzyme is mainly localised in stacked thylakoids in mature chloroplasts and the amount of bound PAs increases with exposure to light. However, the nature of the PA binding has not been clarified.
In the present work, the presence of TGase in the kinetin-stimulated differentiating chloroplasts is demonstrated by its level (Fig. 1), in situ localisation (Fig. 2) and conjugating activity (Fig. 3), and by the identification of plastid proteins acting as substrates (Fig. 4).
Immunodetection of TGases in both etioplasts and chloroplasts of cucumber cotyledons revealed bands at 30, 50 and 77 kDa (Fig. 1). Evidence from other studies has shown that more than one protein with TGase activity could be present in the same tissue, cell or different compartments of the same cell (Falcone et al. 1993; Del Duca et al. 1994; Bernet et al. 1999; Villalobos et al. 2001), or even within the same organelle, as has been shown for chloroplasts by Dondini et al. (2003). In the thylakoid-membrane fraction of cucumber cotyledons, the most common form of TGase is the 77-kDa protein (Sobieszczuk-Nowicka et al. 2007a). This may indicate that it is a thylakoidal isoform and that the 30-kDa fraction, which is abundant in whole plastids (Fig. 1), is a stromal one. The expression of the 50-kDa band seems to be light stimulated (Fig. 1). This agrees with the results described by Dondini et al. (2003) concerning the effect of light on TGase activity.
The data from Figs. 1 and 3 shows that there is a positive correlation between TGase level and enzymatic activity. It is noteworthy that it also demonstrates kinetin regulation of TGase protein expression. The colorimetric assay, performed in the presence of DMC (Fig. 3), showed a TGase activity similar to the catalytic activity of soluble pea root TGase (Lilley et al. 1998). As the exogenous substrate is present in a constant amount throughout all the different light exposure times, the differences in the TGase activity may probably be ascribed to an effect of the enzyme rather than to the substrate.
The suborganellar location of cucumber cotyledon TGase by electron microscopy indicates that the TGase is peculiarly localised (Fig. 2). The variation in the presence of the TGase according to the stage of chloroplast differentiation, and its detection mainly in the appressed grana thylakoid, suggest that the enzyme might be involved in the formation of PSII. It has been previously demonstrated that PAs are bound to the cucumber cotyledon PSIIα fraction (Sobieszczuk-Nowicka et al. 2007a) and to maize LHCII (Della Mea et al. 2004b). The number of gold particles in plastids increased during early stages of the transformation process and this was accelerated by the hormone. However, as the transformation advanced, the number of gold particles began to decrease (Fig. 2). This may suggest that, at the final stage of chloroplast development, the process of PA binding to plastid proteins via TGase is almost over. The low expression of the enzyme observed in mature leaves and in plants receiving continuous light is in agreement with this hypothesis. The Northern blot analysis of maize TGase published by Villalobos et al. (2004) shows that TGase mRNA is expressed shortly after light exposure, when thylakoids and grana start to form. At later stages of chloroplast photodevelopment (between 24 and 48 h), a gradual decrease in RNA accumulation was observed (Villalobos et al. 2004).
One of the best known roles of animal TGase is the production of protein cross-linked nets favouring the stabilisation of new protein structures (Griffin et al. 2002). The data corroborates that [14C]Spd post-translational modification takes place and may favour the stabilisation, aggregation or protein conformational changes of certain plastid proteins of 58, 29, 26 and 12 kDa (Fig. 4). Mono- and bis-glutamyl-Spd derivatives detected in reorganising cucumber cotyledon thylakoid membranes (Sobieszczuk-Nowicka et al. 2007a) may confer an additional positive charge to the modified protein and increase the stabilisation of chloroplast structures by forming ‘‘bridges’’ between different glutamyl residues. This additional charge would cause a conformational rearrangement of the substrate protein that could contribute to the final assembly of these substrates or favour ionic interactions with other molecules. The apoproteins of the chlorophyll a/b antenna complex (LHCII, CP 24, CP 26, CP 29) and the large subunit of Rubisco have been shown to be the substrates of plastid TGases in mature leaves (Margosiak et al. 1990; Del Duca et al. 1994; Dondini et al. 2003; Della Mea at al. 2004b). The TGase substrates of 29 and 26 kDa identified in the present work might correspond to the chlorophyll a/b antenna complex II proteins as well as to the LHCPII. Labelling of the 26, 24 and 12-kDa bands was prominent following light treatment. This suggests that the 12-kDa band might correspond to the small subunit of Rubisco, which is around 12–15 kDa, or to ferredoxin, a small molecule of around 11 kDa, also identified in the stroma fraction by Dondini et al. (2003). The 58-kDa band remains unidentified. This fraction was associated with the etioplast and its level faded when the etioplast-to-chloroplast transformation was in progress.
Taken together, the modification of etioplast or chloroplast proteins could affect the Spd linkage, or conversely, the latter could affect the protein conformation, with a possible effect on photosynthesis and/or its light harvesting efficiency. A significant reduction in TGase activity on purified grana maize proteins treated with an inhibitor of the PSII activity has been observed (Villalobos et al. 2004), which suggests that TGases prefer active proteins of the PSII antenna complex as their substrates.
Concluding remarks
In summary, this manuscript focuses on the involvement of TGase in kinetin-stimulated chloroplast photodevelopment.
TGases bind polyamines to their protein substrates, as a result of a transamidation reaction, and take part in the formation of chloroplast structures, mainly the thylakoid system.
The results suggest that TGases, which are known to be stimulated by light and Ca2+, can also be stimulated by fitohormones such as cytokinin. These findings about the effect of kinetin on the conjugation of PAs via TGases provide a new contribution to the understanding of the mechanism of kinetin action in etioplast-to chloroplast transformation.
Abbreviations
- LHCPII:
-
light-harvesting chlorophyll a/b complex protein
- PA:
-
polyamine
- PSII:
-
photosystem II
- Put:
-
putrescine
- Spd:
-
spermidine
- Spm:
-
spermine
- TGases:
-
transglutaminases
- DMC, N’:
-
N’-dimethylcasein
References
Andreadakis A, Kotzabasis K (1996) Changes in the biosynthesis and catabolism of polyamines in isolated plastids during chloroplast photodevelopment. J Photochem Photobiol B: Biol 33:163–170
Beigbeder A, Vavadakis M, Navakoudis M, Kotzabasis K (1995) Influence of polyamine inhibitors on light-independent and light-dependent chlorophyll biosynthesis and on the photo-synthetic rate. J Photochem Photobiol B: Biol 28:235–242
Bernet E, Claparols I, Dondini L, Santos MA, Serafini-Fracassini D, Torne JM (1999) Changes in polyamine content, arginine and ornithine decarboxylases and transglutaminase activities during light/dark phases of initial differentiation in maize calluses and their chloroplasts. Plant Physiol Biochem 37:1–11
Brown R, Jarvis K, Hyland K (1989) Protein measurement using bicinchoninic acid: elimination of interfering substances. Anal Biochem 180:136–139
Carvajal-Vallejos PK, Campos A, Fuentes-Prior P, Villalobos E, Almeida AM, Barbera E, Torne JM, Santos M (2007) Purification and in vitro refolding of maize chloroplast transglutaminase over-expressed in Escherichia coli. Biotechnol Lett 29:1255–1262
Del Duca S, Serafini-Fracassini D (1993) Bound polyamines in plants. Curr Top Plant Physiol 1:83–102
Del Duca S, Serafini-Fracassini D (2005) Transglutaminases of higher, lower plants and fungi. In: Mehta K, Eckert R (eds) Transglutaminases. Prog Exp Tum Res. Karger, Basel, pp 223–247
Del Duca S, Favali A, Serafini-Fracassini D, Pedrazzini R (1993) Transglutaminase-like activity during greening and growth of Helianthus tuberosus explants. Protoplasma 174:1–9
Del Duca S, Tidu V, Bassi R, Esposito C, Serafini-Fracassini D (1994) Identification of chlorophyll-a/b proteins as substrates of transglutaminase activity in isolated chloroplasts of Helianthus tuberosus L. Planta 193:283–289
Del Duca S, Dondini L, Della Mea M, Munoz de Rueda P, Serafini-Fracassini D (2000) Factors affecting transglutaminase activity catalysing polyamine conjugation to endogenous substrates in entire chloroplasts. Plant Physiol Biochem 38(6):429–439
Della Mea D, Caparros-Ruiz D, Claparolos I, Serafini-Fracassini D, Rigau J (2004a) AtPng1p. The first plant transglutaminase. Plant Physiol 135:2046–2054
Della Mea M, Di Sandro A, Dondini L, Del Duca S, Vantini F, Bergamini C, Bassi R, Serafini-Fracassini D (2004b) A Zea mays 39 kDa thylakoid transglutaminase catalyses Light Harvesting Complex II by polyamines in a light-dependent way. Planta 219:754–764
Dondini L, Bonazzi S, Serafini-Fracassini D (2000) Recovery of growth capacity by polyamines and of chloroplast transglutaminase activity in a polyamine-deficient variant strain of Dunaliella salina. J Plant Physiol 157:473–480
Dondini L, Del Duca S, Dall’Agata L, Bassi R, Gastaldelli M, Della Mea M, Di Sandro A, Claparos I, Serafini-Fracassini D (2003) Suborganellar localisation and effect of light on Helianthus tuberosus chloroplast transglutaminases and their substrates. Planta 217:84–95
Falcone P, Serafini-Fracassini D, Del Duca S (1993) Comparative studies of transglutaminase-like activity and substrates in different organs of Helianthus tuberosus. J Plant Physiol 142:265–273
Griffin M, Casadio R, Bergamini CM (2002) Transglutaminase: nature’s biological glues. Biochem J 368:377–396
Kotzabasis K (1995) A role for chloroplast-associated polyamines? Bot Acta 109:5–9
Kotzabasis K, Fotinou C, Roubelakis-Angelakis KA, Ghanotakis D (1993) Polyamines in the photosynthetic apparatus. Photosynth Res 38:83–88
Kotzabasis K, Strasser B, Navakoudis E, Senger H, Dörnemann D (1999) Charakterization of photoreceptor(s) responsible for the regulation of the intracellular polyamine level and the putative participation of heterotrimetric G-proteins in the signal transduction chain. J Photochem Photobiol B: Biol 50:45–52
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Legocka J, Żarnowska A (1999) Role of polyamines in the cytokinin-dependent physiological processes. I. Effect of benzyladenine on polyamine during chloroplast differentiation in the tissue culture of Dianthus caryophyllus. Acta Physiol Plant 21:349–354
Legocka J, Żarnowska A (2000) Role of polyamines in the cytokinin-dependent physiological processes. II Modulation of polyamine levels during cytokinin-stimulated expansion of cucumber cotyledons. Acta Physiol Plant 22:395–401
Legocka J, Żarnowska A (2002) Role of polyamines in the cytokinin-dependent physiological processes. III Changes in polyamine levels during cytokinin-induced formation of gametophore buds in Ceratodon purpureus. Acta Physiol Plant 24:303–309
Lilley G, Skill J, Griffin M, Bonner P (1998) Detection of Ca2+-dependent transglutaminase activity in root and leaf tissue of monocotyledonous and dicotyledonous plants. Plant Physiol 117:1115–1123
Lorand L, Graham RM (2003) Transglutaminases: crosslinking enzymes with pleiotropic functions. Nature Rev Mol Cell Biol 4:140–156
Margosiak SA, Dharma A, Bruce-Caver MR, Gonzales AP, Louie D, Kuehn GD (1990) Identification of the large subunit of ribulose 1,5-bisphosphate carboxylase/oxygenase as a substrate for transglutaminase in Medicago sativa L. (alfalfa). Plant Physiol 92:88–96
Parthier B (1979) The role of phytohormones (cytokinins) in chloroplast development. Biochem Physiol Pflanz 174:173–214
Signorini M, Beninati S, Bergamini C (1991) Identification of transglutaminase activity in the leaves of silver beet (Beta vulgaris L.). J Plant Physiol 137:547–552
Slaughter TE, Komandor AE, Lai TS, Greenberg CS (1992) A microtiter plate transglutaminase assay utilising 5-(biotinamido)pentylamine as substrate. Annal Biochem 205:167–171
Sobieszczuk-Nowicka E, Di Sandro A, Del Duca S, Serafini-Fracassini D, Legocka J (2007a) Plastid-membrane-associated polyamines and thylakoid transglutaminases during etioplast-to-chloroplast transformation stimulated by kinetin. Physiol Plant 130:590–600
Sobieszczuk-Nowicka E, Rorat T, Legocka J (2007b) Polyamine metabolism and S-adenosylmethionine decarboxylase gene expression during the cytokin-stimulated greening process. Acta Physiol Plant 29:595–502
Villalobos JM, Torne J, Rigau I, Olles I, Claparos M, Santos V (2001) Immunogold localisation of transglutaminase related to grana development in different maize cell types. Protoplasma 216:155–163
Villalobos E, Santos M, Talavera D, Rodrıguez-Falcon M, Torne JM (2004) Molecular cloning and characterization of a maize transglutaminase complementary DNA. Gene 336:93–104
Acknowledgements
The author is very indebted to Prof. Donatella Serafini-Fracassini and her co-workers from Bologna University for methodological support, to Agnieszka Kazikowska for the processing of the figures and to Heidi Nicholl of City University, London for the linguistic correction of the manuscript.
This work was partly supported by the Polish State Committee for Scientific Research grant number 2 PO4C 085 26.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sobieszczuk-Nowicka, E., Krzesłowska, M. & Legocka, J. Transglutaminases and their substrates in kinetin-stimulated etioplast-to-chloroplast transformation in cucumber cotyledons. Protoplasma 233, 187–194 (2008). https://doi.org/10.1007/s00709-008-0002-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-008-0002-y