Abstract
Reliable diagnostic tests that are able to distinguish infected from vaccinated animals are a critical component of regional control programs for foot-and-mouth disease (FMD) in the affected countries. Non-structural protein (NSP) serology based on the 3ABC protein has been widely used for this purpose, and several kits are commercially available worldwide. This report presents the development of a 3ABC-antigen-based indirect ELISA, employing a peroxidase-conjugated protein G secondary antibody that can detect antibodies from multiple species. Recombinant 3ABC protein was expressed in insect cells and purified using affinity column chromatography. Using this protein, an indirect ELISA was developed and validated for the detection of NSP antibodies in serum samples collected from animals with different status of FMD. Diagnostic sensitivity and specificity were found to be 95.8 (95 % CI: 92.8–97.8) and 97.45 % (95 % CI: 94.8–99.0), respectively. The in-house ELISA compared well with the commercially available prioCHECK FMDV NS-FMD kit, with a high agreement between the tests, as determined by the kappa coefficient, which was 0.87. The in-house ELISA showed higher sensitivity for detecting vaccinated and subsequently infected animals, when compared to the reference test. Both of the tests were able to detect NSP antibodies as early as 7–8 days after experimental infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Foot-and-mouth disease (FMD) is endemic in many parts of the world, particularly in Asia and Africa, where it causes substantial economic loss due to trade barriers. In developing countries, where the adverse effects of FMD are often underestimated, the disease undermines food security and economic development, at the level of both village smallholders and the more-organised production chains [1]. FMD is still widespread throughout the world, and by the end of May 2014, more than 100 countries were not FMD-free, and they remain a continuous threat to disease-free countries [1]. This is exemplified by the recent incursion of disease in previously disease-free countries such as Japan and Korea.
Several tools are of critical importance to the global FMD control strategy. These include reliable diagnostics and appropriate vaccines to control FMD in the affected countries, as well as effective surveillance and competent diagnostic services. For the control of FMD, regular vaccination is the most-sought approach in several affected regions. Use of inactivated whole-virus vaccines has been successful in the control and eradication of FMD in several parts of the world, including Europe, South America and South East Asia [2]. Non-structural protein serology combined with structural protein assays are routinely used to monitor virus activity in endemic countries where vaccination is done. Countries that claim to be free of FMD are required to provide evidence of the absence of virus circulation based on NSP serosurveillance (Article 8.7.9 of OIE Terrestrial Animal Health Code). OIE has recognized NCPanaftosa indirect ELISA as the index screening test for DIVA (differentiation of infected from vaccinated animals) [3], and this test, combined with the enzyme-linked immunoelectrotransfer blot (EITB) assay, has been widely applied in South America as a confirmatory test. The use of NSP-based assays that are inexpensively produced and validated are highly useful for close monitoring of the disease situation. This paper describes the development and validation of an in-house indirect ELISA using recombinant 3ABC expressed in insect cells as an antigen and a peroxidase conjugated protein G as the secondary antibody for detection of NSP antibodies in sera from different species.
Materials and methods
Cloning of the FMDV 3ABC region into a transfer vector
The open reading frame (ORF) corresponding to the 3ABC region of foot-and-mouth disease virus (FMDV) type A (IND/40/2000 strain) was amplified by PCR from cDNA synthesized using viral RNA. Briefly, the viral RNA was extracted from FMDV using an RNeasy Mini Kit (QIAGEN), and reverse transcription was performed in a 20-µl reaction volume using an oligo (dT) primer with the ThermoScript™ RT-PCR System (Invitrogen). The cDNA obtained was used as template for PCR amplification of the 3ABC region using Pfu DNA polymerase. Primers targeting the full-length 3ABC region were designed based on the genomic sequence of the virus (GenBank accession no. HM854025). The primers used were HAF2 (5′-TGGGATCCATGCATCACCATCACCATCACTCAATTCCTTCCCAAAAG-3′) and BamHI_3C639 (5′-TGGGATCCTACTCGTGGTGTGGTTCGG-3′). Two additional primers, C163G-F (5′-AGGCTGGCTACGGTGGAGGA-3) and C163G-R (5′-CTCCTCCACCGTAGCCAGCC-3), were used to introduce a mutation (C163G) in the 3C region [4] by overlap PCR.
The PCR product was ligated into the BamHI restriction enzyme recognition site of the PACYM1 transfer vector and sequenced. Recombinant baculovirus was generated by allowing recombination between the linearized baculovirus DNA and recombinant pACYM1 containing the 3ABC sequence. The recombinant virus (Ac-FMDV-3ABC) obtained was plaque purified and amplified to high titres.
Expression and purification of recombinant 3ABC antigen
Two insect cell lines, the Spodoptera frugiperda-derived cell line SF21AE and the Trichoplusia ni-derived cell line BTI TN5B1-4 (Tn5) were used. The cell lines, which were adapted to grow in SF900II™ SFM medium in the presence (SF-21) or absence (Tn5) of 10 % of fetal bovine serum, were used for virus growth and expression studies, respectively.
For expression of the 3ABC protein, Tn5 cells were infected at a multiplicity of infection (MOI) of 5 with Ac-FMDV-3ABC, and cells were collected 3–4 days postinfection by centrifugation at low speed (192 × g) for 10 minutes. Cells were suspended in Tris buffer (10 mM, pH 8.0) containing 0.1 % Triton X-100 and subjected to sonication. The mixture was centrifuged (8000 × g for 10 minutes) to obtain insoluble precipitate containing 3ABC protein. The precipitate was suspended in 10 mM Tris buffer (pH 8.0) containing 8 M urea. This solubilized fraction was subsequently used for purification of protein using a Probond Purification Kit (Invitrogen) under denaturing conditions with minor changes as described previously [5].
ELISA test procedure
Appropriate dilutions of the recombinant 3ABC antigen, antibody and conjugates were determined by checkerboard titration. Serum samples collected from bull calves experimentally infected with FMDV serotype O (IND/R2/75 strain) and adult bovine serum (New Zealand origin) were used as positive and negative control, respectively. An immunoassay plate (Maxisorp, Nunc) was coated with recombinant 3ABC antigen in coating buffer (carbonate-bicarbonate buffer, 0.05 M, pH 9.6) and incubated at 4 °C overnight. On discarding the plate contents, the unbound sites in the wells were saturated with bovine albumin fraction V (BSA-V; 1 % w/v, in 50 mM Tris-buffered saline, pH 7.4) for 30 min at 37 °C. After three washings in wash buffer (50 mM Tris-buffered saline containing 0.1 % Tween-20), the serum samples diluted (1:21) in the diluent buffer (1 % BSA-V made in wash buffer) were dispensed at 50 μl/well in duplicate. After incubation for 1 h at 37 °C, the wells were washed three times. Recombinant protein G peroxidase conjugate (Pierce, Thermo Scientific) was dispensed at 50 μl/well at optimal dilution, and the plate was incubated for 1 h at 37 °C. After four washings, the substrate solution containing ortho-phenylene diamine (OPD) and H2O2 was added at 50 μl/well. The plate was incubated in the dark at 37 °C for 15 min. Colour development was stopped by addition of 50 μl of 1 M H2SO4 to each well. Absorbance was measured at a wavelength of 492 nm and a reference wavelength of 620 nm in an ELISA reader.
The OD values of the positive control (ODpos) and the samples (ODsample) were corrected by subtracting the OD value of the negative control (ODneg). The sample value was calculated as percent positivity using the following [6]: Percent positivity (PP) value = (ODsample − ODneg)*100/(ODpos − ODneg). Samples with a PP value of ≥25 relative to the normalized OD value of the positive sample were considered ‘positive’; and those below 25, ‘negative’.
Determining the cutoff value for the assay
Serum samples maintained in a repository at this laboratory were used in this study. To establish the cutoff value for the assay, serum samples (n = 356) from an unvaccinated, isolated dairy herd with titres of <8 as determined by the liquid-phase blocking ELISA (LPBE) or the virus neutralization test against three current vaccine strains of type O (IND/R2/1975), type A (IND/40/2000) and type Asia 1 (IND/63/1972) were used as negative samples. Based on the OD value obtained for these samples, a cutoff value was determined.
Comparison with prioCHECK FMDV NS test and statistical analysis
The PrioCHECK FMDV NS test (Prionics Lelystad, The Netherlands) was performed as per manufacturer’s instructions. After colour development using TMB substrate, plate readings were taken at 450 nm wavelength, and percent inhibition (PI) values were calculated. Samples showing a PI value of ≥50 % were considered ‘positive’; and those below 50 %, ‘negative’.
Analytical sensitivity
The analytical sensitivity of the in-house 3ABC ELISA was determined using a panel of four positive sera obtained from bull calves experimentally infected with FMDV type A (IND/40/2000 strain), along with a known negative serum (New Zealand origin). The test was carried out in parallel with the prioCHECK FMDV NS test on serial two-fold dilutions of the sera for comparing the endpoint detection limits.
Diagnostic sensitivity and specificity
The indirect ELISA was evaluated with diagnostic performance parameters using sera from animals with a known history of FMDV infection. Samples (n = 274) collected from cattle and buffalo (n = 180), sheep (n = 51) and goats (n = 43) with no previous history of exposure to FMDV or vaccination against FMD were chosen as FMD-negative samples. Sera (n = 286) collected from infected animals, including samples from experimentally infected bull calves, and sheep and goats that were infected during a natural outbreak of FMD, were classified as ‘infected’ samples. Of these, 56 were from unvaccinated bull calves that were experimentally infected with cattle-adapted challenge virus (type O, IND/R2/75 strain) as part of a regular vaccine efficacy trial. Infection was confirmed by the appearance of clear clinical signs, including primary and secondary lesions, and recovery of viral antigen/nucleic acid. Serum samples from these animals were collected from 2 to 4 weeks after experimental infection, except for those from five calves, which were collected at regular intervals for up to 35 weeks postinfection. ‘Infected’ sera also included 64 samples from bull calves that were vaccinated with trivalent oil-adjuvanted FMD vaccine and subsequently challenged with FMDV (type O, R2/75 strain) 28 days post-vaccination. Infection was confirmed by detection of viral nucleic acid by RT-PCR [3]. These animals were bled 3 weeks after challenge infection. Sera (n = 162 samples) collected from sheep (n = 125) and goats (n = 37) from a mixed herd that recovered from an outbreak in 2013, confirmed to be FMDV serotype O based on antigen detection ELISA, were also included as ‘infected’ samples. All experiments involving the use of animals followed the procedures prescribed and approved by the institutional animal ethics committee.
For statistical comparison, agreement between the two tests was measured by calculating the kappa coefficient for the results obtained in both tests across all categories of sera. Diagnostic sensitivity (Ds) and specificity (Dsp) of the assay at different cutoff values were calculated using Medcalc software (11.5.0). For this, percent positivity values of sera along with disease status (0 for no-disease and 1 for disease) were imported into Medcalc software to obtain the Ds and Dsp values.
Detection of early NSP antibodies and their longevity in convalescent animals
Levels of early antibodies following experimental infection with FMDV serotype O (IND/R2/1975) were measured in sheep, goat and cattle by both tests. Representative animals infected via intradermo-lingual inoculation with 10,000 bovine infective dose50 of cattle-adapted virus were sampled at different time points to assess the early detectable NSP antibody response.
To determine the duration of the NSP antibody response, sequential serum samples collected from five calves experimentally infected with the FMDV type O (IND/R2/1975 strain) were used. Sera collected from these animals at various monthly intervals were evaluated using the in-house ELISA and prioCHECK test.
Prevalence of NSP antibodies in the vaccinated population under field conditions
Random serum samples collected from ~5 % of the cattle and buffalo from organized herds in and around Bangalore district were screened using the in-house ELISA in order to determine the prevalence of 3ABC antibodies. These samples collected from animals vaccinated against FMD were available in our laboratory. A total of 3851 sera were screened.
Results
Expression and purification of the 3ABC protein
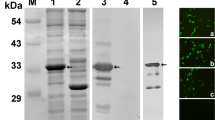
Recombinant virus was amplified to high titres (≥1 × 108 pfu/ml) before expression analysis in Tn5 cells. For this, Tn5 cells were infected at an MOI of 5 with either recombinant virus (Ac-FMDV-3ABC) or a non-recombinant wild-type virus (AcMNPV) and incubated at 27 °C for 3–4 days to allow protein expression. Infected cell lysate was subjected to SDS-PAGE after suspending the cells in sample loading buffer, and the gel stained with Coomassie brilliant blue R-250. A distinct band corresponding to a molecular weight of 52 kDa was seen in the recombinant virus lysate, with no corresponding band in the AcMNPV cell lysate (Fig. 1). The 3ABC protein was found to be insoluble, requiring the use of denaturing conditions to purify it. The recombinant protein reacted specifically in a western blot with a known positive serum obtained from a convalescent animal infected previously with FMDV type A (IND/40/2000 strain) (Fig. 1).
Expression of FMDV 3ABC in insect cells. A). SDS-PAGE (12.5 % polyacrylamide gel) analysis of baculovirus (Ac-FMDV-3ABC)-expressed proteins in insect cells. Lane 1, 3ABC protein (52 kDa) expressed in TN5 cells; lane 2, polyhedrin protein (29 kDa) expressed in AcMNPV (wild-type baculovirus)-infected Tn5; lane M1, prestained protein molecular weight marker (MBI, Fermentas). B). Western blotting for detection of recombinant 3ABC protein. The purified protein showed reactivity, yielding a specific band (lane 1), which was absent in AcMNPV-infected culture (lane 2) when probed with serum from an FMDV (serotype A, IND-40/00 strain)-infected and recovered bull calf. C). SDS-PAGE showing purified recombinant protein. 3ABC purified by Ni-NTA affinity column shows a single band of polyhistidine-tagged protein. Lane M2, pre-stained molecular weight marker (Bio-Rad)
Assay optimisation
The ELISA was optimised using a known set of serum samples. For antigen coating, 50–100 ng of the protein in coating buffer was incubated overnight in each well. The analytical sensitivity was assessed using sera of strong and weak reactivity against the negative control sera, and dilutions in the range of 1:11 to 1:101 were found to be adequate for maximum discrimination between seropositive and negative samples. However, a dilution of 1:21 was found to be appropriate for the test.
Determination of the cutoff value
The mean PP value for negative samples (n = 356) was found to be 8.86 ± 6.59, and the cutoff was determined to be 25 based on the value obtained by summing two times the mean OD of the negative samples and one time the SD value, which corresponds to 24.3. Samples that had PP value of ≥25 were considered positive, and those below that value were considered negative in the indirect ELISA.
Analytical sensitivity and assay repeatability
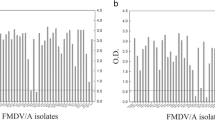
At the given cutoff, the in-house 3ABC ELISA showed higher sensitivity, with detection limits ranging from 1:320 to 1:640 for the three positive sera, while for the strong positive serum, the detection limit was 1:5120. Correspondingly, the endpoint detection limits for the prioCHECK test ranged from 1:80 to 1:160 for three positive sera and 1:1280 for the strong positive serum (Fig. 2).
Analytical sensitivity of the in-house 3ABC ELISA and prioCHECK FMDV NS test. Positive sera (PS) obtained from experimentally FMDV type A (IND-40/00 strain)-infected bull calves were serially diluted and tested at the indicated dilutions in the in-house 3ABC ELISA (A) in comparison with the PrioCHECK FMDV NS kit (B). The convalescent sera were collected at 3 weeks (PS-409), 5 weeks (PS-398), 6 weeks (PS-407) and 18 weeks (PS-411) after experimental infection. An adult bovine serum from an uninfected animal served as a negative serum (NS) control. The assay cutoff in each test system is indicated by a dashed line (–)
Assay repeatability was determined based on the absorbance values obtained for a set of positive and negative control sera used in the assays performed on different days in the laboratory by different operators. Mean OD ± standard deviation values for the positive and negative control samples were 0.89 ± 0.13 and 0.098 ± 0.017, with a coefficient of variation of 14.6 and 17.8 %, respectively.
Diagnostic performance in comparison to the prioCHECK FMDV NS test
The assay was compared with a prioCHECK FMDV NS test system using a panel of over 900 serum samples. Samples representing different profiles with respect to FMDV infection were screened, and the results are summarized in Table 1. Higher concordance was observed between both the assays for the naïve animals and infected animals, while these values were lower (84–91 %) for the vaccinated or vaccinated and infected sera, and for the randomly surveyed samples. About 14 % (20/138) of animals that received two or more vaccinations became positive for 3ABC antibodies when tested by the in-house ELISA, while the prioCHECK kit showed positivity in 8.7 % (12/138) of these animals. A higher proportion of vaccinated and infected animals were found to be positive in the in-house ELISA (86 %) when compared with the prioCHECK test (76.6 %).
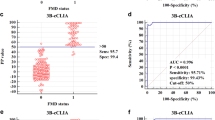
The results obtained in the two tests were compared by calculating the kappa coefficient, which was found to be 0.87 (95 % CI: 0.838–0.902) for the samples across all categories. Analysis based on the Medcalc program gave a cutoff of 24.88, sensitivity of 95.8 (95 % CI: 92.8–97.8) and specificity of 97.45 (95 % CI: 94.8–99.0), as shown in Table 2.
Kinetics of NSP antibody response following experimental infection
Early antibodies were detectable by 7–8 days after experimental infection in sheep, goats and cattle, using the in-house ELISA (Fig. 3) and by prioCHECK test (data not shown). This observation was made in most (18 out of 31) samples, while antibodies were detectable as early as 9 days postinfection in all of the infected animals. Five bull calves, which were monitored to determine the time course of their antibody response, showed varying responses. Two animals showed seropositivity up to 15 weeks postinfection, while in the third animal, antibodies decreased rapidly and were not detectable beyond 8 weeks by either of the tests. In the other two animals, 3ABC antibodies were detectable for up to 35 weeks.
NSP antibodies in randomly collected serum samples from field
Prevalence rates of 24 % in cattle, 20 % in buffalo, 21 % in sheep and 15 % in goats were found in the NSP antibody ELISA when randomly collected serum samples were screened using the in-house 3ABC ELISA (Table 3). Based on chi-square analysis, large ruminants showed a significantly higher seroprevalence (p = 0.044) than small ruminants (sheep and goats).
Discussion
India, with a population of ~520 million domestic livestock, including cattle, buffaloes, sheep, goats and pigs, has a huge challenge to protect this susceptible population from FMD, requiring a large number of quality diagnostic tests and vaccines. Validation of low-cost, in-house assays is thus critical for their large-scale application in the control-and-eradication program.
There are several commercially available diagnostic tests based on FMD NSP antigens, mainly targeting 3ABC (e.g., CHEKIT FMD-3ABC from Idexx Laboratories, PrioCHECK FMDV NS), but a few target either the 3A (UBI, USA) or the 3B protein (UBI, USA), and all of them are in indirect ELISA, blocking ELISA, or competitive ELISA format. Compared to competition ELISA, which targets one or two epitopes of the NSP antigen, indirect ELISA is expected to exhibit higher sensitivity, as it detects all of the antibodies that bind to multiple epitopes on the capture antigen. In the present study, a 3ABC-antigen-based indirect ELISA has been developed to detect NSP antibodies in sera from multiple species by employing a peroxidase-conjugated protein G as the detection antibody. Peroxidase-labeled protein G can substitute for HRP-labeled species-specific secondary antibodies in ELISA, particularly those with the albumin binding site removed from the protein G molecule. The assay was validated by testing different types of serum samples representing an endemic situation, and its performance was compared with that of a commercial kit. We expressed the protein in insect cells as a fusion product with an N-terminal his-tag to enable purification by Ni-NTA affinity chromatography. The major incentive for expressing the protein in an insect cell background was the negligible reactivity of known negative samples, even when the coating protein was not in a completely purified form, unlike E. coli proteins, which require absolute purification to avoid potential false positivity with test samples [7, 8]. The baculovirus system thus greatly helps to eliminate batch-to-batch variation of antigen production, making the test system more reproducible.
The test showed repeatability within the acceptable limits based on the data obtained on a set of control sera in different runs performed on the same day and different days of the assay validation. The analytical sensitivity of the indirect 3ABC ELISA was higher than that of the competition ELISA (prioCHECK test), as shown in Figure 2, although both tests are based on different formats for a realistic comparison. Although the in-house assay works adequately in the dilution range of 1:11 to 1:101 (data not shown), a lower dilution of 1:21 was chosen to avoid potential errors that may arise from higher sample dilutions. The results suggest that the indirect 3ABC ELISA had a similar performance to that of prioCHECK FMDV NS, a widely used test system. Diagnostic sensitivity and specificity were found to be 95.8 and 97.45 %, respectively, based on the serum samples with a known FMD status. The highest concordance between the in-house and the commercial ELISA was observed for naïve vs. infected samples (>98 %), while a lower concordance of 86 % was seen with random samples. The overall agreement between the tests was very good, as shown by a kappa coefficient value of 0.87 (95 % CI: 0.838 to 0.902).
A majority of vaccinated animals seroconvert to NSP antigens following exposure to FMDV [9, 10]. This observation underscores the utility of NSP diagnostics as the basis of the ‘vaccinate to live policy’ [11], a proposal for avoiding the pre-emptive slaughter of animals at risk in the face of an outbreak in disease-free regions, as has been followed in Korea [12]. However, demonstrating the absence of FMDV infection as required in the OIE terrestrial code is impossible in a vaccinated population, as tests are not sensitive enough to detect infection in some animals [3, 12], and there is wide variability in the ability of these tests to identify infected animals [13, 14]. The ability of a given test to detect an NSP antibody response in the vaccinated and subsequently infected animals depends on the potency and protective efficacy of the vaccine in question. Animals that are solidly protected tend to be poor reactors in the NSP antibody detection ELISA (our unpublished data). Our test exhibited relatively a higher sensitivity (86 %) compared to prioCHECK FMD NS (76.6 %) for identifying vaccinated and subsequently infected animals. Similarly the PANAFTOSA indirect ELISA has been shown to be the most sensitive (93.9 %) among the different ELISAs compared on vaccinated and subsequently infected samples [10].
Both the in-house ELISA and the prioCHECK FMDV NS test were sensitive enough to detect antibodies as early as 7–8 days after exposure to FMDV under experimental conditions, in agreement with previous reports [15]. This observation was made for most (18 out of 31) of the samples, while all animals showed positivity by day 9 postinfection. In time course studies, NSP antibodies were detectable in 2 out of 5 convalescent animals for up to 35 weeks, as demonstrated in both of the assays. Brocchi et al. [10] have compared the performance of six NSP ELISAs, and it was demonstrated that prioCHECK FMDV test showed an ability to detect an early antibody response, while NCPanaftosa showed better performance in detecting long-lived NSP antibodies. Although there have been several reports showing detectable NSP antibodies up to or beyond 365 days [8, 16], in the present study, both assays failed to detect the antibodies beyond 15–35 weeks postinfection. This variability can be attributed largely to the immune status of individual animals, previous exposure to infection (endemicity) and possible viral persistence in animals that carry NSP antibodies for a long time after primary exposure.
Earlier studies have shown that sera from animals vaccinated using currently licensed vaccines in India appear to elicit an NSP antibody response after repeated immunizations [17, 18]. Some vaccines, though produced a strong antibody response to structural proteins (data not shown), they apparently were not completely purified from NSP contaminants as evident in this study. It has been shown that administration of FMD vaccine manufactured using appropriate antigen purification methods, even with high antigenic payload, allows differentiation of infected animals using the available NSP antibody detection systems [19]. In India, although the supply of NSP-free vaccines for the publicly funded control program is currently not required to be free from NSP, there is a need to regulate this for the unambiguous interpretation of NSP serology data while assessing the impact of the control program in the long run. This study, though limited by the sample size, shows that animals that received two or more shots of vaccine exhibited an increase in their NSP antibody response, as seen both in the in-house ELISA and the prioCHECK test. The test also showed a significantly higher proportion (p = 0.044) of large animals (20–24 %) being positive for NSP antibodies compared to small ruminants (14–21 %), based on a randomly sampled population. This observation is consistent with previous reports showing a prevalence of 18.4–35 % in the Indian bovine population [20]. The proportion of NSP-antibody-reactive individuals in the bovine population is expected to progressively decrease in the long term, which is achievable through a supply of consistent batches of high-potency vaccines that are free from NSP contaminants. To complement this, the availability of validated and low cost in-house companion diagnostic tests is expected to be potentially useful in the serosurveillance of FMD in the progressive control of the disease.
References
Anon (2012) Global strategy for control of foot-and-mouth disease, Bangkok, pp 27–29
Rweyemamu M, Roeder P, MacKay D, Sumption K, Brownlie J, Leforban Y (2008) Planning for the progressive control of foot-and-mouth disease worldwide. Transbound Emerg Dis 55(1):73–87
OIE (2012) Foot-and-mouth disease. In: Manual of diagnostic tests and vaccines for terrestrial animals. World organization for animal health web. http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/2.01.05_FMD.pdf
Grubman MJ, Zellner M, Bablanian G, Mason PW, Piccone ME (1995) Identification of the active-site residues of the 3C proteinase of foot-and-mouth disease virus. Virology 213(2):581–589
Srisombundit V, Tungthumniyom N, Linchongsubongkoch W, Lekcharoensuk C, Sariya L, Ramasoota P, Lekcharoensuk P (2013) Development of an inactivated 3C(pro)-3ABC (mu3ABC) ELISA to differentiate cattle infected with foot and mouth disease virus from vaccinated cattle. J Virol Methods 188(1–2):161–167
Moonen P, van der Linde E, Chénard G, Dekker A (2004) Comparable sensitivity and specificity in three commercially available ELISAs to differentiate between cattle infected with or vaccinated against foot-and-mouth disease virus. Vet Microbiol 99(2):93–101
Jaworski JP, Compaired D, Trotta M, Perez M, Trono K, Fondevila N (2011) Validation of an r3AB1-FMDV-NSP ELISA to distinguish between cattle infected and vaccinated with foot-and-mouth disease virus. J Virol Methods 178(1–2):191–200
De Diego M, Brocchi E, Mackay D, De Simone F (1997) The non-structural polyprotein 3ABC of foot-and-mouth disease virus as a diagnostic antigen in ELISA to differentiate infected from vaccinated cattle. Arch Virol 142(10):2021–2033
Paton DJ, de Clercq K, Greiner M, Dekker A, Brocchi E, Bergmann I, Sammin DJ, Gubbins S, Parida S (2006) Application of non-structural protein antibody tests in substantiating freedom from foot-and-mouth disease virus infection after emergency vaccination of cattle. Vaccine 24(42–43):6503–6512
Brocchi E, Bergmann IE, Dekker A, Paton DJ, Sammin DJ, Greiner M, Grazioli S, De Simone F, Yadin H, Haas B, Bulut N, Malirat V, Neitzert E, Goris N, Parida S, Sørensen K, De Clercq K (2006) Comparative evaluation of six ELISAs for the detection of antibodies to the non-structural proteins of foot-and-mouth disease virus. Vaccine 24(47–48):6966–6979
Mackay D, Parida S, Paton D, Anderson J (2004) Making a vaccinate-to-live policy a reality in foot-and-mouth disease. Dev Biol (Basel) 119:261–266
Barnett PV, Geale DW, Clarke G, Davis J, Kasari TR (2013) A review of OIE country status recovery using vaccinate-to-live versus vaccinate-to-die foot-and-mouth disease response policies I: benefits of higher potency vaccines and associated NSP DIVA test systems in post-outbreak surveillance. Transbound Emerg Dis. doi:10.1111/tbed.12166
Parida S, Fleming L, Gibson D, Hamblin PA, Grazioli S, Brocchi E, Paton DJ (2007) Bovine serum panel for evaluating foot-and-mouth disease virus nonstructural protein antibody tests. J Vet Diagn Invest 19:539–544
Uttenthal A, Parida S, Rasmussen TB, Paton DJ, Haas B, Dundon WG (2010) Strategies for differentiating infection in vaccinated animals (DIVA) for foot-and-mouth disease, classical swine fever and avian influenza. Expert Rev Vaccines 9(1):73–87
Sørensen KJ, de Stricker K, Dyrting KC, Grazioli S, Haas B (2005) Differentiation of foot-and-mouth disease virus infected animals from vaccinated animals using a blocking ELISA based on baculovirus expressed FMDV 3ABC antigen and a 3ABC monoclonal antibody. Arch Virol 150(4):805–814
Sørensen KJ, Madsen KG, Madsen ES, Salt JS, Nqindi J, Mackay DK (1998) Differentiation of infection from vaccination in foot-and-mouth disease by the detection of antibodies to the non-structural proteins 3D, 3AB and 3ABC in ELISA using antigens expressed in baculovirus. Arch Virol 143(8):1461–1476
Mohapatra JK, Pandey LK, Sanyal A, Pattnaik B (2011) Recombinant non-structural polyprotein 3AB-based serodiagnostic strategy for FMD surveillance in bovines irrespective of vaccination. J Virol Methods 177(2):184–192
Kumar R, Hosamani M, Sreenivasa BP, Kotyal A, Venkataramanan R (2012) Expression of foot-and-mouth disease non-structural protein, 3D in insect cells and its application in detection of anti-FMDV antibodies. Indian J Virol 23(3):326–332
Espinoza AM, Maradei E, Mattion N, Cadenazzi G, Maddonni G, Robiolo B, La Torre J, Bellinzoni R, Smitsaart E (2004) Foot-and-mouth disease polyvalent oil vaccines inoculated repeatedly in cattle do not induce detectable antibodies to non-structural proteins when evaluated by various assays. Vaccine 23(1):69–77
Kumar N, Sharma R, Kakker NK (2007) Non-structural protein 3A for differentiation of foot-and-mouth disease infected and vaccinated animals in Haryana (India). Zoonoses Public Health 54(9–10):376–382
Acknowledgments
The authors thank the Director, Indian Veterinary Research Institute (IVRI), Izatnagar, for funding and facilitating this work.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hosamani, M., Basagoudanavar, S.H., Tamil Selvan, R.P. et al. A multi-species indirect ELISA for detection of non-structural protein 3ABC specific antibodies to foot-and-mouth disease virus. Arch Virol 160, 937–944 (2015). https://doi.org/10.1007/s00705-015-2339-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2339-9