Abstract
The human neurotropic JC virus (JCV) is of significant interest due to its experimental neuro- oncogenic potential. In clinical samples from human central nervous system (CNS) tumors, detection of JCV sequences suggests a possible association with CNS neoplasms, but the results are discrepant worldwide. To assess the prevalence of JCV sequences in Iranian patients with primary and metastatic CNS malignancies, a total of 58 fresh CNS tumors were examined by quantitative real-time PCR targeting the JCV large T antigen (LT-Ag) gene, and JCV DNA load was determined as viral copy number per cell. All patients were immunocompetent, and none of them had received immunosuppressive therapy before surgical operation. JC virus LT-Ag sequences were found in a total of 15 (25.9 %) out of the 58 tested samples. In primary CNS tumors, JCV sequences were identified more frequently in meningiomas (50.0 %) and schwannomas (35.7 %). In metastatic CNS tumors, JCV LT-Ag was identified in one case with brain adenocarcinoma originating from lung cancer. No statistically significant association between JCV positivity and various types of CNS malignancies was observed (P = 0.565). The mean JCV LT-Ag copy number in 15 positive cases was 1.8 × 10−4 ± 4.5 × 10−4 copies per cell (range 1.0 × 10−5-1.78 × 10−3 copies per cell). An inverse correlation between white blood cell (WBC) count and JCV copy number was observed, but this correlation was not statistically significant (R = −0.198, P = 0.480). This study provides the first data on the prevalence of JCV in primary and metastatic CNS tumors from Iranian patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Central nervous system (CNS) tumors are a diverse group of tumors arising from the brain, spine or meninges and comprise 2 % of all human adulthood malignancies [1]. The etiology of CNS tumors is poorly understood, and few well-established risk factors have been recognized. It has been suggested that genetic and environmental risk factors may play a role in CNS cancer development [2]; moreover, the hypothesis that oncogenic viruses may be correlated with the etiology of CNS tumors has also been proposed [3].
With respect to viral etiologies of CNS tumors, the polyomaviruses have been among the most widely investigated [4]. Polyomaviruses are small DNA viruses with experimental oncogenic potential that are ubiquitously found in the human population. Studies of urban sewage samples from various geographical areas have revealed significant shedding of polyomaviruses in human excreta and variable distribution of these viruses among human populations [5, 6]. Moreover, infections with polyomaviruses show signs of association with human populations, and strains of some polyomaviruses sequenced from sewage samples are correlated with previously described viral strains from related populations [5]. To date, 12 human polyomaviruses have been recognized [7]. Primary infection with human polyomaviruses is thought to be asymptomatic and occurs in childhood, after which the viruses maintain lifelong latency in their hosts. The kidneys are considered to play a role in JC virus (JCV) [8] and BK virus (BKV) [9] latency. Periodically, JCV and BKV can be found in the urine of normal individuals, without apparent pathology-associated consequences [10]. The cell type determines the outcome of polyomavirus infection [11]. Infection of permissive cells can lead to cell death and production of progeny virions (lytic infection) [12]. Progressive multifocal leukoencephalopathy (PML) is a neurodegenerative disease caused by lytic infection of JCV in oligodendrocytes [13]. BK polyomavirus lytic infection is an important cause of various urinary tract illnesses [14].
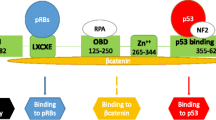
In nonpermissive cells, the virus can enter the cell and express its early genes (tumor antigens), but neither DNA replication nor late gene expression is seen. Therefore, this is a dead-end event that does not result in the production of infectious virions but instead leads to cell transformation and tumorigenesis [15, 16]. The large tumor antigen (LT-Ag) of polyomaviruses has been demonstrated to interfere with cell cycle regulation through binding and inactivation of cellular tumor suppressor proteins, p53 and pRb [12, 17]. The viral LT-Ag protein has also been shown to interact with several important proto-oncogene products such as insulin receptor substrate-1 (IRS-1) and β-catenin [18, 19].
The human neurotropic JC virus is of significant interest for its potential neurooncogenic role, since it has been demonstrated to induce CNS tumors in experimental animals [20–22]. The virus has the ability to transform cells in culture, especially cells of neural origin, including primary hamster brain cells and human fetal glial cells [4, 23, 24]. In immunocompetent individuals, due to immune surveillance mechanisms, JCV establishes and maintains a state of latency in kidneys, lymphoid organs and bone marrow [25–27]. Upon severe immunosuppression, virus can be reactivated and spread to the CNS, probably by infected B lymphocytes [28]. In the CNS, the virus targets oligodendrocytes, leading to demyelination and PML development. It should be noted that there have also been several recent studies reporting JCV persistence in the brains of healthy people [29–31]. Concomitant occurrence of PML and brain tumors provides the first evidence that JCV has a role in the development of these types of malignancies [32]. In the clinical context, JCV genomic sequences have been identified in tumor tissue from patients with a wide variety of CNS tumor subtypes [33–35]. However, the results are discrepant among different research groups, and the presence of JCV sequences in primary and metastatic CNS tumors has been a poorly investigated subject. Therefore, to determine a possible correlation between JCV infection and CNS tumors, more investigation is required. Also, there is little information about the prevalence of JCV in Iran. So far, only one study has evaluated the prevalence of JCV viruria in the Iranian general population [36], and to our knowledge, there are no data on the prevalence of JCV DNA sequences in Iranian patients with CNS tumors.
Hence, the facts reviewed above encouraged us to investigate the prevalence of JCV DNA sequences in tumor samples of Iranian patients diagnosed with primary and metastatic CNS malignancy and evaluate the positive samples in terms of viral copy number per cell.
Materials and methods
Patients and samples
A total of 58 fresh CNS tumor biopsy materials were collected from patients who underwent surgical operations of the CNS at the Neurosurgery Department of Shariati Hospital, affiliated to Tehran University of Medical Sciences. In the operating room, each fresh biopsy specimen was excised and placed immediately into RNALater solution (Sigma R0901, St. Louis, MO, USA). Samples were stored at −80 °C until an extraction procedure was executed. The central nervous system tumor type was confirmed as part of the routine histopathological examination in the Pathology Department of Shariati Hospital. Except for one patient, all of the patients included in this investigation had a white blood cell (WBC) count of more than 4500 cells per microliter prior to surgical operation. None of the patients were positive for antigen or antibody to human immunodeficiency virus type 1 (HIV-1). In addition, none of the patients had received immunosuppressive therapy, except for 8 milligrams of dexamethasone by the intravenous (IV) route prior to surgery to reduce swelling and pressure in the normal tissue around the tumor. Demographic and clinical characteristics for patients in each histopathologic group of CNS tumors are presented in Table 1. This study was approved by the Ethical Committee of Tehran University of Medical Sciences, and for all subjects, written informed consent was obtained.
Cloning of JC virus large T antigen and human RNase P amplicons
Brain sections from an HIV-positive subject with pathological PML were obtained at autopsy. The 247-bp JCV LT-Ag amplicon was amplified by PCR from DNA extracted from sections containing PML lesions, using TaKaRa EX Taq (Takara Bio, Shiga, Japan) with the primers JCTAGP1 (5′-GAGGAATGCATGCAGATCTAC-3′) and JCTAGP2 (5′-TTGCAGGGCATTTTGTTTTTTAC-3′) as described previously [37]. The 65-bp human RNase P gene (RPP30) was amplified by PCR from whole-blood genomic DNA using Taq polymerase and standard PCR conditions with primers RNaseP-F (5′-AGATTTGGACCTGCGAGCG-3′) and RNaseP-R (5′-GAGCGGCTGTCTCCACAAGT-3′). Both JCV LT-Ag and human RNase P amplicons were TA-cloned into pTZ57R/T PCR cloning vector (InsTAclone™ PCR Cloning Kit, Fermentas, MD, USA) according to the manufacturer’s instructions to obtain the plasmids pJCV LT-Ag and pRNase P, respectively. Purified plasmids were submitted for sequencing (Bioneer, Daejeon, South Korea) to verify that the correct sequence was present and used to derive standard curves in the real-time PCR assay for the quantification of JCV LT-Ag and human RNase P amplicons.
DNA extraction
DNA was isolated from the samples using a QIAamp DNA Mini Kit (QIAGEN GmbH, Hilden, Germany) for blood and tissues, according to the manufacturer’s instructions. The quality and quantity of extracted DNA was determined using a NanoDrop spectrophotometer (Thermo Scientific, Wilmington, USA) at the end of the extraction procedure. Sterile tubes containing only reaction mixtures were processed in parallel with the tissue samples as an extraction negative control.
Quantitative real-time polymerase chain reaction
Quantitative real-time PCR was performed using a Rotor-Gene® Q real-time PCR system (QIAGEN GmbH, Hilden, Germany), using primer sets and a TaqMan probe specific for the JCV LT-Ag gene and the human RNase P gene [38, 39]. The primer and probe sequences and cycling conditions are shown in Table 2. All primers and probes were synthesized by Metabion International AG (Martinsried, Germany). Real-time PCR reaction mixes were prepared in a dedicated clean room in a biological hood under stringent sterile conditions. Each reaction consisted of 500 ng of extracted DNA, 12.5 µl Maxima Probe qPCR Master Mix (Fermentas, Glen Burnie, MD, USA), 0.3 µM each primer and 0.2 µM dual-labeled probe in a 25-µl total reaction volume. To generate standard curves, real-time PCR was performed on a tenfold dilution series of each purified plasmid (pJCV LT-Ag and pRNase P) ranging from 2 × 101 to 2 × 106 copies/µl. Samples that were negative for human RNase P amplification were considered to have inadequate DNA integrity and therefore were re-extracted until RNase P amplification could be achieved. Quantitative real-time PCR for JCV LT-Ag was performed on triplicate repeats, and samples were considered positive if two out of the three tests had detectable JCV DNA. Each real-time PCR run included reaction mixtures without DNA template as a negative control and the plasmid pJCV LT-Ag as a positive control. The quality and accuracy of the real-time PCR assay for JCV were confirmed by testing on extracted DNA from clinical samples (brain tissue sections from different brain areas) obtained from a subject with PML. The number of viral gene copies per cell was calculated by dividing the virus copy number by half of the RNase P copy number, because each diploid cell contains two copies of RNase P.
Statistical analysis
Statistical analysis was performed using SPSS version 16 software (SPSS Inc., Chicago, IL, USA). The χ2 test was utilized to assess associations between categorical variables. (The exact P value was estimated by Mont Carlo method.) The normal distribution of the variables was analyzed using the Kolmogorov-Smirnov test. Analysis of continuous variables was carried out using the independent-samples T-test/Mann-Whitney U test and Kruskal-Wallis test. Correlation between quantitative variables was analyzed by Spearman correlation. A P value of ≤0.05 was considered to be statistically significant.
Results
In this investigation, 54 (93.1 %) primary CNS tumors and four (6.9 %) metastatic CNS tumors were investigated. Of the four tumors with CNS metastasis, three cases were diagnosed as adenocarcinoma (originating from lung cancer) and one case was diagnosed as squamous cell carcinoma with unknown primary site of origin. The primary CNS tumor samples in this study were categorized according to histopathologic criteria as follows: Of the 54 samples, 14 (24.1 %) were classified as schwannoma, 12 (20.7 %) as meningioma, and seven (12.1 %) as glioblastoma multiform. The less common tumors included astrocytoma, pituitary adenoma, and epidermoid tumor (each of which was found in three (5.2 %) patients), hemangioblastoma, pineoblastoma and oligodendroglioma (each of which was found in two (3.4 %) patients), and oligoastrocytoma, chordoma, cavernoma, medulloblastoma, xanthoastrocytoma and ependymoma (each of which was found in one (1.7 %) patient). The mean age of the patients (male 28, female 30) was 46.0 ± 17.6 years (range 5–83 years). With respect to WBC count of subjects, the mean was 8438.8 ± 2645.5 cells per microliter (range, 3400-16700 WBC per microliter).
In the current study, specimens from CNS tumors were tested for the presence of JCV LT-Ag sequences by quantitative real-time PCR. The results from quantitative real-time PCR revealed the presence of JCV LT-Ag in a total of 15 (25.9 %) out of the 58 tested samples. Out of 15 subjects who tested positive for JCV, six were male. No significant differences were found between JCV positivity and gender (χ2 test, P = 0.554). The mean age was 48.8 ± 18.6 years for JCV-positive cases (range 18-83 years) and 45.0 ± 17.3 years for patients who tested negative for JCV (range 5-74 years). No statistically significant difference between patients’ mean age and JCV positivity was seen (independent-samples T-test, P = 0.479). The mean WBC count of JCV-positive subjects was 9147.3 ± 3244.2 cells per microliter (range, 4500-16700 WBC per microliter), while the mean WBC count was 8191.7 ± 2397.5 cells per microliter (range, 3400-15490 WBC per microliter) for JCV-negative subjects. There was no significant difference between JCV-positive and negative subjects regarding mean WBC count (Mann-Whitney U test, P = 0.143). In detail, 50 % of meningiomas, 35.7 % of schwannomas, 33.3 % of astrocytomas, 33.3 % of metastatic adenocarcinomas, and 33.3 % of epidermoid tumors contained JCV LT-Ag sequences. Also, the only sample of chordoma that was available for this study was positive for a JCV LT-Ag DNA sequence. No statistically significant association between JCV positivity and the various types of CNS malignancies was observed (χ2 test using the Monte Carlo method, P = 0.565) (Table 3).
In the present study, JCV DNA load was determined as the viral DNA copies per RNase P gene copy (a proven single-copy gene), corresponding to the viral copy number per cell. In addition, amplification of the cellular RNase P gene was used as a control for the presence of sufficient amplifiable DNA. The mean JCV LT-Ag copy number in 15 positive cases was 1.8 × 10−4 ± 4.5 × 10−4 copies per cell (range 1.0 × 10−5-1.78 × 10−3 copies per cell). Among JCV-positive CNS tumors, the mean JCV copy number was higher in the group of “others” (consisting of JCV-positive CNS tumors except for schwannomas and meningiomas, including JCV-positive cases with metastatic adenocarcinoma, astrocytoma, epidermoid tumor and chordoma) (mean = 4.6 × 10−4 ± 8.7 × 10−4 copies per cell) in comparison with schwannoma (mean = 9.0 × 10−5 ± 1.2 × 10−4 copies per cell) and meningioma (mean = 6.0 × 10−5 ± 1.2 × 10−4 copies per cell), but this difference was not statistically significant (Kruskal-Wallis test, P = 0.727) (Fig. 1A).
JC virus DNA loads and WBC count in JCV-positive CNS tumors. (A) The mean JCV LT-Ag DNA load in JCV-positive CNS tumors (B) The mean WBC count per microliter in JCV-positive CNS tumors. The group of others includes JCV-positive cases with metastatic adenocarcinoma, astrocytoma, epidermoid tumor and chordoma. Error bars indicate standard error. The P-value was determined by the Kruskal-Wallis test (A) and Mann-Whitney U test (B)
In JCV-positive CNS tumors, the mean WBC count was higher in the group of “others” (mean = 12,175.0 ± 3160.6 WBC per microliter) in comparison with meningioma (mean = 9185.0 ± 2647.9 WBC per microliter) and schwannoma (mean = 6680.0 ± 1950.0 WBC per microliter), and the WBC counts were significantly higher in the group of “others” than schwannoma (Mann-Whitney U test, P = 0.014) (Fig. 1B). In addition, an inverse correlation between WBC count and JCV copy number per cell was observed, which means that elevation in WBC count correlated with reduction in JCV copy number per cell, but this correlation was not statistically significant (Spearman correlation, R = −0.198, P = 0.480).
Discussion
The most frequent types of CNS tumors in the present cross-sectional study were schwannoma, meningioma and glioblastoma, which is in accordance with the pattern of CNS tumors in developing countries [40]. Reports regarding the presence of JCV sequences in human CNS malignancies vary widely, and it is still a subject of debate whether JCV conclusively plays a role in CNS cancer development. The reason for this discrepancy between the research groups is not entirely clear, but it might be connected to DNA isolation from formalin-fixed or paraffin-embedded tumors or the sensitivity of the different PCR methodologies. In the present study, we used fresh CNS tumor biopsy samples and real-time PCR, based on the quantitation of viral gene copy number per cell to determine the exact JCV copy number in CNS tumor cells. Normalization of viral gene copy numbers to cell numbers might be informative when evaluating viral loads in clinical samples, such as tumor tissues [37].
In the current study, biopsy samples were examined for sequences that are part of the JCV LT-Ag gene. JC virus LT-Ag sequences were found in 25.9 % of the biopsies examined. In primary CNS tumors, JCV genomic sequences were identified more frequently in meningiomas (50.0 %) and schwannomas (35.7 %). In metastatic CNS tumors, JCV LT-Ag was identified in one (33.3 %) out of three adenocarcinoma cases that originated from lung cancer. A recent report demonstrated that JCV may be involved in lung carcinogenesis, and high copy numbers of the virus were found in lymph node metastasis [41]. Also, several studies have provided evidence that different DNA tumor viruses may play a role in the stimulation of tumor cell migration and promote a metastatic phenotype [42, 43]. This concept was also demonstrated for JCV in a recent study showing that viral LT-Ag is associated with increased malignant behavior in colorectal cancers and that JCV LT-Ag is frequently expressed in both primary colorectal cancer and colorectal cancer liver metastasis [44]. In addition, the presence of JC virus in both primary and metastatic CNS tumors may weaken the hypothesis of a pathogenic role for JCV in primary tumor induction.
Several lines of evidence indicate that the product of the LT-Ag gene is a multifunctional protein that has been proposed to play a cardinal role in aberrant stimulation of the cell cycle [45, 46]. Therefore, the presence of LT-Ag sequences in malignant tissues might be indirect evidence in support of a potential role of JCV in oncogenic transformation. In the current investigation, low copy numbers of JCV LT-Ag gene per cell were detected in primary and metastatic CNS tumors. Detection of low copy numbers of JCV LT-Ag in CNS tumors is a matter for debate. If JCV-induced tumors in humans occur as a result of clonal expansion of a virally transformed cell, a single copy of the viral genome would be present in each tumor cell [15]. Alternatively, low copy numbers of the LT-Ag gene may contribute to tumor initiation, but not full progression to malignancy, by the “hit-and-run” mechanism, as proposed for other oncogenic viruses [12, 15].
It has been demonstrated that immunosuppression increases the JCV LT-Ag DNA load in the brain [30]. A similar finding also was made in our study, in which elevation of WBC count was correlated with reduction of JCV copy number in CNS tumor samples. However, our results failed to find a statistical significant correlation between WBC count and JCV LT-Ag DNA load, possibly due to the small number of JCV-positive CNS tumors. Recent evidence has demonstrated JCV latent infection in resident brain cells [29]. In addition, low JCV LT-Ag DNA loads have been detected in the brain tissues of immunocompetent patients without PML [30, 31]. Considering the immunocompetence status of our subjects with respect to WBC count, and the lack of HIV infection, the low copy numbers of JCV LT-Ag might be explained in two ways. The first explanation could be that the low level of JCV contributes to tumor induction by indirect mechanisms. The second could be that the low copy numbers of JCV is due to latent viral infection in the CNS. Taken together, the findings of the current study should be interpreted with caution due to the lack of matching non-neoplastic or normal CNS tissues as a control. In addition, evaluation of the patient’s infection status in plasma and urine may shed more light on results found in CNS tissues, but unfortunately, access to plasma or urine samples from the patients was not possible in our study.
In conclusion, this study provides the first data on the prevalence of JC virus in CNS tumors from Iranian patients. The present study is a preliminary work that reveals a relatively high frequency of JCV LT-Ag sequences in CNS tumors, including schwannomas and meningiomas, but also in metastatic tumors. This is an important aspect that weakens the hypothesis of the pathogenic role of JCV in primary tumor induction. Therefore, further studies should be done to differentiate the possible role of JCV in tumor induction from simple latent viral replication.
References
Siegel R, Naishadham D, Jemal A (2013) Cancer statistics, 2013. CA Cancer J Clin 63:11–30
Preston-Martin S (1996) Epidemiology of primary CNS neoplasms. Neurol Clin 14:273–290
Saddawi-Konefka R, Crawford JR (2010) Chronic viral infection and primary central nervous system malignancy. J Neuroimmune Pharmacol 5:387–403
White MK, Gordon J, Reiss K, Del Valle L, Croul S, Giordano A, Darbinyan A, Khalili K (2005) Human polyomaviruses and brain tumors. Brain Res Brain Res Rev 50:69–85
Bofill-Mas S, Pina S, Girones R (2000) Documenting the epidemiologic patterns of polyomaviruses in human populations by studying their presence in urban sewage. Appl Environ Microbiol 66:238–245
Bofill-Mas S, Rodriguez-Manzano J, Calgua B, Carratala A, Girones R (2010) Newly described human polyomaviruses Merkel cell, KI and WU are present in urban sewage and may represent potential environmental contaminants. Virol J 7:141
Moens U, Van Ghelue M, Ehlers B (2014) Are human polyomaviruses co-factors for cancers induced by other oncoviruses? Rev Med Virol. doi:10.1002/rmv.1798
Padgett BL, Walker DL, ZuRhein GM, Eckroade RJ, Dessel BH (1971) Cultivation of papova-like virus from human brain with progressive multifocal leucoencephalopathy. Lancet 1:1257–1260
Gardner SD, Field AM, Coleman DV, Hulme B (1971) New human papovavirus (B.K.) isolated from urine after renal transplantation. Lancet 1:1253–1257
Markowitz RB, Thompson HC, Mueller JF, Cohen JA, Dynan WS (1993) Incidence of BK virus and JC virus viruria in human immunodeficiency virus-infected and -uninfected subjects. J Infect Dis 167:13–20
Gjoerup O, Chang Y (2010) Update on human polyomaviruses and cancer. Adv Cancer Res 106:1–51
Khalili K, Del Valle L, Otte J, Weaver M, Gordon J (2003) Human neurotropic polyomavirus, JCV, and its role in carcinogenesis. Oncogene 22:5181–5191
Koralnik IJ (2006) Progressive multifocal leukoencephalopathy revisited: Has the disease outgrown its name? Ann Neurol 60:162–173
Arthur RR, Shah KV, Baust SJ, Santos GW, Saral R (1986) Association of BK viruria with hemorrhagic cystitis in recipients of bone marrow transplants. N Engl J Med 315:230–234
Moore PS, Chang Y (2010) Why do viruses cause cancer? Highlights of the first century of human tumour virology. Nat Rev Cancer 10:878–889
zur Hausen H (2001) Oncogenic DNA viruses. Oncogene 20:7820–7823
Poulin DL, Kung AL, DeCaprio JA (2004) p53 targets simian virus 40 large T antigen for acetylation by CBP. J Virol 78:8245–8253
Del Valle L, Wang JY, Lassak A, Peruzzi F, Croul S, Khalili K, Reiss K (2002) Insulin-like growth factor I receptor signaling system in JC virus T antigen-induced primitive neuroectodermal tumors–medulloblastomas. J Neurovirol 8(Suppl 2):138–147
Gan DD, Reiss K, Carrill T, Del Valle L, Croul S, Giordano A, Fishman P, Khalili K (2001) Involvement of Wnt signaling pathway in murine medulloblastoma induced by human neurotropic JC virus. Oncogene 20:4864–4870
London WT, Houff SA, Madden DL, Fuccillo DA, Gravell M, Wallen WC, Palmer AE, Sever JL, Padgett BL, Walker DL, ZuRhein GM, Ohashi T (1978) Brain tumors in owl monkeys inoculated with a human polyomavirus (JC virus). Science 201:1246–1249
London WT, Houff SA, McKeever PE, Wallen WC, Sever JL, Padgett BL, Walker DL (1983) Viral-induced astrocytomas in squirrel monkeys. Prog Clin Biol Res 105:227–237
Walker DL, Padgett BL, ZuRhein GM, Albert AE, Marsh RF (1973) Human papovavirus (JC): induction of brain tumors in hamsters. Science 181:674–676
Frisque RJ, Rifkin DB, Walker DL (1980) Transformation of primary hamster brain cells with JC virus and its DNA. J Virol 35:265–269
Mandl C, Walker DL, Frisque RJ (1987) Derivation and characterization of POJ cells, transformed human fetal glial cells that retain their permissivity for JC virus. J Virol 61:755–763
Atwood WJ, Amemiya K, Traub R, Harms J, Major EO (1992) Interaction of the human polyomavirus, JCV, with human B-lymphocytes. Virology 190:716–723
Randhawa P, Shapiro R, Vats A (2005) Quantitation of DNA of polyomaviruses BK and JC in human kidneys. J Infect Dis 192:504–509
Tan CS, Dezube BJ, Bhargava P, Autissier P, Wuthrich C, Miller J, Koralnik IJ (2009) Detection of JC virus DNA and proteins in the bone marrow of HIV-positive and HIV-negative patients: implications for viral latency and neurotropic transformation. J Infect Dis 199:881–888
Sabath BF, Major EO (2002) Traffic of JC virus from sites of initial infection to the brain: the path to progressive multifocal leukoencephalopathy. J Infect Dis 186(Suppl 2):S180–S186
Perez-Liz G, Del Valle L, Gentilella A, Croul S, Khalili K (2008) Detection of JC virus DNA fragments but not proteins in normal brain tissue. Ann Neurol 64:379–387
Bayliss J, Karasoulos T, McLean CA (2013) Immunosuppression increases JC polyomavirus large T antigen DNA load in the brains of patients without progressive multifocal leukoencephalopathy. J Infect Dis 207:133–136
Tan CS, Ellis LC, Wuthrich C, Ngo L, Broge TA Jr, Saint-Aubyn J, Miller JS, Koralnik IJ (2010) JC virus latency in the brain and extraneural organs of patients with and without progressive multifocal leukoencephalopathy. J Virol 84:9200–9209
Richardson EP Jr (1961) Progressive multifocal leukoencephalopathy. N Engl J Med 265:815–823
Del Valle L, Gordon J, Assimakopoulou M, Enam S, Geddes JF, Varakis JN, Katsetos CD, Croul S, Khalili K (2001) Detection of JC virus DNA sequences and expression of the viral regulatory protein T-antigen in tumors of the central nervous system. Cancer Res 61:4287–4293
Delbue S, Pagani E, Guerini FR, Agliardi C, Mancuso R, Borghi E, Rossi F, Boldorini R, Veggiani C, Car PG, Ferrante P (2005) Distribution, characterization and significance of polyomavirus genomic sequences in tumors of the brain and its covering. J Med Virol 77:447–454
Tsekov I, Ferdinandov D, Bussarsky V, Hristova S, Kalvatchev Z (2011) Prevalence of JC polyomavirus genomic sequences from the large T-antigen and non-coding control regions among Bulgarian patients with primary brain tumors. J Med Virol 83:1608–1613
Bozorgi SM, Tahaei SM, Mohebbi SR, Sahba N, Damavand B, Romani S, Azimzadeh P, Naghoosi H, Milanizadeh S, Mohebbi A, Zali MR (2012) Molecular prevalence of JC virus in Tehran, Iran. Gastroenterol Hepatol Bed Bench 5:84–89
McNees AL, White ZS, Zanwar P, Vilchez RA, Butel JS (2005) Specific and quantitative detection of human polyomaviruses BKV, JCV, and SV40 by real time PCR. J Clin Virol 34:52–62
MacKenzie J, Wilson KS, Perry J, Gallagher A, Jarrett RF (2003) Association between simian virus 40 DNA and lymphoma in the United Kingdom. J Natl Cancer Inst 95:1001–1003
Imajoh M, Hashida Y, Taniguchi A, Kamioka M, Daibata M (2012) Novel human polyomaviruses, Merkel cell polyomavirus and human polyomavirus 9, in Japanese chronic lymphocytic leukemia cases. J Hematol Oncol 5:25
Jazayeri SB, Rahimi-Movaghar V, Shokraneh F, Saadat S, Ramezani R (2013) Epidemiology of primary CNS tumors in Iran: a systematic review. Asian Pac J Cancer Prev 14:3979–3985
Zheng H, Abdel Aziz HO, Nakanishi Y, Masuda S, Saito H, Tsuneyama K, Takano Y (2007) Oncogenic role of JC virus in lung cancer. J Pathol 212:306–315
Behren A, Simon C, Schwab RM, Loetzsch E, Brodbeck S, Huber E, Stubenrauch F, Zenner HP, Iftner T (2005) Papillomavirus E2 protein induces expression of the matrix metalloproteinase-9 via the extracellular signal-regulated kinase/activator protein-1 signaling pathway. Cancer Res 65:11613–11621
Gou XM, Chen Y, Chen XY, Arrand JR (2003) Effects of Epstein-Barr virus latent membrane protein 1 (EBV-LMP1) on related factors of metastasis of nasopharyngeal carcinoma cell line CNE1. Ai Zheng 22:481–485
Link A, Shin SK, Nagasaka T, Balaguer F, Koi M, Jung B, Boland CR, Goel A (2009) JC virus mediates invasion and migration in colorectal metastasis. PLoS One 4:e8146
Krynska B, Gordon J, Otte J, Franks R, Knobler R, DeLuca A, Giordano A, Khalili K (1997) Role of cell cycle regulators in tumor formation in transgenic mice expressing the human neurotropic virus, JCV, early protein. J Cell Biochem 67:223–230
White MK, Khalili K (2006) Interaction of retinoblastoma protein family members with large T-antigen of primate polyomaviruses. Oncogene 25:5286–5293
Acknowledgments
The authors would like to acknowledge the support of the directors and staff of the Neurosurgery Department of Shariati Hospital affiliated to Tehran University of Medical Sciences for their collaboration in sample collection. This study was financially supported by a grant from Tehran University of Medical Sciences (Project code: 92-01-30-20990).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sadeghi, F., Salehi-Vaziri, M., Ghodsi, S.M. et al. Prevalence of JC polyomavirus large T antigen sequences among Iranian patients with central nervous system tumors. Arch Virol 160, 61–68 (2015). https://doi.org/10.1007/s00705-014-2230-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-014-2230-0