Abstract
The complete genome of sweet potato latent virus (SPLV) was determined to be 10081 nucleotides long excluding the 3’ poly (A) tail. The genome contains a single large open reading frame encoding a polyprotein of 3247 amino acids. Its genomic organization is typical of potyviruses and contains motifs conserved in members of the genus Potyvirus. Pairwise comparisons show that SPLV shares identities of 50.0 %-56.3 % to other potyviruses at the genomic sequence level. Phylogenetic analysis shows that SPLV is closely related to four other sweet potato potyviruses in the sweet potato feathery mottle virus lineage, but it lacks the unique PISPO in the P1 region of those viruses. The genome analyses confirm that SPLV is a distinct sweet potato virus in the genus Potyvirus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Members of six species of the genus Potyvirus infect sweet potato and cause significant yield losses in sweet potato production worldwide [1]. Sweet potato latent virus (SPLV), formerly designated as sweet potato virus N, was first reported from Taiwan [2]. The virus was then found in several other countries in Asia and Africa [3–6]. SPLV has flexuous, filamentous particles that are approximately 700-750 nm long, and it induces typical cylindrical inclusions in the cytoplasm of infected cells. The experimental host range of SPLV is wider than that of sweet potato feathery mottle virus (SPFMV), inducing symptoms on some Chenopodium and Nicotiana species [2, 7]. Similar to SPFMV and sweet potato virus G (SPVG), a single infection by SPLV is symptomless in most sweet potato cultivars, but co-infection with sweet potato chlorotic stunt virus (SPCSV) causes synergistic disease [8]. SPLV is serologically related to, but distinct from, SPFMV [7]. Sequence comparison of 3’-partial sequences showed that SPLV is a member of a distinct species in the genus Potyvirus [9]. In this study, the complete genomic sequence of SPLV was determined, and phylogenetic analysis clearly indicated that SPLV is a distinct potyvirus that is most closely related to sweet potato virus 2 (SPV2) and other viruses in the SPFMV lineage.
The SPLV isolate originally collected from a diseased sweet potato cv. Tainung 63 co-infected with SPFMV in Taiwan in 1970s [2] was used in this study. The infected sweet potato plant was transferred from Taiwan to the International Potato Center (CIP) in 1980s. SPLV was transmitted to Nicotiana clevelandii by mechanical inoculation and maintained in this experimental host through mechanical inoculation. Leaf tissue from an infected N. clevelandii plant was obtained under a USDA-APHIS permit and used to inoculate N. benthamiana seedlings. The inoculated N. benthamiana plants were tested by RT-PCR for SPFMV, SPVC, SPVG, SPV2, SPLV and SPMMV, and the results showed that the plants were only infected with SPLV. Total nucleic acids were extracted from an infected N. benthamiana plant by the CTAB method [10]. RT-PCR was performed using a SuperScript TM III One-Step RT-PCR System (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instruction.
The complete genomic sequence of SPLV was determined by a genomic-walking strategy described for ApVY [11] in a series of sequential RT-PCR cloning steps. The 3’-terminal sequence of approximately 1.7 kb was obtained by RT-PCR using a SPLV-specific forward primer designed based on the 3’-partial sequences of SPLV-CY (EF492050) and an oligo (dT) primer. The 3’-terminal sequence was extended to 2950 bp using the degenerate forward primer PotyF1 and an SPLV-specific primer based on the sequence obtained in this study. Sequences of partial CI and HC-Pro fragments were obtained from RT-PCR products using two pairs of degenerate primers (CIFor/CIRev and HPFor/-HPRev, respectively) described by Ha et al. [12]. The HC-Pro–NIb sequence of 5.0 kb was determined from two overlapping RT-PCR clones using SPLV-specific primers based on the sequences of the three fragments (HC-Pro, CI and 3’ terminus) obtained during this study. The 5’-terminal region of 3.3 kb was amplified by RT-PCR using an SPLV-specific reverse primer and a 5’-end forward primer, Poty5endF1 (5’-GTTTTCCCAGTCACGACAAAATATAAAAACTCAACA-3’), based on the 5’-end sequences of some potyviruses. To obtain an accurate 5’-end sequence, a 5’ RACE reaction was conducted using a GeneRacer® Kit (Invitrogen) according to the manufacturer’s instructions. The complete genomic sequence of SPLV was confirmed using nine overlapping fragments obtained by RT-PCR with SPLV-specific primers and another 5’RACE assay.
Contig assembly was performed using the DNAStar 5.01 package (DNAStar Inc., Madison, WI, USA). The putative cleavage sites in the SPLV polyprotein sequence and conserved domains of the putative functional proteins were determined by comparison of its polyprotein sequence with those of representative potyviruses. Pairwise comparisons of nucleotide sequences, deduced polyprotein sequences and individual protein sequences were performed using the EMBOSS Needle Pairwise Sequence Alignment tool at http://www.ebi.ac.uk/Tools/psa/emboss_needle/nucleotide.html. Phylogenetic analysis of the complete genomic sequences of SPLV and 82 other potyviruses was conducted using MEGA 5, and neighbor-joining and maximum-likelihood trees were constructed using 1000 bootstrap replicates. Putative recombination breakpoints were identified using the RDP4 Beta 4.16 program (http://darwin.uvigo.es/rdp/rdp.html).
The complete genomic sequence of SPLV is 10081 nucleotides (nt) long excluding the 3’ poly (A) tail and has been submitted to the GenBank database under accession number KC443039. Its genomic organization is typical of potyviruses, containing a single open reading frame (ORF) (Fig. 1). The putative ORF starts at the first in-frame AUG (nt 144-146) and ends with a UAG termination codon at nt 9885-9887. It encodes a polyprotein of 3247 amino acids (aa) with a calculated molecular weight of 367.8 kDa. The nine putative protease cleavage sites are identified in the SPLV polyprotein, which are predicted to give rise to ten putative mature proteins (P1, HC-Pro, P3, 6K1, CI, 6K2, VPg, NIa-Pro, NIb, and CP). Most motifs typical of potyviruses are found in the amino acid sequences of these mature proteins: 350H-8X-D-31X-G-X-S-G394 and 414F-I/V-V-R-G418 in P1; 623F-R-N-K626 and 786C-72X-H859 in HC-Pro; 1389G-A-V-G-S-G-K-S-T1397, 1409V-L-L-V/L-E-P-T-R-P-L1418 and 1478D-E-X-H1481 in CI; 2239H-34X-D-67X-G-X-C-G-14X-H2360 in NIa-Pro; 2682C-D-A-D-G-S2687 and 2746S-G-3X-T-3X-N-T-30X-G-D-D2789 in NIb [13–15]. In the CP, the D-A-G motif, which is involved in aphid transmission [15], was changed to 2961D-T-G2963. This mutation only occurs in five other isolates of SPLV (accession nos. HQ844128, HQ844129, HQ844131, HQ844147, X84012), and the majority of the SPLV isolates reported to date have the D-A-G motif in the CP. The substitution of threonine in the second position of the D-A-G motif reduces aphid transmissibility in tobacco veinal mosaic virus [16]. A lysine in the first position of another aphid transmissibility motif in the HC-Pro, K-I-T-C [15], is changed to 494E-I-T-C497 in SPLV. The significance of the changes in these two motifs on aphid transmission of this SPLV isolate is not known. A putative P3N-PIPO ORF was identified that include the frame shift sequence 3311GAAAAAA3317 [17].
Schematic representation of the genome organization of sweet potato latent virus. The 5’- and 3’-untranslated regions are represented by solid lines, and the open reading frame (ORF) is depicted by an open box with solid lines. The location of each putative cleavage site on the polyprotein is marked by an arrow, and amino acid residues around each cleavage sites are listed for each arrow. The name and size of each protein are listed inside each mature protein, and number above each cleavage site indicate the number of the first amino acid residue of the mature protein in the polyprotein sequence
A BLAST search in the GenBank database shows that the polyprotein sequence of SPLV is most closely related to that of SPV2. Pairwise comparisons show that SPLV shares identities of 51.0 % (onion yellow dwarf virus, OYDV) to 56.3 % (SPV2) at whole genomic sequence level and 39.6 % (narcissus degeneration virus) to 49.8 % (SPV2) at the polyprotein sequence level, respectively, with 82 other members of the genus Potyvirus. These values fall under the species demarcation thresholds [18], confirming that SPLV is a distinct species in the genus Potyvirus. Detailed comparisons of individual protein sequences show that SPLV has the highest degree of identity to viruses in the SPFMV lineage (SPFMV, sweet potato virus C, SPVG and SPV2) [19] for HC-Pro, P3, 6K1, CI, 6K2, NIa and NIb, reflecting the relatedness of the whole genome. The aa sequences of the P1, VPg and CP of SPLV share the highest degree of identity with Brugmansia suaveolens mottle virus, Moroccan watermelon mosaic virus and pepper veinal mottle virus, respectively.
Despite similarities in genomic and aa sequences of seven central proteins (HC-ProNIb), SPLV is different from viruses of the SPFMV lineage, especially in the P1 region. The P1 region of SPLV does not contain the third ORF encoding the putative PISPO unique to the SPFMV lineage [19]. The SPLV P1 protein of 442 aa is shorter than that of the SPFMV lineage (618-724 aa) due to four deletions at the N-terminal and central regions. The P1 protein also does not contain WG/GW(Y) motifs and zinc-finger-like structure unique to the SPFMV lineage and sweet potato mild mottle virus (SPMMV). The sizes of all individual mature proteins of SPLV are similar to those of most potyviruses. The sequence identities of these proteins between SPLV and the viruses of the SPFMV lineage are also similar to those of most members of the genus Potyvirus.
A phylogenetic analysis was initially conducted using the deduced polyprotein sequences of SPLV and 82 other members of the genus Potyvirus. Both neighbor-joining and maximum likelihood trees clearly place SPLV in a distinct branch adjacent to the SPFMV lineage in the PVY group [19, 20]. The tree was then simplified to include the representative members of five groups of the genus Potyvirus (Fig. 2). With the exception of P1 and CP, a similar clustering of SPLV and the SPFMV lineage is maintained in phylogenetic trees based on the other proteins and the complete genomic sequences of all available potyviruses (data not shown). The results indicate that SPLV and the SPFMV lineage may share a common ancestor, with separation of SPLV and the SPFMV occurring before speciation of the viruses in the SPFMV lineage. However, how and where these potyviruses have evolved is not clear, since they are very closely related to the viruses of subgroup 1 in the PVY group that have evolved in the Asia-Mediterranean-Europe region. Furthermore, the distribution of SPLV is mainly restricted to Asian production areas [6, 7], suggesting that the virus might have evolved there. Analysis of the complete genomic sequences of SPLV, four viruses in the SPFMV lineage, and SPMMV using the RDP4 program did not reveal any positive recombination breakpoints in SPLV (data not shown).
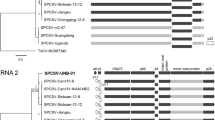
Maximum-likelihood tree based on the deduced polyprotein sequences of sweet potato latent virus and representative members of the genus Potyvirus. Bootstrap analysis was applied using 1000 bootstrap replicates. The scale bar represents a genetic distance of 0.2. Sugarcane streak mosaic virus, a member of the genus Poacevirus, was used as an outgroup
Phylogenetic analysis based on the CP aa sequences of this isolate and 30 other isolates of SPLV (27 from China, two from Taiwan and one from Japan) available in GenBank showed that this Taiwan isolate was most closely related to two other Taiwan isolates (Taiwan [accession no. X84012] and CY [EF492090]), and they are placed in the major cluster [7]. Comparison of the 3’ partial sequences of 1076 nt showed that there were only 14 nt differences (1.3 %) between the two Taiwan isolates. The difference is very similar to those among different clones from a given region (approximately 1 %). Therefore, the two Taiwan isolates might come from the same source, representing the same isolate. Serial passage of this isolate in tobacco plants through continuous mechanical inoculation might not have much effect on the viral population structure. This study provides the first report of the complete genomic sequence of SPLV and will assist in the development of molecular diagnostic tests and effective disease management strategies.
References
Clark CA, Davis JA, Abad JA, Cuellar WJ, Fuentes S, Kreuze JF, Gibson RW, Mukasa SB, Tugume AK, Tairo FD, Valkonen JPT (2012) Sweetpotato viruses: 15 years of progress on understanding and managing complex diseases. Plant Dis 96:168–185
Liao CH, Chien IC, Chung ML, Chiu RJ, Han YH (1979) A study of sweetpotato virus disease in Taiwan. I. Yellow spot virus. J Agric Res China 28:127–137
Wang Q, Zhang L, Wang B, Yin Z, Feng C, Wang Q (2010) Sweetpotato viruses in China. Crop Protect 29:110–114
Qin Y, Zhang Z, Qiao Q, Zhang D, Tian Y (2013) Molecular variability of sweet potato chlorotic stunt virus (SPCSV) and five potyviruses infecting sweet potato in China. Arch Virol 158:491–4955
Yun WS, Lee YH, Kim KH (2002) First Report of Sweet potato latent virus and Sweet potato chlorotic stunt virus Isolated from Sweet Potato in Korea. Plant Pathol J 18:126–129
Njeru RW, Bagabe MC, Nkezabahizi D, Kayiranga D, Kajuga J, Butare L, Ndirigue J (2008) Viruses infecting sweet potato in Rwanda: occurrence and distribution. Ann Appl Biol 153:215–221
Rossel HW, Thottappilly G (1988) Complex virus diseases of sweet potato. In: Exploration, maintenance and utilisation of sweet potato genetic resources. Report of first Sweet Potato planning Conference (1987). International Potato Centre, Lima, Peru, pp 291–302
Untiveros M, Fuentes S, Salazar LF (2007) Synergistic interaction of Sweet potato chlorotic stunt virus (Crinivirus) with carla-, cucumo-, ipomo-, and potyviruses infecting sweet potato. Plant Dis 91:669–676
Colinet D, Kummert J, Lepoivre P (1997) Evidence for the assignment of two strains of SPLV to the genus Potyvirus based on coat protein and 3’ non-coding region sequence data. Virus Res 49:91–100
Li R, Mock R, Huang Q, Abad J, Hartung J, Kinard G (2008) A reliable and inexpensive method of nucleic acid extraction for the PCR-based detection of diverse plant pathogens. J Virol Methods 154:48–55
Xu D, Liu HY, Koike ST, Li F, Li R (2011) Biological characterization and complete genomic sequence of Apium virus Y infecting celery. Virus Res 155:76–82
Ha C, Coombs S, Revill PA, Harding RM, Vu M, Dale JL (2008) Design and application of two novel degenerate primer pairs for the detection and complete genomic characterization of potyviruses. Arch Virol 153:25–36
Adams MJ, Antoniw JF, Beaudoin F (2005) Overview and analysis of the polyprotein cleavage sites in the family Potyviridae. Mol Plant Pathol 6:471–487
Revers F, Le Gall O, Candresse T, Maule AJ (1999) New advances in understanding the molecular biology of plant/potyvirus interactions. Mol Plant Microbe Interact 12:367–376
Urcuqui-Inchima S, Haenni AL, Bernardi F (2001) Potyvirus proteins: a wealth of functions. Virus Res 74:157–175
Atreya PL, Lopez-Moya JJ, Chu M, Atreya CD, Pirone TP (1995) Mutational analysis of the coat protein N-terminal amino acids involved in potyvirus transmission by aphids. J Gen Virol 76:265–270
Chung BYW, Miller WA, Atkins JF, Firth AE (2008) An overlapping essential gene in the Potyviridae. Proc Natl Acad Sci USA 105:5897–5902
Adams MJ, Antoniw JF, Fauquet CM (2005) Molecular criteria for genus and species discrimination within the family Potyviridae. Arch Virol 150:459–479
Li F, Xu D, Abad J, Li R (2012) Phylogenetic relationships of closely related potyviruses infecting sweet potato determined by genomic characterization of Sweet potato virus G and Sweet potato virus 2. Virus Genes 45:118–125
Gibbs A, Ohshima K (2010) Potyviruses and the digital revolution. Ann Review Phytopathol 48:205–223
Acknowledgments
The authors thank Crindi Loschinkohl, Whitney Hymes and Sam Grinstead for excellent technical assistance, and Dr. Gary Kinard for critical review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, M., Abad, J., Fuentes, S. et al. Complete genome sequence of the original Taiwanese isolate of sweet potato latent virus and its relationship to other potyviruses infecting sweet potato. Arch Virol 158, 2189–2192 (2013). https://doi.org/10.1007/s00705-013-1705-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1705-8