Abstract
We determined complete 1D gene sequences for one serotype A and seven additional serotype O Korean foot-and-mouth disease viruses (FMDV) and then analyzed them together with published sequences for 180 type A and 300 type O isolates from throughout the world using a Bayesian coalescent approach. Here, Korean serotype A virus was linked with those from Laos. Korean serotype O viruses were divided into three clades and were closely related to isolates from Japan, Thailand, the UK, France, Ireland, South Africa, and Singapore, as well as Laos. There was no apparent correlation between time, country, or host species and the evolution of global FMDVs. Additionally, our results showed that purifying selection acts on the overall 1D sequences and there was no evidence of recombination among the FMDV sequences. The evolutionary rates were 5.77 × 10−3 substitutions/site/year for serotype A and 4.81 × 10−3 substitutions/site/year for serotype O. Serotype A viruses diverged approximately 110 years ago, while serotype O isolates segregated approximately 127 years before the present. In both serotype isolates, the effective number of infections remained constant until the late 1990 s, after which the virus population size underwent a rapid, sharp decline until the present.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Foot-and-mouth disease (FMD) is highly infectious and causes serious economic losses in the global livestock industry. The notorious pathogen, foot-and-mouth disease virus (FMDV), belongs to the genus Aphthovirus of the family Picornaviridae and is divided into seven antigenically distinct serotypes: A, O, C, Asia 1, and SAT 1–3. The virus has a linear single-stranded RNA genome of about 8.5 kb that consists of 12 protein-coding genes encoding a leader peptide (L), structural proteins P1 (1A, 1B, 1C, 1D), and non-structural proteins P2 (2A, 2B, 2C), P3 (3A, 3B, 3C, 3D) [1]. 1D encodes VP1, which is the most surface-exposed capsid protein and contains two important immunogenic sites at amino acid positions 140–160 (the G-H loop) and residues 200–213 (the C-terminus). Variation at specific amino acid sites within those two immunogenic regions contributes to the antigenic variability of the virus [2]. Since the first report of foot-and-mouth disease (FMD) in South America in 1870 [3], 1D sequences have been regarded as the most useful markers for determining the genetic relationships of these viruses [4–6].
Korea had been free of FMD for 66 years, since 1934; then, there were three FMD outbreaks, in 2000, 2002, and 2010. Two outbreaks, reported in 2000 and 2002, were caused by serotype O viruses, whereas the one in 2010 was due to serotype A viruses. Despite many global studies based on 1D sequences [7–9], the molecular epidemiology of FMDV with an emphasis on Korean isolates has not been well resolved. Only three studies [10–12] have been published, although some pathological [13] and antigenic [14–16] studies of Korean FMDVs have been reported. Shin et al. [10] determined the first partial 1D sequences (162 nucleotides in length) from seven Korean FMDV isolates in 2000 and then compared them with the sequences of 67 other foreign isolates. Their results indicated that Korean isolates could be divided into two distinct clusters within the Pan Asian clade. Cluster 1 was most closely related to the strains prevalent in the Far East, and cluster 2 was further diverged from cluster 1 in their tree. Oem et al. [11] reported two additional Korean FMDV genome sequences (O/SKR/2000, O/SKR/2002) in 2000 and 2002 and analyzed their polyprotein gene sequences phylogenetically, together with 15 other Pan-Asian isolates, including two published Korean isolates. In their studies, Korean FMDVs did not appear as an independent clade, and it was unclear how and when this virus was introduced into Korea. Subsequently, Oem et al. [12] also performed a phylogenetic analysis based on 26 partial 1D sequences (165 nucleotides in length) from FMDV isolates from various parts of the world, including 10 Korean isolates from 2000 and 2002. Their results indicated that the Korean viruses could be separated into three clusters. Here, Korean isolates from 2000 were divided into two clusters, clusters 2 and 3, but the isolates from 2002 belonged to cluster 1. In their tree, the Korean FMDVs of cluster 1 were closely related to samples from Mongolia, whereas Korean members of cluster 2 were linked to samples from Russia. However, they did not further characterize the phylogeny of FMDV with an emphasis on Korean isolates. Unfortunately, previous studies on the phylogeny of Korean FMDV used obscure analytical methods and did not present details of the assumptions used [10–12]. Bootstrap values outlining the robustness of the phylogenetic analysis were not shown in these three publications. Additionally, they analyzed FMDV sequences for only one serotype, serotype O.
There has been only one study related to the time scale of FMDVs based on 1D sequences [9]. In that report, the mean substitution rates were 4.26 × 10−3 substitutions/site/year for serotype A FMDVs and 3.14 × 10−3 substitutions/site/year for serotype O viruses. Here, serotype A and O viruses diversified approximately 178 and 92 years ago, respectively. However, the population dynamics of both serotype A and O viruses were not presented in that study.
To further investigate the molecular epidemiology and evolution of FMD virus serotypes A and O with an emphasis on Korean isolates, we determined the complete 1D sequences of the first Korean A-type and seven additional Korean O-type isolates. Additionally, we then analyzed them along with published sequences for 180 type A and 300 type O (seven from Korea) isolates from throughout the world. The specific aims of this study were: (1) to determine whether the Korean FMDVs have a common origin, (2) to assess the phylogenetic relationships between Korean FMDVs and those from other countries, (3) to test the influences of time periods, countries, and host species on global FMDV evolution, (4) to examine the selection pressures and recombination of global FMDV evolution, and (5) to estimate substitution rates, divergence times, and population size changes in global FMDV evolution.
Materials and methods
Samples and data collection
FMDV isolates were selected from four pigs and four cows showing clinical symptoms from seven different Korean regions (two from Jinchon, one each from Yongin, Anseong, Pyoungtaek, Boryeong, Hongseong, and Pocheon) in 2000, 2002, and 2010. The eight sequences obtained (one for serotype A and seven for serotype O) were analyzed together with published sequences from seven Korean isolates (all serotype O) and from 480 non-Korean FMDV isolates (180 for serotype A and 300 for serotype O). For all analyses in the present study, two datasets were made: one consisted of the 1D sequences of 181 type-A viruses collected from various parts of the world over 67 years (1943–2010), and the other consisted of the sequences of 307 serotype O isolates circulating globally over the last 70 years (1939-2009).
RNA extraction, RT-PCR, and sequencing
Viral genomic RNAs were extracted from clinical epithelium or vesicular fluid samples using a BioRobot M48 Workstation (QIAGEN) and a MagAttract Viral RNA M48 Kit (QIAGEN). cDNAs were synthesized from the viral RNAs using Superscript III with random hexamers (Invitrogen) according to the manufacturer’s protocol. Aliquots of 4 μL of cDNA were mixed with 10 μL of buffer (5×), 8 μL of MgCl2 (7.5 mM), 1 μL of dNTPs (10 mM), 2 μL of primers (10 mM), 0.5 μL of Go Taq (Promega), and 22.5 μL of nuclease-free water. The primer sequences and PCR amplification conditions used were reported previously [12]. The amplified fragments were sequenced in both the forward and reverse orientations using an ABI Prism® 3730xi DNA sequencer at the Macrogen Institute (Macrogen Co., Ltd.) using the corresponding 1D PCR primers. Complete 1D sequences of each isolate determined in this study were submitted to GenBank, with accession numbers GU082465, GU082466, GU082471, GU082476, GU082477, GU082480, GU082481, and GU441855.
Sequence analysis, selection pressure analysis, and detection of recombination
Both nucleotide and amino acid sequence identities of the 1D region were calculated using BIOEDIT 7.053 [17]. Base frequencies, ts/tv ratios, and the numbers of total sites (including gaps), conserved sites, and variable sites were calculated using Modeltest 3.7 [18].
To evaluate the selective pressure driving FMDV evolution, nonsynonymous/synonymous substitution ratio (ω = dN/dS) values were estimated using ClustalX 1.81 [19], PAL2NAL [20], and the codeml program in the PAML 3.14.1 package [21, 22].
To detect putative recombinants, identification of likely parental sequences and localization of possible recombination breakpoints were performed using the RDP 3.0b41 package [23] with the default parameters. This package consists of six different recombination detection programs: RDP [24], GENECONV [25], MaxChi [26], BOOTSCAN [27], Chimeara [28], and SiScan [29]. To exclude the possibility of detecting false-positive recombinants, we considered only putative recombinant sites detected by at least three methods. In all analyses, P < 0.05 was taken to indicate statistical significance.
Phylogeny inference and molecular dating
Phylogenetic trees, rates of nucleotide substitutions, times of the most recent common ancestor (TMRCA), and changes in population size were coestimated using the Bayesian Markov Chain Monte Carlo (MCMC) approach, as implemented in BEAST 1.5.3 [30]. For both datasets (serotypes A and O), GTR+I+G was selected as the best-fit model using Akaike’s Information Criterion (AIC) in Modeltest 3.7 [18]. We set the parameters of nst=6 and rates=gamma as the likelihood settings for both data matrices.
We employed both strict and relaxed (both uncorrelated exponential and uncorrelated log-normal) molecular clocks [31] with five different demographic models (constant size, exponential growth, expansion growth, logistic growth, and Bayesian skyline). By comparison of Bayes factors (log10 Bayes factors > 2 in all cases) based on the relative marginal likelihoods of the models [32], the relaxed uncorrelated exponential clock and exponential population size model were selected as showing the best fit for both FMDV datasets. The changes in effective population size over time were examined using the Bayesian skyline plot (BSP). The datasets were each run for 200,000,000 generations to ensure convergence of all parameters (ESSs > 200), with discarded burn-in of 10%. The resulting convergence was analyzed using Tracer 1.5 (http://beast.bio.ed.ac.uk/Tracer), and the statistical uncertainties were summarized in the 95% highest probability density (HPD) intervals. Trees were summarized as maximum clade credibility trees using the TreeAnnotator program, which forms part of the BEAST package, and visualized using FigTree 1.3.1 [33].
Results
Sequence analysis, selection pressure analysis, and detection of recombination
We determined the complete 1D sequences for the first Korean serotype A isolate from 2010 as well as for seven additional Korean serotype O viruses from 2000 and 2002. All of the 1D sequences of the Korean FMDVs were the same length (633 nucleotides). Thus, the deduced amino acid sequences were 211 residues in length.
The variability in the complete 1D sequences of FMDVs from throughout the world is summarized in Table 1. The overall lengths of both serotype alignments, including gaps, were 642 base pairs, and the deduced amino acid sequences were 214 residues in length. The 1D sequences of the serotype A sequences revealed considerable genetic variation: 236 (36.8%) of the nucleotide positions and 90 (42.3%) of the amino acid positions were conserved. The nucleotide sequence identities among the serotype A isolates ranged from 74.4 to 99.8%, corresponding to 78.4–99.5% identity at the amino acid level. The transition/transversion ratio estimated from the dataset was 4.25. In comparison with serotype A sequences, serotype O sequences were more variable: 171 (26.6%) of the nucleotide positions and 56 (26.3%) of the amino acid positions were conserved. Pairwise comparisons showed that the identities between serotype O isolates ranged from 73.2 to 99.8% for the nucleotide sequences and from 76.5 to 99.5% for the amino acid sequences. The transition/transversion ratio calculated from the serotype O virus dataset was 5.00.
The nonsynonymous/synonymous substitution ratio (ω = dN/dS) value of serotype A isolates was 0.1029, while that of serotype O viruses was 0.0685. Accordingly, our analyses indicated that purifying selection acted on the 1D sequences of both serotype A and O FMDVs. Additionally, no recombination events were detected among the viruses.
Phylogenetic analysis
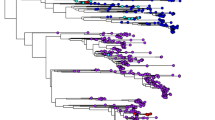
For both FMDV datasets (serotypes A and O), we conducted phylogenetic analysis using the Bayesian method. Figure 1 shows the maximum clade credibility tree using alignment of the complete 1D sequences of 181 global serotype A isolates. Bayesian analysis indicated that all of the A-type FMDVs could be divided into one of three groups or were unclassified (one isolate, EU553852) with very high posterior probabilities (all ≥1.00). Group 1 of serotype A viruses consisted of 120 isolates (66.3% of all A-type viruses) collected in 1965–2010. All members, except three European viruses (one each from Azerbaijan, Macedonia, and Albania), were from Asian countries. Here, Korean FMDV had a sister-group relationship with isolates from Laos. All isolates of group 2 originated from African countries (Egypt, Kenya, Ethiopia, Gambia, Mali, and Cameroon) between 1972 and 2007. Members of group 3 were derived from three different continents, Europe (Greece, Italy, Netherlands, Germany, Spain, and Malta), South America (Argentina, Uruguay, Brazil, and Venezuela), and Africa (Morocco and Libya) during 1943–2002. The host species of all A-type FMDVs, excluding two isolates (FJ755026, sheep; FJ755061, goat), was cattle.
Bayesian maximum clade credibility phylogenetic tree obtained from complete 1D sequences of 181 serotype A FMDVs from around the world. The Korean isolate is indicated in italic and bold type. Bayesian posterior probabilities (≥0.80) are shown above the nodes. Divergence times (in years) are shown below the nodes, and the 95% HPD intervals are indicated in brackets. Groups are marked by “G”
Figures 2 and 3 show the maximum clade credibility trees derived from the complete 1D sequences of 307 global serotype O isolates. The topology of the trees indicated that all of the global O-type FMDVs belonged to one of seven groups (all, posterior probabilities ≥ 0.92) or were unclassified (6 isolates: AJ303500, AJ303501, AJ303502, AJ303503, AJ303509, and AY593825). Group 1 consisted of 169 viruses, which accounted for 55% of the global O-type FMDVs. They were from 32 countries of Asia, Europe, and Africa, isolated during 1988–2007, and their hosts included cattle, pigs, sheep, goats, and buffaloes. In particular, all Korean O-type isolates appeared to be members of this group. They were not monophyletic but were divided into three clusters: K1, K2, and K3. Here, both K1 and K2 consisted of isolates sampled in 2000, while K3 contained viruses collected in 2002. FMDVs of K1 had close relationships with the clade including viruses from Northeast Asia (Japan), Southeast Asia (Thailand), Europe (UK, France, and Ireland), and South Africa, while the K2 members were closely related to samples from Laos. Isolates of K3 were linked with viruses from Singapore. Group 2 consisted of FMDV viruses collected from Southwest Asia (Israel and Oman), South Asia (Pakistan), Europe (Italy), and Africa (Egypt) during 1972–1997. Viruses of group 3 were collected from 10 African countries (Ethiopia, Sudan, Algeria, Cote d’Ivoire, Guinea, Ghana, Kenya, Uganda, Malawi, and Tanzania) during 1978–2007 and had only one host species (cattle). Group 4 was grouped with isolates from Southeast Asia (Malaysia, Myanmar, Thailand, Laos, and Vietnam) and South Asia (India) during 1962–2003. Their hosts included cattle, pigs, and buffaloes. Members of group 5 were collected from Northeast Asia (Hong Kong) and Southeast Asia (Taiwan, China, Philippines, and Vietnam) during 1958–2009. They had bovine and porcine hosts. Group 6 contained FMDV viruses obtained from South America (Brazil, Bolivia, Paraguay, Argentina, and Uruguay) during 1983–2006, while group 7 consisted of isolates from Europe (UK, Denmark, Germany, and Belgium) during 1963–2007. Group 6 had bovine hosts.
Bayesian maximum clade credibility phylogenetic tree obtained from complete 1D sequences of 307 serotype O FMDVs from around the world. Korean isolates are indicated in italic and bold type. Bayesian posterior probabilities (≥0.80) are shown above the nodes. Divergence times (in years) are shown below the major nodes, and the 95% HPD intervals are indicated in brackets. Groups are marked by “G”
Substitution rates, divergence times, and population size changes
The estimated mean evolutionary rates of serotype A and O FMDVs were 5.77 × 10−3 (95% HPD = 4.81 × 10−3–6.74×10−3) and 4.81 × 10−3 (95% HPD = 4.04 × 10−3–5.46 × 10−3) substitutions/site/year, respectively. The TMRCA estimated for the current diversity of serotype A viruses was 110.4 (95% HPD = 84.4–150.3) years ago, while the TMRCA value for the serotype O isolates was estimated at 126.5 (95% HPD = 81.3–193.9) years before the present. On the other hand, Bayesian skyline plot analysis of serotype A isolates (Fig. 4) showed that the viruses appeared to have been evolving under almost constant population size until the late 1940 s, when they experienced a sharp population expansion until the late 1950 s. Their population size then remained constant again until the late 1990 s, when a rapid, sharp decline in the effective number of infections occurred. Subsequent analysis of the population dynamics of serotype O isolates (Fig. 5) indicated that the population size remained constant until the mid-1990 s, when a rapid, sharp decline in the effective population size began.
Discussion
FMD, caused by foot-and-mouth disease virus (FMDV), is still one of the most severe problems affecting cloven-hoofed animals worldwide. Accordingly, both national and global strategies are necessary to prevent and control this acute disease. The expanding database and evolutionary information on FMDV sequences obtained from this study could also be useful for increasing our understanding of its emergence and epidemiology and may aid in the prevention and control of this virus.
In sequence comparison of global FMDVs, 1D sequences of both serotype A and serotype O showed a very high degree of genetic diversity, with dissimilarities between global A-type isolates ranging from 0.2 to 25.6% and from 0.5 to 21.6% at the nucleotide and amino acid sequence level, respectively. Those between global O-type FMDVs were from 0.2 to 26.8% and from 0.5 to 23.5% at the nucleotide and amino acid sequence level, respectively. This confirmed the viewpoints of some FMDV workers, such as Batschelet et al. [34] and Drake and Holland [35], who reported that FMDV is characterized by high mutation rates, of 10−3 to 10−5 per replication cycle. Carrillo et al. [36] reported that 1D is the most variable of the 12 individual genes. To date, 1D has been regarded as a target gene for PCR detection of FMDV, and the sensitivity of PCR detection relies on sequence variability. Additionally, the capsid protein encoded by this gene is immunologically important because it contains cell receptor sites on the GH-loop (residues 140–160) as well as at the C-terminus (residues 200–213) [2]. Accordingly, continuous molecular monitoring of the variability of the FMDV 1D sequence is necessary to update primers and for studies of the major antigenic genes.
Regarding selection pressure on FMDVs, our findings indicated that the 1D regions of serotypes A and O are both under strong purifying selection; the ω (dN/dS) value of serotype A isolates was 0.1029 and that of serotype O viruses was 0.0685. There have been previous studies related to selection pressure. Tully and Fares [37] suggested that positive selection has governed the evolution of the major antigenic regions of the corresponding VP1 protein of serotypes A, Asia 1, O, SAT 1, and SAT 2, but not C or SAT 3. Lewis-Rogers et al. [38] noted that 29% of residues in the capsid proteins were under positive selection. Additionally, Cooke and Westover [39] reported that both antigenic and non-antigenic regions were subject to purifying selection, although several antigenic sites showed a pattern of nucleotide substitution suggesting repeated positive selection. On the other hand, exchange of different genetic sites between viruses by recombination contributes to FMDV genetic diversity and evolution. In our analyses, no evidence of a recombination event was observed within the 1D region in serotype A or O.
In the global phylogeny of serotype A FMDVs resulting from our Bayesian approach using complete 1D sequences, the Korean isolate was linked to samples from Laos. It is interesting that serotype A virus was newly introduced to Korea from Southeast Asia in 2010, although only serotype O viruses occurred in Korea in 2000 and 2002. Globally, among FMDV type O viruses, the Korean FMDVs did not emerge as an independent clade but were divided into three clades: K1, K2, and K3 (K1 and K2 for 2000; K3 for 2002). This supported the suggestion of Oem et al. [12], based on partial 1D sequences. Here, K1 isolates were closely related to the clade consisting of viruses from Japan, Thailand, the UK, France, Ireland, and South Africa, while K2 samples had a common origin with those from Laos. K3 viruses were linked with the Singapore isolates. In contrast to the present study, however, Oem et al. [12] suggested that Korean type O FMDVs originated from Mongolia and Russia. These sister-group relationships of Korean FMDVs with many different countries may be largely due to frequent international trade in animals.
We found no apparent correlation between time, country, or host species and the evolution of FMDVs of serotypes A or O. These findings were consistent with the views of other groups [8, 38, 40], and this may be largely due to the explosive and wide-ranging divergence of FMDVs over a relatively short period of time as well as their rapid spread via the frequent international trade in livestock. The mixed population structure can make vaccine development more difficult. Thus, it is essential to continuously screen for changes in the mixed population structure of this virus.
Regarding the evolutionary rates and divergence times of FMDV serotypes A and O, there has been one study [9], based on 1D sequences of 37 serotype A and 47 serotype O viruses. These authors calculated mean evolutionary rates of 4.26 × 10−3 for serotype A and 3.14 × 10−3 for serotype O isolates and suggested that the common ancestors of serotype A and O FMDVs occured about 178 and 92 years ago, respectively. In contrast, the mean evolutionary rates estimated in the present study were higher: 5.77 × 10−3 (95% HPD = 4.81 × 10−3–6.74 × 10−3) substitutions/site/year for serotype A and 4.81 × 10−3 (95% HPD = 4.04 × 10−3–5.46 × 10−3) substitutions/site/year for serotype O. Additionally, the TMRCA estimated here did not support these previous suggestions. Our findings indicated that serotype O and A diverged approximately at the same time; serotype A viruses diverged 110.4 (95% HPD = 84.4–150.3) years ago, and the serotype O isolates were segregated 126.5 (95% HPD = 81.3–193.9) years before the present.
In terms of dynamics in the population size of FMDVs, our Bayesian skyline plot analyses showed similar configurations for serotype A and O isolates; the effective number of infections remained constant until the late 1990s, after which the virus population size underwent a rapid, sharp decline until the present day. The sharp decrease of effective FMDV population size since the late 1990s may be due to the effective use of vaccination to control FMD worldwide for many years [41]. This feature could also be the result of many other events, such as a bottleneck event or a reduced substitution rate.
Here, we report the first Korean serotype A virus sequence and the phylogeny of Korean FMDV serotypes A and O. We also explain the evolutionary and past epidemiological dynamics of FMDV serotypes A and O from throughout the world using a Bayesian coalescent approach.
References
Domingo E, Baranowski E, Escarmis C, Sobrino F (2002) Foot and-mouth disease virus. Comp Immunol Microbiol Infect Dis 25:297–308
Sobrino F, Saiz M, Jimenez-Clavero MA, Nunez JI, Rosas MF, Baranowski E, Ley V (2001) Foot-and-mouth disease virus: a long known virus, but a current threat. Vet Res 32:1–30
Melo EC, López A (2002) Control of foot and mouth disease: the experience of the Americas. Rev Sci Tech Off Int Epiz 21:695–698
Beck E, Strohmaier K (1987) Subtyping of European foot-and-mouth disease virus strains by nucleotide sequence determination. J Virol 61:1621–1629
Tosh C, Hemadri D, Sanyal A, Bandyopadhyay SK (2003) Genetic and antigenic analysis of two recently circulating genotypes of type A foot-and-mouth disease virus in India: evidence for positive selection in the capsid-coding genes. Arch Virol 148:853–869
König GA, Palma EL, Maradei E, Piccone ME (2007) Molecular epidemiology of foot-and-mouth disease virus types A and O isolated in Argentina during the 2000–2002 epizootic. Vet Microbiol 124:1–15
Mohapatra JK, Priyadarshini P, Pandey L, Subramaniam S, Sanyal A, Hemadri D, Pattnaik B (2009) Analysis of the leader proteinase (L(pro)) region of type A foot-and-mouth disease virus with due emphasis on phylogeny and evolution of the emerging VP3(59)-deletion lineage from India. Virus Res 141:34–46
Schumann KR, Knowles NJ, Davies PR, Midgley RJ, Valarcher JF, Raoufi AQ, McKenna TS, Hurtle W, Burans JP, Martin BM, Rodriguez LL, Beckham TR (2008) Genetic characterization and molecular epidemiology of foot-and-mouth disease viruses isolated from Afghanistan in 2003–2005. Virus Genes 36:401–413
Tully DC, Fares MA (2008) The tale of a modern animal plague: tracing the evolutionary history and determining the time-scale for foot and mouth disease virus. Virology 382:250–256
Shin JH, Sohn HJ, Choi KS, Kwon BJ, Ko YJ, An DJ, Cha SH, Park JH, Jeong WS, Park JY, Choi CU, Kweon CH, Song JY, Kim JY, An SH, Kim SJ, Joo YS (2003) Molecular epidemiological investigation of foot-and-mouth disease virus in Korea in 2000. J Vet Med Sci 65:9–16
Oem JK, Lee KN, Cho IS, Kye SJ, Park JH, Joo YS (2004) Comparison and analysis of the complete nucleotide sequence of foot-and-mouth disease viruses from animals in Korea and other Panasia strains. Virus Genes 29:63–71
Oem JK, Lee KN, Cho IS, Kye SJ, Park JY, Park JH, Kim YJ, Joo YS, Song HJ (2005) Identification antigenic site analysis of foot-and-mouth disease virus from pigs and cattle in Korea. J Vet Sci 6:117–124
Oem JK, Yeh MT, McKenna TS, Hayes JR, Rieder E, Giuffre AC, Robida JM, Lee KN, Cho IS, Fang X, Joo YS, Park JH (2008) Pathogenic characteristics of the Korean 2002 isolate of foot-and-mouth disease virus serotype O in pigs and cattle. J Comp Pathol 138:204–214
Park JH, Kim SJ, Oem JK, Lee KN, Kim YJ, Kye SJ, Park JY, Joo YS (2006) Enhanced immune response with foot and mouth disease virus VP1 and interleukin-1 fusion genes. J Vet Sci 7:257–262
Heo EJ, Lee HS, Jeoung HY, Ko HR, Kweon CH, Ko YJ (2010) Development of a blocking ELISA using a recombinant glycoprotein for the detection of antibodies to vesicular stomatitis New Jersey virus. J Virol Methods 164:96–100
Ko YJ, Lee HS, Jeoung HY, Heo EJ, Ko HR, Chang BS, Joo HD, Gerelmaa U, Dashzeveg B, Tserendorj S, Sodnomdarjaa R, Park JH, Kweon CH, Cho IS, Paik SG (2010) Use of a baculovirus-expressed structural protein for the detection of antibodies to foot-and-mouth disease virus type A by a blocking enzyme-linked immunosorbent assay. Clin Vaccine Immunol 17:194–198
Hall TA (1999) BIOEDIT: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acid Symp Ser 41:95–98
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal-windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acid Res 22:4673–4680
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acid Res 34:W609–W612
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 48:929–936
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Martin DP, Williamson C, Posada D (2005) RDP2: recombination detection and analysis from sequence alignments. Bioinformatics 21:260–262
Martin D, Rybicki E (2000) RDP: detection of recombination amongst aligned sequences. Bioinformatics 16:562–563
Padidam M, Sawyer S, Fauquet CM (1999) Possible emergence of new geminiviruses by frequent recombination. Virology 265:218–225
Smith JM (1992) Analyzing the mosaic structure of genes. J Mol Evol 34:126–129
Martin DP, Posada D, Crandall KA, Williamson C (2005) A modified bootscan algorithm for automated identification of recombinant sequences and recombination breakpoints. AIDS Res Hum Retroviruses 21:98–102
Posada D, Crandall KA (2001) Evaluation of methods for detecting recombination from DNA sequences: computer simulations. Proc Natl Acad Sci USA 98:13757–13762
Gibbs MJ, Armstrong JS, Gibbs AJ (2000) Sister-scanning: a Monte Carlo procedure for assessing signals in recombinant sequences. Bioinformatics 16:573–582
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Drummond AJ, Ho SY, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Bio 4:e88. doi:10.1371/journal.pbio.0040088
Suchard MA, Weiss RE, Sinsheimer JS (2001) Bayesian selection of continuous-time Markov chain evolutionary models. Mol Biol Evol 18:1001–1013
Rambaut A (2006) Available at http://evolve.zoo.ox.ac.uk/beast
Batschelet E, Domingo E, Weissmann C (1976) The proportion of revertant and mutant phage in a growing population, as a function of mutation and growth rate. Gene 1:27–32
Drake JW, Holland JJ (1999) Mutation rates among RNA viruses. Proc Natl Acad Sci USA 96:13910–13913
Carrillo C, Tulman ER, Delhon G, Lu Z, Carreno A, Vagnozzi A, Kutish GF, Rock DL (2005) Comparative genomics of foot-and-mouth disease virus. J Virol 79:6487–6504
Tully DC, Fares MA (2006) Unravelling selection shifts among foot-and-mouth disease virus (FMDV) serotypes. Evol Bioinform 2:237–251
Lewis-Rogers N, McClellan DA, Crandall KA (2008) The evolution of foot-and-mouth disease virus: impacts of recombination and selection. Infect Genet Evol 8:786–798
Cooke JN, Westover KM (2008) Serotype-specific differences in antigenic regions of foot-and-mouth disease virus (FMDV): a comprehensive statistical analysis. Infect Genet Evol 8:855–863
Valarcher J-F, Leforban Y, Rweyemamu M, Rweywmamu M, Roeder PL, Gerbier G, Mackay DKJ, Sumption KJ, Paton DJ, Knowles NJ (2008) Incursions of foot-and-mouth disease virus into Europe between 1985 and 2006. Transbound Emerg Dis 55:14–34
Sutmoller P, Barteling SS, Casas-Olascoaga R, Sumption KJ (2003) Control and eradication of foot-and-mouth disease. Virus Res 91:101–144
Acknowledgments
This study was supported by a grant from the Rural Development Administration of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yoon, S.H., Lee, KN., Park, JH. et al. Molecular epidemiology of foot-and-mouth disease virus serotypes A and O with emphasis on Korean isolates: temporal and spatial dynamics. Arch Virol 156, 817–826 (2011). https://doi.org/10.1007/s00705-011-0921-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-011-0921-3