Summary.
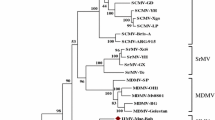
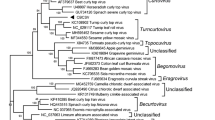
We have characterised two distinct geminiviruses that infect cucurbit cultivars in Vietnam. The genomes of both viruses consisted of two circular ssDNA components (DNA-A and DNA-B), with a genome arrangement and coding sequence typical of viruses in the Begomovirus genus in the family Geminiviridae. The sequence of DNA-A of one of the viruses was approximately 97% similar to Squash leaf curl virus-China (SLCV-Ch), for which a DNA-B has yet to be identified. We have named this virus Squash leaf curl virus-Vietnam (SLCV-Vn). The intergenic region of the SLCV-Vn DNA-B contained a 40 nt deletion between the putative AC1 TATA box and the stem loop. A second virus isolated from loofa in southern Vietnam was only 80% similar to SLCV-Vn over the complete DNA-A sequence, however the nucleotide sequence in the coat protein coding regions was 95% similar. We have named this virus Loofa yellow mosaic virus-Vietnam (LYMV-Vn). Other regions of the SLCV-Vn and LYMV-Vn genomes differed markedly, suggesting the coat protein coding region was recombinant. The DNA-B of both viruses were only 60% similar over the complete nucleotide sequence, although the encoded amino acid sequence of the BC1 gene was 90% identical.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Received September 4, 2002; accepted March 7, 2003 Published online June 5, 2003
Rights and permissions
About this article
Cite this article
Revill, P., Ha, C., Porchun, S. et al. The complete nucleotide sequence of two distinct geminiviruses infecting cucurbits in Vietnam. Arch Virol 148, 1523–1541 (2003). https://doi.org/10.1007/s00705-003-0109-6
Issue Date:
DOI: https://doi.org/10.1007/s00705-003-0109-6