Abstract
Parkinson’s disease (PD) etiology is heterogeneous, genetic, and multi-factorial, resulting in a varied disease from a mild slow progression to a more severe rapid progression. Prognostic information on the nature of the patient’s disease at diagnosis aids the physician in counseling patients on treatment options and life planning. In a cohort of PD patients from the PPMI study, the relative gene expression levels of SKP1A, UBE2K, ALDH1A1, PSMC4, HSPA8 and LAMB2 were measured in baseline blood samples by real-time quantitative PCR. At baseline PD patients were up to 2 years from diagnosis, H&Y scale ≤ 2 and PD treatment naïve. PD-Prediction algorithm comprised of ALDH1A1, LAMB2, UBE2K, SKP1A and age was created by logistic regression for predicting progression to ≤ 70% Modified Schwab and England Activities of Daily Living (S&E-ADL). In relation to patients negative for PD-Prediction (n = 180), patients positive (n = 30) for Cutoff-1 (at 82% specificity, 80.0% sensitivity) had positive hazard ratio (HR+) of 10.6 (95% CI, 2.2–50.1), and positive (n = 23) for Cutoff-2 (at 93% specificity, 47% sensitivity) had HR+ of 17.1 (95% CI, 3.2–89.9) to progress to ≤ 70% S&E-ADL within 3 years (P value < 0.0001). Likewise, patients positive for PD-Prediction Cutoff-1 (n = 49) had HR+ 4.3 (95% CI, 1.6–11.6) for faster time to H&Y 3 in relation to patients negative (n = 170) for PD-Prediction (P value = 0.0002). Our findings show an algorithm that seems to predict fast PD progression and may potentially be used as a tool to assist the physician in choosing an optimal treatment plan, improving the patient’s quality of life and overall health outcome.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the first description of Parkinson’s disease (PD) as a neurological disorder by James Parkinson (1755–1824) in 1817, many important discoveries have been made during the last two centuries (Braak et al. 2003; Singleton et al. 2017; Li and Le 2017; Van den Eaden et al. 2003). PD is the second most common neurodegenerative disorder (after Alzheimer dementia) affecting approximately 30 million people worldwide. PD prevalence is expected to double within the next two decades due to population aging and improved life expectancy (Rossi et al. 2018).
PD is a complex neurodegenerative disorder. The etiology of PD is heterogeneous, genetic, multi-factorial, resulting in a highly variable clinical course, spanning from a slow progressive, benign disease to a rapid progressive, disabling disease (Lawton et al. 2015; Birkmayer et al. 1979). The disease is characterized by rest tremor, rigidity, bradykinesia and postural instability (Fahn et al. 2011; Matsumoto 2000).
Symptomatic treatment can provide benefit for many years (Ahlskog 2000). When PD is first diagnosed the patient and their family require information on the nature of the patient’s disease. “To be or not to be a PD patient?” This is the first basic question patients ask.
The patients next ask “Doctor what is going to be my future. How will the disease progress? What are the treatment options and what can be expected concerning the quality of life? A good prediction of the rate of PD progression would impact on the capacity to choose the most beneficial treatment with the least number of side effects”.
PD can involve late or early cognitive and behavioral changes, early or late autonomic disturbances, and various manifestations of pain or sleep disturbances (Wickremaratchi et al. 2011; van Rooden et al. 2011).
Although symptomatic therapy can provide benefit for many years, PD is a progressive disorder that will eventually result in significant morbidity. Improvement in the ability to predict the rate of progression would impact on the ability to prescribe a more personalized treatment strategy considering specific patient needs, clinical phenotype and prognostic biomarkers (Titova and Chaudhuri 2017). The need for objective prognostic information was clearly expressed by PD patients who took part in the Muhammed Ali survey on medical care satisfaction, which found that patients were highly dissatisfied with the availability of prognostic information on their disease course (Dorsey et al. 2010). Objective biomarkers that predict the rate of PD progression would empower clinicians to better counsel patients regarding prognosis, treatment and life expectancy. Importantly, novel biomarkers can also aid in the discovery of new therapeutic targets (Singleton et al. 2013), increase the accuracy of patient enrollment and drug dosage in clinical trials, and detect patients who would benefit from novel therapies (Titova et al. 2017; Lang and Espay 2018).
Our group has investigated the prognosis ability of the blood expression of genes previously found to be associated with PD (Grünblatt et al. 2004, 2010). The Michael J Fox Foundation (MJFF) led Parkinson’s Progression Markers Initiative (PPMI) cohort (Marek et al. 2011) is a landmark study with the main aim of identifying biomarkers of PD progression. This 5-year observational, international, multi-center study comprised of ~ 400 recently diagnosed PD and ~ 200 healthy subjects, are followed longitudinally and comprehensively for biomarker assessment using standardized data acquisition protocols at twenty-one clinical sites. Several findings from the PPMI study (Marek et al. 2011) included the association of plasma ApoA1 with early age PD onset and greater motor severity (Swanson et al. 2015; Nazeri et al. 2015), prediction of disease severity by serum urate level combined with a genetic profile, prediction of cognitive impairment by measuring CSF β-amyloid 1–42 level in early PD (Fullard et al. 2016; Terrelonge et al. 2016) and prediction of cognitive decline by assessment of olfactory impairment.
We have shown in previous studies, that a gene expression profile was able to distinguish PD patients (early and advanced) from healthy controls and Alzheimer patients, as well as between early and advanced PD patients. Therefore, we decided to investigate the ability of the expression levels of the same genes to predict the rate of PD progression according to Modified Schwab and England Activities of Daily Living Scale (S&E-ADL)(Schwab and England 1969) and Hoehn and Yahr scale (Hoehn and Yahr 1967).
The tested genes were S-phase kinase-associated protein 1A (SKP1A), huntingtin interacting protein-2/ubiquitin conjugating enzyme E2K (HIP2/UBE2K), aldehyde dehydrogenase family 1 subfamily A1 (ALDH1A1), proteasome (prosome, macropain) 26 S subunit, ATPase 4 (PSMC4), heat shock 70-kDa protein 8 (HSPA8) and laminin subunit beta 2 (LAMB2). High score of the biomarker algorithm, comprised of low expression of ALDH1A1, SKP1A, UBE2K and high expression of LAMB2, predicted a faster rate of PD progression.
Methods
Study design and participants
This study includes a sub-cohort of newly diagnosed, untreated PD patients from The Parkinson’s Progression Marker Initiative (PPMI, https://www.ppmi-info.org/, accessed June 19, 2018), a multicenter international collaborative effort. At its core, PPMI is a longitudinal observational study that aims to identify markers of progression for PD.
Gene expression levels were screened in available baseline blood samples from PD patients (n = 279) whose mean age was 59.4 years (range is 30–83 years), 96 females and 183 males. The patients were not prescribed PD medicine at the time of baseline blood collection and were not expected to require PD medication within at least 6 months from baseline. The patients were enrolled in the study if they had at least two of the following: resting tremor, bradykinesia, rigidity (must have either resting tremor or bradykinesia); or either asymmetric resting tremor or asymmetric bradykinesia. Patients’ were Hoehn and Yahr (H&Y) stage I or II. Blood samples were taken. These patients progressed to H & Y stage III. Blood samples were taken again. Dopamine (DA) transporter deficit was confirmed (or for sites where DaTSCAN™ was not available, VMAT-2 PET scan was consistent with VMAT deficit). The study was approved by the institutional review board at each site, and participants provided written informed consent.
Blood collection, storage, RNA quality, quality assurance and analysis
Venous whole blood was collected from the patients in PAXgene tubes (PreAnalytiX, Switzerland) and messenger RNA (mRNA) was extracted by the MJFF led PPMI study team, according to manufacturer’s instructions and study protocol. RNA samples with a concentration of at least 20 ng/μL, and RNA integrity number (RIN) > 5.0 were selected for the study. Quality of RNA samples was assured by the MJFF PPMI study team. RNA samples (1 μg RNA, variable volume and concentration) were shipped frozen (− 70 °C). We were blinded to the clinical identity and time of collection of the RNA samples. Relative expression of SKP1A, HIP2/UBE2K, ALDH1A1, PSMC4, HSPA8 and LAMB2 were measured. Briefly, mRNA was reversed transcribed to cDNA. The cDNA was diluted to the testing concentration. Real-time quantitative PCR (RT-qPCR) was performed in a 96-well format, 25 μL total reaction volume using RT-qPCR master mix with SYBR Green florescence. A single PCR reaction was performed in each well. To avoid any operator biases or inaccuracies, a dedicated automatic pipetting system (Ezmate™ 400 by ARIS Biotech, USA) was used for mixing the reagents (primers, master mix, cDNA samples, positive controls, calibrator, and water) and dispensing them onto the PCR plates. In each plate, we tested four cDNA samples for the expression of six target genes and two reference genes. Measurement of the samples and calibrator was performed in duplicates. Each plate included negative control (no template), specific positive control of each gene (comprised of synthetic amplicon in a predefined concentration), calibrator/reference sample comprised of a mix of the synthetic amplicons to normalize plate to plate variations and cDNA samples of four patients.
Following plate preparation, the assay was performed using the StepOne™ PCR machine (Applied Biosystems, USA). Each run included amplification cycles and melt curve analysis for quality control. Relative expression of each of the target genes in each sample was calculated by the ddCT method using the StepOne™ dedicated software. All run parameters were exported to the study database. Quality assurance parameters were calculated and evaluated regarding precision between duplicates dCt SD (< 0.5), absolute Ct values of controls (positive, negative and calibrator) within a predefined range, and absence of multiple peaks in the melt curve. If the results did not match QC criteria, the sample was re-run. Identification of samples clinical data was done only after sending gene expression values to PPMI database. Data on PPMI are accessible at https://www.ppmi-info.org/access-data-specimens/download-data/.
Statistical methods
Statistical analysis was performed using MedCalc Statistical Software version 16.2.1 (MedCalc Software bvba, Ostend, Belgium; https://www.medcalc.org; 2016). Multivariable logistic regression analysis was performed for determining the composition of the predictive classifier algorithm (PD-Prediction). Variables were kept in the model if their P value was less than 0.15 and the variable improved the model performance. The discriminant performance and cutoff values of PD-Prediction were determined by receiver operating characteristic (ROC) curve. Time-to-event curves were drawn up by the Kaplan–Meier method with the log-rank test for comparisons, reporting also hazard ratios. Correlation was tested using Spearman’s rho rank test. Continuous values were compared by Mann–Whitney U. Uncertainty of results was expressed by 95% confidence intervals P values of less than 0.05 were considered to be statistically significant.
Results
PD-Prediction classifier construction
This analysis was performed on PD patients with recorded S&E-ADL scale from their third-year study visit (n = 241 of 279), mean age was 61.4 years (range was 33–82 years, 82 female and 159 male). The thirty-eight patients not included in the final analysis did not have recorded S&E-ADL scale from their third-year study visit. PD patients (n = 241) baseline gene expression blood levels of ALDH1a1 and PSMC4 correlated to 3 years post-baseline S&E-ADL. Baseline ddCT levels of ALDH1A1 (P value = 0.001, rho = − 0.208, 95% CI − 0.326 to − 0.0839) (Fig. 1a) and PSMC4 (P value = 0.006, rho = − 0.178, 95% CI − 0.298 to − 0.0522) (Fig. 1b) significantly decreased with higher S&E-ADL at 3 years after blood collection. A trend was observed for LAMB2 (P value = 0.075, rho = − 0.115, 95% CI − 0.238 to 0.0116).
Scatter plots showing correlation of 3 years post-baseline S&E-ADL with baseline blood expression levels of a ALDH1a (n = 241) and b PSMC4 (n = 239). S&E-ADL Modified Schwab & England (1969) Activities of Daily Living, ALDH1A1 Aldehyde dehydrogenase family 1 subfamily A1, PSMC4 proteasome (prosome, macropain) 26 S subunit, ATPase 4
Logistic Regression analyses were performed to construct a classifier comprised of the gene expression baseline values, age and gender for identifying PD patients who reached S&E-ADL ≤ 70% by 3 years post-baseline. Starting from Model 1 to Model 3 (Table 1), variables with P values < 0.15 were included in subsequent models until reaching a model with all variables having a P value < 0.15. The final model (Model 3) included ALDH1A1, LAMB2, SKP1A, UBE2K and age (overall model fit P value = 0.0004). Model 3 coefficients were used to build the classifier’s algorithm (PD-Prediction) to calculate the predictive probability (PP) values (Two patients not included due to missing gene expression data).
Receiver operator characteristics (ROC) curve (AUC = 0.852, 95% CI 0.800–0.894, P value < 0.0001) was used to determine two cutoff values for positivity of the PD-Prediction algorithm PP values (Fig. 2a). Cutoff-1 (0.0877 PP value) was at the Youden Index (J = max (sensitivity-c + specificity-c − 1, graphically J is the maximum vertical distance between the ROC curve and the diagonal) having a specificity of 81.7% (95% CI 76.0–86.5%) and sensitivity of 80.0% (95% CI 51.9–95.7%) and Cutoff-2 (0.180 PP value) having a high specificity of 93.3% (95% CI 89.2–96.2%) and moderate sensitivity of 46.7% (21.3–73.4%). As of interest, the AUC for age alone for detecting patients who reached S&E-ADL ≤ 70% by 3 years post-baseline was 0.731 (95% CI, 0.670–0.786, P value = 0.0005).
a ROC curve of PD-Prediction baseline values and age for detecting PD patients who reached S&E-ADL ≤ 70% at 3 years post-baseline. Diagnostic accuracy of Cutoff-1 and Cutoff-2 are shown. b–d are Kaplan–Meir curves of PD-Prediction for predicting PD patients who have faster time to ≤ 70% S&E-ADL in b is Cutoff-1, in c is Cutoff-2 and in d is both Cutoff-1 and Cutoff-2. ROC receiver operator characteristics, S&E-ADL Modified Schwab & England (1969) Activities of Daily Living, AUC area under the curve
PD-Prediction classifier time to event analysis for ≤ 70% S&E-ADL
Time-to-event analyses were performed for testing the prognostic ability of PD-Prediction. Time from baseline to the endpoint was calculated for each patient, i.e. if the patient reached ≤ 70% S&E-ADL by the 2nd year visit then time to the endpoint was calculated as time (days) from baseline to the 2nd year visit. For patients that did not reach the endpoint the last reported visit was included in the analysis. Patients who reached endpoint, but then in the subsequent visit improved to above the endpoint were not included in the analysis (n = 5) and 1 patient with missing time to event data was not included.
PD patients positive for PD-Prediction Cutoff-1 had a faster time to ≤ 70% S&E-ADL than patients who were negative for PD-Prediction (P value < 0.0001); the positive hazard ratio (HR+) of reaching the endpoint was 13.6 (95% CI, 4.1–45.4), negative hazard ratio (HR−) was 0.074 (95% CI, 0.022–0.25). Median time to the endpoint of patients who were positive for the marker was 1218 days (95% CI, 1127). Patients who were negative for the marker did not reach the median time to endpoint (Fig. 2b).
At the high specificity cutoff, Cutoff-2, PD patients positive for PD-Prediction had a faster time to ≤ 70% S&E-ADL than patients who were negative for PD-Prediction (P value < 0.0001); the HR+ of reaching the endpoint was 7.4 (95% CI, 1.4–38.5), HR− was 0.13 (95% CI, 0.026–0.70). Median time to the endpoint of patients who were positive for the classifier was 1218 days (95% CI, 1096). Patients who were negative for the classifier did not reach the median time to endpoint (Fig. 2c).
In a time-to-event analysis using both Cutoff-1 and Cutoff-2 the PD-Prediction classifier was able to predict PD patients who progressed faster to ≤ 70% S&E-ADL (P value < 0.0001). In relation to negative, the HR+ at Cutoff-2 was 17.1 (95% CI, 3.2–89.9) and the HR+ at Cutoff-1 was 10.6 (95% CI, 2.2–50.12), and the HR− was 0.059 (95% CI, 0.11–0.31) compared to Cutoff-2 and 0.095 (95% CI, 0.020–0.45) compared to Cutoff-1 (Fig. 2d).
PD-Prediction baseline levels detects PD patient who reach H&Y ≥ 3 at 3 years post-baseline
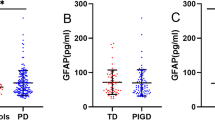
This analysis included PD patients with recorded H&Y scale from their third-year study visit (n = 220 of 279) and PD-Prediction baseline level (3 patients had missing gene expression data) mean age was 62 years (range was 33–82 years), 76 female and 144 male. PD-Prediction PP baseline values were significantly higher (P value = 0.0001) in the blood of PD patients who reached H&Y stage ≥ 3 (median 0.351 PP value, 95% CI 0.0621–0.174) than PD patients whose H&Y stage < 3 (median 0.029 PP value, 95% CI 0.020–0.036) at 3 years post-baseline (average days ± SD was 1106 ± 29 days). It is notable that the PD-Prediction levels of 11 of 162 PD patients with H&Y stage 2 were considered far-outliers in a box plot depicting each H&Y stage separately (Fig. 3a).
Time-to-event analysis was performed using Cutoff-1 and Cutoff-2. PD-Prediction significantly distinguished patients (P value = 0.0002) with faster time to H&Y stage ≥ 3 using Cutoff-1 (Fig. 3b) with an HR+ of 4.3 (95% CI, 1.6–11.6) and HR− of 0.24 (95% CI, 0.085–0.65). The higher cutoff (Cutoff-2) also predicted faster progression to H&Y stage ≥ 3 (P value = 0.02), with an HR+ of 3.1 (95% CI 0.70–13.6) and HR− 0.33 (95% CI 0.074–1.43). The patient groups did not reach the median time to H&Y stage ≥ 3 for both cutoffs. In the combined cutoff analysis, in relation to negative (P value = 0.001) the HR + at Cutoff-1 was 4.5 (95% CI, 1.01–19.9) and the HR+ at Cutoff-2 was 4.1 (95% CI, 1.2–14.1), and the HR− was 0.22 (95% CI, 0.05–0.99). Due to the crossing over of the confidence intervals and minimal contribution of the Cutoff-2 performance, only Cutoff-1 is taken in consideration for prediction of H&Y stage ≥ 3.
Discussion
Here we report on a gene expression classifier for the prediction of the rate of Parkinson’s disease progression defined by S&E-ADL ≤ 70% and H&Y ≥ 3. In a cohort of patients from the PPMI study, gene expression levels were measured in blood samples collected from PD patients who were up to 2 years from diagnosis, H&Y scale ≤ 2 and PD treatment naïve. The PD-Prediction algorithm was comprised of ALDH1A1, LAMB2, SKP1A, UBE2K and age. Using logistic regression, we created the multifactorial PD classifier by deciphering the weighted relationships between the baseline genes’ expression and 3 years-post disease progression (S&E-ADL). Since the classifier was created using a cross-sectional approach, we also demonstrated the classifier’s predictive ability along a “time interval” using Kaplan–Meier curves for S&E-ADL, and for H&Y, proving the robustness of the genetic signature for different PD endpoints. It should be emphasized that this is the first study demonstrating the predictive ability of this specific combination of the expressed genes and the accumulation of further patient data may lead to a better-defined algorithm and cutoffs.
These results extend findings depicting the difference between early and advanced PD in the predicted probability values of a five-gene expression panel, including ALDH1A1, SKP1A, UBE2K, in addition to PSMC4 and HSPA8, which were not included in the final algorithm of the current study. The main focus of this previous study was the diagnostic performance of the five-gene molecular signature differentiating PD from healthy controls and Alzheimer patients (Molochnikov et al. 2012).
In another independent study, a four-gene panel comprised of ALDH1A1 and LAMB2 (together with PSMA2 and HIST1H3E) was found to differentiate PD from healthy controls and Alzheimer patients (Grünblatt et al. 2010). Biomarkers from these studies were chosen from genes that were differentially expressed in post-mortem substantia nigra of sporadic parkinsonian patients compared to non-diseases controls.
Peripheral blood gene expression analysis, reflecting genetic, physiological and environmental changes is a rational approach for the development of biomarkers for a disease with multi-factorial etiology such as PD. Another research group found an association between the early development of postural instability in PD patients and the expression of genes related to nucleic acid metabolism, mitochondria and immune response (Pinho et al. 2016). The PD-Prediction classifier reported here is comprised of the expression of genes coding for proteins involved in the metabolism of toxins, the ubiquitin–proteasome protein degradation system and a component of the basal lamina. In the brain, ALDH1A1 participates in the metabolism of DA by degrading 3,4-dihydroxyphenylacetaldehyde (DOPAL), an aldehyde metabolite of DA generated by the action of monoamine oxidase (MAO)-A and -B (Anderson et al. 2011) DOPAL destroys nigrostriatal DA terminals and contributes to the profound putamen DA deficiency that characterizes Parkinson’s disease (Hirsch et al. 2013). LAMB2 is a subunit of laminin, the major non-collagenous component of the basal lamina. Laminin injection, a neurite-outgrowth promoting factor, was reported as improving neurochemical and behavioral consequences of Parkinsonism in a rat PD model (Sever et al. 2016). Both SKP1A and UBE2K are involved in the ubiquitin–proteasome protein degradation system. Dysfunction of the ubiquitin–proteasome system is one of the major mechanisms of Parkinson’s disease pathogenesis. SKP1A is a component of the E3 ubiquitin ligase protein, found to protect against parkinsonism neurotoxin induction (Fishman-Jakob et al. 2009). UBE2K accepts ubiquitin from E1 complex and catalyzes its link to proteins. UBE2K was demonstrated as being reduced in the striatum of brain in mice with the MPTP induced early symptomatic stages (Filatova et al. 2014).
Our present blood study demonstrates a potential method for differentiating PD patients with slow disease course from patients with fast disease course that can be used as an aid in therapeutic decisions. Until now, the treatment of PD is not curative, but only palliative accompanied by a myriad of side-effects that are difficult to manage. Among the main drugs used to treat PD are levodopa, dopamine agonists, amantadine, MAO B inhibitors (selegiline, rasagiline, safinamide), catechol-O-methyl transferase (COMT) inhibitors (entacapone, tolcapone, obicapone) and anticholinergics (Poewe and Mahlknecht 2020). Surgical approaches such as deep brain stimulation (DBS) is another treatment option (Mitchell and Ostrem 2020). Though the high cost of DBS is an important issue (Becerra et al. 2016), the treatment is cost-effective for advanced PD. Novel therapies under research will most likely further extend the physician’s treatment choices for PD patients (Foltynie and Athauda 2020). Exenatide, a glucagon-like peptide-1 agonist for treating insulin resistance in type 2 diabetes, delayed Parkinson’s disease motor and cognitive ability in an early proof of concept trial (Athauda et al. 2017). Anti-α-synuclein immunotherapy clinical trials are being initiated after pre-clinical trials demonstrated therapeutic promise in treating PD (George and Brundin 2015; Wang et al. 2019; Brys et al. 2019).
It has been almost four decades since authors from our research alliance have published findings pointing out the heterogeneous disease course of PD patients who had either a slow or fast disease progression. In the last two decades, the growing number of clinical trials for new PD therapies and potential treatment choices for PD emphasizes the need of minimally invasive tests for PD progression prognosis. The creation of a gene expression classifier for the prediction of PD rate of progression can provide additional information improving the design and enrollment of therapeutic clinical trials and aid the physician in achieving a more personalized approach in treatment decisions and patient lifestyle counselling. Therefore, ensuring the Parkinson’s disease patient receives the best possible treatment needed to improve their health outcome, while minimizing unnecessary and sometimes debilitating treatment side-effects.
Conclusion
An algorithm comprised of ALDH1A1, LAMB2, UBE2K, SKP1A and age is predictive of faster decrease S&E-ADL ≤ 70% and H&Y ≥ 3. These findings should be confirmed in another independent cohort.
References
Ahlskog E (2000) Approach to the patient with a movement disorder: basic principles of neurologic diagnosis. From Parkinson’s disease and movement disorders : diagnosis and treatment guidelines for the practicing physician (edited by Adler CH and Ahlskog JE). Humana Press, Totowa, New Jersey, pp 3–34
Anderson DW, Schray RC, Duester G, Schneider JS (2011) Functional significance of aldehyde dehydrogenase ALDH1A1 to the nigrostriatal dopamine system. Brain Res 1408:81–87
Athauda D, Maclagan K, Skene SS, Bajwa-Joseph M, Letchford D, Chowdhury K, Hibbert S, Budnik N, Zampedri L, Dickson J, Li Y, Aviles-Olmos I, Warner TT, Limousin P, Lees AJ, Greig NH, Teb S, Foltynie T (2017) Exenatide once weekly versus placebo in Parkinson’s disease: a randomised, double-blind, placebo-controlled trial. Lancet 390:1664–1675
Becerra JE, Zorro O, Ruiz-Gaviria R, Castañeda-Cardona C, Otálora-Esteban M, Henao S, Navarrete S, Acevedo JC, Rosselli D (2016) Economic analysis of deep brain stimulation in Parkinson disease: systematic review of the literature. World Neurosurg 93:44–49
Birkmayer W, Riederer P, Youdim MBH (1979) Distinction between benign and malignant type of Parkinson’s disease. Clin Neurol Neurosurg 81:158–164
Braak H, Rub U, Gai WP, Del Tredici K (2003) Idiopathic Parkinson’s disease: possible routes by which vulnerable neurons types may be subject to neuroinvasion by an unknown pathogen. J Neural Trans 110:510–537
Brys M, Fanning L, Hung S, Ellenbogen A, Penner N, Yang M, Welch M, Koenig E, David E, Fox T, Makh S, Aldred J, Goodman I, Pepinsky B, Liu Y, Graham D, Weihofen A, Cedarbaum JM (2019) Randomized phase I clinical trial of anti-α-synuclein antibody BIIB054. Mov Disord 34(8):1154–1163. https://doi.org/10.1002/mds.27738 (Epub 17 Jun 2019)
Dorsey TS, Voss DR, Shprecher LM, Deuel CA, Beck IF, Gardiner MA, Coles RS, Burns FJ, Marshall KMB (2010) A U.S. survey of patients with Parkinson’s disease: satisfaction with medical care and support groups. Mov Disord 25:2128–2135
Fahn S, Jankovic J, Hallet M (2011) Principles and practice of movement disorders. Elsevier Publication, Amsterdam
Filatova EV, Shadrina MI, Alieva AKh, Kolacheva AA, Slominsky PA, Ugrumov MV (2014) Expression analysis of genes of ubiquitin-proteasome protein degradation system in MPTP-induced mice models of early stages of Parkinson’s disease. Dokl Biochem Biophys 456:116–118
Fishman-Jakob T, Reznichenko L, Youdim MBH, Mandel SA (2009) A sporadic Parkinson disease model via silencing of the ubiquitine-proteasome/E3 ligase component SKP1A. J Biol Chem 284:32835–32932
Foltynie T, Athauda D (2020) Repurposing anti-diabetic drugs for the treatment of Parkinson’s disease: rationale and clinical experience. Prog Brain Res 252:493–523. https://doi.org/10.1016/bs.pbr.2019.10.008 (Epub 27 Nov 2019)
Fullard ME, Tran B, Xie S, Toledo JB, Scordia C, Linder C, Purri R, Weintraub D, Duda JE, Chahine LM, Morley JF (2016) Olfactory impairment predicts cognitive decline in early Parkinson’s disease. Parkinsonism Relat Disord 25:45–51
George S, Brundin P (2015) Immunotherapy in Parkinson’s disease: micromanaging alpha-synuclein aggregation. J Parkinsons Dis 5:213–222
Grünblatt E, Mandel S, Jacob-Hirsch J, Zeligson S, Amariglo N, Rechavi G, Li J, Ravid R, Roggendorf W, Riederer P, Youdim MB (2004) Gene expression profiling of parkinsonian substantia nigra pars compacta; alterations in ubiquitin-proteasome, heat shock protein, iron and oxidative stress regulated proteins, cell adhesion/cellular matrix and vesicle trafficking genes. J Neural Transm 111:1543–1573
Grünblatt E, Zehetmayer S, Jacob CP, Müller T, Jost WH, Riederer P (2010) Pilot study: peripheral biomarkers for diagnosing sporadic Parkinson’s disease. J Neural Transm 117:1387–1393
Hirsch E, Jenners P, Przedborski S (2013) Pathogenesis of Parkinson’s disease. Mov Dis 28(1):24–30
Hoehn M, Yahr M (1967) Parkinsonism: onset, progression and mortality. Neurology 17:427–442
Lang AE, Espay AJ (2018) Disease modification in Parkinson’s disease: current approaches, challenges, and future considerations. Mov Disord 33(2018):660–677
Lawton M, Baig F, Rolinski M, Ruffman C, Nithi K, Hu MT (2015) Parkinson’s disease subtypes in the Oxford Parkinson disease centre (OPDC) discovery cohort. J Parkinsons Dis 5:269–279
Li S, Le W (2017) Biomarker discovery in Parkinson’s disease: present challenges and future opportunities. Neurosci Bull 33(5):481–482
Marek K, Jennings D, Lasch S et al (2011) The Parkinson progression marker initiative (PPMI). Prog Neurobiol 95:629–635. https://doi.org/10.1016/j.pneurobio.2011.09.005
Matsumoto JY (2000) Tremor disorders overview. Chapter 21. From Adler CH, Ashlskog JE Editors (2000). From Parkinson’s disease and movement disorders. Diagnosis and treatment guidelines for the practicing physician (edited by Adler CH and Ashlkog JE). Humana press, Totowa, New Jersey
Mitchell KT, Ostrem JL (2020) Surgical treatment of Parkinson disease. Neurol Clin 38(2):293–307. https://doi.org/10.1016/j.ncl.2020.01.001 (Epub 22 Feb 2020)
Molochnikov L, Rabey JM, Dobronevsky E, Bonucelli U, Ceravolo R, Frosini D, Grünblatt E, Riederer P, Jacob C, Aharon-Peretz J, Bashenko Y, Youdim MB, Mandel SA (2012) A molecular signature in blood identifies early Parkinson’s disease. Mol Neurodegener 31:7–26
Nazeri A, Roostaei T, Sadaghiani S, Chakravarty MM, Eberly S, Lang AE, Voineskos AN (2015) Genome-wide variant by serum urate interaction in Parkinson’s disease. Ann Neurol 78:731–741
Pinho R, Guedes LC, Soreq L, Lobo PP, Mestre T, Coelho M, Rosa MM, Gonçalves N, Wales P, Mendes T, Gerhardt E, Fahlbusch C, Bonifati V, Bonin M, Miltenberger-Miltényi G, Borovecki F, Soreq H, Ferreira JJ, Outeiro F (2016) Gene expression differences in peripheral blood of Parkinson’s disease patients with distinct progression profiles. PLoS ONE 11(6):e0157852
Poewe W, Mahlknecht P (2020) Pharmacologic treatment of motor symptoms associated with Parkinson disease. Neurol Clin 38(2):255–267. https://doi.org/10.1016/j.ncl.2019.12.002 (Epub 20 Feb 2020)
Rossi A, Berger K, Chen H et al (2018) Projection of the alence of PD in the coming decades. Revis Mov Disord 33(1):156–161
Schwab RS, England AC (1969) Projection technique for evaluating surgery in Parkinson’s disease. In: Gillingham FJ, Donaldson MC (eds) Third symposium on Parkinson’s disease. Livingston, Edinburgh, pp 152–157
Sever M, Turkyilmaz M, Sevinc C, Cakir A, Ocalan B, Cansev M, Guler MO, Tekinay AB (2016) Regenerative effects of peptide nanofibers in an experimental model of Parkinson’s disease. Acta Biomater 46:79–90
Singleton AB, Farrer MJ, Bonifati V (2013) The genetics of Parkinson’s disease: progress and therapeutic implications. Mov Disord 28:14–23
Singleton AB, Hardy JA, Gasser T (2017) The birth of the modern era of Parkinson’s disease genetics. J Parkinsons Dis 7(Suppl 1):58–59
Swanson CR, Berlyand Y, Xie SX, Alcalay RN, Chahine LM, Chen-Plotkin AS (2015) Plasma apolipoprotein A1 associates with age at onset and motor severity in early Parkinson’s disease patients. Mov Disord 30:1648–1656
Terrelonge M, Marder KS, Weintraub D, Alcalay RN (2016) CSF β-amyloid 1–42 predicts progression to cognitive impairment in newly diagnosed Parkinson disease. J Mol Neurosci 58:88–92
Titova N, Chaudhuri KR (2017) Personalized medicine in Parkinson’s disease: time to be precise. Int Rev Neurobiol 134:33–54
Titova N, Qamar MA, Chaudhuri KR (2017) Biomarkers of Parkinson’s disease: an introduction. Int Rev Neurobiol 132:183–196
van den Eaden S, Tanner C, Bernstein A et al (2003) Incidence of Parkinson’s disease: variations by age, gender and race/ethnicity. Am J Epidemiol 157:1015–1022
van Rooden SM, Colas F, Martínez-Martín P, Visser M, Verbaan D, Marinus J, Chaudhuri RK, Kok JN, van Hilten JJ (2011) Clinical subtypes of Parkinson’s disease. Mov Disord 26(1):51–58
Wang Z, Gao G, Duan C, Yang H (2019) Progress of immunotherapy of anti-α-synuclein in Parkinson’s disease. Biomed Pharmacother 115:108843. https://doi.org/10.1016/j.biopha.2019.108843 (Epub 2 May 2019)
Wickremaratchi MM, Knipe MD, Sastry BS, Morgan E, Jones A, Salmon R, Weiser R, Moran M, Davies D, Ebenezer L, Raha S, Robertson NP, Butler CC, Ben-Shlomo Y, Morris HR (2011) The motor phenotype of Parkinson’s disease in relation to age at onset. Mov Disord 26:457–463
Acknowledgements
The Michael J Fox Foundation supported this work (Gift ID: 10866). The clinical data and blood samples were from the Parkinson’s Progression Marker Initiative (PPMI) study, which is supported by the Michael J Fox Foundation for Parkinson’s Research and is co-funded by the Michael J Fox Foundation for Parkinson’s Research, Abbvie, Avid Radiopharmaceuticals, Biogen Idec, Bristol-Myers Squibb, Covance, Eli Lilly and Co, F Hoffman-La Roche, GE Healthcare, Genentech, GlaxoSmithKline, Lundbeck, Merck, MesoScale, Piramal, Pfizer, and UCB. Data used in the preparation of this article were obtained from the Parkinson’s Progression Markers Initiative (PPMI) database (https://www.ppmi-info.org/data). For up-to-date information on the study, https://www.ppmi-info.org.
Funding
PPMI is sponsored by the Michael J. Fox Foundation for Parkinson’s Research (MJFF) and is co-funded by MJFF, Abbvie, Avid Radiopharmaceuticals, Biogen Idec, Bristol-Myers Squibb, Covance, Eli Lilly & Co., F. Hoffman-La Roche, Ltd., GE Healthcare, Genentech, GlaxoSmithKline, Lundbeck, Merck, MesoScale, Piramal, Pfizer and UCB.
Author information
Authors and Affiliations
Contributions
JMR is a senior adviser and shareholder of BioShai Ltd. JSY was employed by Bioshai and had received share options of BioShai Ltd. ND was employed by Bioshai and had received share options of BioShai Ltd. DM was an employee of Bioshai and had received share options of BioShai Ltd. MBHY and PR are advisors and shareholders of BioShai Ltd. The patent—USA provisional 62/583,132, related to this work was assigned to BioShai Ltd.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rabey, J.M., Yarden, J., Dotan, N. et al. Creation of a gene expression classifier for predicting Parkinson’s disease rate of progression. J Neural Transm 127, 755–762 (2020). https://doi.org/10.1007/s00702-020-02194-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-020-02194-y