Abstract
Iran is reported to be a center of diversity for Juglans regia and wild walnut trees are found in virtually every corner of the country. Thus Iran is considered a rich natural pool of walnut germplasm for developing improved genotypes. Kerman province is the most important Iranian province for walnut culture and has the largest walnut plantations in Iran. Genetic structure and gene flow were analyzed in six walnut populations of this province using 17 microsatellite loci. The number of alleles per locus ranged from 4 to 11, with a total of 147 alleles and 5.16 effective alleles per locus. The polymorphism information content for the loci ranged from 0.56 to 0.82. The expected heterozygosity (H e) for the populations ranged from 0.65 to 0.87. There were differences between populations regarding the number of effective alleles and Shannon’s information index (I). In all populations, observed heterozygosity (H o) was lower than expected, but diversity within the populations was high (I = 1.5) and many of the private alleles were present at relatively high frequency. The average F st value was 0.08. The level of gene flow based on F st was high (N m = 3.01), which meant that the high level of genetic diversity maintained within each population was less susceptible to genetic drift. The geographical proximity of the populations was not correlated with their level of genetic relatedness. These results imply the high potential of walnut populations of Kerman province for breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Persian walnut, Juglans regia L., is the most economically important member of its genus. It is cultivated for its timber and edible nuts throughout temperate regions of the world (Vahdati 2000; Bayazit et al. 2007). Persian walnut was probably domesticated in the region of Iran and Afghanistan and subsequently introduced to China, Russia and Eastern Europe by ancient tribes (Bayazit et al. 2007). According to the FAO (2011), Persian walnut grows on 64,000 ha in Iran, producing 165,508 tons of nuts in shell, ranking second globally. Kerman province is the leading area for walnut production in Iran, with about 17,095 ha under cultivation. This province, with varied eco-geographical regions, is one of the major centers for Persian walnut diversity, and walnut populations are widely scattered in this region (Vahdati 2000).
Genetic variation among walnut populations has been studied using a large number of marker systems, including morphological indices (Malvolti et al. 1994), isozymes (Arulsekar et al. 1986; Malvolti et al. 1993; Solar et al. 1994; Fornari et al. 1999), restriction fragment length polymorphisms (RFLPs) (Fjellstrom et al. 1994; Fornari et al. 2001), randomly amplified polymorphic DNA (RAPD) (Nicese et al. 1998), amplified fragment length polymorphisms (AFLPs) (Bayazit et al. 2007) and inter-simple sequence repeats (ISSRs) (Potter et al. 2002). These studies reflect the need to identify cultivars accurately (genetic fingerprints) and verify paternity and genealogy. Woeste et al. (2002) developed a panel of 30 nuclear microsatellites (SSR) for a wide range of genetic investigations in Juglans, including clonal identification (Dangl et al. 2005; Foroni et al. 2007), a broad-scale study of the genetic structure of J. nigra populations in the Central Hardwood Region of the United States of America (Victory et al. 2006) and the identification of hybridogenic walnut plants (Pollegioni et al. 2009). In a recent study, Gunn et al. (2010) evaluated 220 walnut trees from six Tibetan villages in China using 14 SSR markers.
Microsatellites or simple sequence repeats (SSRs) have proved to be suitable markers for variety characterization, and a few have already been developed for J. regia (Dangl et al. 2005; Foroni et al. 2007; Pollegioni et al. 2011). Microsatellites are ideal markers for characterizing relationships among individuals because of the co-dominant inheritance, hyper-variability, high information content and the reproducibility of genotyping results among laboratories (Foroni et al. 2006). The published data on genetic diversity and structure for Persian walnut are mostly from breeding populations and European populations, and there are no data for the species in either its native range or in the region of its domestication (Malvolti et al. 1993, 2010; Fjellstrom and Parfitt 1994; Nicese et al. 1998; Fornari et al. 2001; Dangl et al. 2005).
The objective of the present study was to use microsatellite markers to determine genetic structure and gene flow among six landraces of J. regia from different valleys of Iran’s Kerman province, which is home to one of the world’s largest populations of Persian walnut.
Materials and methods
Plant materials
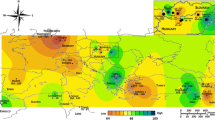
Six populations of walnut, namely Hanza (HA, lat. 57°11′N), Gogher (GO, lat. 56°24′N), Kiskan (KI, lat. 56°38′N), Rayen (RI, lat. 57°19′N), Bidkhan (BI, lat. 56°30′N) and Bezenjan (BE, lat. 56°41′N), located in different valleys of Kerman province in southeastern Iran, were selected for study (Fig. 1). The sampled populations inhabit disjunctive mountainous areas with a narrow geographic range of longitude (29°14′–29°37′E). Populations consisted of old walnut trees from open pollinated seedlings (70- to 150-year-old trees with terminal or lateral bearing habit). Summary of morphological and phenological traits in studied populations are shown in Table 1. Plants within 15 km of each other were considered to belong to the same population (Malvolti et al. 1993). The number of samples collected per site ranged from 6 to 18 based on the density of the plants found at each site. In total, 66 genotypes in six populations were analyzed.
The trees cultivated in these populations represent local populations (seedlings) that were randomly planted by humans or birds (mostly crows) and now grow across wide areas at a spacing of about 4–8 m. Moreover, the number of individuals collected is not same in each population because of the different plant density found in each valley. This is the reason of the low number of trees sampled in some populations. The sampled populations, located in different valleys of Kerman province, were separated by 14–100 km (approximately 45 km in average).
DNA extraction
For each mother tree, at least six young leaves were collected in summer and used for DNA extractions. A 150- to 200-mg sample was ground in 2-mL Eppendorf tubes with 1,800 mL of extraction buffer (2 % CTAB, 100 mM Tris, 1.4 M NaCl, and 20 mM EDTA, pH 8.3). Then DNA was extracted following the CTAB method (Doyle and Doyle 1987). DNA quantity and concentrations were determined spectrophotometrically at 260 and 280 nm and by electrophoresis on 0.8 % (w/v) agarose gel. The agarose gel was stained by ethidium bromide and visualized with UV light.
SSR primers and PCR amplification
Seventeen SSR paired primers with the prefix WGA designed from the sequence of clones from an enriched (GA/CA)n library of black walnut (J. nigra) (Woeste et al. 2002; Dangl et al. 2005), were used to amplify genomic DNA of each individual plant from the six populations in order to identify polymorphic SSR loci (Table 2). Polymerase chain reaction (PCR) amplifications were performed in 20 µL reaction solution containing 2 µL 10× buffer (100 mM Tris–HCl, pH 8.0 and 500 mM KCl), 2 mM MgCl2, 0.2 mM deoxyribonucleoside triphosphate (dNTP), 50 ng of each forward and reverse primer, 1 unit Taq polymerase (Cinnagene,Tehran, Iran) and 60 ng template DNA. Reactions were performed in a Biorad MJ Mini thermocycler according to the following procedure: an initial denaturation step at 94 °C for 5 min, followed by 35 cycles at 94 °C for 45 s, an annealing step at 57 °C for 40 s and an extension at 72 °C for 1 min, with a final extension step at 72 °C for 10 min; the final product was kept at 4 °C. A negative control reaction without DNA template was included in each amplification. The product was run on a 1-mm-thick, 6 % non-denaturing polyacrylamide gel. Gels were pre-run at 1,500 V for 30 min. The samples were loaded and run at 1,500 V for about 1 h. Fragments were visualized by silver staining (Bassam et al. 1991). Each gel had a 100–1000-bp DNA ladder (Cinnagene, Tehran, Iran) and a standard sample to estimate molecular weight and control gel-to-gel variation. In all cases, PCR reactions were performed at least twice to ensure that allele sizes were consistent.
Evaluation of polymorphisms and statistical analysis
Polymorphic alleles were scored as AA, BB, CC, etc. for homozygous individuals and AB, AC, BC, etc. for heterozygous individuals. For each locus, genotypes showing one and two bands were scored as homozygous and heterozygous, respectively. Observed heterozygosity (H o) was calculated as the ratio between the number of heterozygous individuals and the total number of genotypes per locus. Expected heterozygosity (H e) was estimated according to the formula \( H_{\text{e}} = 1 - \varSigma p_{i}^{2} \), where p i is the frequency of the ith allele for the studied locus (Levene 1949). Population structure was analyzed using Wright’s (Wright 1978), F-statistics (F IT, F IS and F ST), where F IT represents the overall inbreeding coefficient, F IS represents the level of inbreeding due to nonrandom mating within populations, and F ST represents the population subdivision. Polymorphism information content PIC and I values are frequently used to estimate the genetic diversity of genotypes (Shannon and Weaver 1949). The POPGENE (ver. 1.31) software (Yeh et al. 1999), was used to determine percentage of polymorphic loci, observed number of alleles (N a), effective number of alleles (N e), Shannon’s information index (I), Wright’s (Wright 1978), F-statistics (F IT, F IS and F ST) and Nei’s (Nei 1978). The H e, H o and I values for each locus and population were compared separately by analysis of variance using a completely randomized design by SAS software (version 9.0, SAS Institute, Inc., Cary, North Carolina, USA). The Neighbor-Joining dendrogram using genetic distance measures on SSR markers data among the populations was computed with NTSYSpc (ver 2.02) (Rohlf 1998) software.
Results and discussion
Genetic diversity
All the tested primers produced satisfactory amplification products and were multiallelic (Table 2). The total number of alleles per locus ranged from 4 (in WGA225) to 11 (in WGA001, WGA009, WGA071, WGA069 and WGA202), with a total of 147 alleles for all loci.
The average N a and N e per locus were 8.65 and 5.16, respectively (Table 3). Allele numbers were relatively high compared to the levels of variability detected in 15 European walnut populations (Fornari et al. 1999), 48 J. regia cultivars (Dangl et al. 2005) and five J. regia populations from central and southwestern China (Wang and Pei 2008). Although descriptive data for WGA69 included in Table 3, this locus showed significant deviations from the other loci and was not included in subsequent analyses.
The peculiar results of WGA069 for Juglans have been observed in several studies. Lewontin and Krakauer (1973) and Luikart et al. (2003) observed that selection and mutation have locus-specific effects while genetic drift and gene flow act at a genome-wide scale. By analyzing the number of alleles per locus and the large number of alleles in common between J. nigra and J. regia, Pollegioni et al. (2009) postulated a low mutation rate at locus WGA69. Several studies report that the interruption of perfect microsatellites is related to DNA stability in the region (Taylor et al. 1999). These authors suggested that the purity of a repeat region influences its mutation rate and consequently, the level of polymorphism in SSR loci. Interrupted microsatellites, such as WGA69, appear to have lower mutation rates than pure microsatellites. As reported by Cornuet and Luikart (1996), this feature makes WGA69 a useful marker for detecting a bottleneck. Storz (2005) also indicated that the risk of detecting false positives is high using Beaumont and Nichols (1996) because bottlenecks can produce effects similar to natural selection. Pollegioni et al. (2011) in study of genetic structure of Italian walnut by SSR markers suggested that the atypical behavior of WGA69 may be a consequence of its low rate of mutation and a human-mediated domestication bottleneck.
The alleles present with the highest frequency (0.50) were the 190- and 300-bp alleles at the WGA032 and WGA331 loci, respectively, whereas the alleles with the lowest frequency (0.07) were the 260- and 270-bp alleles at the WGA202 locus (data not shown). The H e values ranged from 0.65 for WGA032 to 0.87 for WGA071 and WGA118, with an average of 0.79 for all loci (Table 3). The analysis of variance showed that H e differed significantly among loci (F = 3.45, P ≤ 0.01). The H o values ranged from 0.00 for WGA027 and WGA071 to 0.85 for WGA069, with an average of 0.23 for all loci. The H o values differed significantly among loci (F = 18.64, P ≤ 0.01). The difference between H e and H o was high that it can be result from selection of the best genotypes by growers and increase inbreeding coefficient in populations. The average number of alleles per locus was 8.05, much higher than the 1.3, 3.9 and 5.5 detected in J. regia with RAPDs (Nicese et al. 1998), ISSRs (Potter et al. 2002), and SSRs (Foroni et al. 2007), respectively. Dangl et al. (2005) also observed 3–8 alleles per locus in their genetic analysis and cultivar identification of walnut using 14 SSR markers. In the present study, the number of alleles for WGA032 and WGA279 was similar to results published by Foroni et al. (2006) and Dangl et al. (2005), respectively.
In the present study, at the level of populations, the PIC and I of the populations averaged 72 % and 1.79, respectively. PIC values provide an estimate of the discriminatory power of any locus by considering the number of alleles per locus and their relative frequencies in the population (i.e. gene diversity values, essentially) (Rongwen et al. 1995). The PIC values for the populations in the present study ranged from 0.56 to 0.82 and classified six loci as informative markers (PIC >0.5) and eleven loci as suitable for mapping (PIC >0.7) (Table 3). These results indicate that all of the markers could contribute substantial information to walnut genetics and breeding research. The results also show that the walnut populations of Kerman province have relatively high levels of neutral genetic diversity. Pollegioni et al. (2011) observed that except WGA004 (0.355) and WGA331 (0.382), all markers had PIC >0.50. The I values for loci were highly variable: they averaged 1.79, with the maximum (2.22) for WGA071 and the minimum (1.18) for WGA225. The PIC and I values differed significantly among loci and populations (data not shown).
On the other hand, gene flow is the movement of genes within and between populations (Grant 1991). In this study, the levels of gene flow (N m) ranged from 1.83 to 6.10 with a mean of 3.03 for all loci which can be calculated indirectly by F ST (Table 3), which meant that the high level of genetic diversity maintained within each population was less susceptible to genetic drift. Seeds and pollen are the two main vectors of gene flow for seed plants (Hamrick et al. 1991). Seeds of wild walnut trees spread mainly by gravity and by animal movement over short distances. In long distance, seed dispersal by animal movement is rare, but dispersal of pollen is likely a main way of gene flow between populations. Another possibility is seed movement by growers in order to selection of the best genotypes or increasing cultivation areas. Dispersal of pollen is likely the main mechanism of gene flow among the studied populations, a conclusion that was also reached in other studies of similar large-seeded species (Victory et al. 2006). The high amount of gene flow (N m = 3.03, Table 3) would reduce the disjunction between these populations. In study of genetic variation in Korean populations of J. sinensis, gene flow among populations was reported relatively low (N m = 1.80, Lee and Lee 1997). The number and frequency of private alleles can provide insight into levels of gene flow among populations. The present study found that, among the Persian walnuts in Kerman province, 14 % of all detected alleles were private (19/136 alleles; locus WGA69 was excluded from this analysis), a figure that is not a great departure from the number of private alleles observed in a large study of J. nigra populations in the U.S. (about 10 % private; Victory et al. 2006) (Data not shown). The HA, GO and KI populations had the largest number of private alleles, with six, five and six, respectively (data not shown); RI and BE had no private alleles, but this was probably because of our small sample size from this population. It was surprising that several of the private alleles were present at relatively high frequencies, in three cases exceeding 10 %. Because the detection of rare and private alleles is highly dependent on sample sizes, which were small in this study, few conclusions can be drawn from these data. F ST statistics at the loci assayed ranged between 0.04 and 0.12, with the average 0.08. Positive values of F ST are usually interpreted as indicators of inbreeding and in general of assertive mating. The observed negative figures could be ascribed to the combination of a high degree of outcrossing and to some mechanism of heterozygote advantage (Hartel and Clark 2007). The positive values of F ST were in agreement with high level of gene flow (N m) between the populations. Because seed movement would increase the similarity of the genotypes, then sib-mating or inbreeding would enlarge in the populations. In wild populations of J. sinensis in Korea, F ST was 0.122 (Lee and Lee 1997); among J. nigra populations in the U.S., F ST was 0.017 (Victory et al. 2006); and among Asian and European populations of J. regia, F ST was 0.108 (Fornari et al. 2001).
There were differences between the six studied J. regia populations regarding H e (genetic diversity), N e and I (Table 4). The average I value for all populations was 1.49, with the maximum (1.67) for the HA population and the minimum (1.21) for the RI population. The HA (N = 18) and KI (N = 14) populations also had the highest number of alleles (both N a and N e) and RI population (N = 6) had the lowest, reflecting correlation allele number with the sample size of each population. The mean number of alleles per polymorphic locus (N a = 5.69) was higher than the typical values reported for widespread plant species (N a = 3.70) and European and Asian walnut populations (N a = 2.27) (10, 16). In all the studied Iranian populations, H o was low, but diversity within the populations was high (I = 1.5). The maximum H o was found in the KI and BI populations (H o = 0.25) (Table 4), and there were significant differences among populations (F = 3.45, P ≤ 0.01); the average H o in the samples in the present study was 0.23, lower than the averages in the J. regia populations studied by (H o = 0.50, Malvolti et al. 1994), (H o = 0.698, Foroni et al. 2007), (H o = 0.585, Foroni et al. 2006) and Dangl et al. (2005) (H o = 0.597, Pollegioni et al. 2011) and in the J. nigra populations studied by (H o = 0.781, Victory et al. 2006) based on SSR marker data. The H o observed in the present study was closer to that of European and Asian populations of J. regia (H o = 0.39, Fornari et al. 2001) and populations of J. nigra (H o = 0.31; Hamrick et al. 1991) based on isozyme markers. The H e ranged from 0.63 (RI) to 0.76 (HA). In study of Pollegioni et al. (2011) the value of H e in populations was 0.644. The levels of H o was significantly lower than H e. This result is probably due to sample size, Wahlund effect or inbreeding.
Population variation
As mentioned previously, the F ST estimates for each locus assayed ranged between 0.04 and 0.12, with an average of 0.08 (Table 5). Positive F ST values are usually interpreted as an indicator of divergence among subpopulations or spatial genetic structure that could be the result of drift, assortative mating and natural selection, processes reinforced by restriction of gene flow among populations. In most of the wind-pollinated tree species studied, F ST values tend to be lower than 0.10, demonstrating that more than 90 % of the neutral genetic variation is maintained within populations (Malvolti et al. 1993). The inbreeding coefficient (F IS = 0.68, Table 5) indicated that all of the polymorphism loci showed significant heterozygote deficiency. This phenomenon was attributed to high inbreeding rate at each of the examined locus. In addition, the overall inbreeding coefficient (F IT=0.71, Table 5) is greater than that within populations (F IS = 0.68). The result implies that the mean genetic variability were under-estimated due to the Wahlund effect. Wahlund effect occurs when populations are subdivided due to restrict gene flow (Hartel and Clark 2007). F IS and F IT values are unusually high for a wind-pollinated species and indicate a considerable amount of inbreeding. The large degree of inter-population differentiation and the high within population variability could be attributed to the homogenizing effect of wind pollination or more in general of efficient out crossing (Hamrick et al. 1991). On the other hand, these stands are human origin, and anthropogenic effects may be relevant as well. The average F ST value is 0.08, indicating that 92 % of genetic variability occurred among the six populations studied (Table 5). According to (Wright 1978), F ST values of 0.05–0.15 indicate moderate differentiation among populations. The relatively low H o within Persian walnut populations in Kerman province valleys could be explained by unusual levels of self-pollination or sib-mating (or anthropic effect). In a study of the genetic variation in Italian populations of J. regia, the means of F IS, F IT and F ST were −0.004, 0.08 and 0.085, respectively (Malvolti et al. 1993). In another study, which analyzed the genetic structure of 21 stands of walnut in central Italy, F IS, F IT and F ST were −0.22, −0.6 and 0.15, respectively (Malvolti et al. 1994). The F ST, F IS and F IT values in J. sinensis in Korea were 0.122, 0.156 and 0.258, respectively (Lee and Lee 1997). In study of Pollegioni et al. (2011) the means of F IS, F IT and F ST were 0.022, 0.074 and 0.054 respectively.
Genetic distance among populations
Analysis of the pairwise genetic similarity of the populations (Table 6) showed that the range of S (similarity) as defined by Nei (1978), was from 0.62 to 0.85. The RI population was, on average, the most genetically distant from the other populations (S = 0.67), and KI was the least distant (S = 0.78). The neighbor-joining dendrogram using genetic distance measures on SSR markers data among the populations (Fig. 2) showed that the RI population lay relatively far from the HA (S = 0.67) and BE (S = 0.62) populations (Table 6). Furthermore, the HA, BI, KI and GO populations were in the same group, the BE and RI populations lay in the second and third group, respectively (Fig. 2). The most genetic similarity was observed between BI and HA populations (S = 0.85) (Table 6). Because HA is an ancient walnut-growing region that contains trees about 2,500 years (as the growers mention), it is possible that walnut growers of the BI valley obtained walnut seeds from this area.
The topology of the dendrogram suggests geographical isolation. This isolation could be due to the presence of hills or mountains as geographical barriers between the valleys of the area. This could be the case in the HA population, which is geographically close but genetically distant from the RI population, moreover small sample size of RI population may be reason of these results. Consequently, any correlation was not between genetic distance and geographical distribution of populations from Kerman. In study of genetic variation of J. sinensis in Korea, there was little relationship between geographic and genetic distance between pair populations (Lee and Lee 1997).
It is possible that selection based on unexamined ecological or environmental factors (e.g. rainfall, soil type or also manipulation) has contributed to the divergence and similarity of the populations that were sampled.
The results of this study illustrate the effectiveness of SSR markers for discriminating among genotypes. The results also show that the populations of the valleys of Kerman province have a high level of diversity that can be useful for future breeding programs. Additional collections and further analysis could improve and confirm the above results.
References
Arulsekar S, McGranahan GH, Parfitt DE (1986) Inheritance of phosphoglucomutase and esterase isozymes in Persian walnut. J Hered 77:220–221
Bassam BJ, Caetano-Anolles G, Gresshoff PT (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Bayazit S, Kazan K, Golbitti S, Cevik V, Ayanogla H, Ergul A (2007) AFLP analysis of genetic diversity in low chill requiring walnut (Juglans regia L.) genotyping from Hatay Turkey. Sci Hortic 111:394–398
Beaumont MA, Nichols RA (1996) Evaluating loci for use in genetic analysis of population structure. Proc R Soc Lond B Biol Sci 263:1619–1626
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetic 144:2001–2014
Dangl GS, Woeste K, Aradhya MK, Koehmstedt A, Simon C, Potter D, Leslie C, McGranahan GH (2005) Characterization of 14 microsatellite markers for genetic analysis and cultivar identification of walnut. J Am Soc Hort Sci 130:348–354
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bulletin 19:11–15
Fjellstrom RG, Parfitt DE (1994) Walnut (Juglans spp.) genetic diversity determined by restriction fragment length polymorphisms. Genome 37:690–700
Fjellstrom RG, Parfitt DE, McGranahan GH (1994) Genetic relationship and characterization of Persian walnut (Juglans regia L.) cultivars using restriction fragment length polymorphisms (RFLPs). J Amer Soc Hort Sci 119:833–839
Food and Agriculture Organization (FAO) (2011) FAO statistics division. http://faostat.fao.org
Fornari B, Cannata F, Spada M, Malvolti ME (1999) Allozyme analysis of genetic diversity and differentiation in european and asiatic walnut (Juglans regia L.) populations. Forest Genet 6:115–127
Fornari B, Malvolti ME, Taurchini D, Fineschi S, Beritognolo I, McCaglia E, Cannata F (2001) Isozyme and organellar DNA analysis of genetic diversity in natural/naturalised european and asiatic walnut (J. regia) populations. Acta Hort 544:167–178
Foroni I, Rao R, Woeste K (2006) Molecular characterization of Juglans regia L. cultivar with SSR markers. Acta Hort 705:207–213
Foroni I, Woeste K, Monti LM, Rao R (2007) Identification of ‘Sorrento’ walnut using simple sequence repeats (SSRs). Genet Resour Crop Evol 54:1081–1094
Grant V (1991) The evolutionary process: a critical study of evolutionary theory. Columbia University Press, New York
Gunn BF, Aradhya M, Salick JM, Miller AJ, Yongping Y, Lin L, Xian H (2010) Genetic variation in walnuts (Juglans regia and J. sigillata; Juglandaceae): species distinctions, human impacts, and the conservation of agrobiodiversity in Yunnan, China. Am J Bot 97((4):660–671
Hamrick JL, Godt MJW, Murawski DA, Loveless MD (1991) Correlation between species traits and allozyme diversity: implications for conservation biology. In: Falk DA, Holsinger KE (eds) Genetic and conservation of rare plants. Oxford University Press, New York, pp 75–86
Hartel DL, Clark AG (2007) Principle of population genetics, 4th edn. Sinauer Associates Inc, Sunderland
Lee SW, Lee MH (1997) Genetic variation of Juglans sinensis in Korea. Silvae Genet 46:102–107
Levene H (1949) On a matching problem arising in genetics. Ann Math Stat 20:91–94
Lewontin RC, Krakauer J (1973) Distribution of gene frequency as a test of the theory of the selective neutrality of polymphisms. Genetic 74:175–195
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003) The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4:981–994
Malvolti ME, Paciucci M, Cannata F, Fineschi S (1993) Genetic variation in Italian populations of Juglans regia L. Acta Hort 311:86–94
Malvolti ME, Fineschi S, Pigliucci M (1994) Morphological integration and genetic variability in Juglans regia L. J Hered 85:389–394
Malvolti ME, Pollegioni P, Mapelli S, Cannata F (2010) Research of Juglans regia provenances by molecular, morphological and biochemical markers: a case study in Italy. Biorem Biodiv Bioava 4:84–92
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetic 89:583–590
Nicese FP, Hormaza JI, McGranahan GH (1998) Molecular characterization and genetic relatedness among walnut (Juglans regia L.) genotypes based on RAPD markers. Euphytica 101:199–206
Pollegioni P, Woeste K, Major A, Scarascia Mugnozza G, Malvolti ME (2009) Characterization of Juglans nigra L., Juglans regia L. and Juglans×intermedia Carr. by SSR markers: a case study in Italy. Silvae Genet 58:68–78
Pollegioni P, Woeste K, Olimpieri I, Marandola D, Cannata F, Malvolti ME (2011) Long term human impacts on genetic structure of Italian walnut inferred by SSR markers. Tree Genet Genomes 7:707–723
Potter D, Fangyou G, Aiello G, Leslie G, McGranahan GH (2002) Intersimple sequences repeat markers for fingerprinting and determining genetic relationships of walnut (Juglans regia) cultivars. J Hort Sci Biotech 127:75–81
Rohlf FJ (1998) NTSYSpc numerical taxonomy and multivariate analysis system version 2.0 user guide. Applied Biostatistics Inc., Setauket
Rongwen J, Akkaya MS, Bhagwat AA, Lavia U, Cregan PB (1995) The use of microsatellite DNA marker for soybean genotype identification. Theor Appl Genet 90:43–48
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Solar A, Smole J, Stampar F, Virscek-Marn M (1994) Characterization of isozyme variation in walnut (Juglans regia L.). Euphytica 77:105–112
Storz JF (2005) Using genome scans of DNA polymorphism to infer adaptative population divergence. Mol Ecol 14:671–688
Taylor JS, Durkin JMH, Breden F (1999) The death of a microsatellite: a phylogenetic perspective on microsatellite interruptions. Mol Biol Evol 16:567–572
UPOV (1999) International union for the protection of new varieties of plants. Guidelines for the conduct of tests for distinctness, uniformity and stability. Walnut (Juglans regia). Geneva. p34
Vahdati K (2000) Walnut situation in iran. Nucis Newsl 9:32–33
Victory ER, Glaubitz JC, Rhodes OEJ, Woeste K (2006) Genetic homogeneity in Juglans nigra (Juglandacea) at nuclear microsattelites. Am J Bot 93:118–126
Wang H, Pei D (2008) Genetic diversity and structure of walnut populations in central and southwestern China revealed by microsatellite markers. J Am Soc Hort Sci 133:197–203
Woeste K, Burns R, Rhodes O, Michler C (2002) Thirty polymorphic nuclear microsatellite loci from black walnut. J Hered 93:58–60
Wright S (1978) Evolution and the genetics of populations, variability within and among natural populations, vol 4. University of Chicago Press, Chicago
Yeh F, Yang RC, Boyle TJ (1999) Pop Gene version 1.31. Microsoft window based freeware for population genetic analysis. University of Alberta and Centre for International Forestry Research
Acknowledgments
The authors thank the University of Tehran and the Iran National Science Foundation (INSF) for their support of this research. The authors also acknowledge Dr. Thierry Marcel (Wageningen UR Plant Breeding, Wageningen, The Netherlands), Dr. Pedro Martínez Gómez (Department of Plant Breeding, CEBAS-CSIC, Murcia, Spain) and Dr. Mohsen Ebrahimi (Department of Agronomy and Crop Breeding Sciences, College of Aburaihan, University of Tehran, Tehran, Iran) for pre-reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vahdati, K., Mohseni Pourtaklu, S., Karimi, R. et al. Genetic diversity and gene flow of some Persian walnut populations in southeast of Iran revealed by SSR markers. Plant Syst Evol 301, 691–699 (2015). https://doi.org/10.1007/s00606-014-1107-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-014-1107-8