Abstract
The hybrid origin of Miscanthus purpurascens has previously been proposed, primarily because of its intermediate morphology. In this study, phylogenies based on the DNA sequences from the internal transcribed spacer region of nuclear ribosomal DNA (nrDNA ITS), on the DNA sequences of the trnL intron and trnL-F intergenic spacer of chloroplast DNA, and on amplified fragment length polymorphism (AFLP) fingerprinting confirm that M. purpurascens originated through homoploid hybridization between M. sinensis and M. sacchariflorus. Two different types of ITS sequences were identified from almost all plants of M. purpurascens. One type was found to be closely related to M. sinensis and the other to M. sacchariflorus. Miscanthus purpurascens was found to possess many M. sinensis- and M. sacchariflorus-specific AFLP bands but no band specific to itself. Clustering with the Unweighted Pair Group Method with Arithmetic Mean and principal coordinate analysis based on the AFLP data also demonstrated that M. purpurascens is an approximate intermediate of the two species. In addition, M. purpurascens has the plastid genome of M. sinensis or M. sacchariflorus, suggesting that either species could be its maternal parent. All specimens of M. purpurascens and its coexisting parental species are identified as diploids (2n = 2x = 38). Possible mechanisms of natural hybridization, hybrid status, chloroplast DNA recombination, and evolutionary implications of this hybridization are also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Spontaneous interspecific hybridization is widespread in plants due to incomplete reproductive isolation between closely related species. It has long been recognized as an evolutionary force capable of creating genetic novelties that promote adaptive evolution and speciation (Arnold 1997, 2004; Rieseberg 1997; Seehausen 2004). It is estimated that approximately 50 % of angiosperm species are of hybrid origin (Arnold 1997). Usually, hybrid speciation is accomplished through either of two different mechanisms: allopolyploid hybrid speciation (in which the hybrid species has a higher ploidy level than either parent) or homoploid hybrid speciation (in which the hybrid species has the same number of chromosomes as either parent). Homoploid hybrid speciation involves hybridization between plant taxa at the same ploidy level and is considered rarer than allopolyploid hybrid speciation. It is achieved through ecological and spatial reproductive isolations or chromosomal rearrangement within the hybrid (reviewed in Abbott et al. 2010). Recently, more and more examples of natural homoploid hybrid species in plants have been reported (Ferguson and Sang 2001; Gammon et al. 2007; Gross et al. 2003; Nagamitsu et al. 2006; Pan et al. 2007). These provide opportunities for investigation of the different ways in which they originate.
Miscanthus Andersson (Poaceae) is a perennial rhizomatous grass with a C4 photosynthetic pathway. Most Miscanthus species are native to East Asia. In recent years, Miscanthus has attracted considerable attention for its potential as a bioenergy crop (Heaton et al. 2004; Zub and Brancourt-Hulmel 2010). Miscanthus x giganteus, which was introduced to Europe from Japan as an ornamental grass in 1935, has been extensively cultivated and studied in Europe and North America for biomass feedstock production because of its high biomass yield and relatively low energy and financial input requirements (Bullard et al. 1997; Collura et al. 2006; Heaton et al. 2008; Lewandowski et al. 2000). Based on its morphological and cytological characteristics, it has been hypothesized that M. x giganteus may be an allotriploid hybrid (2n = 3x = 57) that originated from natural hybridization between M. sinensis and M. sacchariflorus (Greef and Deuter 1993; Hodkinson and Renvoize 2001; Linde-Laursen 1993). Evaluation of the interrelationship between M. x giganteus and other Miscanthus taxa using amplified fragment length polymorphism (AFLP) fingerprinting supported this hypothesis (Hodkinson et al. 2002a). Analyses of the DNA sequences from the internal transcribed spacer region of nuclear ribosomal DNA (nrDNA ITS) and the trnL intron and trnL-F intergenic spacer of chloroplast DNA (cpDNA) confirmed the hybrid origin of M. x giganteus, suggesting that M. sacchariflorus was the maternal species of M. x giganteus (Hodkinson et al. 2002b). However, fluorescent in situ hybridization (FISH) and genomic in situ hybridization (GISH) failed to differentiate the different parental genomes present in M. x giganteus, suggesting pronounced similarity in the repetitive DNA sequences (Hodkinson et al. 2002b), and it remains unclear which parental species provides two genomes to M. x giganteus. Until recently, no other natural hybrid in Miscanthus has been recognized or reported.
Here, we report molecular evidence supporting the origin of a diploid hybrid through homoploid hybridization between M. sinensis and M. sacchariflorus. Miscanthus purpurascens was described as a separate species by Liu (1997) in Flora of China, and it is mainly characterized by the purple callus hairs at the base of spikelet, different from the white or yellow callus hairs in other Miscanthus taxa. But Chen and Renvoize (2006) believed that difference in callus hair color is not a sufficient evidence for distinguishing M. purpurascens from M. sinensis. Recently, Sun et al. (2010) further considered that callus hair color is not an ideal taxonomic character, because it can become white from purple gradually during Miscanthus panicle development. Thus, the taxonomic status of M. purpurascens is in doubt. During our previous field investigations of wild Miscanthus germplasms in 2006–2009, we found M. purpurascens to be mainly spread in Liaoning and southern Jilin areas and to be sympatric with M. sinensis and M. sacchariflorus. These three taxa usually coexist in small, mixed populations at some locations. This is because the ability of Miscanthus to reproduce sexually is weak. Miscanthus sinensis and M. sacchariflorus are distinct in their morphology (Chen and Renvoize 2006). However, M. purpurascens exhibits several intermediate morphological characters suggestive of both M. sinensis and M. sacchariflorus. This allows us to speculate that M. purpurascens may be of hybrid origin. In this study, molecular data of nrDNA ITS sequences and AFLP markers from the above-mentioned three Miscanthus taxa were analyzed to confirm the hybrid origin of M. purpurascens. The maternally inherited chloroplast trnL-F intergenic spacer and rpl16 intron region were also sequenced to identify the putative maternal parent species for M. purpurascens. In addition, the conventional root tip squashing technique was used to determine the chromosome number and ploidy level of the Miscanthus samples. The purpose of this study was to determine the hybrid nature of M. purpurascens and assess its hybrid status.
Materials and methods
Plant materials
Twenty-nine specimens including nine M. purpurascens, ten M. sinensis, and ten M. sacchariflorus were sampled from one pure M. purpurascens population, three pure M. sinensis populations, three pure M. sacchariflorus populations, and five mixed populations located in Liaoning and Jilin Provinces in China (Fig. 1; Table 1). Miscanthus sinensis has short vertically oriented rhizomes, clustered culms without nodal buds, and a long lemma awn of 10–17 mm. Miscanthus sacchariflorus is awnless but has long horizontally oriented rhizomes and scattered culms with nodal buds that usually develop into branches. Miscanthus purpurascens exhibits a short lemma awn of 1–6 mm and has medium-sized rhizomes and clustered culms with nodal buds that rarely produce branches. Rhizomes from individuals were collected from sampling locations and grown in the Miscanthus Germplasm Nursery of Hunan Agricultural University. Voucher specimens were deposited in the Herbarium of Miscanthus Research Institute, Hunan Agricultural University.
Map of the locations from which Miscanthus purpurascens and its putative parental species were collected for this study. Numbers correspond to location numbers of the populations given in Table 1
DNA sequencing
Total genomic DNA was extracted from young leaves using a CTAB procedure (Doyle and Doyle 1987). The ITS1-5.8S-ITS2 region of nuclear ribosomal DNA was amplified from all specimens using the universal primers “ITS4” (5′-TCC TCC GCT TAT TGA TAT GC-3′) and “ITS5” (5′-GGA AGT AAA AGT CGT AAC AAG G-3′) (White et al. 1990). The chloroplast trnL-F intergenic spacer region was amplified with the primers “c” (5′-CGA AAT CGG TAG ACG CTA CG-3′) and “f” (5′-ATT TGA ACT GGT GAC ACG AG-3′) described by Taberlet et al. (1991). The chloroplast rpl16 intron region was amplified as described by Jordan et al. (1996) with the primers “rpl16-F71” (5′-GCT ATG CTT AGT GTG TGA CTC GTT G-3′) and “rpl16-R1516” (5′-CCC TTC ATT CTT CCT CTA TGT TG-3′). PCR products were separated through 1 % agarose gel electrophoresis and then purified using the E.Z.N.A.™ Gel Extraction Kit (Omega Bio-Tek, Doraville, GA, USA). Direct sequencing was conducted from both strands to produce consensus sequences on an ABI 3730 automated DNA sequencer (Applied Biosystems, Foster City, CA, USA) by Shanghai Sunny Biotechnology, Inc. Considering the possible heterogeneity within ITS sequences, a T-A cloning step was also performed for ITS PCR products using the InsTAclone™ Cloning Kit (Fermentas, Burlington, Ontario, Canada) and 10–20 positive bacterial colonies for each individual were sequenced. Corresponding DNA fragments in M. paniculatus, M. nepalensis, and M. nudipes were also sequenced and used as outgroups in the phylogenetic analyses. All DNA sequences for this study have been deposited in GenBank (Accession numbers JN544243–JN544347).
DNA sequences were aligned using ClustalX 1.83 (Thompson et al. 1997) and BioEdit 7.0.8 (Ibis Biosciences, Carlsbad, CA, USA). Gap information in the sequences was coded and added to the data matrix using the method described by Simmons and Ochoterena (2000) using the Gapcoder software (Yong and Healy 2003). Chloroplast trnL-F and rpl16 sequences were aligned individually and then concatenated into a single, contiguous sequence for each specimen. Molecular phylogenies between taxa were reconstructed using the neighbor-joining (NJ) and Bayesian inference (BI) methods. NJ analysis was conducted using MEGA 4 (Tamura et al. 2007) with a Kimura 2-parameter distance model (Kimura 1980) and bootstrap analysis used 1,000 replicates. BI analysis was performed in MrBayes 3.1 (Huelsenbeck and Ronquist 2001). jModelTest 0.1.1 (Posada 2008) was used to estimate the best-fit model of nucleotide substitution including rate heterogeneity under the Akaike Information Criterion (AIC; Akaike 1974). The best-fit model for ITS, trnL-F and rpl16 was TIM3+I, F81 and F81+G, respectively. Chloroplast concatenated dataset was partitioned by locus, with the previously inferred best-fit model for each locus. The Markov Chain Monte Carlo (MCMC) analysis consisted of two independent runs starting from random trees with four chains that were run for 2,000,000 generations and sampled every 1,000 generations. Topological convergence between runs was assessed using the online program AWTY (Nylander et al. 2008). The samples generated before reaching stationarity in each run were discarded as burn-in and the remaining samples were used to calculate posterior probabilities.
AFLP analysis
AFLP analysis was performed using a fluorescent AFLP Kit (Dingguochangsheng Biotechnology, Inc., Beijing, China) in accordance with the manufacturer’s instruction. Genomic DNA (200 ng) was double-digested with HindIII and MseI restriction enzymes and then ligated to HindIII and MseI adapters. The digestion-ligation mixture was preamplified by PCR using a HindIII adapter primer (5′-AGA CTG CGT ACC AGC TTA-3′) and an MseI adapter primer (5′-GAT GAG TCC TGA GTA AC-3′). A 1:10 diluted aliquot of the pre-amplified mix was used as the template for selective amplification. Combinations of four HindIII primers (H-AAC, -ACA, -ACG, and -AGC) and two FAM fluorescence-labeled MseI primers (M-CAA and -CAC) were employed in selective amplification. The selective PCR products were separated through 4 % denaturing polyacrylamide gel electrophoresis in an ABI PRISM 377 DNA sequencer (Applied Biosystems, Foster City, CA, USA). AFLP analysis was repeated twice, and only reproducible DNA fragments ranging from 50 to 500 base pairs were scored with GeneScan 3.1 software (Applied Biosystems, Foster City, CA, USA) and Binthere 1.0 software (Garnhart 2001). AFLP data were analyzed using NTSYSpc 2.1e software (Exeter Software, Setauket, NY, USA) to determine the Dice similarity coefficient matrix. Using this matrix, a dendrogram was constructed using clustering with the Unweighted Pair Group Method with Arithmetic Mean (UPGMA). A principal coordinate analysis (PCoA) was performed with the MVSP 3.1 software (Kovach Computing Services, Anglesey, Wales, UK).
Chromosome number and ploidy level
Root tips 0.5–1.0 cm long were pretreated at 10 °C for 2 h with a mixture of 0.2 % colchicine and 0.002 mol L−1 8-hydroxyquinoline. They were then fixed in 3:1 alcohol-acetic acid solution at 4 °C for 20 h. The fixed samples were hydrolyzed with 1 mol L−1 hydrochloric acid at 60 °C for 15 min and 2 % cellulase at 28 °C for 1 h and then squashed in 45 % acetic acid and stained with 0.5 % modified carbol fuchsin. Over 30 well-spread cells were examined to determine the number of chromosomes for each sample.
Results
Phylogenetic analyses based on DNA sequence data
The topology of phylogenetic trees from the neighbor-joining and Bayesian analyses of ITS or combined chloroplast DNA (trnL-F + rpl16) data set is consistent. Only the NJ trees are shown here. NJ bootstrap support >50 % and BI posterior probabilities >0.7 are indicated by numbers above and below the branches, respectively.
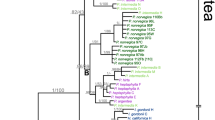
Of the 661 characters in the aligned ITS matrix, 34 singleton variable sites, 16 parsimony informative sites, and 5 indel sites were identified. In the ITS phylogenies, the cloned sequences from samples of M. sinensis and M. sacchariflorus clustered into two distinct species-clades with a strong branch support (Fig. 2). However, two divergent types of ITS sequences were identified from all M. purpurascens samples except one individual (7F). One sequence was found to be closely related to the M. sinensis type and another to the M. sacchariflorus type. The ITS clones of M. purpurascens were divided between the M. sinensis clade and the M. sacchariflorus clade in the phylogenetic tree but they did not form a monophyletic subclade (Fig. 2). In the clones from 7F, only M. sacchariflorus ITS sequences were detected.
The phylogeny of Miscanthus purpurascens, M. sinensis, and M. sacchariflorus obtained from DNA sequences of the internal transcribed spacer region of nuclear ribosomal DNA. Numbers above and below branches represent the relative bootstrap values >50 % and the posterior probability multiplied by 100 and >70, respectively. Accession numbers follow their corresponding taxon names. Clone numbers follow hyphens
The aligned combined chloroplast sequence (trnL-F + rpl16) had a length of 2,083 characters, with 18 singleton variable sites, 11 parsimony informative sites, and 148 indel sites. In the chloroplast DNA phylogeny, the accessions of M. sinensis and M. sacchariflorus were also grouped into two well-supported species-clades, as did the ITS phylogeny (Fig. 3). Two specimens of M. purpurascens from populations 6 (6B) and 7 (7H) formed a monophyletic group with M. sinensis, and the other six specimens of M. purpurascens (2A, 4A, 5C, 6C, 7B, and 7F) were clustered together with M. sacchariflorus (Fig. 3).
The phylogeny of Miscanthus purpurascens, M. sinensis, and M. sacchariflorus obtained from combined chloroplast trnL-F intron spacer and rpl16 intron sequences. Numbers above and below branches represent relative bootstrap values >50 % and the posterior probability multiplied by 100 and >70, respectively
One specimen of M. purpurascens (7D) was excluded from the phylogenetic analysis of chloroplast DNA because its chloroplast genome was a recombinant of M. sinensis and M. sacchariflorus (data not shown). This chloroplast DNA recombinant possessed a single-nucleotide polymorphism (SNP) unique to M. sinensis and one SNP and one indel unique to M. sacchariflorus in its trnL-F sequence. It also had one SNP and one indel unique to M. sinensis and three SNP unique to M. sacchariflorus in its rpl16 sequence. Chloroplast DNA homologous recombination events have been reported in interspecific hybridization of Chlamydomonas (Lemieux et al. 1981, 1984) and in somatic hybridization of Nicotiana (Medgyesy et al. 1985).
Phylogenetic analyses based on AFLP data
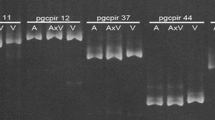
AFLP analysis was conducted to assess the hybrid status of M. purpurascens. A total of 657 bands were identified using eight selective primer combinations that discriminated 29 specimens of M. purpurascens, M. sinensis, and M. sacchariflorus. Among those, 140 bands were found to be common to all specimens analyzed. There were 65 monomorphic, species-specific bands present in all M. sinensis specimens and 68 monomorphic species-specific bands in all M. sacchariflorus specimens, but no bands unique to M. purpurascens were detected. Miscanthus purpurascens specimens possessed 28–36 (mean = 32.1) M. sinensis-specific bands and 22–36 (mean = 28.7) M. sacchariflorus-specific bands (Table 2). Two M. sinensis-specific bands and four M. sacchariflorus-specific bands were absent from all M. purpurascens specimens.
The average genetic similarity index between M. sinensis and M. purpurascens was 0.6333, but the average genetic similarity index between M. sacchariflorus and M. purpurascens was 0.6157. The UPGMA tree based on AFLP data comprised three distinct primary clusters corresponding to the three taxa recognized by morphology. The cluster representing M. purpurascens was approximately equidistant between the M. sinensis cluster and the M. sacchariflorus cluster (Fig. 4). The intermediacy of M. purpurascens was also demonstrated by PCoA, in which the three taxa were completely discriminated along the first axis (Fig. 5).
Chromosome number and ploidy level analysis
Karyotyping of root tip cells revealed that all specimens of M. purpurascens, M. sinensis, and M. sacchariflorus were diploids with a chromosome number of 2n = 2x = 38 (Fig. 6).
Discussion
The hybrid origin of M. purpurascens from hybridization between M. sinensis and M. sacchariflorus has been confirmed in this study. First, both ITS sequence types from parental species are present in the same individuals of M. purpurascens plants, suggesting that the hybrids originated so recently that concerted evolution has not yet been able to homogenize their nuclear ribosomal loci. The absence of M. sinensis ITS sequences from M. purpurascens 7F may have been caused by an insufficient number of sequenced clones, but an alternative explanation could be biased gene conversion (Hillis et al. 1991). In a hybrid scallop system (Chlamys farreri × Argopecten irradians), rapid, concerted gene evolution of the ITS regions via maternally biased gene conversion was observed, causing the absence of paternal ITS sequences from the hybrids (Wang et al. 2010). In other cases, the ribosomal repeats were homogenized to one parental type or a recombinant intermediate following hybridization (Franzke and Mummenhoff 1999; Hughes and Petersen 2001; Wendel et al. 1995). Second, M. purpurascens was found to have the chloroplast genome of either M. sinensis or M. sacchariflorus, indicating reciprocal hybridization between the two species. Third, M. purpurascens additively inherited 28–36 M. sinensis-specific and 22–36 M. sacchariflorus-specific AFLP bands. Both the UPGMA and PCoA analyses indicated that all M. purpurascens specimens showed values approximately halfway between those of M. sinensis and those of M. sacchariflorus.
Miscanthus sinensis and M. sacchariflorus are sympatric in northern and northeastern China, but M. sacchariflorus populations are denser and this species is distributed farther north than M. sinensis because its long, robust rhizomes help it to survive through freezing, drought, and alkaline conditions. Northern China is a heavily populated plains region and most of its land has been reclaimed for agriculture use. Agricultural encroachment has almost driven M. sinensis to extinction, but M. sacchariflorus can survive and grow along the edges of fields. In contrast, both species can survive and coexist in hilly regions of northeastern China (especially Southeastern Liaoning and southern Jilin), which provides an opportunity for natural hybridization. Another factor contributing to natural hybridization is their overlapping flowering periods. Miscanthus sacchariflorus blooms from late August to September and M. sinensis blooms slightly later, and there is about 1 month of overlap between the two species. This suggests that hybridization can occur. Southeastern Liaoning and southern Jilin areas belong to a wider Changbaishan Mountain system, with a very rich forest. Miscanthus species are pioneer plants that are able to colonize the vacant or newly reclaimed lands, and in the process of ecological succession they are gradually pushed away by shade trees, which overshadow them. In addition, Miscanthus species have low sexual reproductive capacity and low seed fitness. These characteristics limited the dispersal and then the population size of M. purpurascens in most habitats. An exception is the population 7 from Qinghe, Liaoning, which is situated on a gradual slope at the edge of a westward trend of the Changbaishan Mountain system toward the Northeast Plain. The slope is an uncultivated south-facing land with dwarf shrubs, which facilitates pollination and hybridization of Miscanthus. Therefore, this population comprises dozens of M. purpurascens plants. Maybe more populations like this can be found in plain regions, but these populations are still fragile because of possible human disturbance.
All specimens of M. purpurascens and coexisting parental species presented in this study were detected as diploids (2n = 2x = 38), suggesting that M. purpurascens originated through hybridization between the diploid species M. sinensis and M. sacchariflorus. M. x giganteus, which was previously the only known Miscanthus hybrid, is an allotriploid hybrid (2n = 3x = 57) between M. sinensis and M. sacchariflorus (Greef and Deuter 1993; Linde-Laursen 1993; Hodkinson and Renvoize 2001). The present study revealed a novel form of hybrid speciation in Miscanthus, one other than allopolyploid hybrid speciation. So far, the genome composition of M. x giganteus is still unknown because neither FISH nor GISH can differentiate the parental genomes present in M. x giganteus because of the pronounced similarity of repetitive DNA sequences between the two parental species (Hodkinson et al. 2002b; Takahashi and Shibata 2002). Compared to M. purpurascens, M. x giganteus has a morphology bias toward M. sacchariflorus, suggesting that the allotriploid should have two sets of M. sacchariflorus genomes, that it is a hybrid between 4x M. sacchariflorus and 2x M. sinensis. This conclusion is also supported by the flow cytometry analysis conducted by Rayburn et al. (2009). That analysis indicated that the hypothetical hybrid between 4x M. sinensis and 2x M. sacchariflorus was estimated to have a nuclear DNA content of 7.75 pg, and the hybrid between 4x M. sacchariflorus and 2x M. sinensis would have a DNA content of 7.2 pg, which corresponds more closely to the observed nuclear DNA content of M. x giganteus (7.0 pg).
AFLP analysis proved to be a reliable method for assessing hybrid status (Kameyama et al. 2005; Teo et al. 2002; Zhou et al. 2005). Because AFLP is a dominant marker, a real hybrid will additively express the bands specific to either of its parental species. For example, an F1 generation will always be heterogeneous for each set of parental species-specific AFLP bands, and 100 % of these would be expressed. With one backcross, the proportion of parental species-specific bands observed in BC1 generation would be 100 % for one parental species and 50 % for the other. Similarly, the proportion of parent-specific AFLP bands for either parental species would be 75 % in F2 generation, 50 % in F3 generation, and so on. The AFLP analysis in this study showed that only 43.1–55.4 % of M. sinensis-specific bands and 32.4–52.9 % of M. sacchariflorus-specific bands expressed in each genotype of M. purpurascens and, therefore, M. purpurascens would most likely be the hybrids of F3 generation. However, Miscanthus is self-incompatible, resulting in highly heterozygous alleles. Because we cannot distinguish between homozygous and heterozygous status of the dominant AFLP markers, the absence of some parental species-specific bands from these hybrids may be caused by parental heterozygosity and consequent homozygous recessive status of hybrids (Jug et al. 2004). For this reason, the possibility that an individual M. purpurascens would be an F1 or F2 hybrid cannot be excluded. The most likely hypothesis is that M. purpurascens is a sterile F1 hybrid. This is because a hybrid swarm (a continuum of morphological variation between two parental species) created by intercrossing among the hybrids was not observed in the population. In this case, the diploid hybrid M. purpurascens cannot be established as a novel species from the viewpoint of sexual reproduction. At present, it is impossible to test the fertility of M. purpurascens because the Miscanthus plants collected from northeastern China cannot flower normally when grown in Hunan. However, some pre- and post-zygotic barriers may be involved in the reproductive isolation between the hybrids and their parental species. For example, the seed-set rate of M. purpurascens, which was tested using several panicles collected from original sampling locations, ranged only from 1.7 to 3.2 %. Almost none of the seeds were able to germinate, most of them being empty or shriveled (data not shown).
In general, F1 hybrids of closely related species tend to exceed their parents in vegetative growth vigor, also called robustness. The transgressive changes in phenotypic traits and gene expression are of great importance in the adaptation of homoploid hybrid species to habitats that are ecologically and spatially different from those of their parents, allowing the hybrids to compete with or even replace their parental species (reviewed by Les and Philbrick 1993). The reproduction of Miscanthus mainly depends on clonal propagation via branched rhizomes, and M. purpurascens has a more robust rhizome system than M. sinensis, implying that it is well adapted to the increasingly serious drought and alkaline conditions in northeastern China, as reflected in the fact that M. purpurascens outnumbers M. sinensis in mixed populations. However, a more detailed study is required to determine the relationship between the hybrid vigor of M. purpurascens and habitat adaptation.
The inheritance of chloroplast DNA in Miscanthus was assumed to be maternal (Hodkinson et al. 2002b). However, the chloroplast genome of the M. purpurascens 7D was confirmed to involve recombination between two parental species. Because of strict uniparental inheritance of chloroplast DNA in most plant species, homologous chloroplast DNA recombination is believed to be absent (Corriveau and Coleman 1988; Nagata 2010; Zhang and Sodmergen 2003). So far, it has only been reported in hybrids of Chlamydomonas eugametos and C. moewusii and in the somatic hybrids of Nicotiana plumbaginifolia and N. tabacum (Lemieux et al. 1981, 1984; Medgyesy et al. 1985). However, mitochondrial DNA homologous recombination has been confirmed in many animals and plants (Burzyński et al. 2003; Ciborowski et al. 2007; Gantenbein et al. 2005; Hoarau et al. 2002; Jaramillo-Correa and Bousquet 2005; Kraytsberg et al. 2004; Städler and Delph 2002; Tsaousis et al. 2005; Ujvari et al. 2007). These chloroplast DNA and mitochondrial DNA recombination events were predominantly discovered in interspecific hybrids, and there are two possible explanations: (1) The frequency of intraspecific homologous recombination in chloroplast or mitochondrial genomes may be much higher than reported, possibly because the absence of diagnostic DNA markers makes it difficult to detect. (2) Molecular recognition systems that result in destruction of chloroplast DNA or mitochondrial DNA may be relaxed during interspecific hybridizations, increasing the frequency of parental leakage (the survival of chloroplast DNA or mitochondrial DNA in sperms or zygotes) in such crosses (Rokas et al. 2003).
References
Abbott RJ, Hegarty MJ, Hiscock SJ, Brennan AC (2010) Homoploid hybrid speciation in action. Taxon 59:1375–1386
Akaike H (1974) A new look at the statistical model identification. IEEE T Automat Contr 19:716–723
Arnold ML (1997) Natural hybridization and evolution. Oxford University Press, New York
Arnold ML (2004) Transfer and origin of adaptations through natural hybridization: were Anderson and Stebbins right? Plant Cell 16:562–570
Bullard MJ, Nixon PMI, Heath MC (1997) Quantifying the yield of Miscanthus x giganteus in the UK. Asp Appl Biol 49:199–206
Burzyński A, Zbawicka M, Skibinski DO, Wenne R (2003) Evidence for recombination of mtDNA in the marine mussel Mytilus trossulus from the Baltic. Mol Biol Evol 20:388–392
Chen SL, Renvoize SA (2006) MISCANTHUS Andersson, Öfvers. Kongl. Vetensk.-Akad. Förh. 12:165. 1855. Flora of China, 2nd edn, vol 22, pp 581–583
Ciborowski KL, Consuegra S, de Leániz CG, Beaumont MA, Wang J, Jordan WC (2007) Rare and fleeting: an example of interspecific recombination in animal mitochondrial DNA. Biol Lett 3:554–557
Collura S, Azambre B, Finqueneisel G, Zimny T, Weber JV (2006) Miscanthus x giganteus straw and pellets as sustainable fuels: combination and emission tests. Environ Chem Lett 4:75–78
Corriveau JL, Coleman AW (1988) Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 angiosperm species. Am J Bot 75:1443–1458
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ferguson D, Sang T (2001) Speciation through homoploid hybridization between allotetraploids in peonies (Paeonia). Proc Natl Acad Sci USA 98:3915–3919
Franzke A, Mummenhoff K (1999) Recent hybrid speciation in Cardamine (Brassicaceae)—conversion of nuclear ribosomal ITS sequences in statu nascendi. Theor Appl Genet 98:831–834
Gammon MA, Grimsby JL, Tsirelson D, Kesseli R (2007) Molecular and morphological evidence reveals introgression in swarms of the invasive taxa Fallopia japonica, F. sachalinensis, and F. x bohemica (Polygonaceae) in the United States. Am J Bot 94:948–956
Gantenbein B, Fet V, Gantenbein-Ritter IA, Balloux F (2005) Evidence for recombination in scorpion mitochondrial DNA (Scorpiones: Buthidae). Proc R Soc B 272:697–704
Garnhart NJ (2001) Binthere v1.0, A program to bin AFLP data. University of New Hampshire
Greef JM, Deuter M (1993) Syntaxonomy of Miscanthus x giganteus Greef et Deu. Angew Bot 67:87–90
Gross BL, Schwarzbach AE, Rieseberg LH (2003) Origin (s) of the diploid hybrid species Helianthus deserticola (Asteraceae). Am J Bot 90:1708–1719
Heaton EA, Long SP, Voigt TB, Jones M, Clifton-Brown J (2004) Miscanthus for renewable energy generation: European Union experience and projections for Illinois. Mitig Adapt Strat Glob Change 9:433–451
Heaton EA, Dohleman FG, Long SP (2008) Meeting US biofuel goals with less land: the potential of Miscanthus. Global Change Biol 14:2000–2014
Hillis DM, Moritz C, Porter CA, Baker RJ (1991) Evidence for biased gene conversion in concerted evolution of ribosomal DNA. Science 251:308–310
Hoarau G, Holla S, Lescasse R, Stam WT, Olsen JL (2002) Heteroplasmy and evidence for recombination in the mitochondrial control region of the flatfish Platichthys flesus. Mol Biol Evol 19:2261–2264
Hodkinson TR, Renvoize SA (2001) Nomenclature of Miscanthus x giganteus (Poaceae). Kew Bull 56:759–760
Hodkinson TR, Chase MW, Renvoize SA (2002a) Characterization of a genetic resource collection for Miscanthus (Saccharinae, Andropogoneae, Poaceae) using AFLP and ISSR PCR. Ann Bot 89:627–636
Hodkinson TR, Chase MW, Takahashi C, Leitch IJ, Bennett MD, Renvoize SA (2002b) The use of DNA sequencing (ITS and trnL-F), AFLP, and fluorescent in situ hybridization to study allopolyploid Miscanthus (Poaceae). Am J Bot 89:279–286
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Hughes KW, Petersen RH (2001) Apparent recombination or gene conversion in the ribosomal ITS region of a Flammulina (Fungi, Agaricales) hybrid. Mol Biol Evol 18:94–96
Jaramillo-Correa JP, Bousquet J (2005) Mitochondrial genome recombination in the zone of contact between two hybridizing conifers. Genetics 171:1951–1962
Jordan WC, Courtney MW, Neigel JE (1996) Low levels of intraspecific genetic variation at a rapidly evolving chloroplast DNA locus in North American duckweeds (Lemnaceae). Am J Bot 83:430–439
Jug T, Dovc P, Pohar J, Snoj A (2004) RAPD analysis as a tool for discriminating marble trout from hybrids (marble trout × brown trout) in the zones of hybridization. J Anim Breed Genet 121:156–162
Kameyama Y, Toyama M, Ohara M (2005) Hybrid origins and F1 dominance in the free-floating, sterile bladderwort, Utricularia australis f. australis (Lentibulariaceae). Am J Bot 92:469–476
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kraytsberg Y, Schwartz M, Brown TA, Ebralidse K, Kunz WS, Clayton DA, Vissing J, Khrapko K (2004) Recombination of human mitochondrial DNA. Science 304:981–983
Lemieux C, Turmel M, Lee RW (1981) Physical evidence for recombination of chloroplast DNA in hybrid progeny of Chlamydomonas eugametos and C. moewusii. Curr Genet 3:97–103
Lemieux C, Turmel M, Seligy VL, Lee RW (1984) Chloroplast DNA recombination in interspecific hybrids of Chlamydomonas: linkage between a nonmendelian locus for streptomycin resistance and restriction fragments coding for 16S rRNA. Proc Natl Acad Sci USA 81:1164–1168
Les DH, Philbrick CT (1993) Studies of hybridization and chromosome number variation in aquatic angiosperms: evolutionary implications. Aquat Bot 44:181–228
Lewandowski I, Clifton-Brown JC, Scurlock JMO, Huisman W (2000) Miscanthus: European experience with a novel energy crop. Biomass Bioenerg 19:209–227
Linde-Laursen I (1993) Cytogenetic analysis of Miscanthus ‘Giganteus’, an interspecific hybrid. Hereditas 119:297–300
Liu L (1997) Miscanthus Anderss. Flora of China, 1st edn, vol 10, pp 4–9. (in Chinese)
Medgyesy P, Fejes E, Maliga P (1985) Interspecific chloroplast recombination in a Nicotiana somatic hybrid. Proc Natl Acad Sci USA 82:6960–6964
Nagamitsu T, Kawahara T, Kanazashi A (2006) Endemic dwarf birch Betula apoiensis (Betulaceae) is a hybrid that originated from Betula ermanii and Betula ovalifolia. Plant Spec Biol 21:19–29
Nagata N (2010) Mechanisms for independent cytoplasmic inheritance of mitochondria and plastids in angiosperms. J Plant Res 123:193–199
Nylander JAA, Wilgenbusch JC, Warren DL, Swofford DL (2008) AWTY (are we there yet?): a system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics 24:581–583
Pan J, Zhang D, Sang T (2007) Molecular phylogenetic evidence for the origin of a diploid hybrid of Paeonia (Paeoniaceae). Am J Bot 94:400–408
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Rayburn AL, Crawford J, Rayburn CM, Juvik JA (2009) Genome size of three Miscanthus species. Plant Mol Biol Rep 27:184–188
Rieseberg LH (1997) Hybrid origins of plant species. Annu Rev Ecol Syst 28:359–389
Rokas A, Ladoukakis E, Zouros E (2003) Animal mitochondrial DNA recombination revisited. Trends Ecol Evol 18:411–417
Seehausen O (2004) Hybridization and adaptive radiation. Trends Ecol Evol 19:198–207
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381
Städler T, Delph LF (2002) Ancient mitochondrial haplotypes and evidence for intragenic recombination in a gynodioecious plant. Proc Natl Acad Sci USA 99:11730–11735
Sun Q, Lin Q, Yi ZL, Yang ZR, Zhou FS (2010) A taxonomic revision of Miscanthus s.l. (Poaceae) from China. Bot J Linn Soc 164:178–220
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Takahashi C, Shibata F (2002) Analysis of Miscanthus sacchariflorus and M. sinensis chromosomes by fluorescence in situ hybridization using rDNA and total genomic DNA probes. Chromosome Sci 6:7–11
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Teo LL, Kiew R, Set O, Lee SK, Gan YY (2002) Hybrid status of kuwini, Mangifera odorata Griff. (Anacardiaceae) verified by amplified fragment length polymorphism. Mol Ecol 11:1465–1469
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tsaousis AD, Martin DP, Ladoukakis ED, Posada D, Zouros E (2005) Widespread recombination in published animal mtDNA sequences. Mol Biol Evol 22:925–933
Ujvari B, Dowton M, Madsen T (2007) Mitochondrial DNA recombination in a free-ranging Australian lizard. Biol Lett 3:189–192
Wang S, Zhang LL, Hu JJ, Bao ZM, Liu ZJ (2010) Molecular and cellular evidence for biased mitotic gene conversion in hybrid scallop. BMC Evol Biol 10:6
Wendel JF, Schnabel A, Seelanan T (1995) Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium). Proc Natl Acad Sci USA 92:280–284
White TJ, Bruns TL, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yong ND, Healy J (2003) Gapcoder automates the use of indel characters in phylogenetic analysis. BMC Bioinformatics 4:6
Zhang Q, Sodmergen LY (2003) Examination of the cytoplasmic DNA in male reproductive cells to determine the potential for cytoplasmic inheritance in 295 angiosperm species. Plant Cell Physiol 44:941–951
Zhou R, Shi S, Wu C (2005) Molecular criteria for determining new hybrid species—An application to the Sonneratia hybrids. Mol Phylogenet Evol 35:595–601
Zub HW, Brancourt-Hulmel M (2010) Agronomic and physiological performances of different species of Miscanthus, a major energy crop.A review. Agron Sustain Dev 30:201–214
Acknowledgments
This research was funded by the National High-tech R&D Program (863 Program) of China (2011AA10020903, 2012AA10180104) and the Mendel Biotechnology Inc., USA
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, J., Zhu, M., Ai, X. et al. Molecular evidence for a natural diploid hybrid between Miscanthus sinensis (Poaceae) and M. sacchariflorus . Plant Syst Evol 299, 1367–1377 (2013). https://doi.org/10.1007/s00606-013-0801-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-013-0801-2