Abstract
Cattleya elongata is a rupicolous orchid species spread throughout and endemic to outcrop islands in campo rupestre vegetation of the Chapada Diamantina, northeastern Brazil. We scored nine natural populations of C. elongata for morphological and genetic variability, covering the whole distribution area of the species, using allozymes and ISSR markers and morphometric multivariate analyses. Genetic variability in allozimes was relatively high (H e = 0.12–0.25), and unexpectedly higher than the values based on ISSR (H e = 0.16–0.19). The populations present moderate structuring (allozymes, ΦPT = 0.14; ISSR, ΦPT = 0.18) and low inbreeding (allozymes, F IS = 0.06). Genetic similarity among the populations was high in both markers, in spite of the discontinuity of the outcrops of the Chapada Diamantina. We found no particular biogeographical pattern to the distribution of the genetic and morphologic similarity among the populations of C. elongata. We found high morphological variability with moderate differentiation among the populations. We did not find any correlation among genetic, morphological, and geographical distances, and among the variability found in the morphological and genetic markers. The differences observed between the two genetic markers and the various morphological markers examined here indicated that the isolated use of any single parameter of these different populations for conservation planning or management would not consider all of the variability to be found in the species, as found in other Brazilian campos rupestres plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Chapada Diamantina is an important center of diversity for the Brazilian flora and one of the regions of greatest diversity in all of South America (Giulietti et al. 1997). The vegetation there has a very high degree of endemism and many genera demonstrating extraordinary degrees of diversification, but it suffers with high anthropogenic degradation caused by substitution of native vegetation by pasture lands and cultivation (Giulietti and Pirani 1988; Giulietti et al. 1997). The Chapada Diamantina is located in the northern portion of the Cadeia de Espinhaço mountain range that stretches approximately 1,000 km from the Serra de Jacobina in the northern part of Bahia state to Serra do Ouro Branco in Minas Gerais state to the south. The mountains in the highest regions of the Chapada Diamantina (generally above 1,000 m) host herbaceous and sub-shrub plants growing on shallow sandy or rocky-sandy soils, as well as herbaceous and shrub formations growing directly on rock outcrops that are known as campos rupestres (Giulietti and Pirani 1988). Areas of campos rupestres are also found in southwestern and southern Minas Gerais, Goiás, and in northern São Paulo states, and in the Federal District as isolated floristic islands surrounded by cerrado (savanna) or caatinga (dryland) vegetation (Giulietti and Pirani 1988; Giulietti et al. 1997). The isolation of these campos rupestres “islands” has been suggested as being responsible for their notable diversity and high degree of endemism (Giulietti and Pirani 1988; Borba et al. 2001; Jesus et al. 2001; Lambert et al. 2006a, b; Pereira et al. 2007; Ribeiro et al. 2008).
According to Giulietti and Pirani (1988), the current phytogeographical patterns seen in the Chapada Diamantina (and other portions of the Cadeia do Espinhaço Range) reflect a long history of climatic fluctuations during which populations were alternately united and separated and geographical barriers were formed that created physical obstacles to gene flow—resulting in a large contingent of endemic species. These authors also concluded that each population has a speciation rate highly dependent on the specific environment in which it grows, resulting in the formation of local floras at individual sites that are extremely small. Significant efforts have been made in the last three decades to describe the flora of the campos rupestres, but information about the reproductive biology of its component species is still scarce, and even less is known about their genetic variability. Borba et al. (2001) noted an elevated genetic variability and moderate to high genetic differentiation between conspecific populations of species of Acianthera (Orchidaceae) in areas of campos rupestres, indicating that geographical distribution contributes to the genetic differentiation of populations found in the Cadeia do Espinhaço. More recent studies with other groups of plants (Jesus et al. 2001, 2009; Lambert et al. 2006a, b; Pereira et al. 2007) as well as other Orchidaceae (Azevedo et al. 2007; Ribeiro et al. 2008) have strengthened this view. Lambert et al. (2006a, b) and Pereira et al. (2007) identified high local genetic differentiation in groups of populations of Cactaceae and Eriocaulaceae species, respectively, occurring in some particular mountains of the Chapada Diamantina. However, no investigations have yet been undertaken using a species widely distributed throughout all of the Chapada Diamantina.
Cattleya elongata Barb. Rodr. (Orchidaceae) is the only species of the genus that is exclusively rupiculous, and its distribution is restricted to the Chapada Diamantina (Cruz et al. 2003). This species is widely distributed in the region, occurring on mountain peaks along the main axis of this range. This species is pollinated by bees and self-compatible and may hybridize with C. tenuis, probably facilitated by habitat disturbance (Smidt et al. 2006). As this is one of the most beautiful species of orchids in the region, some of the populations that are relatively easy to reach have been heavily harvested (or even decimated) through commercial collecting activities. Studies of the genetic and morphological variability in populations of C. elongata were undertaken throughout its range to increase our understanding about the degree of differentiation observed in populations of endemic plants due to fragmentation of campos rupestres occurring in the Chapada Diamantina. Most studies to date with Brazilian species of campos rupestres have used the codominant allozyme marker, but recently studies using more variable dominant markers have been reported. In this study we used a codominant (allozymes) and a dominant (ISSR) genetic marker with the purpose of comparing the response of these two markers in the same plant species, and we correlated the patterns of genetic with morphological variability found in the populations.
Materials and methods
Populations sampled
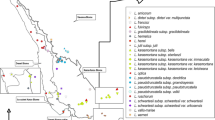
Plant material was collected from nine native populations (180 individuals) of Cattleya elongata along Chapada Diamantina. Populations sampled occur in the municipalities of Ibicoara, Igatu, Lençóis, Morro do Chapéu, Mucugê, Palmeiras, Piatã, Rio de Contas, and Seabra, in the Bahia state, northeastern Brazil (Table 1; Fig. 1). The distance between populations ranges from 9 (Igatu–Mucugê) to 223 (Morro do Chapéu–Rio de Contas) km, with an average distance of 86.5 km. All these populations are usually quite large, having hundreds of individuals. For allozyme analyses, leaf tissue was collected and stored in liquid nitrogen. DNA was extracted from petals dried in silica gel. For morphometric analyses, flowers of the same individuals used in the genetic analyses were collected and preserved in glycerined ethanol 70%. Voucher specimens are deposited in the herbarium HUEFS.
Map of distribution of the nine populations of Cattleya elongata occurring in the Chapada Diamantina, Bahia state, Brazil, used in this study. The regions marked in gray indicates areas where the campo rupestre is the predominant vegetation above 1,000 m. See Table 1 for the names of the populations
Allozyme analysis
The study of genetic variability was carried out using allozyme horizontal electrophoresis in 8.5% starch gel. Small pieces of leaf tissue were crushed in 500 μl of grinding buffer as described in Lambert et al. (2006a). We used the buffer systems 1, 2, 3, and 4 of Lambert et al. (2006a), applying the same running conditions. Eight enzymatic systems presented good resolution and were used in the analyses: buffer system 1—diaphorase (DIA; EC 1.8.1.4); buffer system 2—isocitrate dehydrogenase (IDH; EC 1.1.1.42), malate dehydrogenase (MDH; EC 1.1.1.37); buffer system 3—shikimate dehydrogenase (SKDH; EC 1.1.1.25), phosphoglucoisomerase (PGI; EC 5.3.1.9), phosphoglucomutase (PGM; EC 2.7.5.1); and buffer system 4—esterase (EST; EC 3.1.1.1.), acid phosphatase (ACP; EC 3.1.3.2). The staining procedures were similar to but slightly adjusted from Brune et al. (1998; DIA, SKDH, EST), Corrias et al. (1991; IDH), and Soltis et al. (1983; MDH).
Genetic variability for every population was estimated by the following parameters: proportion of polymorphic loci (P; 0.95 criterion), mean number of alleles per locus (A), mean number of effective alleles per locus (A e), and observed (H o) and expected (H e) mean heterozygosity per locus using GenAlex 6.4 (Peakall and Smouse 2006). Deviations from the expected mean heterozygosity under Hardy-Weinberg (HW) equilibrium were tested. A test for linkage disequilibrium was performed using 100 batches of 1,000 iterations per batch with 1,000 dememorization steps using GENEPOP software (Raymond and Rousset 1995) followed by Bonferroni procedure (Rice 1989). Inbreeding was estimated with Wright’s F IS and partitioning of genetic diversity among conspecific populations was estimated by ΦPT. Analysis of molecular variance (AMOVA) was carried out with 999 permutations to detect variation within and among populations. Cluster analysis was performed with the genetic distance matrix (Nei’s unbiased genetic distance; Nei 1978) of the populations with UPGMA as clustering algorithm in STATISTICA 6.1 (StatSoft 2003).
ISSR analysis
DNA was extracted from dried petals of 15–20 samples of each population using cetyltrimethylammonium bromide (CTAB) protocols (Doyle and Doyle 1987). Amplifications were performed in 20 µl of reaction volumes containing 1× buffer, 2.5 mM MgCl2, 0.2 mM dNTPs, 0.5 mM of primer, 1 U of Taq DNA polymerase (Invitrogen), and DNA template. We tested 20 primers from the ISSR Resources web site (http://www.biosci.ohio-state.edu/~awolfe/ISSR/ISSR.html) and selected 8 with satisfactory resolution for analysis: AW3 (5′-GTG TGT GTG TGT RG-3′), CHRIS (5′-CAC ACA CAC ACA CAY G-3′), DAT (5′-GAG AGA GAG AGA GAR G-3′), UBC 898 (5′-CAC ACA CAC ACA RY-3′), UBC 899 (5′-CAC ACA CAC ACA RG-3′), UBC 901 (5′-GTG TGT GTG TGT YR-3′), UBC 902 (5′-GTG TGT GTG TGT AY-3′), and UBC 944 (5′-CTC TCT CTC TCT CTC TRC-3′). Amplification was performed as follows: initial denaturation at 94°C for 4 min, and then 37 cycles of 1 min at 94°C, 1 min at 46°C (primers UBC844, UBC898, UBC899, and UBC902) or 47°C (primers UBC901, CHRIS, DAT, and AW3), 2 min at 72°C, with a final extension at 72°C for 5 min. PCR reactions were visualized on a 1.4% agarose gel electrophoresis in 1× SB buffer (Brody and Kern 2004) stained with ethidium bromide. The genetic variability of each population was estimated by allele frequency, private alleles, percentage of polymorphic loci (P; 0.95 criterion), expected mean heterozygosity per locus (H e), and Nei’s genetic distance (“unbiased genetic identity”; Nei 1978).
A matrix of genetic distance among populations was obtained, and a principal coordinates analysis (PCA) was carried out. AMOVA was carried out with 999 permutations to detect variation within and among populations. The analyses of variability were performed using GenAlex 6.4. A phenetic analysis using matrix of Nei (1978) unbiased genetic distance with UPGMA as clustering algorithm was performed in STATISTICA 6.1.
Morphometric multivariate analyses
We analyzed 29 floral morphological continuous quantitative characters (Fig. 2; Table 2) in the same individuals used in the genetic analyses. Patterns of morphological similarity/difference were analyzed by multivariate statistical methods using STATISTICA 6.1. The analyses included discriminant analysis, canonical variate analysis (CVA), and cluster analysis for the calculation of variability parameters and morphological structuring. CVA and discriminant analysis were performed with population as the categorical variable (individuals were grouped according to the population to which they belonged). The standardized coefficients for canonical variables resulting from CVA were used to identify the characteristics that most significantly contribute to the resulting patterns observed. In the discriminant analysis we obtained a matrix of squared Mahalanobis distances of individuals to the centroid of the group (D2); the morphologic variability of populations was calculated as the median of these distances (D2m) (Goldman et al. 2004). We used the median of the squared Mahalanobis distances instead of an average of these distances because of the non-normal distribution of the data. Cluster analysis was carried out on a matrix of morphological distance among populations using Mahalanobis Generalized Distance as the distance coefficient, and UPGMA was used as clustering algorithm in STATISTICA 6.1.
Outline of flower parts indicating the morphological characters used in the morphometric analysis of nine populations of Cattleya elongata occurring in the Chapada Diamantina, Bahia state, Brazil. See Table 2 for character codes
A multiresponse permutation procedure (MRPP) analysis was carried out with the PC-ORD 4.10 program (McCune and Mefford 1999) to calculate the chance-corrected within-group agreement (A MRPP) among populations of the studied species. The A MRPP values may be considered comparable with the indexes of genetic differentiation among conspecific populations (ΦPT) (Borba et al. 2002; Lambert et al. 2006a, b). The average Euclidian distance (ED) between the individuals of each population resulting from the MRPP analysis was also utilized as a measure of variability within populations (Borba et al. 2002; Lambert et al. 2006a, b). The two indices of morphological variability are essentially different, as D2m is more affected by form and ED is more affected by size of the characters (Lambert et al. 2006a, b).
Correlation analyses between the markers
The correlation between the matrices of genetic distances (based on ISSR and allozymes markers), between the matrices of genetic and geographic distances in order to test for isolation by distance, between the matrices of genetic and morphological distances, and between matrices of morphological and geographical distances, was tested using the Mantel test with the method of randomization (Monte Carlo, 1,000 randomizations) in PC-ORD 4.10. The pair-wise geographical distances between the populations were computed with geodetic distances on WGS84 earth ellipsoid calculated using the INVERSE 2.0 program (National Geodetic Survey 2002). A Spearman rank correlation analysis between the morphological (ED and D2m) and genetic (H e) variability of populations was carried out using STATISTICA 6.1.
Results
Allozymes
Ten loci with good resolution were obtained for the eight enzyme systems used. No characteristics were observed that indicated the occurrence of polyploidy among the populations, such as individuals with three or more alleles per locus and/or heterozygosity fixed in the populations. No disequilibrium linkage was detected in any loci. Only 10 (5.6%) of the 64 tests carried out with the fixation indices (F) demonstrated significant deviations in relation to that expected by the Hardy-Weinberg (HW) equilibrium. The values of F IS (0.058) and ΦPT (0.143) indicated low inbreeding levels and moderate structuring within the species. An elevated F IS was observed, however, in the Ibicoara population, and moderate values in Rio de Contas (Table 3).
Most of the loci were polymorphic in most of the populations. The loci PGM-1 and MDH-1 were considered monomorphic in all of the populations (0.95 criteria). The number of alleles per locus ranged from two to five. Three populations presented one (Seabra and Mucugê) or two (Palmeiras) exclusive alleles. No population, however, had exclusive fixed alleles or diagnostic loci, although some did show elevated frequencies of alleles that were considered rare for most of the other populations (e.g., allele MDH-2/111 in LE and MC). Intrapopulational variability was relatively high (Table 3): the percentage of polymorphic loci (P; 0.95 criteria) ranged from 50 to 80%; the average number of alleles per locus (A) ranged from 2.0 to 2.4; and the average expected heterozygosity (H e) ranged from 0.12 (Ibicoara) to 0.25 (Morro do Chapéu).
The genetic distance between populations ranged from 0.004 to 0.091 and pairwise F ST from 0.021 to 0.100, with the lowest values being observed in the populations from Seabra and Mucugê, and the highest in the populations from Lençóis and Mucugê. The dendrogram based on Nei’s genetic distances indicated the formation of two main groups, one composed of the populations from Lençóis and Morro do Chapéu, and another larger group composed of the remaining populations (Fig. 3a). This larger group was in turn divided into three subgroups, the first formed by the populations from Ibicoara, Rio de Contas, and Igatu; this larger subgroup was linked to a second formed by the populations from Mucugê, Seabra, and Palmeiras; the Piatã population was externally united with these two.
Dendrogram showing the phenetic relationships among nine populations of Cattleya elongata occurring in the Chapada Diamantina, Bahia state, Brazil, with UPGMA as clustering algorithm. Constructed using the matrix of genetic distances (Nei 1978; unbiased estimate) based on 10 allozymic loci (a) and on 119 ISSR loci (b) and using the matrix Mahalanobis generalized distance based on 29 morphological floral characters (c). Cophenetic correlation = 0.942 (a), 0.957 (b), and 0.872 (c). See Table 1 for the names of the populations
ISSR
The eight primers utilized produced 119 loci with good resolution. The size of the fragments analyzed ranged from approximately 120 to 2,100 base pairs (bp). The majority of the loci were polymorphic. The number of fragments generated per primer ranged from 9 to 24, with an average of 14.9 bands per primer. The AMOVA demonstrated high variation within the populations (82%) but low divergence between them (18%). The value of the genetic structuring as calculated by ΦPT was 0.18. From one (Mucugê, Seabra, and Palmeiras) to four (Igatu) specific bands were found in the populations, although these fragments occurred only at low frequency. The proportions of polymorphic loci ranged from 52.1 (Morro do Chapéu) to 61.3% (Igatu and Palmeiras). The populations demonstrated similar mean expected heterozygosity, with the highest value being observed in the Morro do Chapéu population (0.19) and the lowest in Mucugê (0.16) (Table 3).
The genetic distance between populations ranged from 0.017 to 0.153 and pairwise F ST from 0.000 to 0.230, with the lowest values being observed in the populations from Mucugê and Ibicoara, and the highest in the populations from Mucugê and Palmeiras. In the cluster analysis, the nine populations were subdivided into three groups, and the resulting dendrogram demonstrated a different topology from that observed with the allozyme analysis (Fig. 3b). One group was composed only of the population from Palmeiras, which demonstrated high divergence in relation to the other populations. This large group is divided into a subgroup composed of Piatã, Lençóis, and Morro of Chapéu and another subgroup composed of the remaining populations. The first three axes of the principal coordinates analysis (PCA) explained 72.76% of the observed variability. The Palmeiras population could be distinguished from the others on axis 1 (37.32% of the variability). Axis 1 together with axis 2 (22.28% of the variability) indicated a structuring of the populations into the same three groups observed in the cluster analysis. The PCA indicated that the relationship between the three subgroups observed in the grouping analysis was not stable.
Multivariate morphometric analysis
Only the first four axes were statistically significant in the canonic variance analysis (CVA). The first axis accumulated 51.89% of the variance and was principally correlated with variables #10, 16, 20, and 23 (Table 2). The second axis accumulated 14.73% of the variance and was principally correlated with variables #1, 14, 23, and 28. The third axis accumulated 14.27% of the variance and was correlated with variables #15, 20, and 24. Axis 4 accumulated 8.28% of the variance and was correlated with variables 3, 15, and 27. The scatterplot of the individual scores within these populations in the CVA revealed on axis 1 the separation of the Rio de Contas population from the others, as well as a lesser separation of the Morro do Chapéu population from Piatã, Ibicoara, Lençóis, and Palmeiras (Fig. 4a). On axis 2, the Igatu population was seen to be separate from the Palmeiras and Morro do Chapéu populations; on the third canonic axis the Ibicoara population was seen to be separate from the Seabra population (although there was overlap on both axes). The fourth axis demonstrated a separation of the Morro do Chapéu population from that of Seabra (Fig. 4b).
Representation of the scores on the four first canonical axes of the CVA using 29 morphological floral characters in nine populations of Cattleya elongata occurring in the Chapada Diamantina, Bahia state, Brazil. a Canonical axes 1 and 2. b Canonical axes 3 and 4. Percentage of variance accumulated on the axes: axis 1 = 51.89%; axis 2 = 14.73%; axis 3 = 14.27%; axis 4 = 8.28%. See Table 1 for the names of the populations
In the cluster analysis of the centroids of the populations, the nine populations were subdivided into three groups (Fig. 3c). The first group was composed only of the Rio de Contas population, which demonstrated a high differentiation, linking externally to the large group that was formed by the remaining populations. This large group was, in turn, subdivided in two groups, one composed of the Seabra and Morro of Chapéu populations, and the other composed of the remaining populations (but with the Igatu population linking externally with the latter). The floral diagrams of those individuals considered to be most representative of each population (those that demonstrated the least distance from the centroids of their respective populations) can be seen in Fig. 5.
Outline of flower parts of the individual closest to the centroid of each of the nine populations of Cattleya elongata occurring in the Chapada Diamantina, Bahia state, Brazil, in the canonical variate analysis based on 29 morphological characters. a Igatu (IG), b Mucugê (MU), c Ibicoara (IB), d Rio de Contas (RC), e Seabra (SE), f Palmeiras (PA), g Piatã (PI), h Lençóis (LE), i Morro do Chapéu (MC)
The populations demonstrated moderate morphological structuring (A MRPP = 0.2454). The Rio de Contas population had the greatest values in both analyses of morphological variability (Table 3). The average Euclidian distances (ED) between all of the members of the populations had wide amplitudes, ranging from 23.52 (Palmeiras population) to 44.8 (Rio de Contas population). The median of the generalized Mahalanobis distance (D2m) ranged from 20.27 (Seabra population) to 31.81 (Rio de Contas population).
Correlation analyses between the markers used
The Mantel test indicated the absence of any significant correlation between the genetic distances obtained from the two molecular markers, the allozymes and ISSR (r = 0.0034, p = 0.2994); between the allozyme genetic distance and the geographical distance (allozymes r = 0.2499, p = 0.1730); between the genetic distance based on ISSR and the geographical distance (ISSR r = −0.0742, p = 0.4924); between the morphological and geographical distances (r = 0.4827, p = 0.0794); or between the genetic and morphological distances (allozymes r = −0.1379, p = 0.3298; ISSR r = −0.2321, p = 0.1376). The Spearman correlation analyses between genetic variability based on allozymes and ISSR (H e) with the morphological data demonstrated an absence of correlation between genetic and morphological variability (D2m × allozymes: r = 0.0450, p = 0.2242; D2m × ISSR: r = 0.5000, p = 0.1704; ED × allozymes: r = 0.5166, p = 0.1543; ED × ISSR: r = 0.2500, p = 0.5164) as well as between the two measures of genetic variability utilized in this study (allozymes × ISSR; r = 0.4666, p = 0.2000).
Discussion
The populations of Cattleya elongata examined here demonstrated moderate to high levels of allozyme genetic variability when compared with other groups of endemic plants from campos rupestres vegetation in the Cadeia do Espinhaço Range (Jesus et al. 2001; Lambert et al. 2006a, b; Pereira et al. 2007), but average values when compared to other plants with similar characteristics (long-lived perennial with narrow distribution, herbaceous, wind-dispersed, outcrossing plants; Hamrick and Godt 1990). In general, these values were similar or superior to those observed in the majority of studies undertaken in orchids (e.g., Case et al. 1998; Chung and Chung 1999; Gustafsson 2000; Chung et al. 2004; Trapnell et al. 2004). However, this genetic variability was less than that encountered in other species of Orchidaceae that occur in the same vegetation formation (Borba et al. 2001; Azevedo et al. 2007; Ribeiro et al. 2008), including other species of Cattleya in the Chapada Diamantina (Borba et al. 2007a, b). The indices of genetic variability based on ISSR were lower than the average values encountered in plants with the same characteristics of C. elongata (Nybom 2004), including other orchid species (e.g., Smith et al. 2002). The genetic markers utilized here demonstrated heterozygosity values considerably different from the expected patterns, with variability as measured using allozymes being superior to that observed using ISSR (Hamrick and Godt 1990; Nybom 2004). The low observed genetic diversity values as estimated by the ISSR markers, however, may be related to the primer choices utilized in the present analyses (Smith et al. 2002). A similar situation was observed with Tipularia discolor (Orchidaceae), in which intra- and interpopulational genetic variability levels measured using ISSR markers were also found to be low (Smith et al. 2002).
The Ibicoara and Mucugê populations demonstrated the lowest genetic variability values, while the most geographically isolated population, Morro do Chapéu, demonstrated the highest values as estimated using both markers. Stochastic events, such as genetic drift and genetic bottlenecks, as well as a decrease (or interruption) of gene flow can produce geographically isolated populations that show reduced genetic variability, such as that seen in the Ibicoara population. However, this would appear not to be the case with the Mucugê population, which is geographically very near to other populations that demonstrated much greater genetic variability (such as Igatu). The Mucugê population is located at the edge of an important local highway, which favors intense harvesting, and this population should be monitored to avoid an even greater depletion of its variability, which could jeopardize its viability.
The F IS values were low in all of the allozyme systems, with only small differences between the observed and expected heterozygosity values in the populations. The populational genetic structure of plants is directly related to factors such as their reproduction systems and their mechanisms of pollen and seed dispersal, their effective population sizes, and their capacity to colonize new sites (Hamrick and Godt 1990, 1996). Cattleya elongata is pollinated by queens of Bombus (Fervidobombus) brevivillus Franklin 1913 (Smidt et al. 2006). Although C. elongata is self-compatible, this species presents a mechanism of pollination by deception (the flowers produce no nectar) that favors cross-pollination (Smidt et al. 2006) and is probably responsible for the low observed F IS values. Little is known about seed dispersal in the Orchidaceae, but the genetic structures of the populations of some species are known to be low, demonstrating that gene flow driven by either seed or pollen dispersal is quite high (Tremblay and Ackerman 2001; Trapnell and Hamrick 2004).
Cattleya elongata is distributed in disjunct populations in the Chapada Diamantina mountains due to the discontinuity of the outcrops there, and the genetic structuring found was moderate, but higher than that normally observed with orchid species in campos rupestres areas (Borba et al. 2001, 2007b; Azevedo et al. 2007; Ribeiro et al. 2008). There is an increased probability that the genes of plants with long life cycles (due to their capacity for vegetative propagation) will be transported to more distant localities and thus resist genetic drift (Loveless and Hamrick 1984). The interpopulational variability estimated from analysis of molecular variance demonstrated most variation is found within the populations, and low divergence between them, which is consistent with an allogamic reproductive pattern of species with wind-dispersed seeds. Although most of the genetic variation of the species is found within populations, some populations presented exclusive alleles, such as Igatú, Lençóis, Palmeiras, Seabra, and Mucugê, indicating their uniqueness.
The phenetic analyses based on the Nei’s distance (1978) obtained from genetic data indicated no single biogeographical pattern that could describe the distribution of the genetic similarity of the populations of C. elongata. It was to be expected that the closest populations, or those that occupied contiguous mountain chains in the Chapada Diamantina, would demonstrate higher genetic similarity. The two closest populations (Mucugê and Igatu—only 26 km apart) demonstrated intermediate levels of genetic similarity. Nonetheless, it was observed that genetic distance between populations was in general considerably reduced, especially as evaluated by using allozyme markers. As such, it cannot be categorically stated that there is high degree of differentiation between populations in the same region of the Chapada Diamantina or even between quite different regions. In general, the Morro do Chapéu population appears to be more similar to the Lençóis population than to the others, indicating any decrease in gene flow between these two populations must have only occurred recently. These two populations diverge from the others when evaluated using allozyme markers, but remain in the same small group when analyzed using ISSR markers, in spite of the fact that the Lençóis population is geographically closer to the Palmeiras population (which was the most genetically divergent group in ISSR data). This divergence of the Palmeiras population was a surprising result, as this population is actually located very close to the other populations examined here. The most plausible explanation for this situation is that ecological factors must be acting to diminish gene flow between this population and the others. Additionally, reports of hybridization of this species with C. tenuis in this region have been reported (Cruz et al. 2003; Borba et al. 2007b). The groupings observed here may also reflect adaptations to different ecological conditions, because in spite of their geographical proximity, many of these populations are found (for example) at different altitudes. As observed for genetic markers, the high A MRRP value indicates moderate morphological structuring, which was mainly due to differentiation in characteristics such as the angle of the apex of the lateral sepal and of the petal, the width of the labellum, and the width of the lobe terminal. Morphological data also demonstrated no well-defined biogeographical patterns to the distribution of morphological similarities among the populations, although these were better defined than those observed with the molecular markers. The Morro do Chapéu and Rio de Contas populations, located in the extreme northern and southern distribution limits of this species, and the Seabra population (disjunct from the others by lowlands) were observed to be most morphologically divergent from the populations in the core area of the species.
The differentiation of populations isolated by depressions or disjunct of C. elongata, such as Morro do Chapéu in allozymes, Palmeiras in ISSR, and Rio de Contas, Seabra and Morro do Chapéu in morphometrics, may be related to climate changes during the Pleistocene in the tropical region of Brazil (Salgado-Labouriau et al. 1998), according to the theory of refugia (Prance 1982). During the Pleistocene climate changes, the decrease in temperature and humidity led to the reduction of forests and favored the expansion of the distribution of plant species from drier areas. This process could have resulted in expansion of areas of campos rupestres vegetation, currently restricted to altitudes above 800 m (see Fig. 1), to lowlands occupying large quartzite areas, environment suitable for C. elongata. At the end of glaciation, with the increase in temperature and humidity and expansion of forest areas, there was a retraction of the campos rupestres, with the extinction of intermediate populations occurring at lower altitudes, leading to allopatric fragmentation of the species. This model has been used to explain the patterns of species differentiation in these and adjacent formations (Collevatti et al. 2009; Ramos et al. 2007; Feres et al. 2009; Lage-Novaes et al. 2010) as well as the Atlantic forest in Brazil (e.g., Palma-Silva et al. 2009), and may be a major factor in high endemism (Giulietti and Pirani 1988) and high differentiation in disjunct populations of species of campos rupestres (e.g., Jesus et al. 2001; Pereira et al. 2007).
The absence of any correlations between any of the genetic and morphological variability indices was unexpected. However, correlations between genetic and morphological data have not been very frequently observed in many groups of organisms (Mitton 1978; Gilles 1984; Elisens et al. 1992; Avise 1994), including plants of campos rupestres (Borba et al. 2001; Lambert et al. 2006a, b; Conceição et al. 2008). This result is apparently due to the fact that changes at the genetic level are not always reflected in morphological variations as mutations may occur in many different regions of the genome (either in coding regions or not), and DNA polymorphism markers do not necessarily correspond to regions related to the expression of morphological characters. Likewise, no correlation was detected between genetic and geographical distances, nor between morphological and geographical distances, even though various disjunctions were observed in the geographical distributions of the analyzed populations that have previously been associated with the genetic differentiation of populations in other groups of plants (Borba et al. 2001; Jesus et al. 2001; Lambert et al. 2006a, b; Pereira et al. 2007; Ribeiro et al. 2008).
In spite of past anthropogenic influences in the last two centuries, it could be seen that these populations still demonstrated moderate levels of genetic and morphological variability. This could easily and quickly deteriorate, however, as there is still no control over the commercial exploitation of this species—a situation that deserves special attention from environmental authorities. Knowledge of the level of genetic diversity of a taxon, not simply its geographical distribution, is an important consideration when planning conservation strategies for threatened species (Hamrick 1989; Lambert et al. 2006a, b). The differences observed between the two genetic markers and the various morphological markers examined here indicated that the isolated use of any single parameter of these different populations for conservation planning or management would not consider all of the variability to be found in the species—a situation that has been observed with other Brazilian campos rupestres plants (Borba et al. 2002, 2007a; Lambert et al. 2006a, b; Pereira et al. 2007; Ribeiro et al. 2008). As such, it is fundamental that as much information as possible be used in conservation planning.
References
Avise JC (1994) Molecular markers, natural history and evolution. Chapman & Hall, New York
Azevedo MTA, Borba EL, Semir J, Solferini VN (2007) High genetic variability in Neotropical myophilous orchids. Bot J Linn Soc 153:33–40
Borba EL, Felix JM, Solferini VN, Semir J (2001) Fly-pollinated Pleurothallis (Orchidaceae) species have high genetic variability: evidence from isozyme markers. Am J Bot 88:419–428
Borba EL, Shepherd GJ, van den Berg C, Semir J (2002) Floral and vegetative morphometrics in five Pleurothallis (Orchidaceae) species: correlation with taxonomy, phylogeny, genetic variability and pollination systems. Ann Bot 90:219–230
Borba EL, Funch RR, Ribeiro PL, Smidt EC, Silva-Pereira V (2007a) Demography, genetic and morphological variability of the endangered Sophronitis sincorana (Orchidaceae) in the Chapada Diamantina, Brazil. Plant Syst Evol 267:129–146
Borba EL, Funch RR, Ribeiro PL, Smidt EC, Silva-Pereira V (2007b) Demografia, variabilidade genética e morfológica e conservação de Cattleya tenuis (Orchidaceae), espécie ameaçada de extinção da Chapada Diamantina. Sitientibus Sér Ciênc Biol 7:211–222
Brody JR, Kern SE (2004) Sodium boric acid: a tris-free, cooler conductive medium for DNA electrophoresis. Biotechniques 36:2–4
Brune W, Alfenas AC, Junghans TG (1998) Identificações específicas de enzimas em géis. In: Alfenas AC (ed) Eletroforese de isoenzimas e proteinas afins: fundamentos e aplicações em plantas e microorganismos. Editora Universidade Federal de Viçosa, Viçosa, pp 201–328
Case MA, Mlodozeniec HT, Wallace LE, Weldy TW (1998) Conservation genetics and taxonomic status of the rare Kentucky lady’s slipper: Cypripedium kentuckiense (Orchidaceae). Am J Bot 89:843–853
Chung MY, Chung MG (1999) Allozyme diversity and population structure in Korean population of Cymbidium goeringii (Orchidaceae). J Plant Res 112:139–144
Chung MY, Nason JD, Chung MG (2004) Spatial genetic structure in populations of the terrestrial orchid Cephalanthera longibracteata (Orchidaceae). Am J Bot 91:52–57
Collevatti RG, Rabelo SG, Vieira RF (2009) Phylogeography and disjunct distribution in Lychnophora ericoides (Asteraceae), an endangered cerrado shrub species. Ann Bot 104:655–664
Conceição AS, Queiroz LP, Lambert SM, Pereira ACS, Borba EL (2008) Biosystematics of Chamaecrista sect. Absus subsect. Baseophyllum (Leguminosae-Caesalpinioideae) based on allozyme and morphometric analyses. Plant Syst Evol 270:183–207
Corrias B, Rossi W, Arduino P, Cianchi R, Bullini L (1991) Orchis longicornu Poiret in Sardinina: genetic, morphological and chorological data. Webbia 45:71–101
Cruz DT, Borba EL, van den Berg C (2003) O gênero Cattleya Lindl. (Orchidaceae) no estado da Bahia, Brasil. Sitientibus Sér Ciênc Biol 3:26–34
Doyle JJ, Doyle JL (1987) A rapid isolation procedure for small quantities of fresh tissue. Phytochem Bull 19:11–15
Elisens WJ, Boyd RD, Wolfe AD (1992) Genetic and morphological divergence among varieties of Aphanostephus skirrhobasis (Asteraceae-Asterae) and related species with different chromosome numbers. Syst Bot 17:380–394
Feres F, Zucchi MI, Souza AP, Amaral MCE, Bittrich V (2009) Phylogeographic studies of Brazilian “campo-rupestre” species: Wunderlichia mirabilis Riedel ex Baker (Asteraceae). Biotemas 22:17–26
Gilles BE (1984) A comparison between quantitative and biochemical variation in the wild barley Hordeum murinum. Evolution 38:34–41
Giulietti AM, Pirani JR (1988) Patterns of geographic distribution of some plant species from the Espinhaço Range, Minas Gerais and Bahia, Brazil. In: Vanzolini PE, Heyer WR (eds) Proceedings of a workshop on neotropical distribution patterns. Academia Brasileira de Ciências, Rio de Janeiro, pp 36–69
Giulietti AM, Pirani JR, Harley RM (1997) Espinhaço Range, eastern Brazil. In: Davis SD, Heywood VH, Herrera-Macbyde O, Villa-Lobos J, Hamilton AC (eds) In centres of plant diversity. A guide and strategy for their conservation, v.3. The Americas IUCN Publication Unity, Cambridge, pp 397–404
Goldman DH, van den Berg C, Griffith MP (2004) Morphometric circumscription of species and infraspecific taxa in Calopogon R.Br. (Orchidaceae). Plant Syst Evol 247:37–60
Gustafsson S (2000) Patterns of genetic variation in Gymnadenia conopsea, the fragrant orchid. Mol Ecol 9:1863–1872
Hamrick JL (1989) Isozymes and the analysis of genetic structure in plant population. In: Soltis DE, Soltis PS (eds) Isozymes in plant biology. Dioscorides Press, Portland, pp 87–105
Hamrick JL, Godt MJW (1990) Allozyme diversity in plant species. In: Brown ADH, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, pp 43–63
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Phil Trans R Soc London B 351:1291–1298
Jesus FF, Solferini VN, Semir J, Prado PI (2001) Local genetic differentiation in Proteopsis argentea (Asteraceae), a perennial herb endemic in Brazil. Plant Syst Evol 226:59–68
Jesus FF, Abreu AG, Semir J, Solferini VN (2009) Low genetic diversity but local genetic differentiation in endemic Minasia (Asteraceae) species from Brazil. Plant Syst Evol 277:187–196
Lage-Novaes RM, Lemos-Filho JP, Ribeiro RA, Lovato MB (2010) Phylogeography of Plathymenia reticulata (Leguminosae) reveals patterns of recent range expansion towards northeastern Brazil and southern Cerrados in eastern Tropical South America. Mol Ecol 19:985–998
Lambert SM, Borba EL, Machado MC, Andrade SCS (2006a) Allozyme diversity and morphometrics of Melocactus paucispinus (Cactaceae) and evidence for hybridization with M. concinnus in the Chapada Diamantina, north-eastern Brazil. Ann Bot 97:389–403
Lambert SM, Borba EL, Machado MC (2006b) Allozyme diversity and morphometrics of the endangered Melocactus glaucescens (Cactaceae), and investigation of the putative hybrid origin of Melocactus × albicephalus (Melocactus ernestii × M. glaucescens) in north-eastern Brazil. Plant Species Biol 21:93–108
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Annu Rev Ecol Syst 15:65–95
Mccune B, Mefford MJ (1999) PC-Ord—multivariate analysis of ecological data, version 4.10. MjM Software, Gleneder Beach
Mitton JB (1978) Relationship between heterozygosity for enzyme loci and variation of morphological characters in natural populations. Nature 273:661–662
National Geodetic Survey (2002) INVERSE version 2.0. http://ngs.noaa.gov/. 18 April 2006
Nei M (1978) Estimation of average heterozigosity and genetic distance from a small number of individuals. Genetics 89:583–590
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Palma-Silva C, Lexer C, Paggi GM, Barbará T, Bered F, Bodanese-Zanettini MH (2009) Range-wide patterns of nuclear and chloroplast DNA diversity in Vriesea gigantea (Bromeliaceae), a neotropical forest species. Heredity 103:503–512
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pereira ACS, Borba EL, Giulietii AM (2007) Genetic and morphological variability of the endangered Syngonanthus mucugensis Giul. (Eriocaulaceae), from the Chapada Diamantina, Brazil: implications for conservation and taxonomy. Bot J Linn Soc 153:401–416
Prance GT (1982) Forest refuges: evidence from wood angiosperms. In: Prance GT (ed) Biological diversification in the tropics. Columbia University Press, New York, pp 137–158
Ramos ACS, Lemos-Filho JP, Ribeiro RA, Santos FR, Lovato MB (2007) Phylogeography of the tree Hymenaea stigonocarpa (Fabaceae: Caesalpinioideae) and the influence of quaternary climate changes in the Brazilian cerrado. Ann Bot 100:1219–1228
Raymond M, Rousset F (1995) GENEPOP (version 1–2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Ribeiro PL, Borba EL, Smidt EC, Lambert SM, Schnadelbach AS, van den Berg C (2008) Genetic and morphological variation in the Bulbophyllum exaltatum (Orchidaceae) complex occurring in the Brazilian campos rupestres: implications for taxonomy and biogeography. Plant Syst Evol 270:109–137
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Salgado-Labouriau ML, Barberi M, Ferraz-Vicentini KR, Parizzi MG (1998) A dry climatic event during the late quaternary of tropical Brazil. Rev Paleobot Palynol 99:115–129
Smidt EC, Silva-Pereira V, Borba EL (2006) Reproductive biology of two Cattleya (Orchidaceae) species endemic to north-eastern Brazil. Plant Species Biol 21:85–91
Smith JL, Hunter KL, Hunter RB (2002) Genetic variation in the terrestrial orchid Tipularia discolour. South Nat 1:17–26
Soltis DE, Haufler CH, Darrow DC, Gastony GJ (1983) Starch gel electrophoresis of ferns: a compilation of grinding buffers, gel and electrode buffers, and staining schedule. Am Fern J 73:9–27
StatSoft Inc. (2003) STATISTICA (data analysis software system), version 6. StatSoft Inc., Tulsa
Trapnell DW, Hamrick JL (2004) Partitioning nuclear and chloroplast variation at multiple spatial scales in the neotropical epiphytic orchid, Laelia rubescens. Mol Ecol 13:2655–2666
Trapnell DW, Hamrick JL, Nason JD (2004) Three-dimensional fine-scale structure of the neotropical epiphytic orchid, Laelia rubescens. Mol Ecol 13:1111–1118
Tremblay RL, Ackerman JD (2001) Gene flow and effective population size in Lepanthes (Orchidaceae): a case for genetic drift. Biol J Linn Soc 72:47–62
Acknowledgments
We thank Ricardo Villas Boas Gomes for technical support. This work was supported by a grant from the Fundação de Apoio à Pesquisa do Estado da Bahia (FAPESB), the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and Programa de Pesquisa em Biodiversidade do Semi-Árido (PPBio). D.T.C. received a scholarship from CNPq. E.L.B. is supported by a productivity grant (PQ2) from CNPq.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
da Cruz, D.T., Selbach-Schnadelbach, A., Lambert, S.M. et al. Genetic and morphological variability in Cattleya elongata Barb. Rodr. (Orchidaceae), endemic to the campo rupestre vegetation in northeastern Brazil. Plant Syst Evol 294, 87–98 (2011). https://doi.org/10.1007/s00606-011-0444-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-011-0444-0