Abstract
Multivariate analysis of both endocarp and leaf morphometrics is combined with cluster analysis and Bayesian inference of AFLP markers to assess the morphologic and genetic variation of five European members of Prunus section Prunus (P. cerasifera, P. domestica, P. insititia, P. spinosa, and P. × fruticans). Endocarp morphometrics separate most Prunus taxa studied, but overlap remains between P. domestica and P. cerasifera, and P. spinosa and P. × fruticans. Leaf morphometrics yield better separation of P. domestica and P. cerasifera, but do not allow distinction between P. spinosa and P. × fruticans. Both cluster analysis and PCoA of AFLP markers equally produce three distinct clusters. A first consists of all P. cerasifera samples and the sole P. cocomilia; a second cluster includes all individuals of P. domestica and P. insititia; and a third group comprises all P. spinosa and P. × fruticans samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Prunus L. (Rosaceae) is naturally distributed in the temperate regions of the Northern Hemisphere with some species occurring in the tropical and subtropical regions (Rehder 1940; Krüssmann 1978; Scholz and Scholz 1995; Kalkman 2004; Mabberley 2008). The genus comprises around 200 species and is traditionally divided into five subgenera: Amygdalus (L.) Focke, Cerasus Pers., Laurocerasus Koehne, Padus (Moench) Koehne, and Prunus (Rehder, 1940). The subgenus Prunus, consists of three sections: Armeniaca (Lam.) Koch. (Apricots), Prunocerasus Koehne (North American plums), and Prunus (Eurasian plums) (Rehder, 1940). Following Mason (1913) and Rehder (1940), the section Prunus consists of P. cerasifera Ehrh. (Cherry plum or Myrobalan plum), P. divaricata Ledeb. [=Prunus cerasifera Ehrh. var. divaricata (Ledeb.) L. H. Bailey], P. domestica L. (European plum), P. salicina Lindl. (Japanese plum), P. simonii Carr. (Apricot plum), and P. spinosa L. (Hawthorn or Sloe). Based on sequence analysis of ITS and trnL–trnF Spacer DNA, Bortiri et al. (2001) partitioned the section Prunus in two clades: one consisting of P. salicina and P. simonii and one including P. cerasifera, P. domestica, and P. spinosa. This division is consistent with results from isozyme data (Mowrey and Werner 1990) and cpDNA sequence analysis (Shaw and Small, 2004). The close relationship between P. cerasifera, P. domestica (including P. insititia), and P. spinosa has been demonstrated by a number of morphologic studies (Hanelt 1997; Kühn 1999; Woldring 2000; Nielsen and Olrik 2001) and has been confirmed in several genetic analyses (Bortiri et al. 2001; Aradhya et al. 2004; Shaw and Small 2004; Katayama and Uematsu 2005). Still, taxonomic delineation of these taxa is problematic, and their mutual relationships are far from well resolved. Several factors may have contributed to the complexity of this polymorphic group. Most probably, polyploidisation events (e.g., Reynders and Salesses 1991; Zohary 1992; Scholz and Scholz 1995; Körber-Grohne 1996; Woldring 2000), extensive hybridisation and introgression (e.g., see Stace 1975; Woldring 2000), and centuries of domestication and cultivation (e.g., see Behre 1978; Zohary 1992; Zohary and Hopf 1994; Woldring 2000; Nielsen and Olrik 2001; Frary and Doğanlar 2003) resulted in a wide range of variation and transitional states between and within the different taxa. Still, the delineation of the different taxa involved and knowledge on their morphological and genetic diversity is indispensable basic information for their proper conservation.
Endocarp dimension and shape characteristics are very useful for identification of the taxa of Prunus section Prunus (Röder, 1940; Behre, 1978; Van Zeist and Woldring, 2000; Nielsen and Olrik, 2001; Depypere et al., 2007). Still, Depypere et al. (2007) illustrated that a single endocarp characteristic is not sufficient to discriminate between the taxa of Prunus section Prunus because of the overlapping ranges of endocarp size and shape parameters. Similarly, Nielsen and Olrik (2001) state that it is possible to separate well developed specimens of P. spinosa and P. insititia by means of a single character, but that evaluation of several characters is necessary to obtain an exact identification when other taxa or Prunus subgenus Prunus hybrids are involved. Therefore, multivariate analysis of both endocarp and leaf morphometrics is performed in the present study.
Since morphological characteristics are subject to environmental variability and selection pressure and since hybrids do not always show an intermediate morphology (Thórsson et al. 2001), we combined the morphometric study with an AFLP-based analysis. AFLP has a number of advantages: polymorphic markers are generated at a high frequency, the technique has a high reproducibility and involves genome wide sampling and the markers are not under direct selection pressure (Jones et al. 1997; McGregor et al. 2000; Russell et al. 1997). In the clustering methods of the phenetic analyses the trees are calculated without the assumptions of an evolutionary model (Koopman et al. 2008). Since hybrids usually violate such assumptions, these clustering methods are appropriate to study species complexes that incorporate hybrids (Koopman et al. 2008). UPGMA has been demonstrated to produce trees with the highest cophenetic correlation, i.e. with the best representation of the data (e.g., Mace et al. 1999a, b; Koopman et al. 2001; Van Droogenbroeck et al. 2002; Drossou et al. 2004). Still, trees yielded by clustering methods depict phenetic relationships (reflecting similarity) rather than phylogenetic relationships (reflecting evolution). However, AFLP is a valuable tool for both phenetic studies of closely related taxa (e.g., apricot in Hagen et al. 2002; Malus in Coart et al., 2003; Rosa in De Cock et al., 2008) and for phylogeny of evolutionary complex groups (e.g., Rosa in Koopman et al., 2008; Betula in Schenk et al., 2008). Bayesian inference based on AFLP data is suitable to provide phylogenetic information of evolutionary complex groups (e.g., Rosa in Koopman et al., 2008; Betula in Schenk et al., 2008). For these reasons, both UPGMA and Bayesian inference is used on the AFLP data. Principal Co-ordinate Analysis was performed to visualize the genetic diversity. The endocarp and leaf morphometric studies provide complementary information that contributes to a more detailed understanding of the taxonomic status of the taxa. To our knowledge, this is the first report that combines a morphological and an AFLP-based variation study of five closely related Eurasian plum taxa (P. cerasifera, P. domestica, P. insititia, P. spinosa, and P. × fruticans).
Materials and methods
Plant material
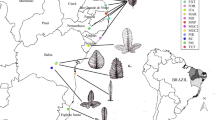
A total of 82 Prunus section Prunus accessions were used. Except for P. cerasifera_COMM01 that was purchased commercially, and P. cocomilia_MEISE1 that originated from Makedonia (Greece) and was provided by the National Botanic Garden of Belgium (Meise), all other plant material belonging to Prunus cerasifera, P. domestica, P. insititia, P. spinosa, and P. × fruticans was sampled on nine locations in three regions of Flanders (Belgium, see Fig. 1) based on the inventories of autochthonous trees and shrubs (Maes 1993; Maes and Rövekamp 1998, 1999; Rövekamp and Maes 2000; Rövekamp et al. 2000; Opstaele 2001; Maes et al. 2003). Field identifications are based on Scholz and Scholz (1995), Körber-Grohne (1996), and Lambinon et al. (1998).
Geographic distribution of the sampling sites (locations) in three regions of Flanders. For each location, the numbers of samples per taxon are mentioned between brackets. Locations: DEF Deftinge, HAA Haaltert, HEE Heers, LOR Lo-Reninge, OUD Oudenaarde, RIE Riemst, SIN Sint-Truiden, TON Tongeren, WES Westouter. Regions: LIM Zuid-Limburg, VAR Vlaamse Ardennen, WVH West-Vlaams Heuvelland
Endocarp and leaf morphometrics
For all individuals, except for P. cerasifera_COMM01 and P. cocomilia_MEISE1 of which no endocarps and no leaves were available, a maximum of five endocarps and five leaves per individual were digitized and dimensions and shape characteristics were analysed (Table 1a, b) with TomatoAnalyzer (Brewer et al. 2006). For visualization and calculation methods of the listed characters, we refer to Brewer et al. (2006). Principal component analysis (PCA) was run several times in PAST (Version 1.82, Hammer et al. 2001) for both the endocarp and the leaf characteristics to determine diagnostic and mutually independent characters. The PCA based on endocarp width (SW), endocarp area (S_Area), endocarp shape triangle (S_Triangle), and endocarp shape circular (S_Circular) and the PCA based on leaf length/leaf width (LL/LW), leaf area (L_Area), leaf shape ellipse (L_Ellipse), and leaf shape ovate (L_Ovate) were most discriminating and were used to visualize and investigate the relationships between the individuals studied. Correlations between endocarp and leaf characteristics were analyzed using the Spearman’s rho correlations coefficient, as most characteristics were not normally distributed.
DNA isolation and AFLP analysis
DNA extraction was performed on 10–20 mg dried leaf tissue grinded with a Retchmill shredder. Total genomic DNA was extracted using the Invisorb® Spin Plant Mini Kit (Invitek, Berlin, Germany). AFLP analysis is performed according to Vos et al. (1995) with minor modifications. Restriction-ligation of the template was performed in one single step, amplification in two steps. Preselective amplification was performed with primers complementary to the adaptors with an extra selective base on each primer (EcoRI-A/MseI-C). Selective amplification, in which primers contained two additional selective nucleotides, was carried out with four primer combinations (EcoRI-ACT/MseI-CTG, EcoRI-ATC/MseI-CAT, EcoRI-ACT/MseI-CAG, EcoRI-ATC/MseI-CAC). After amplification, the DNA fragments were separated on the Global Edition IR² system of LI-COR following the procedure of the Genetic Analysis Manual—Global Edition IR² system (LI-COR). The automatically generated TIFF-files were analyzed in SAGA-MX version 3.0 (LI-COR) according to the standard procedure. A couple of samples were included several times for verification of reproducibility and only the clearly visible and unambiguously scorable bands were used. In total, 103 fragments between 66 and 784 bp were scored. Afterwards the scoring results were transformed into a matrix with values 1 (present), 0 (absent), and 9 (failed or missing). Samples with ≥10 missing data in the binary AFLP matrix were excluded from the analysis.
AFLP-based data analysis
Five members of Prunus section Prunus were analysed using the unweighted pair group method with arithmetic mean (UPGMA, Sokal and Michener 1958) and the Nei and Li (1979) distance estimation in TreeCon (Version 1.3b, Van de Peer and De Wachter 1994). Principal co-ordinate analysis (PCoA; Gower 1966) based on the Dice (1945) similarity was performed in PAST (Version 1.82, Hammer et al. 2001). A Bayesian inference of phylogenetic relationships was performed using Mr. Bayes (Version 3.1, Ronquist and Huelsenbeck 2003) including two independent runs of 120,000,000 generations, each consisting of 10 independent chains, with a temperature of 0.005 for the heated chain, a sample frequency of 5,000, and a burnin of 5,000 samples. In this Bayesian approach, the sole sample of P. cocomilia was set as outgroup. To calculate the correlation between the genetic markers and the morphometric endocarp and leaf features, Manteltests (Mantel 1967) were performed between the genetic distance matrix calculated from all genetic markers and the morphologic distance matrices calculated from either the endocarp or leaf characteristics.
Results
Principal component analysis
In the PCA with endocarp characteristics SW, S_Area, S_Triangle, and S_Circular (represented by the lines in Fig. 2a), the first two principal components explain 78% of the total variation. The first axis allows separation of most of the taxa studied. The characteristics S_Area and SW form the smallest angles with the first principal component, which indicates their high (but negative) correlation with this axis (quantified as correlation coefficients in Table 2a). P. spinosa can be easily distinguished from P. × fruticans by its more circular (i.e. closer to the end point of the line representing S_Circular) and smaller (i.e. further away from the end point of the line visualizing S_Area) endocarps. Separation between endocarps of P. cerasifera, P. domestica, and P. insititia is less obvious. The endocarps of P. cerasifera are located between two small clusters of P. domestica endocarps, whereas samples of P. insititia are spread out over the biplot without forming a clearly delineated group. One sample of P. × fruticans (P. × fruticans_SIN294, arrow in Fig. 2a) clusters together with two P. domestica samples from Sint-Truiden (P. domestica_SIN287 and P. domestica_SIN288).
Principal component analysis of a endocarp morphometrics, and b leaf morphometrics. The first and the second axis account for, respectively, 51 and 27% (a), 54 and 27% (b) of the total variation. Parameter abbreviations are explained in Table 1. Filled triangle: P. cerasifera; filled square: P. domestica; plus symbol: P. insititia; open diamond: P. spinosa; inverted triangle: P. × fruticans
The first two principal components of the PCA with leaf morphometrics L_Area, L_Ellipse, LL/LW, and L_ovate (Fig. 2b), together account for 81% of the total variation. The lines of L_ellipse and LL/LW form the smallest angles and are thus highest correlated with the first axis (quantified in Table 2b). In contrast to the endocarp grouping, separation between individuals of P. spinosa and P. × fruticans is not possible. Individuals of P. insititia are spread out over the biplot without forming a clearly distinct group. But, individuals of P. cerasifera and P. domestica can easily be distinguished based on L_Area and L_ovate. Based on leaf characteristics, P. × fruticans_SIN294 (arrow in Fig. 2b) clusters together with the other P. × fruticans samples.
All dimensional endocarp characteristics (SL, SW, ST, S_Perimeter, S_Area) and the index values 100SW/SL, 100ST/SW are significantly correlated at the 0.05 level with the dimensional leaf characteristics (LW, LL, L_Perimeter, L_Area) and with the leaf shape characteristics LL/LW, L_Ellipse, L_Circular, L_Ovate, and L_Vasym (Table 3). The endocarp shape characteristics S_Circular and S_Rectangular are significantly but lowly correlated at the 0.05 level with the leaf dimensional characteristics.
AFLP polymorphism and cluster analysis
The genetic diversity of the 82 Prunus accessions was analysed using 103 (37 monomorphic and 66 polymorphic) AFLP markers generated from four primer combinations (Table 4). Monomorphic bands were kept in the analysis to avoid an underestimation of the overall similarity between the samples.
In the UPGMA phenogram (Fig. 3), the studied individuals are split up into three major clusters: a first cluster (I) consists of all P. cerasifera samples and the sole P. cocomilia sample; a second cluster (II) includes all individuals of P. domestica and P. insititia; and a third group (III) comprises all P. spinosa and P. × fruticans samples. Within cluster II, samples of P. domestica and P. insititia are mixed up. Analogously, within cluster III, P. spinosa and P. × fruticans samples do not form clearly separate groups. In this last cluster (III), individuals originating from one location tend to group together (small accolades in Fig. 3). Still, for some locations, several distinct clusters can be distinguished. Remarkably, when focusing on each population (location) separately, individuals of P. × fruticans either form a sister group of a P. spinosa cluster (e.g. P. × fruticans_WES078 in the WES1 group and P. × fruticans_WES093 in the WES2 group) or are attributed to a clearly distinct cluster (e.g., the P. × fruticans individuals in the TON1 group). Thus, individuals from the two taxa P. spinosa and P. × fruticans seem to be more related, genetically, to each other when growing together at one location than they are related to their congeners in other growing sites.
UPGMA phenogram showing the genetic relationship among 82 Prunus section Prunus accessions based on AFLP data, using the Nei and Li (1979) distance estimation. Location abbreviations are explained in the caption of Fig. 1. Numbers shown at the different nodes indicate bootstrap confidence values (2,000 bootstrap replicates). Bootstraps values above 30% are indicated on the branches
Principal co-ordinate analysis
The first two eigenvectors of the PCoA plot (Fig. 4) together account for 28% of the total variation and are able to separate clusters I, II, and III as recognized in the cluster analyses. Similar to the cluster analysis, the PCoA plot reveals the close relationship between P. cocomilia and P. cerasifera. Also, P. spinosa and P. × fruticans overlap completely. In contrast, while P. insititia and P. domestica are mixed up in the cluster analysis (Fig. 2), these taxa form more or less distinct groups in the principal coordinate analysis.
PCoA diagram showing the genetic relationship between 82 Prunus accessions using the Dice similarity coefficient. The first and the second axis account for, respectively, 17 and 11% of the variation. Parameter abbreviations are explained in Table 1. Filled triangle: P. cerasifera; open square: P. cocomilia; filled square: P. domestica; plus symbol: P. insititia; open diamond: P. spinosa; inverted triangle: P. × fruticans
The Bayesian approach
In the Bayesian approach, the consensus tree consists of two clearly distinguishable and well-supported clusters (Fig. 5). The first one (A), has a posterior probability of 0.87 and comprises all P. cerasifera individuals. Therefore, it corresponds to cluster I of the UPGMA and principal co-ordinate analyses. The second cluster (B) has a 0.90 posterior probability and encloses cluster II and cluster III of the previous analyses. Except for two P. insititia samples from Tongeren (P. insititia_TON337 and P. insititia_TON341 in the TON3 group), within cluster B, all other P. insititia and P. domestica samples group together (C). Analogous to the cluster analysis, samples of P. spinosa and P. × fruticans are mixed up. Also, in the Bayesian analysis, individuals from one location tend to group together (small accolades in Fig. 5) and individuals of P. × fruticans either form a sister group with the P. spinosa individuals from the same location (e.g., P. × fruticans_WES078 in the WES1 group) or they are attributed to a separate cluster (the P. × fruticans individuals in the TON2 group).
Consensus tree of the Bayesian inference. Location abbreviations are explained in the caption of Fig. 1. Posterior probabilities ≥0.50 are indicated above the branches (as probabilities × 100); clusters I, II, and III correspond with those of the UPGMA and PCoA analyses
Correlation between endocarp morphology and genetics
The distance matrix based on all endocarp characteristics studied is significantly correlated with the genetic matrix based on all AFLP markers (Table 5a). The characteristics Area (0.41), perimeter (0.40), endocarp thickness (ST; 0.35), and endocarp length (SL; 0.30) are highest correlated with the AFLP data. The index values 100SW/SL and 100ST/SL, and characteristics triangle, ellipse, circular, and rectangular are not significantly correlated with the genetic data. The overall leaf morphometry is not significantly correlated with the genetic matrix (Table 5b). However, L_Ovate and L_Circular are significantly correlated with the AFLP markers, together with L_Hasym, LL/LW, and LW.
Discussion
Within Prunus section Prunus, the mutual relationships between P. cerasifera, P. domestica, P. insititia, P. spinosa, and P. × fruticans are far from well resolved. Still, delineation of the different taxa involved and knowledge on their morphological and genetic diversity is indispensable basic information for their proper conservation. As mentioned in the introductory chapter polyploidisation, hybridization, introgression and cultural history have turned the European section Prunus taxa into a complex in which the biological species concept and species relationships become expired. The finding that intra-populational coherence is more pronounced than interspecific coherence, as shown also for dog-roses (subgenus Rosa section Caninae) by De Cock et al. (2008), raises the question whether it is further reasonable to distinguish several species. In their phylogeographic study of North American plums (Prunus section Prunocerasus), Shaw and Small (2005) demonstrated similar high levels of geographically localized sharing of genetic material. The authors conclude: “because of the level of para- and polyphyly observed in many of the taxa in section Prunocerasus and the lack of distinctive morphological characters in the section, we feel that taxonomic “lumping” may be in order.” This may also be the appropriate approach for the Eurasian plum taxa studied here. Similar as in dog-roses (Wissemann and Hellwig 1997; Ritz and Wissemann 2003; De Cock et al. 2008), well-defined species groups are delimited and spontaneous hybrids often are morphologically indistinguishable from their parents. The apparent morphological similarities allow the taxa to be merged into fewer but more diverse species-groups according to the preferences of the taxonomists (De Cock et al. 2008).
As a consequence, nature conservation should focus on preserving these species complexes rather than sticking on to the individual species. Most probably, the evolutionary phenomena of polyploidisation, hybridisation, and domestication led recently to an irreversible and ongoing merging of species, which constrains conservationists to focus on the current patterns instead of aiming to get back to the original situation.
Because of the difficult morphologic delineation of specimens that are intermediate between P. spinosa and P. insititia, the taxonomic status of these intermediates is uncertain (Körber-Grohne 1996; Woldring 2000; Hübner and Wissemann 2004). Some authors consider the intermediates as large-fruited varieties (e.g. Hegi, 1906 ff., cite in Woldring 2000), large-fruited subspecies (e.g. Werneck 1961) or intra-specific hybrids (Fournier 1977) of P. spinosa. In contrast, Webb (1968) regards the intermediate specimens as hybrids of Sloe and Damson plums (P. × fruticans). Other authors (e.g., Peyre 1945; Körber-Grohne 1996) distinguish large-fruited forms of P. spinosa next to P. × fruticans. In our study the intermediates were nominated P. × fruticans. PCA revealed that their endocarps are larger and more elongated than those of P. spinosa. However, leaf morphometrics could not discriminate both taxa. In addition, the genetic analysis could not differentiate them. These findings support the classification of P. × fruticans as large-fruited form of P. spinosa instead of hybrid product of crossings between P. spinosa and P. insititia. However, if P. × fruticans hybrids have originated several times at different locations, they may resemble more their local parents than hybrids of other locations. In both the UPGMA clustering and the Bayesian approach, P. × fruticans individuals of a certain population (location) either clustered as a sister group of P. spinosa individuals at the same location or they were attributed to a clearly distinct cluster. This points to a certain degree of genetic dissimilarity at the local population level. It is known that selection pressure may result in character similarity for evolutionary divergent species adapting to similar conditions, and in striking morphological differences between related species adapting to different conditions (Koopman et al. 2008). Here, we find morphological differences (character differentiation) that are expressed in an AFLP-based approach only at the local population level. Similar growth conditions probably conserved an evolutionary or man-made (domesticated) difference between the two taxa. But, divergent growth conditions (at different sites) resulted in a differentiation in such a way that taxa-specific markers faded. A possibly similar location effect was recently observed in Flemish dog roses (De Cock et al. 2008).
According to Woldring (2000), both the morphological similarities between P. insititia and P. domestica and the occurrence of a wide range of forms with many overlapping features point to a very close relationship. Overlap in characters between P. domestica subsp. insititia (=P. insititia in this study) and related species or subspecies are probably due to centuries of cultivation (Gleason 1958). In our analysis, both taxa exhibited high levels of morphological variation despite the relatively low sample sizes, and a tendency towards morphological differentiation. Still, the AFLP-based UPGMA clustering and the Bayesian inference both attribute individuals of P. insititia and P. domestica to the same cluster. This high genetic similarity is in accordance with the cpDNA studies of Shaw and Small (2004) and Katayama and Uematsu (2005), in which both taxa are shown to have very similar cpDNA patterns. Still, in our principal co-ordinate analysis, the individuals of P. insititia and P. domestica tend to form separate clusters. Even though P. insititia is older than P. domestica, both taxa originated most probably from an identical or very similar ancestral species by human selection and domestication (e.g., Woldring 2000), which may explain their high genetic similarity. These arguments favour inclusion of P. insititia in P. domestica.
Stace (1975) and Zohary (1992) insinuate a close relationship between P. cerasifera and P. domestica as hybrids between both taxa are described. Also, Zohary (1992) states that the 6× P. domestica plums, and particularly subsp. insititia (=P. insititia in this study), are morphologically very similar to P. cerasifera. Our study revealed a high similarity of endocarp morphology between individuals of P. domestica and P. cerasifera. However, leaves of P. domestica are generally larger and differently shaped than those of P. cerasifera.
Recently, Katayama and Uematsu (2005) showed that P. cerasifera belongs to the same cpDNA type as P. domestica and P. insititia, which is in accordance with the hypothesis of a polyploid P. cerasifera origin for P. domestica (including subsp. insititia = P. insititia in this study) (see Decroocq et al. 2004). In contrast, in our study both the cluster analyses and the principal co-ordinate analysis clearly separate the P. cerasifera samples from the P. domestica and P. insititia individuals. Our result corresponds to the findings of Shaw and Small (2004) who illustrated in their cpDNA study that P. domestica and P. insititia share three autapomorphic positions compared to P. cerasifera and P. spinosa. Similarly, Aradhya et al. (2004) demonstrated based on AFLP that within the section Prunus, P. cerasifera and P. spinosa formed (along with P. speciosa and one plum × apricot accession) a separate group beside a cluster consisting of P. domestica. Still, in our study, both the endocarp morphometric and the genetic analyses attribute P. spinosa and P. cerasifera to clearly separated clusters. This is surprising as Reynders and Salesses (1991) showed based on restriction maps of the ribosomal genes that P. spinosa exhibits next to specific units of the species also units identical to those of P. cerasifera.
Our study revealed significant correlations between endocarp and leaf characteristics. Furthermore, some parallel patterns of morphological and genetic diversity were noticeable. Both morphologically and genetically, P. spinosa and P. domestica exhibited clear dissimilarities. However, for the other taxa studied, incongruity existed between the endocarp morphometric and the genetic data. Although genetically identical, P. spinosa and P. × fruticans had differently sized and differently shaped endocarps. This was also true for P. insititia and P. domestica. In contrast, despite the fact that P. cerasifera belonged to a distinct genetic cluster without interference of the other taxa studied, this taxon was hard to separate from P. domestica based on endocarp dimensions and shape characteristics. Notwithstanding these disagreements between endocarp morphometrics and genetic data, we noticed some statistically significant correlations between the observed patterns of genetic diversity and six endocarp morphometric features: area, perimeter, endocarp thickness, endocarp length, endocarp width, and the index value 100ST/SW (100 × endocarp thickness/endocarp width). Remarkably, no correlations were observed between the endocarp shape characteristics (triangle, ellipse, circular, rectangular) and the genetic data. In contrast, next to the LW, also leaf shape (L_Circular, L_Ovate, and L_Hasym) was significantly correlated with the AFLP-based markers. The correlations between morphologic and genetic markers point toward some degree of genetic determinism for the morphological features. However, additional experiments (e.g. mapping studies) are needed to identify specific genes or genome regions that might have a direct influence on the observed morphometric variation.
In conclusion, we divided the European section Prunus taxa P. cerasifera, P. domestica, P. insititia, P. spinosa, and P. × fruticans into three genetic clusters: a first consisting of P. cerasifera (and the very closely related P. cocomilia), a second including P. domestica and P. insititia, and a third comprising P. spinosa and P. × fruticans. Within these three clusters, separation of taxa is less obvious and further research such as hybrid experiments, chromosome counts, and (sequence) analysis of other genetic markers is needed to completely resolve this complex Prunus group. The finding that intra-populational coherence often is more pronounced than interspecific coherence raises the question whether it is further reasonable to distinguish several species or whether instead fewer but more diverse species-groups should be considered.
In contrast to the AFLP-based analyses, multivariate analysis of endocarp morphometrics allows separation of P. spinosa endocarps from those of P. × fruticans. Based on both endocarp and leaf morphometrics, P. insititia individuals do not clearly cluster together with those of P. domestica. Endocarps of P. domestica and P. cerasifera are hard to distinguish based on endocarp dimensions and shape characteristics. Dimensional endocarp features (S_Area, S_Perimeter, SL, ST) are highest and significantly correlated with the genetic data, while for the leaves, especially leaf shape is to a certain extent determined by the genetic data.
References
Aradhya MK, Weeks C, Simon CJ (2004) Molecular characterization of variability and relationships among seven cultivated and selected wild species of Prunus L. using amplified fragment length polymorphism. Scientia Horticulturae 103:131–144
Behre KE (1978) Formenkreise von Prunus domestica L. von der Wikingerzeit bis in die frühe Neuzeit nach Fruchtsteinen aus Haithabu und Alt-Schleswig. Berichte der Deutschen Botanischen Gesellschaft 91:161–179
Bortiri E, Oh S-H, Jiang J, Baggett S, Granger A, Weeks C, Buckingham M, Potter D, Parfitt DE (2001) Phylogeny and systematics of Prunus (Rosaceae) as determined by sequence analysis of ITS and the chloroplast trnL–trnF spacer DNA. Systematic Botany 26(4):797–807
Brewer MT, Lang L, Fujimura K, Dujmovic N, Gray S, van der Knaap E (2006) Development of a controlled vocabulary and software application to analyze fruit shape variation in tomato and other plant species. Plant Physiology 141:15–25
Coart E, Vekemans X, Smulders MJM, Wagner I, Van Huylenbroeck J, Van Bockstaele E, Roldán-Ruiz I (2003) Genetic variation in the endangered wild apple (Malus sylvestris (L.) Mill.) in Belgium as revealed by amplified fragment length polymorphism and microsatellite markers. Molecular Ecology 12:845–857
De Cock K, Vander Mijnsbrugge K, Breyne P, Van Bockstaele E, Van Slycken J (2008) Morphological and AFLP-based differentiation within the taxonomical complex section Caninae (subgenus Rosa). Annals of Botany 102(5):685–697
Decroocq V, Hagen LS, Favé M-G, Eyquard J-P, Pierronnet A (2004) Microsatellite markers in hexaploid Prunus domestica species and parentage lineage of three European plum cultivars using nuclear and chloroplast simple-sequence repeats. Molecular Breeding 13:135–142
Depypere L, Chaerle P, Vander Mijnsbrugge K, Goetghebeur P (2007) Stony endocarp dimension and shape variation in Prunus L. section Prunus. Annals of Botany 100:1585–1597
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Drossou A, Katsiotis A, Leggett JM, Loukas M, Tsakas S (2004) Genome and species relationships in genus Avena based on RAPD and AFLP molecular markers. Theoretical and Applied Genetics 109:48–54
Fournier P (1977) Les Quatres Flores de la France. Lechevalier, Paris
Frary A, Doğanlar S (2003) Comparative genetics of crop plant domestication and evolution. Turk J Agric For 27:59–69
Gleason HA (1958) Illustrated flora, vol 2. Lancaster Press, Lancaster
Gower JC (1966) Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53:325–338
Hagen LS, Khadari B, Lambert P, Audergon J-M (2002) Genetic diversity in apricot revealed by AFLP markers: species and cultivar comparisons. Theoretical and Applied Genetics 105:298–305
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4(1):9 pp. http://palaeo-electronica.org/2001_1/past/issue1_01.htm
Hanelt P (1997) European wild relatives of Prunus fruit crops. Bocconea 7:401–408
Hübner S, Wissemann V (2004) Morphometrische Analysen zur Variabilität von Prunus spinosa L.—Populationen (Prunoideae, Rosaceae) im Mittleren Saaletal, Thüringen. Forum Geobotanicum 1:19–51
Jones CJ, Edwards KJ, Castaglione S, Winfield MO, Sala F, Van De Wiel C, Bredemeijer G, Vosman B, Matthes M, Daly A, Brettschneider R, Bettini P, Buiatti M, Maestri E, Malcevschi A, Marmiroli N, Aert R, Volckaert G, Rueda J, Linacero R, Vazquez A, Karp A (1997) Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Molecular Breeding 3:381–390
Kalkman C (2004) Rosaceae. In: Kubitzki K (ed) The families and genera of vascular plants, vol VI. Springer, Berlin
Katayama H, Uematsu C (2005) Structural analysis of chloroplast DNA in Prunus (Rosaceae): evolution, genetic diversity and unequal mutations. Theoretical and Applied Genetics 111:1430–1439
Koopman WJM, Zevenbergen MJ, Van den Berg RG (2001) Species relationships in Lactuca s.l. (Lactuceae, Asteraceae) inferred from AFLP fingerprints. American Journal of Botany 88:1881–1887
Koopman WJM, Wissemann V, De Cock K, Van Huylenbroeck J, De Riek J, Sabatino GJH, Visser D, Vosman B, Ritz CM, Maes B, Werlemark G, Nybom H, Debener T, Linde M, Smulders MJM (2008) AFLP markers as a tool to reconstruct complex relationships: a case study in Rosa (Rosaceae). American Journal of Botany 95(3):353–366
Körber-Grohne U (1996) Pflaumen, Kirschpflaumen, Schlehen—Heutige Pflanzen und ihre Geschichte seit der Frühzeit. Thesis, Stuttgart
Krüssmann G (1978) Prunus. In: Handbuch der Laubgehölze, Band III. Verlag Paul Parey, Berlin, pp 13–56
Kühn F (1999) Alte Pflaumen, lebende Zeugen mittelalterlichen Obstbaus. Hamburger Werkstattreihe zur Archäologie 4:70–77
Lambinon J, De Langhe J-E, Delvosalle J, Duvigneaud J (1998) Flora van België, het Groothertogdom Luxemburg, Noord-Frankrijk en de aangrenzende gebieden (Pteridofyten en Spermatofyten), derde druk. Uitgave van de Nationale Plantentuin van België, Meise, pp 1091
Mabberley DJ (2008) Mabberley’s plant-book. A portable dictionary of plants, their classification and uses, 3rd edn. Cambridge University Press, Cambridge
Mace ES, Gebhardt CG, Lester RN (1999a) AFLP analysis of genetic relationships in the tribe Datureae (Solanaceae). Theoretical and Applied Genetics 99:634–641
Mace ES, Lester RN, Gebhardt CG (1999b) AFLP analysis of genetic relationships among the cultivated eggplant, Solanum melongena L., and wild relatives (Solanaceae). Theoretical and Applied Genetics 99:626–633
Maes N (1993) Genetische kwaliteit van inheemse bomen en struiken. Deelproject: Randvoorwaarden en knelpunten bij behoud en toepassing van inheems genenmateriaal. IBN-rapport 20, IKC-NBLF, IBN-DLO, Wageningen
Maes N, Rövekamp C (1998) Oorspronkelijk inheemse bomen en struiken in Vlaanderen. Een onderzoek naar autochtone genenbronnen in de Ecologische Impulsgebieden. Ministerie van de Vlaamse Gemeenschap, Afdeling Bos en Groen, Brussel
Maes B, Rövekamp C (1999) Cursus autochtone bomen en struiken Vlaanderen (een productie van het Ekologisch Adviesburo Maes en BRONNEN). Educatief bosbouwcentrum Groenendaal, Brussel
Maes N, Rövekamp C, Opstaele B, Zwaenepoel A (2003) Oorspronkelijke inheemse bomen en struiken in de houtvesterijen Antwerpen en Turnhout. Ministerie van de Vlaamse Gemeenschap, afdeling Bos en Groen, Brussel
Mantel NA (1967) The detection of disease clustering and a generalized regression approach. Cancer Research 27:209–220
Mason SC (1913) The pubescent-fruited species of Prunus of the Southwestern States. J Agr Res 1:147–179
McGregor CE, Lambert CA, Greyling MM, Louw JH, Warnich L (2000) A comparative assessment of DNA fingerprinting techniques (RAPD, ISSR, AFLP and SSR) in tetraploid potato (Solanum tuberosum L.) germplasm. Euphytica 113:135–144
Mowrey BD, Werner DJ (1990) Phylogenetic relationships among species of Prunus as inferred by isozyme markers. Theoretical and Applied Genetics 80:129–133
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. In: Proceedings of the National Academy of Sciences of the United States of America, vol 76, pp 5269–5273
Nielsen J, Olrik DC (2001) A morphometric analysis of Prunus spinosa, P. domestica ssp. insititia, and their putative hybrids in Denmark. Nordic Journal of Botany 21(4):349–363
Opstaele B (2001) Autochtone bomen en struiken in de houtvesterijen Leuven en Hasselt. Ministerie van de Vlaamse Gemeenschap, afdeling Bos en Groen, Brussel
Peyre P (1945) Les pruniers sauvages cultivés. Clermont-Ferrand, Paris
Rehder A (1940) Manual of cultivated trees and shrubs hardy in North America, 2nd edn. The Macmillan Company, New York
Reynders S, Salesses G (1991) Study of the genetic relationships within the subgenus Prunophora. Restriction maps of the ribosomal genes in P. cerasifera and P. spinosa. Acta Horticulturae 283:17–25
Ritz CM, Wissemann V (2003) Male correlated non-matroclinal character inheritance in reciprocal hybrids of Rosa section Caninae (DC.) Ser. (Rosaceae). Plant Systematics and Evolution 241:213–221
Röder K (1940) Sortenkundliche Untersuchungen an Prunus domestica. Kühn Archiv 54:1–132
Ronquist F, Huelsenbeck JP (2003) MRBAYES: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rövekamp C, Maes N (2000) Oorspronkelijk inheemse bomen en struiken in het Regionaal Landschap West-Vlaamse Heuvels. Een onderzoek naar autochtone genenbronnen. Ministerie van de Vlaamse Gemeenschap, Afdeling Bos en Groen, Brussel
Rövekamp C, Maes N, Zwaenepoel A (2000) Oorspronkelijk inheemse bomen en struiken en cultuurwilgen in de Vlaamse Vallei. Een onderzoek naar autochtone genenbronnen in Gent en omgeving. Brussel, Ministerie van de Vlaamse Gemeenschap, Afdeling Bos en Groen
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theoretical and Applied Genetics 95:714–722
Schenk MF, Thienpont C-N, Koopman WJM, Gilissen LJWJ, Smulders MJM (2008) Phylogenetic relationships in Betula (Betulaceae) based on AFLP markers. Tree Genetics and Genomes 4:911–924
Scholz H, Scholz I (1995) Prunoideae. In: Conert HJ, Jäger EJ, Kadereit JW, Schultze-Motel W, Wagenitz G, Weber HE (eds) Gustav Hegi’s Illustrierte Flora von Mitteleuropa, Band IV Teil 2B. Blackwell Wissenschafts-Verlag, Berlin, pp 446–510
Shaw J, Small RL (2004) Addressing the “hardest puzzle in American pomology:” Phylogeny of Prunus sect. Prunocerasus (Rosaceae) based on seven noncoding chloroplast DNA regions. American Journal of Botany 91:985–996
Shaw J, Small RL (2005) Chloroplast DNA phylogeny and phylogeography of the North American plums (Prunus subgenus Prunus section Prunocerasus, Rosaceae). American Journal of Botany 92:2011–2030
Sokal RR, Michener CD (1958) A statistical method for evaluating systematic relationships. University of Kansas Science Bulletin 28:1409–1438
Stace CA (1975) Hybridization and the Flora of the British Isles. Academic Press, London
Thórsson ÆT, Salmela E, Anamthawat-Jónsson K (2001) Morphological, cytogenetic, and molecular evidence for introgressive hybridization in birch. Journal of Heredity 92:404–408
Van de Peer Y, De Wachter R (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Computer Applications in Biosciences 10:569–570
Van Droogenbroeck B, Breyne P, Goetghebeur P, Romeijn-Peeters E, Kyndt T, Gheysen G (2002) AFLP analysis of genetic relationships among papaya and its wild relatives (Caricaceae) from Ecuador. Theoretical and Applied Genetics 105:289–297
Van Zeist W, Woldring H (2000) Plum (Prunus domestica L.) varieties in late- and post-medieval Groningen: the archaeobotanical evidence. Palaeohistoria 39(40):563–576
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Research 23:4407–4414
Webb DA (1968) Prunus. In: Tutin TG, Heywood VH, Burges NA, Moore DM, Valentine DH, Walters SM, Webb DA (eds) Flora Europaea. Rosaceae to Umbelliferae, vol 2. Cambridge University Press, Cambridge
Werneck HL (1961) Die Wurzel- und Kernechten Stammformen der Pflaumen in Oberösterreich. Naturkundliches Jahrbuch der Stadt Linz
Wissemann V, Hellwig FH (1997) Reproduction and hybridisation in the genus Rosa, section Caninae (Ser.) Rehd. Botanica Acta 110:251–256
Woldring H (2000) On the origin of plums: a study of sloe, damson, cherry plum, domestic plums and their intermediates. Palaeohistoria 39(40):535–562
Zohary D (1992) Is the European plum, Prunus domestica L., a P. cerasifera Ehrh. × P. spinosa L. allo-polyploid? Euphytica 60:75–77
Zohary D, Hopf M (1994) Domestication of plants in the Old World, 2nd edn. Clarendon press, Oxford, p 279
Acknowledgments
Research funded by a PhD grant of the Institute for the Promotion of Innovation through science and Technology in Flanders (IWT-Vlaanderen). We are grateful to Stefaan Moreels and Leen Verschaeve for their assistance during sampling, Tine Paessens, Joel Dockx, Leen Verschaeve, David Halfmaerten, and Tina Kyndt for guidance with DNA isolation and/or AFLP analysis, Bert Maes for help with the plant identifications.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Depypere, L., Chaerle, P., Breyne, P. et al. A combined morphometric and AFLP based diversity study challenges the taxonomy of the European members of the complex Prunus L. section Prunus . Plant Syst Evol 279, 219–231 (2009). https://doi.org/10.1007/s00606-009-0158-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-009-0158-8