Abstract
Eight variable regions (microsatellites, insertion/deletion and duplication regions) from the plastid DNA genome were analyzed for 91 populations belonging to Dactylorhiza majalis ssp. traunsteineri and closely related taxa. A total of 36 composite plastid haplotypes were found. The two dominating haplotypes had a clear geographic distribution suggesting at least two separate immigration routes into Scandinavia after the last glaciation: one southwestern route and one or two southeastern routes. D. majalis ssp. traunsteineri could not be clearly separated from any of the other taxa included in the study except for D. majalis ssp. sphagnicola. The morphologically similar taxa D. majalis ssp. traunsteineri, D. majalis ssp. lapponica and D. majalis ssp. russowii showed no genetic differentiation, and therefore we suggest an amalgamation of the three taxa into one broadly circumscribed subspecies; D. majalis ssp. lapponica. The plastid data also revealed incidents of hybridization and possible introgression between D. majalis ssp. lapponica and other members of the genus, e.g., D. incarnata.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fennoscandia was covered with ice during the last glaciation (c. 22,000 to 17,000 cal yrs BP) and was not completely ice-free until approximately 8,000 years ago. Gradually, plants hibernating in refugial areas outside the ice sheet recolonized the open habitats left by the ice. Immigration routes and histories of Fennoscandian populations have been described for various species (e.g., Picea abies in Lagercrantz and Ryman 1990; Picea abies in Kullman 1996; Silene dioica in Malm and Prentice 2002; Calluna vulgaris in Rendell and Ennos 2002; Carex digitata and Melica nutans in Tyler et al. 2002; Betula pendula in Palmé et al. 2003; Dactylorhiza maculata s.l. in Ståhlberg 2007). Populations in previously glaciated areas may be genetically depleted as a consequence of repeated bottlenecks during stepwise migration, a pattern that has been described for several species of deciduous forest trees (e.g., Ferris et al. 1998; King and Ferris 1998). Such species seem to have been located in distant southern refugia during the last ice age. Plants with a more temperate distribution have, however, been shown to be equally genetically variable in Fennoscandia as populations of the same taxon in other parts of Europe (Tyler et al. 2002; Borgen and Hultgård 2003; Palmé et al. 2003; Skrede et al. 2006; Ståhlberg 2007). These temperate species may have had refugia close to the ice sheet (Stewart and Lister 2001) and seem to have kept most of their genetic variation during the migration process. Additionally, many plant species have colonized Fennoscandia through more than one immigration route (Hultén 1950; King and Ferris 1998; Nordal and Jonsell 1998; Malm and Prentice 2005; Ståhlberg 2007), which may have further contributed to comparatively high levels of genetic diversity.
Another factor affecting the genetic diversity of immigrated temperate species is the occurrence of polyploidy. Polyploid plant species are thought to increase in number with latitude in the Northern Hemisphere (Löve and Löve 1974; Grant 1981; Otto and Whitton 2000; Brochmann et al. 2004) and, for example, 44% of the plant taxa in boreal areas of the Arctic are polyploids compared to 82% in the polar desert further north (Brochmann et al. 2004). Several studies have shown that polyploids have higher levels of genetic diversity than their related diploids (e.g., Soltis and Rieseberg 1986; Lumaret and Barrientos 1990; Luttikhuizen et al. 2007) and due to their multiple chromosome complements they might have a higher potential for storing their genetic diversity during repeated bottle-neck episodes associated with recolonization of previously glaciated areas.
Dactylorhiza Necker ex Nevski (Orchidaceae) is an example of a temperate plant genus with members of different ploidy levels. It is dominated by a polyploid complex consisting of diploid and tetraploid taxa. The complex must have originated well before the Weichselian glaciation (Hedrén et al. 2007) and is now found in large parts of Europe and Asia Minor (Pridgeon et al. 2001; Delforge 1995). During the ice ages, the Balkans (Hedrén et al. 2007), central Europe, and parts of central Russia (Ståhlberg 2007) acted as the most important refugia for the complex. Subsequent migration to formerly glaciated areas probably happened rapidly due to efficient seed dispersal by small seeds (Dressler 1993) and availability of suitable habitats (Adams 1997; Adams and Faure 1997).
D. maculata (L.) Soó s.l. and D. incarnata (L.) Soó s.l. are the present-day representatives of the parental lineages that built up the polyploid complex of Dactylorhiza in Europe (Hedrén 1996). Repeated hybridizations between the parental lineages have given rise to several allotetraploid derivatives (Hedrén 2003). This reticulate evolution has yielded a large amount of morphological variation in the complex which makes species delimitation difficult. Hybridization between present day representatives of the complex has often been considered as an additional factor contributing to further taxonomic confusion (Mossberg and Nilsson 1987; Baumann and Künkele 1988; Delforge 2001). Moreover, it could be discussed whether morphologically distinct populations represent independent evolutionary lineages arising from separate polyploidization events, or they have differentiated from widespread taxa as a consequence of genetic drift or selection. In the former case it may be motivated to recognize deviating forms as species, whereas in the latter case they may merely be seen as slightly deviant local forms without taxonomic value. Depending on underlying principles of species delimitation, different authors have recognized between one (Pedersen et al. 2003) and 23 (Delforge 1995) tetraploid species. Here we mainly follow the taxonomic delimitation of taxa as in Delforge (2001) but treat all allotetraploid taxa as subspecies of D. majalis (Rchb.) P. F. Hunt & Summerh. as suggested by Pedersen et al. (2003). One of the most common and variable allotetraploid is D. majalis (Rchb.) P. F. Hunt & Summerh. ssp. traunsteineri (Saut.) H. Sund., which was originally described on basis of material collected at Schwarzsee, Kitzbühel, Austria. Populations referable to this taxon have subsequently been reported from a wide area of northern Europe, including the Fennoscandian-Baltic region and the British Isles, although the taxonomic status of material from these latter regions has often been discussed and there is no consensus among present authors. It is characterized by having bluish-green leaves that are usually spotted on the upper surface, acute leaf apex, and a few-flowered, lax inflorescence with fairly large purple flowers provided with a prominent median lip lobe (Delforge 2001; Mossberg and Stenberg 2003). The value of leaf spotting as key character has however been questioned as it often varies within and between populations (Hylander 1966). D. traunsteineri has repeatedly been divided into subspecies and varieties (Hylander 1966; de Soó 1980) and morphologically distinct forms from certain areas have often been recognized (Hylander 1966; Hansson 1994; Andersson 1995). In the Fennoscandian-Baltic area two further taxa have been recognized that are clearly linked to D. majalis ssp. traunsteineri; D. majalis (Rchb.) P. F. Hunt & Summerh. ssp. lapponica (Laest.) H. Sundermann (Hylander 1966; Senghas 1968; de Soó 1980; Andersson 1996) and D. majalis (Rchb.) P. F. Hunt & Summerh. ssp. russowii (Klinge) H. Sund. (Senghas 1968; Mossberg and Nilsson 1987; Andersson 1994). D. majalis ssp. lapponica has a northern and alpine distribution whereas D. majalis ssp. russowii can be found in the Baltic countries and in Russia. The three taxa are not only morphologically similar but also grow in similar habitats. For instance, both D. majalis ssp. traunsteineri from southern Sweden and ssp. russowii from the East of the Baltic sea are found in calcareous fens growing in association with, e.g., Carex lepidocarpa Tausch ssp. lepidocarpa, Schoenus ferrugineus L., Primula farinosa L. and Epipactis palustris Crantz. The above-mentioned members of Dactylorhiza are the central focus for this study and subsequently denoted “the core complex”. The core complex may sometimes be hard to delimit from other allotetraploid taxa in the Fennoscandian-Baltic area. For example, D. majalis (Rchb.) P. F. Hunt & Summerh. ssp. sphagnicola (Höppner) H. A. Pedersen & Hedrén (Hylander 1966; Ekman 1985; Bjurulf 2005) has some flower characters in common with ssp. traunsteineri but is normally found in poor fens and is not associated with indicator species of calcareous fens (Mossberg and Nilsson 1987; Baumann et al. 2006).
As mentioned above several morphological studies have been made to clarify the relationships between the different taxa in the core complex but no fine scale genetic analyses have been performed up to now. To be able to elucidate variation patterns and taxonomic limits within this group of closely related taxa we used fast-evolving plastid markers of microsatellite type for this study. The plastid genome is uniparentally inherited (presumably maternally in orchids, Corriveau and Coleman 1988; Cafasso et al. 2005) and markers thereof are commonly used in phylogeographic and phylogenetic studies (summarized in Lowe et al. 2004). Another important aspect of plastid markers, especially when studying allopolyploids, are the infrequence of recombination (Wolfe and Randle 2004), a problem associated with data sets from the nuclear genome.
The specific aims of this study were (1) to describe the genetic differentiation within the core complex in Fennoscandia and the Baltic area, (2) to describe limits between the core complex and other allotetraploid members of Dactylorhiza occurring in the area, and (3) to describe geographic variation patterns and relate these patterns to potential recolonization routes into Scandinavia after the last ice age.
Materials and methods
Plant material
Six hundred and fifty individuals from 91 populations were included in this study (Appendix). Two or more floral buds or apparently un-pollinated flowers with bracts were collected from each specimen and immediately dried in silica gel (Chase and Hills 1991). A majority of the populations (79) belonged to the core complex (D. majalis ssp. traunsteineri, ssp. russowii and ssp. lapponica) and the sampling covered most of its distribution in Fennoscandia (mainly Sweden, Norway and Finland) and Estonia. Additionally, five populations of D. majalis ssp. traunsteineri were sampled in Russia, Lithuania and Austria. One of the Austrian populations was from the type locality of ssp. traunsteineri near Kitzbühel. We did not separate out the subordinate taxon sometimes associated with D. majalis ssp. traunsteineri, ssp. curvifolia, but plants in agreement with this form were included from populations in both Finland and Sweden.
Most of the material in the core complex was easy to identify whereas some populations were more or less intermediate between the three subspecies. Accordingly, there are both typical D. majalis ssp. traunsteineri populations and ambiguous populations within the core complex included in this study. Moreover, 11 populations from Norway and middle Sweden were classified as D. majalis ssp. traunsteineri in local floras or by field botanists, but approached ssp. sphagnicola in our opinion. Morphologically, the two subspecies can be separated by differences in the spur: ssp. sphagnicola is characterized by a narrow, cylindrical and decumbent spur, whereas ssp. traunsteineri is characterized by a slightly wider, conical and straight spur (Delforge 2001; Bjurulf 2005; Baumann et al. 2006). The ambiguity of the populations motivated the inclusion of some populations of D. majalis ssp. sphagnicola as reference material. As further reference material, populations of all the remaining allotetraploid species of Fennoscandia were included: D. majalis (Rchb.) P. F. Hunt & Summerh. sspp. majalis, praetermissa (Druce) D. M. Moore & Soó, purpurella (T. Stephenson & T. A. Stephenson) D. M. Moore & Soó and baltica (Klinge) H. Sund. Earlier studies (Devos et al. 2003; Hedrén 2003; Shipunov et al. 2005; Pillon et al. 2007) of plastid DNA variation in Dactylorhiza have shown that allotetraploid Dactylorhiza have inherited their plastid genomes from D. maculata s.l, which means that D. maculata s.l and not D. incarnata s.l. is the seed parent of the allotetraploids. If more recent hybridization between an allotetraploid and D. incarnata has occurred, and D. incarnata has served as the seed parent, the D. incarnata plastid haplotype can be seen in the allotetraploid, but probably only in a fraction of plants within populations. In order to discover such secondary hybridization we also included one population of D. incarnata as a reference. The taxonomic classifications of the plant material in this study were based on scientific floras (Hylander 1966; de Soó 1980), field floras (Krok and Almquist 1994; Mossberg and Stenberg 2003) and local expertise. Voucher material of all populations (dried flowers) has been deposited at the Lund botanical museum (LD).
Molecular methods
Total DNA was extracted by the CTAB (cetyltrimethyl ammonium bromide) method (Doyle and Doyle 1990). Eight polymorphic plastid loci were analyzed for each sample. Five of the loci were mononucleotide repeats, two were duplicated regions and one was an insertion/deletion region. All loci were selected because of their high polymorphism for the genus, (previously tested by Hedrén et al. unpublished). All the alleles were defined according to size and combined into haplotypes. Detailed information of the loci and primers is given by Table 1. PCR conditions for all fragments were as follows: 5.6 ng DNA was amplified using 0.26 μM Cy5 labeled primer, 0.26 μM unlabeled primer, 1.6 mM MgCl2, 210 μM dNTP’s, 1× PCR buffer (Applied Biosystems) and 0.12 U Amplitaq Gold polymerase (Applied Biosystems), in a total volume of 5 μL. The PCR profile was: 94°C 1 min, T a (Table 1) 1 min, 72°C 1 min 30 s for 40 cycles. PCR fragments were labeled and separated on an ALF Express II automated sequencer (Amersham Biosciences). Size determination was performed using ALFwin Fragment Analyser 1.03.01 software.
Data analysis
Population differentiation patterns were summarized in two multidimensional scaling diagrams (MDS; Kruskal 1964a, b) using the computer program NT-SYSpc 2.2 (Rohlf 2005). In both calculations Nei’s average number of differences between populations (Nei and Li 1979) were calculated between all pairs of populations in the computer program Arlequin 3.01 (Excoffier et al. 2005) and used as input matrix for the multidimensional scaling analysis.
Relationships between all haplotypes were illustrated in a Median-joining network assisted by the program NETWORK 4.5 (Bandelt et al. 1999) with all options set to default.
To test whether genetic distances were correlated to geographic distances between populations a Mantel test (Mantel 1967) was performed using NT-SYSpc 2.2 (Rohlf 2005). A geographic distance matrix based on Euclidian distances was compared with a genetic distance matrix based on Nei’s average number of differences between populations (Nei and Li 1979). The geographically distant Russian and Austrian populations were excluded from the test, as we were primarily interested in the correlation between genetic and geographic distances in the Scandinavian/Baltic area.
Results
Characterization of haplotypes
Using eight variable loci, 36 haplotypes were found among the 650 analyzed individuals (annotated H1–H36; Table 2). Two of the haplotypes (H1 and H4) counted for more than 60% of the individuals and 13 of the haplotypes (H1, H2, H4–H8, H10–H12, H17, H18, and H21) counted for more than 91% of the individuals. The relationships between haplotypes and their relative frequencies can be seen in the haplotype network (Fig. 1). Three distinct haplotype groups appeared in the network. The first group, denoted group I, included most of the haplotypes and was divided into two subgroups, dominated by haplotypes 1 and 4, respectively. The second group, to the left in Fig. 1, consisted mainly of haplotype 6 to haplotype 10 and was denoted group II. In the right end of the network several rare haplotypes, e.g., H11 and H12, clustered into the third group, denoted incarnata group. Haplotype 1 was the most common haplotype and a lot of rare haplotypes were connected to it, e.g., H15, H22 and H23. In contrast, haplotype 4, which was the second most common haplotype, only connected to a few rare haplotypes, e.g., H20. Haplotype 21 had a somewhat distant position in relation to the other haplotypes of the second group with only haplotype 13 connected to it.
Population differentiation
The MDS diagram based on average pairwise differences between haplotypes and including all populations is given as Fig. 2. Three main clusters appeared in the MDS. The cluster to the upper left included the reference population of D. incarnata and five allotetraploid populations fixed for incarnata haplotypes (H11 and H12). Less than four samples were examined from any of these allotetraploid populations. The cluster to the lower left contained the three reference populations of D. majalis ssp. sphagnicola and eight other allotetraploid populations dominated by group II haplotypes. The larger cluster to the lower right contained more than 60% of the examined populations. Populations observed in between the three main clusters were either constituted by two or more different haplotypes from the three clusters or carried some of the odd haplotypes described below. Two dense subclusters that appeared in the large cluster to the lower right were included in a separate MDS given as Fig. 3. Haplotype 1 was prevalent in the populations in the left half of the diagram in Fig. 3, whereas haplotype 4 dominated to the right.
Multidimensional scaling analysis of populations of the right cluster of Fig. 2. Only populations of the core complex are included. Stress = 0.10
Taxonomic differentiation in haplotypes
The MDS diagram of all populations (Fig. 2) revealed no separation of populations labeled as D. majalis ssp. traunsteineri from other allotetraploid populations. The D. majalis ssp. traunsteineri populations were evenly distributed along the axes and were interspersed by other taxa. Other members of the core complex were also not distinct. The populations of D. majalis ssp. lapponica did not form a separate group in any of the MDS ordinations (Figs. 2, 3), although all populations were included in the right cluster of Fig. 2. The same result appeared for the ssp. russowii populations (Figs. 2, 3). Similarly, the three D. majalis ssp. majalis populations as well as the two populations of ssp. purpurella were located among ssp. traunsteineri populations (Figs. 2). However, the reference population of D. majalis ssp. praetermissa was located below the right cluster and the populations of ssp. baltica were placed within the upper left cluster or half-way between this cluster and the right cluster (Fig. 2). All the populations of D. majalis ssp. sphagnicola were situated in the lower left cluster in Fig. 2. This cluster also contained the 11 ambiguous populations with uncertain affinity to D. majalis ssp. traunsteineri from Norway and middle Sweden (populations 24, 26, 28, 29–32, 42, 49–51), which all carried some of the typical ssp. sphagnicola haplotypes from haplotype group II. The three populations of D. majalis ssp. majalis had different haplotypes including H4, H21 and H25. Haplotype 21 was also present in four populations of D. majalis ssp. traunsteineri and two of ssp. lapponica, all from the mixed region in Northern Lapland. Haplotype 25 is a rare haplotype unique to one of the D. majalis ssp. majalis populations and has not been found elsewhere. The single population of D. majalis ssp. praetermissa from Denmark contained the unique haplotype H13.
Geographic variation within the core complex
The MDS ordination given as Fig. 3 was based on 52 populations of the core complex. (All the populations in the two groups of the lower right cluster in Fig. 2 plus the 21 adjacent populations, populations of D. majalis ssp. praetermissa and ssp. majalis were excluded). Groups separated along the horizontal axis had different geographic origins. Notably, the left side contained all populations from the southern Swedish mainland whereas all populations from Gotland but one were located on the right side. The right group was more geographically widespread than the left and contained populations from Russia, Austrian Alps and all but one of the Finnish populations.
Some populations were located between the main clusters, including populations from Finland, Sweden and Estonia. The Swedish populations were all from north of the southern border of Lapland (64°15′N) and the Finnish populations were north of the Gulf of Bothnia.
The geographic pattern seen in Fig. 3 was also evident in Fig. 4a, where core complex populations containing haplotypes 1 and 4 were marked:
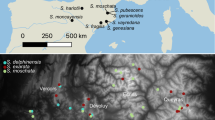
Geographic distribution of haplotypes. a Core complex populations containing haplotype 1 and/or 4. Black symbols populations containing haplotype 4. White symbols populations containing haplotype 1. Grey symbols populations with individuals carrying haplotype 1 as well as individuals carrying haplotype 4. b Populations containing haplotype 10. Black symbols populations carrying haplotype 10. c Populations containing group II haplotypes. Black symbols populations totally dominated by individuals with group II haplotypes (100% carry the haplotypes). Grey symbols populations with a lesser content of individuals carrying group II haplotypes (less than 50% carry the haplotypes). White symbols populations with no individuals of group II haplotypes. Populations 68, 70 and 71 are allopatric D. majalis ssp. sphagnicola populations, whereas the other populations with black symbols are identified as D. majalis ssp. traunsteineri populations
Haplotype 1 was totally dominating on the Swedish mainland apart from northern Scandinavia. It was not found in central Finland and only rarely encountered in Norway. On Gotland we only found two individuals containing haplotype 1 in one single population (out of seven populations examined). This haplotype was also present in Estonia and Lithuania. Haplotype 4 had a more eastern distribution and was most common in Finland and on Gotland. Additionally, it was found in Estonia and Austria. There were also populations from northern Sweden and northern Norway containing haplotype 4 and the haplotype could be found as far south as southern Lapland. This distribution of haplotypes created a mixed zone in northern Fennoscandia where both haplotypes 1 and 4 were present and sometimes observed in the same populations. There were other meeting zones of the two haplotypes in Estonia (Hiiumaa and Saaremaa) and on Gotland (Sweden). Only one population (D. majalis ssp. traunsteineri) fixed for haplotype 1 was found on Gotland while two such populations were encountered in Estonia (ssp. russowii). Moreover, there was a population of D. majalis ssp. russowii on Saaremaa carrying this haplotype together with haplotype 4.
The map given as Fig. 4b shows the distribution of a less common haplotype, H10, which was found in D. majalis ssp. traunsteineri populations on Gotland and in ssp. russowii populations on Saaremaa and Hiiumaa. Haplotype 10 was strongly divergent from the other haplotypes encountered in the core complex and most closely related to the D. majalis ssp. sphagnicola haplotypes (Fig. 1). Except for Gotland and Estonia it was also found in Austria.
Almost all populations initially labeled as D. majalis ssp. traunsteineri from Norway and four populations from central Sweden were fixed for typical D. majalis ssp. sphagnicola haplotypes (Fig. 4c).
A weak but highly significant positive correlation, 0.18975, was observed between geographic and genetic distances among populations from the Fennoscandic-Baltic area (Mantel test; p < 0.001).
Discussion
Haplotype evaluation
Plastid markers are commonly used in phylogeographic studies due to their relatively slow mutation rate and uniparental origin. However, for the same reasons, plastid markers may not be suitable for distinguishing between closely related taxa. Introgression of plastid DNA as a result of interspecific hybridization may further complicate phylogeographic studies based on plastid data. However, the mutation rate may differ substantially between plastid microsatellite loci and when combining several loci with different mutation rates into combined haplotypes a more structured pattern may appear. Furthermore, the fact that orchids have very small seeds and thus have the capacity to spread over long distances suggests that the gene flow by seeds may be more important than pollen flow (discussed in Cozzolino et al. 2003). In this study, the eight chloroplast markers proved to be very informative when answering our questions on both taxonomy and phylogeography.
Thirty-six haplotypes were found in the distribution area out of which two (H1 and H4) were present in more than 60% of the samples. Despite the high frequencies of these two haplotypes, we found variation within populations and often variation within a small defined area.
Gene flow by hybridization
Most allotetraploid populations in this study had plastid haplotypes previously characterized as typical D. maculata s.l. haplotypes. This finding confirms the common notion that D. maculata s.l. has acted as donor of the plastid genome at the polyploidization event (Devos et al. 2003; Hedrén 2003; Shipunov et al. 2005; Pillon et al. 2007). Further, the finding that two clearly defined and large subgroups of haplotypes dominate in the core complex (characterized by H1 and H4, respectively) supports earlier results suggesting multiple origins of allotetraploids in Dactylorhiza (Hedrén 1996, 2003; Devos et al. 2006). However, no present day representatives of D. maculata s.l. carrying haplotype 4 have been found in spite of a large sample surveyed from the major part of the European distribution area (Ståhlberg 2007), and a D. maculata s.l. ancestor carrying this haplotype should have been found if it still existed. The high number of allotetraploid populations carrying haplotype 4 and the wide distribution of this haplotype in the allotetraploids suggest that haplotype 4 might be an old haplotype originating well before the last glaciations and that different allotetraploid taxa carrying this haplotype share a common history. The alternative explanation to the observed pattern is that haplotype 4 has differentiated from another allotetraploid haplotype, for example haplotype 1 which is also widespread in present day D. maculata s.l. However, 4 to 12 mutations at four loci out of eight studied is required for that conversion and we consider this a less likely scenario.
Some of the allotetraploid populations had haplotypes typical of D. incarnata (Fig. 2). It is hypothetically possible that these populations have inherited the incarnata plastids at some polyploidization events. However, since the number of individuals carrying the incarnata haplotypes is low, only eight individuals in altogether five populations, the presence of incarnata haplotypes rather indicate backcrossing between the allotetraploids and D. incarnata at a later stage. Introgression with D. maculata s.l. may also take place, but is more difficult to detect since the allotetraploids normally inherit the plastids from this lineage. Nevertheless, the traunsteineri population 78 in southern Sweden may show such a history. It contains two haplotypes, the common traunsteineri haplotype 1 and one individual with H7, a group II haplotype. In addition, individuals of D. maculata s.str. that grow at the same locality are dominated by haplotype 7 (Hedrén et al. unpublished). It is however difficult to separate between hybridization with D. maculata s.str. and hybridization with D. majalis ssp. sphagnicola since they both carry group II haplotypes. The same conclusions about introgression could be drawn for the two traunsteineri populations 48 and 59 which carry a few individuals with typical sphagnicola haplotypes. Previous studies on Dactylorhiza have also reported introgression of allotetraploids by their parental lineages (Hedrén 2003; Aagaard et al. 2005; Shipunov et al. 2005) as well as hybridization between allotetraploids (Heslop-Harrison 1954; Hedrén 2003).
Taxonomy
We conclude that the pattern of plastid haplotype composition found in this study does not support the subdivision of the core complex into three taxa. This conclusion is strongly supported by previous (Andersson 1996) and unpublished (Pedersen unpublished) findings on variation in morphology of the core complex as well as earlier genetic studies of the taxa using allozymes, AFLPs and chloroplast data (Hedrén 1996; Hedrén et al. 2001; Hedrén 2003; Pillon et al. 2007). Therefore we suggest an amalgamation of the three taxa into one with the oldest name applied: D. majalis ssp. lapponica.
Core complex individuals with the two most common haplotypes (H1 and H4) sometimes grew at adjacent sites or even in mixed populations. When examining such populations in the field we found no obvious differences in morphology between the two haplotypes. However, our genetic data is in congruence with previous findings on differentiation in morphology (Andersson 1994; Hansson 1994) and allozymes (Hedrén 1996) between populations of D. majalis ssp. traunsteineri on Gotland and mainland Sweden. Our results are also congruent with the finding that D. majalis ssp. traunsteineri on Gotland and ssp. russowii in Estonia are closely similar in morphology (Andersson 1994). All individuals of the taxonomically ambiguous populations (belonging to either D. majalis ssp. traunsteineri or ssp. sphagnicola) from the area around Oslo (Norway) and Gästrikland and Hälsingland (Sweden), showed typical sphagnicola haplotypes. Therefore we suggest that they should be treated as this taxon. In general, all populations of D. majalis ssp. sphagnicola in this study are well separated from the rest of the allotetraploids (Fig. 2) which is in agreement with earlier studies (Hedrén 2003; Devos et al. 2006) and further strengthens the position of ssp. sphagnicola as a separate subspecies. In contrast, our data indicates no clear differentiation between D. majalis ssp. traunsteineri on the one hand and ssp. majalis, ssp. purpurella or ssp. praetermissa on the other. To confirm this finding additional analysis including more material of all the different tetraploid taxa is needed.
As the interpretation of plastid markers could be hampered by chloroplast capture and that the results only show genetic variation from one of the parents, nuclear markers would better describe the reticulate evolution of polyploid complexes. Consequently, newly developed nuclear microsatellites (Nordström and Hedrén 2007) might be useful for future studies of the problematic species delimitations in Dactylorhiza.
Geographic pattern
The distribution of the common haplotypes 1 and 4 in this study shows a clear geographic pattern (Fig. 4a). The most common haplotype (H1) is dominating the Swedish mainland, particularly the southern parts, while haplotype 4 is prevalent in Finland, on Gotland (Sweden) and in northern Scandinavia. The high frequencies of these two haplotypes may be explained by rapid expansion after the last glaciations of populations carrying these haplotypes (cf. Hewitt 1996). On the basis of the present distribution of the two haplotypes the immigration history for the Scandinavian populations can be described. Two immigration routes are seen, carrying haplotype 1 and 4, respectively: One southwestern route of haplotype 1 originating in continental Europe and entering the Swedish and Norwegian mainland probably via Denmark, and one path of haplotype 4 with a more eastern direction through Estonia and Finland to northern Sweden and Norway. This immigration scenario is similar to results from earlier studies of phylogeography in Scandinavia on both plants (Nordal and Jonsell 1998; Berglund and Westerbergh 2001; Malm and Prentice 2005) and animals (Jaarola and Tegelström 1995; Taberlet et al. 1995).
Interestingly, the distribution of haplotype 4 also reveals a connection between the Estonian islands Saaremaa and Hiiumaa, and Gotland (Sweden) (Fig. 4a). The same connection is revealed by the distribution of the rare haplotype 10 (Fig. 4b). We interpret this pattern such that Gotland, as well as Saaremaa and Hiiumaa are all colonized by the same refugial population located somewhere to the southeast of the Baltic Sea. The dispersal to Gotland over the Baltic Sea seems surprising since Gotland is located more than 100 km from the eastern shore. However, the small air-filled seeds of orchids may travel thousands of kilometers by wind (Jersáková and Malinová 2007) and supposedly also by epizoochory. Haplotype 4 is also found in D. majalis ssp. majalis from southernmost Sweden and it may be hypothesized that the haplotype has spread to the core complex on Gotland from these populations. However, the flowering times of D. majalis ssp. majalis and ssp. traunsteineri are different and together with the fact that no other population of ssp. traunsteineri on the Swedish mainland carries the haplotype, such a spreading route does not seem likely.
Recent studies on immigration history in Nordic plants have revealed eastern refugia during the last glaciations (e.g., Lagercrantz and Ryman 1990; Skrede et al. 2006; Ståhlberg 2007). Haplotype 4 may have such an origin, or alternatively it may have had refugia further south since it is present in Austrian populations. Haplotype 1 is ubiquitous in Europe including Russia (Ståhlberg 2007). A study at a larger geographical scale may have the potential to identify the refugial areas of the haplotypes found in this study.
Haplotypes 1 and 4 meet in northern Fennoscandia. Similar contact zones between colonizing genotypes are seen for other species with southeastern and northwestern immigration routes into Scandinavia (Berglund and Westerbergh 2001; van Rossum and Prentice 2004; Ståhlberg 2007) and could be explained by glacial history. The ice cover in northern Scandinavia melted slowly and was completely gone much later than in the adjacent areas (Berglund 2004). This led to an accumulation of immigrating species on both sides of the ice sheet and as the last remains of the ice melted populations of different origin came into contact in approximately the same area. Another meeting zone of haplotypes 1 and 4 is in Estonia, where both haplotypes occur in close vicinity together with several others. Perhaps the genetic variation in Estonia could be explained by its geographic location close to refugia.
To conclude, we did not find any support in plastid haplotypes for separating D. majalis ssp. traunsteineri, ssp. lapponica and ssp. russowii into three taxa. Considering also the high degree of morphological similarity and results from previous studies (Andersson 1994; Hedrén 1996; Hedrén et al. 2001; Hedrén 2003; Pillon et al. 2007) we suggest an amalgamation of the three taxa into D. majalis ssp. lapponica. Furthermore, this broadly defined subspecies was not separated from other allotetraploids in the study area why additional studies should be made on a larger scale and with additional taxa. Newly developed nuclear microsatellites (Nordström and Hedrén 2007) may be used to bring clarity to the relationships of the genus. The genetic variation in the core complex had a clear geographic component and we have identified a minimum of two immigration routes into Scandinavia after the last glaciation.
References
Aagaard SMD, Sastad SM, Greilhuber J, Moen AA (2005) Secondary hybrid zone between diploid Dactylorhiza incarnata ssp cruenta and allotetraploid D. lapponica (Orchidaceae). Heredity 94:488–496
Adams JM (1997) Global land environments since the last interglacial. Oak Ridge National Laboratory, TN. http://www.esd.ornl.gov/ern/qen/nerc.html
Adams JM, Faure H (eds) (1997) QEN members. Review and atlas of palaeovegetation: preliminary land ecosystem maps of the world since the last glacial maximum. Oak Ridge National Laboratory, TN. http://www.esd.ornl.gov/ern/qen/adams1.html
Andersson E (1994) On the identity of orchid populations: a morphometric study of the Dactylorhiza traunsteineri complex in eastern Sweden. Nord J Bot 14:269–275
Andersson E (1995) Age-related morphological differentiation among populations of Dactylorhiza traunsteineri (Orchidaceae) in eastern Sweden. Nord J Bot 15:127–137
Andersson E (1996) Morphological variation in the orchid Dactylorhiza traunsteineri. Scale and systematic implications. Doctoral dissertation, Uppsala University, Uppsala
Bandelt H-J, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Molec Biol Evol 16:37–48. http://www.fluxus-engineering.com
Baumann H, Künkele S (1988) Die Orchideen Europas. Kosmos Naturführer, Stuttgart
Baumann H, Künkele S, Lorenz R (2006) Orchideen Europas. Mit angrenzenden Gebieten. Ulmer Naturführer, Stuttgart
Berglund M (2004) Holocene shore displacement and chronology in Ångermanland, eastern Sweden, the Scandinavian glacio-isostatic uplift centre. Boreas 33:48–60
Berglund AN, Westerbergh A (2001) Two postglacial immigration lineages of the polyploid Cerastium alpinum (Caryophyllaceae). Hereditas 134:171–183
Bjurulf P (2005) Morphological variation in Dactylorhiza majalis ssp. sphagnicola and ssp. traunsteineri in different habitats. Svensk Bot Tidskr 99:124–132
Borgen L, Hultgård UM (2003) Parnassia palustris: a genetically diverse species in Scandinavia. Bot J Linn Soc 142:347–372
Brochmann C, Brysting AK, Alsos IG, Borgen L, Grundt HH, Scheen A-C, Elven R (2004) Polyploidy in arctic plants. Biol J Linn Soc 82:521–536
Cafasso D, Widmer A, Cozzolino S (2005) Chloroplast DNA inheritance in the orchid Anacamptis palustris using single-seed polymerase chain reaction. Heredity 96(1):66–70
Chase MW, Hills HG (1991) Silica gel: an ideal material for field preservation of leaf samples for DNA studies. Taxon 40:215–220
Corriveau JL, Coleman AW (1988) Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 angiosperm species. Amer J Bot 75(10):1443–1458
Cozzolino S, Cafasso D, Pellegrino G, Musacchio A, Widmer A (2003) Fine-scale phylogeographical analysis of Mediterranean Anacamptis palustris (Orchidaceae) populations based on chloroplast minisatellite and microsatellite variation. Molec Ecol 12:2783–2792
Delforge P (1995) Orchids of Britain and Europe. Harper Collins Publishers, London
Delforge P (2001) Guide des Orchidées d′Europe, d′Afrique du Nord et du Proche-Orient. Delachaux et Niestlé, Lausanne
Demesure B, Sodzi N, Petit RJ (1995) A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Molec Ecol 4:129–131
de Soó R (1980) Dactylorhiza. In: Tutin TG et al. (eds) Flora Europea 5. Cambridge University Press, Cambridge, pp 333–337
Devos N, Tyteca D, Raspé O, Wesselingh RA, Jacquemart A-L (2003) Patterns of chloroplast diversity among western European Dactylorhiza species (Orchidaceae). Pl Syst Evol 243:85–97
Devos N, Raspé O, Oh S-H, Tyteca D, Jacquemart A-L (2006) The evolution of Dactylorhiza (Orchidaceae) allotetraploid complex: insights from nrDNA sequences and cpDNA PCR-RFLP data. Molec Phylogenet Evol 38:767–778
Doyle JJ, Doyle JH (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Dressler RL (1993) Phylogeny and classification of the orchid family. Cambridge University Press, Cambridge
Ekman S (1985) Dactylorhiza sphagnicola and D. traunsteineri with unspotted leaves found in E Uppland. Svensk Bot Tidskr 79:85–91
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online 1:47–50
Ferris C, King RA, Väinölä R, Hewitt GM (1998) Chloroplast DNA recognizes three refugial sources of European oaks and suggests independent eastern and western immigrations to Finland. Heredity 80:584–593
Grant V (1981) Plant speciation. Colombia University press, New York
Hansson S (1994) A study of Dactylorhiza lapponica. Svensk Bot Tidskr 88:17–28
Hedrén M (1996) Genetic differentiation, polyploidization and hybridization in northern European Dactylorhiza (Orchidaceae): evidence from allozyme markers. Pl Syst Evol 201:31–55
Hedrén M (2003) Plastid DNA variation in the Dactylorhiza incarnata/maculata polyploid complex and the origin of allotetraploid D. sphagnicola (Orchidaceae). Molec Ecol 12:2669–2680
Hedrén M, Fay MF, Chase MW (2001) Amplified fragment length polymorphisms (AFLP) reveal details of polyploid evolution in Dactylorhiza (Orchidaceae). Amer J Bot 88:1868–1880
Hedrén M, Nordström S, Hovmalm HAP, Pedersen HÆ, Hansson S (2007) Patterns of polyploid evolution in Greek marsh orchids (Dactylorhiza; Orchidaceae) as revealed by allozymes, AFLPs, and plastid DNA data. Amer J Bot 94:1205–1218
Heslop-Harrison J (1954) A synopsis of the dactylorchids of the British Isles. Berichte des Geobotanischen Forschungsinstituts Rübel 1953:53–82
Hewitt G (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hultén E (1950) Atlas of the distribution of vascular plants in NW Europe. Generalstabens litografiska anstalts förlag, Stockholm
Hylander N (1966) Nordisk kärlväxtflora II. Almquist och Wiksell, Stockholm
Jaarola M, Tegelström H (1995) Colonization history of north European field voles (Microtus agrestis) revealed by mitochondrial DNA. Molec Ecol 4:299–310
Jersáková J, Malinová T (2007) Spatial aspects of seed dispersal and seedling recruitment in orchids. New Phytol 176:448–459
Jordan WC, Courtney MW, Neigel JE (1996) Low levels of intraspecific genetic variation at a rapidly evolving chloroplast DNA locus in North American duckweeds (Lemnaceae). Amer J Bot 83:430–439
King RA, Ferris C (1998) Chloroplast DNA phylogeography of Alnus glutinosa (L) Gaertn. Molec Ecol 7:1151–1162
Krok ThOBN, Almquist S (1994) Svensk flora: Fanerogamer och ormbunksväxter. Liber, Stockholm
Kruskal JB (1964a) Multidimensional scaling by optimizing goodness of fit to a nonmetric hypothesis. Psychometrika 29:1–27
Kruskal JB (1964b) Nonmetric multidimensional scaling: a numerical method. Psychometrika 29:28–42
Kullman L (1996) Norway spruce present in the Scandes mountains, Sweden at 8000 BP: new light on holocene tree spread. Glob Ecol Biogeogr lett 5:94–101
Lagercrantz U, Ryman N (1990) Genetic structure of Norway spruce (Picea abies): concordance of morphological and allozymic variation. Evolution 44:38–53
Löve A, Löve D (1974) Origin and the evolution of the arctic and alpine floras. In: Ives JD, Barry RG (eds) Arctic and alpine environments. Methuen, London, pp 571–603
Lowe A, Harris S, Ashton P (2004) Ecological genetics—design, analysis and application. Blackwell Publishing, Oxford
Lumaret R, Barrientos E (1990) Phylogenetic relationships and gene flow between sympatric diploid and tetraploid plants of Dactylis glomerata (Gramineae). Pl Syst Evol 169:81–96
Luttikhuizen PC, Stift M, Kuperus P, van Tienderen PH (2007) Genetic diversity in diploid vs. tetraploid Rorippa amphibia (Brassicaceae). Molec Ecol 16:3544–3553
Malm U, Prentice HC (2002) Immigration history and gene dispersal: allozyme variation in Nordic populations of the red campion, Silene dioica (Caryophyllaceae). Biol J Linn Soc 77:23–34
Malm U, Prentice HC (2005) Chloroplast DNA haplotypes in Nordic Silene dioica: postglacial immigration from the east and the south. Pl Syst Evol 250:27–38
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mossberg B, Nilsson S (1987) Orkidéer- Europas vildväxande arter. Wahlström & Widstrand, Stockholm
Mossberg B, Stenberg L (2003) Den nya nordiska floran. Wahlström & Widstrand, Stockholm
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Nordal I, Jonsell B (1998) A phylogeographic analysis of Viola rupestris: three post-glacial immigration routes into the Nordic area? Bot J Linn Soc 128:105–122
Nordström S, Hedrén M (2007) Development of polymorphic nuclear microsatellite markers for polyploid and diploid members of the orchid genus Dactylorhiza. Molec Ecol Notes 7:644–647
Otto SP, Whitton J (2000) Polyploid incidence and evolution. Ann Rev Genet 34:401–437
Palmé AE, Su Q, Rautenberg A, Manni F, Lascoux M (2003) Postglacial recolonization and cpDNA variation of silver birch, Betula pendula. Molec Ecol 12:201–212
Pedersen HÆ, Hedrén M, Bateman RM (2003) Proposal to conserve the name Orchis majalis against O. elata, O. vestita and O. sesquipedalis (Dactylorhiza: Orchidinae: Orchidaceae). Taxon 52:633–634
Pillon Y, Fay MF, Hedrén M, Bateman RM, Devey DS, Shipunov AB, van der Bank M, Chase MW (2007) Evolution and temporal diversification of western European polyploid species complexes in Dactylohiza (Orchidaceae). Taxon 56:1185–1208
Pridgeon AM, Cribb PJ, Chase MW, Rasmussen FN (2001) Genera orchidacearum, 2: 1. Oxford University Press, Oxford
Rendell S, Ennos RA (2002) Chloroplast DNA diversity in Calluna vulgaris (heather) populations in Europe. Molec Ecol 11:69–78
Rohlf FJ (2005) NTSYSpc: numerical taxonomy system, ver. 2.20. Exeter Publishing, Ltd., Setauket
van Rossum F, Prentice HC (2004) Structure of allozyme variation in Nordic Silene nutans (Caryophyllaceae): population size, geographical position and immigration history. Biol J Linn Soc 81:357–371
Senghas K (1968) Taxonomische Übersicht der gattung Dactylorhiza Necker ex Nevski. In: Senghas K, Sundermann H (eds) Probleme der Orchideengattung Dactylorhiza. Jahresber Naturwiss Vereins Wuppertal 21–22:32–67
Shipunov AB, Fay MF, Chase MW (2005) Evolution of Dactylorhiza baltica (Orchidaceae) in European Russia: evidence from molecular markers and morphology. Bot J Linn Soc 147:257–274
Skrede I, Eidesen PB, Portela RP, Brochmann C (2006) Refugia, differentiation and postglacial migration in arctic-alpine Eurasia, exemplified by the mountain avens (Dryas octopetala L.). Molec Ecol 15:1827–1840
Soliva M, Widmer A (1999) Genetic and floral divergence among sympatric populations of Gymnadenia conopsea s.l. (Orchidaceae) with different flowering phenology. Int J Pl Sci 160:897–905
Soltis DE, Rieseberg LH (1986) Autopolyploidy in Tolmiea-menziesii (Saxifragaceae): Genetic insights from enzyme electrophoresis. Amer J Bot 73:310–318
Ståhlberg D (2007) Systematics, phylogeography and polyploid evolution in the Dactylorhiza maculata complex (Orchidaceae). Doctoral dissertation, Lund University, Lund
Stewart JR, Lister AM (2001) Cryptic northern refugia and the origins of the modern biota. Trends Ecol Evol 16:608–613
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Pl Molec Biol 17:1105–1109
Taberlet P, Swenson JE, Sandegren F, Bjärvall A (1995) Localization of a contact zone between two higly divergent mitochondrial DNA lineages of the brown bear (Ursus arctos) in Scandinavia. Conserv Biol 9:1255–1261
Tyler T, Prentice HC, Widén B (2002) Geographic variation and dispersal history in Fennoscandian populations of two forest herbs. Pl Syst Evol 233:47–64
Wolfe AD, Randle CP (2004) Recombination, heteroplasmy, haplotype polymorphism, and paralogy in plastid genes: implications for plant molecular systematics. Syst Bot 29:1011–1020
Acknowledgments
We would like to thank Sunniva Aagaard, Åse Bøilestad Breivik, Stefan Ericsson, Sven Hansson, Ursula Malm, Lennart Nordström, Tarmo Pikner, Mikko Piirainen, Mari Reitalu, David Ståhlberg, Taavo Tuulik, Kai Vahtra and Finn Wischmann for field assistance and/or collecting material. We also thank two anonymous reviewers for valuable comments. The study was supported by grants from Lunds Botaniska Förening to SN and from the Crafoord Foundation and the Swedish research council for environment, agricultural sciences and spatial planning, FORMAS (grant 2002-102) to MH.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Table 3
Rights and permissions
About this article
Cite this article
Nordström, S., Hedrén, M. Genetic differentiation and postglacial migration of the Dactylorhiza majalis ssp. traunsteineri/lapponica complex into Fennoscandia. Plant Syst Evol 276, 73–87 (2008). https://doi.org/10.1007/s00606-008-0084-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-008-0084-1