Abstract
MicroRNAs (miRNAs), found in blood and body fluids, have emerged as potential non-invasive biomarkers for disease and injury. miRNAs are quantitatively evaluated using typical RNA analysis methods such as the quantitative reverse transcription polymerase chain reaction, microarrays, and Northern blot, all of which require complex procedures and expensive reagents. To utilize miRNAs as practical biomarkers, it will be helpful to develop simple and user-friendly sensors. In this study, a paper-based miRNA sensor was developed by combining two methods: (1) target-recycled DNAzyme (Dz) amplification and (2) graphene oxide-assisted Dz blotting on paper. The Dz spots on paper caused a miRNA-dependent color change in presence of colorimetric reagents and facilitated the quantification of absolute amount of the target miRNA, irrespective of the volume, with high reproducibility. This approach is technologically straightforward and enables quantification of as low as 7.75 fmol miRNA using a portable smartphone.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs), a type of small non-coding RNAs, are approximately 21–25 nucleotides in length and play an important role in the post-transcriptional regulation of gene expression [1,2,3,4,5]. For decades, many researchers have reported that abnormal miRNA expression levels are related to the occurrence and development of cancers [6, 7], viral infections [8], and organ or tissue injury [9]. Interestingly, miRNAs in serum and other body fluids also show distinctive expression patterns in response to several serious diseases, such as cancer and organ injury [10, 11]. This suggests that miRNAs are potential non-invasive biomarkers. For example, miRNA-122 (miR-122), one of the well-studied miRNA, is the most abundant and specific miRNA expressed in the liver, constituting 70% of the total hepatic miRNA pool [9]. When injured, hepatocytes release miR-122 into the bloodstream, and its level strongly correlates with the traditional serum markers ALT and AST [12]. Thus, miR-122 is a promising indicator of liver function.
In general, miRNAs have been quantitatively evaluated using RNA-based methods, such as the reverse transcription polymerase chain reaction [13], microarrays [9], and Northern blot [14], all of which require complicated procedures and expensive reagents. To utilize miRNAs as non-invasive biomarkers, it is necessary to develop simple and user-friendly sensors. Recently, various technical and scientific attempts have been made to adopt nanotechnology for establishing facile and simple detection methods for next-generation miRNA sensing [15, 16]. The large surface area and unique properties of nanomaterials make them suitable to be designed and operated at the molecular level, thereby enhancing their sensing performances and broadening their practical applicability. To date, several miRNA sensing platforms have been reported based on diverse techniques, such as optical [17, 18], surface plasmon resonance [19], and electrochemical [20] approaches. Among them, colorimetric methods are considered the most suitable for personal diagnostics, owing to their simplicity and low cost [21, 22].
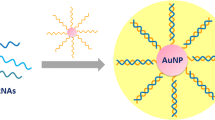
Graphene oxide (GO), an oxidized graphene sheet, is a water-dispersible derivative that has been employed in electronics [23], environmental [24], energy storage [25], and biomedical [26] applications. GO has a large surface area and many unique properties, including superior mechanical, electronic, and thermal properties [27]. GO possesses sp3 domains with oxygen-containing groups, as well as the sp2 domains of the original graphene structure [28]. It is suitable for biomedical applications typically performed in aqueous conditions. Significantly, GO preferentially adsorbs single-stranded DNA (ssDNA) over double-stranded DNA via pi–pi interactions and hydrogen bonding, enabling the development of DNA probe-based biosensors [29]. Among diverse carbon nanoallotropes that display the properties mentioned above, GO offers the benefits of cost-effective larger-scale production, and more efficient ssDNA binding due to its lower curvature, thereby expanding the range of its practical uses [30,31,32]. In our previous work, we utilized GO as a net (designated as GONET) for capturing ssDNA probes [33]. GONET effectively interacted with the ssDNA probes, facilitating the enrichment of the DNA probes by easily gathering micro-sized GONET complexes. Furthermore, we provided a proof of principle for its use in miRNA sensing systems by spinning down and concentrating the GONET complex in solution.
In this study, we established an advanced paper-based sensor for miRNAs by combining GONET and the target-recycled signal amplification strategy. This is the first study to report a GO-based paper sensor for quantitative miRNA sensing. This new amplification strategy provides a facile method for accessing the results with naked eye. The amplified colorimetric signal in solution was easily transferred on to a paper by GONET blotting, which could be quantified using a smartphone.
Experimental methods and materials
Materials
DNA strands were purchased from Genotech (Daejeon, Korea), and RNA strands were purchased from Bioneer (Daejeon, Korea). GO, hemin, ABTS, and H2O2 were purchased from Sigma-Aldrich (St. Louis, MO, USA). Whatman™ Grade 2 qualitative filter paper was purchased from GE Healthcare Life Sciences (Pittsburg, PA, USA). Capillary glass tubes were purchased from WPI (Sarasota, FL, USA). Temperature was regulated using a thermocycler (Bio-Rad, Hercules, CA, USA). Absorbance and fluorescence were measured using a Cytation 5 multimode reader (Biotek, Winooski, VT, USA). Images were captured using the built-in camera of an iPhone 8 smartphone, under a controlled light source from a fixed distance.
miR-122 recycled amplification of Dz
To prepare the Dz/lock probe, 2.5 μL of Dz strand (100 μM) was mixed with an equal amount of lock strand in 50 μL of 1 × PBS. The mixture was annealed by heating to 90 °C for 5 min, followed by slow cooling at room temperature for 1 h. For miRNA amplification, target (miR-122) was mixed with 1 μL of Dz/lock probe (5 μM) and 1 μL of fuel strand (5 μM) in 100 μL of the 1 × Dz buffer (20 mM Tris buffer (pH 7.2), 20 mM KCl, and 100 mM NaCl) for 30 min.
For developing the green color, the sensing solution was mixed with 1 μL of hemin (400 nM), 5 μL of ABTS (50 mg/mL), and 1 μL of H2O2 (25 mM). After 10-min incubation, the absorbance was measured at 420 nm.
GONET-assisted sensor in solution
One hundred microliters of the sensing solution was mixed with 2 μL of GO (1 mg/mL) and incubated for 10 min at room temperature, after which the mixture was centrifuged at 15,000 rpm for 30 min. After separating the supernatant, the pellet was re-dispersed in 100 μL of the 1 × Dz buffer. For developing the green color, each solution was mixed with 1 μL of hemin (400 nM), 5 μL of ABTS (50 mg/mL), and 1 μL of H2O2 (25 mM). After 10-min incubation, the absorbance was measured at 420 nm.
GONET-assisted paper sensor
One hundred microliters of the sensing solution was mixed with 2 μL GO (1 mg/mL) and incubated for 10 min at room temperature. Next, 15-μL mixture was aspirated using a glass capillary tube and applied to one side of the filter paper. Then, a mixture containing 2 μL hemin (160 μM), 5 μL ABTS (50 mg/mL), and 2 μL H2O2 (12.5 mM) was applied on the entire spot. The target-dependent green color developed within 2 min, the images were captured using the smartphone at the highest possible resolution, and each color signal was quantitated using ImageJ software.
Statistical analysis
Each experiment was performed in triplicate and the data are presented as mean ± the standard error of the mean (SEM). Differences were evaluated using Student’s t-tests. A p-value < 0.05 indicated a statistically significant result.
Results and discussion
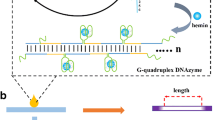
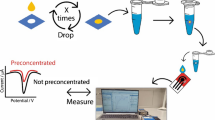
The paper-based miRNA sensor developed in this study was based on a two-step process comprising the following steps: (1) target-responsive DNAzyme (Dz) amplification and (2) GONET-assisted color development on paper (Fig. 1). Guanine (G)-quadruplex Dz is a functional DNA with stacked G-quartets that can bind hemin, thereby showing peroxidase-mimicking activity [34]. The G-rich Dz sequence offers several advantages over protein enzymes, such as thermal stability and ease of modification; thus, it has been widely employed to generate signals in colorimetric assays. In step 1, the Dz/lock duplex probe, in which the quadruplex folding of Dz is blocked by a lock strand, recognizes the target miRNA using the toehold region (indicated in red in Fig. 1a). Target binding initiates sequence-specific, toehold-mediated strand displacement (TMSD) [35] among the three strands, resulting in the formation of a new lock/miRNA duplex and free Dz. At this moment, the new toehold of the lock/miRNA (indicated in yellow in Fig. 1a) is recognized by the fuel strand, followed by TMSD-based target displacement. The free miRNA can participate in unlocking Dz again, thereby repeatedly producing free Dz, which results in a G-quadruplex structure with a long, single-stranded tail (Fig. 1a). In step 2, upon adding GO to the solution obtained from step 1, the amplified Dz strands are adsorbed onto the GO surface, followed by the formation of micro-sized Dz/GO composites [33]. Finally, the Dz/GO composite can be easily accumulated onto an intended spot on the paper, which amplifies Dz-catalyzed colorimetric reaction in the presence of colorimetric reagents, including hemin, 2,2-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) diammonium salt (ABTS), and hydrogen peroxide (H2O2) (Fig. 1b).
To confirm the target-recycled amplification based on TMSD, we compared the miRNA sensing performance with and without the fuel strands in a buffer solution containing colorimetric reagents. Additionally, miR-122 was chosen as the representative miRNA biomarker. The probes and fuel strands were prepared as detailed in Table S1. The toehold region on the lock strand was 7 nt long, which was sufficient for initiating TMSD [35]. As shown in Fig. 2, the basic assay without the fuel strands and the amplified assay with the fuel strands exhibited target-dependent color development. In the basic assay using a 50-nM duplex probe, the absorbance at 420 nm steadily increased up to 50 nM miRNA. The limit of detection (LOD) was evaluated to be 7.59 nM, according to the equation LOD = 3.3(SD/S), where SD is the standard deviation and S is the slope of the calibration curve. This value was similar to that obtained in a previous study [33]. By contrast, in the presence of the fuel strands, the colorimetric signal was highly amplified compared to that obtained using the basic method, which demonstrated a significantly lower LOD (0.2 nM). The amplified color changes were discernible to the naked eye, whereas those resulting from the basic method were not (Fig. 2b). The TMSD-mediated amplification was re-confirmed by performing polyacrylamide gel analysis (Fig. S1). Our results showed that in the presence of fuel strands, miRNA consumed the Dz/lock probe at a higher rate than in the absence of the fuel strands and produced a lock/fuel duplex. These results indicated that the well-designed fuel strands induce target-recycled Dz amplification, strengthen ABTS oxidation reaction, and shift the valid detectable range, thereby enabling color change discernible to the naked eye.
Signal-amplified colorimetric assay for miRNA in the concentration range of 0–50 nM. Target miRNA concentration-dependent (a) increase in absorbance at 420 nm and (b) colorimetric appearance of the sensing solutions with and without fuel strands. Absorbance spectra of the sensing solutions in the (c) absence and (d) presence of the fuel strands. Each sample solution was treated with hemin, ABTS, and H2O2 for color development. Data is presented as the mean ± SEM of triplicate samples (n = 3)
Prior to GONET application, we optimized the amount of GO using a fluorescein-labeled Dz strand (FAM-Dz) (Fig. 3a). We used 5-nM miRNA as the representative target concentration to establish sufficient difference between the basic and amplified assays. The interaction of GO with FAM-Dz in the sensing solution induced fluorescence quenching via fluorescence resonance energy transfer, which implies that Dz was captured by GO [36]. A sensing solution containing miR-122, FAM-Dz/lock probe, and the fuel strand was mixed with various amounts of GO. The fluorescence intensity of FAM-Dz gradually decreased with increasing GO and was quenched by up to 95% in presence of 2 µg GO (Fig. 3b). Moreover, when GO composites were separated by centrifugation, the fluorescence of the supernatant was similar to that of the total mixture. These data indicate that fluorescence quenching occurred by the actual adsorption of FAM-Dz onto GO, and the interaction was strong enough to separate the Dz strands from the solution. Therefore, we used 2 µg GO to collect the amplified Dz in the GONET application.
(a) Relative fluorescence intensities of FAM-labeled Dz/Lock (5 pmol) incubated with 0.5 pmol miRNA and 5 pmol fuel strand in total sensing mixtures or supernatant with various amounts of GO. (b) Fluorescence spectra of the total sensing mixtures with 0 or 2 μg GO. Data is presented as the mean ± SEM of triplicate samples (n = 3)
Based on these results, we applied the GONET strategy for signal-amplified miRNA sensing. In the case of the solution, the Dz/GO composite, separated by spinning down, showed dramatic increase in absorbance at 420 nm in the presence of the fuel strand, even with low levels of miRNA that could not be detected using the basic assay (Fig. 4a and Fig. S2). The supernatants, irrespective of the presence of fuel strands, exhibited insignificant increase in absorbance at 420 nm upon miRNA addition. These results indicate that GONET can successfully collect activated Dz and enrich the target-responsive signal in the amplified condition. Next, we applied the amplified GONET to the paper. Instead of spinning down in solution, the Dz/GO composite was directly spotted onto the paper using capillary glass tubes, generating a pale brown circle of ~ 2 mm diameter. In presence of colorimetric reagents, the amplified GONET assay led to color development visible to the naked eye, whereas the basic GONET assay did not (Fig. 4b). The color changes were quantitatively evaluated using the smartphone camera and ImageJ software. The relative values were calculated using the negative (GO only) and positive (pre-adsorbed Dz/GO) control groups (Fig. 4c). These paper-based results were consistent with the solution-based results of GONET, thereby suggesting that GONET enables the transfer of the amplified miRNA-responsive signal from solution to paper.
(a) Comparison of absorbance at 420 nm between the supernatant (Sup.) and pellet separated by centrifuging the miRNA sensing solution with or without fuel strand in the presence of GO. (b) Colorimetric appearance and (c) extracted color intensities of the paper onto which the GONET sensing solution was applied (scale bar: 2 mm). All sample solutions and spots were treated with hemin, ABTS, and H2O2 for color development. Data is presented as the mean ± SEM of triplicate samples (n = 3)
To support the blotting of GONET onto paper, we evaluated the paper-based miRNA sensing by adjusting the dilution factors and volume with a fixed miRNA amount. We prepared a series of miRNA sensing solutions of increasing volumes and decreasing concentrations that all contained 37.5 fmol of miR-122. When each solution was mixed with GO and blotted onto the paper, similar pale brown spots were observed at the intended position on the paper. With the common colorimetric reagent treatment applied, the spots turned light or dark green depending on the absence or presence of miR-122 (Fig. 5a). Upon digitizing each signal using ImageJ software, statistically significant signals were obtained in four sets of miRNA sensing solutions in the presence of miRNA (p < 0.01) (Fig. 5b). Moreover, similar color intensities were obtained regardless of the volume and concentration. This finding was confirmed by further analysis using a broad range of solutions of various concentrations and volumes that contained different miRNA varieties (Fig. S3). These data indicate that the GONET on paper can maintain the collected yield of Dz even with increased volumes and lower concentrations; thus, the colorimetric signal on paper originates from the absolute quantity of the target miRNA rather than the concentration.
(a) Colorimetric appearance and (b) extracted color intensities of each spot on paper obtained by applying a series of amplified GONET solutions prepared in various volumes and concentrations that equally contained 37.5 fmol of miR-122. Each spot was treated with hemin, ABTS, and H2O2 for color development. Data is presented as the mean ± SEM of triplicate samples (n = 3). The p-values were calculated using the Student’s t-test (**p < 0.01, ***p < 0.001)
Finally, we quantified miRNA using our GONET paper sensor. The sensing solution was prepared in 15 µL of 1 × Dz buffered solution containing the Dz/lock probe, a fuel strand, and miR-122 ranging from 0 to 150 fmol. The color steadily changed from pale brown to green, which was easily digitized using the smartphone camera and ImageJ (Fig. 6 and Fig. S4). Based on the data set, the LOD was 7.75 fmol, as determined using the equation LOD = 3.3(SD/S). This value was converted to 0.52 nM under the corresponding experimental conditions, which could be lowered when the sample volume was greater than 15 μL. This sensing performance was comparable to that of previous GO-based colorimetric sensors (Table S2). Therefore, the amplified GONET paper sensor can quantify sub-nanomolar amounts of miRNA without the need for sophisticated instruments.
(a) Colorimetric appearance and (b) extracted color intensities of spots obtained by applying an amplified GONET solution containing various concentrations of miRNA, ranging from 0 to 150 fmol in 15-μL volume. Each spot was treated with hemin, ABTS, and H2O2 for color development. Data is presented as the mean ± SEM of triplicate samples (n = 3)
CONCLUSIONS
In this study, a sensitive, colorimetry-based paper sensor was developed for detecting miRNAs. This sensor utilized miRNA-recycled amplification and GONET-based paper blotting to achieve its superior performance. The target recycling exhibited significantly higher sensitivity with a 38-fold improvement in the LOD. The GONET, as a blotting agent, transferred the amplified miRNA-responsive signal from solution to paper, which was sufficient to be quantified using a smartphone, and showed high reproducibility. The amplified GONET paper sensor relies on the determination of the absolute amount of target miRNA rather than its concentration. This permits experimental flexibility in the parameters of the present sensor, such as variable detection-spot size and adjustable sample volume, according to the sample condition or sensing purpose. This system is versatile and can be broadly applied to other genes and proteins simply by changing the target binding sequence, as well as other measuring techniques based on signal generator elements other than peroxidase mimicking Dz. This amplified GONET sensor is technologically straightforward and satisfies the requirements of point of care testing, including simplicity, low cost, and ease of use. This is the first step toward GO application in the form of a paper-based sensor for quantitative miRNA sensing; however, several limitations remained to be overcome for real-world application. The sensing specificity and sensitivity of this system may be influenced by co-existing biomolecules in biological samples. To solve this limitation, we plan to introduce chemically modified GO and artificial probes in our future studies. We anticipate that this novel strategy will be a valuable resource for miRNA-based disease diagnosis and applied research in nano-biotechnology.
Supplementary information
The Supporting Information is available online.
Sequences of probes and miRNA (Table S1); PAGE analysis of TMSD-assisted target recycling (Fig. S1); absorbance spectra in the absence and presence of fuel strand in miRNA sensing solution (Fig. S2); colorimetric appearance of the paper-based sensor (Figs. S3 and S4); GO-based colorimetric sensors for nucleic acid detection (Table S2).
Data availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
Abbreviations
- ABTS :
-
2,2-Azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) diammonium salt
- Dz :
-
DNAzyme
- FAM-Dz :
-
Fluorescein-labeled Dz strand
- FRET :
-
Fluorescence resonance energy transfer
- G :
-
Guanine
- GO :
-
Graphene oxide
- LOD :
-
Limit of detection
- miR-122 :
-
MiRNA-122
- miRNA :
-
MicroRNA
- S :
-
Slope
- SEM :
-
Standard error of the mean
- SD :
-
Standard deviation
- ssDNA :
-
Single-stranded DNA
- Sup :
-
Supernatant
- TMSD :
-
Toehold-mediated strand displacement
References
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114. https://doi.org/10.1038/nrg2290
Hwang HW, Mendell JT (2006) MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 94:776–780. https://doi.org/10.1038/sj.bjc.6603023
Jovanovic M, Hengartner MO (2006) miRNAs and apoptosis: RNAs to die for. Oncogene 25:6176–6187. https://doi.org/10.1038/sj.onc.1209912
Inui M, Martello G, Piccolo S (2010) MicroRNA control of signal transduction. Nat Rev Mol Cell Biol 11:252–263. https://doi.org/10.1038/nrm2868
Sayed D, Abdellatif M (2011) MicroRNAs in development and disease. Physiol Rev 91:827–887. https://doi.org/10.1152/physrev.00006.2010
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med 20:460–469. https://doi.org/10.1016/j.molmed.2014.06.005
Reddy KB (2015) MicroRNA (miRNA) in cancer. Cancer Cell Int 15:38. https://doi.org/10.1186/s12935-015-0185-1
Skalsky RL, Cullen BR (2010) Viruses, microRNAs, and host interactions. Annu Rev Microbiol 64:123–141. https://doi.org/10.1146/annurev.micro.112408.134243
Jopling C (2012) Liver-specific microRNA-122: biogenesis and function. RNA Biol 9:137–142. https://doi.org/10.4161/rna.18827
Kosaka N, Iguchi H, Ochiya T (2010) Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci 101:2087–2092. https://doi.org/10.1111/j.1349-7006.2010.01650.x
Bala S, Petrasek J, Mundkur S, Catalano D, Levin I, Ward J, Alao H, Kodys K, Szabo G (2012) Circulating microRNAs in exosomes indicate hepatocyte injury and inflammation in alcoholic, drug-induced, and inflammatory liver diseases. Hepatology 56:1946–1957. https://doi.org/10.1002/hep.25873
Sharapova T, Devanarayan V, LeRoy B, Liguori MJ, Blomme E, Buck W, Maher J (2016) Evaluation of miR-122 as a serum biomarker for hepatotoxicity in investigative rat toxicology studies. Vet Pathol 53:211–221. https://doi.org/10.1177/0300985815591076
Schmittgen TD, Lee EJ, Jiang J, Sarkar A, Yang L, Elton TS, Chen C (2008) Real-time PCR quantification of precursor and mature microRNA. Methods 44:31–38. https://doi.org/10.1016/j.ymeth.2007.09.006
Várallyay E, Burgyán J, Havelda Z (2008) MicroRNA detection by Northern blotting using locked nucleic acid probes. Nat Protoc 3:190–196. https://doi.org/10.1038/nprot.2007.528
Degliangeli F, Pompa PP, Fiammengo R (2014) Nanotechnology-based strategies for the detection and quantification of microRNA. Chemistry 20:9476–9492. https://doi.org/10.1002/chem.201402649
Chandrasekaran AR, Punnoose JA, Zhou L, Dey P, Dey BK, Halvorsen K (2019) DNA Nanotechnology approaches for microRNA detection and diagnosis. Nucleic Acids Res 47:10489–10505. https://doi.org/10.1093/nar/gkz580
Yang L, Liu C, Ren W, Li Z (2012) Graphene surface-anchored fluorescence sensor for sensitive detection of microRNA coupled with enzyme-free signal amplification of hybridization chain reaction. ACS Appl Mater Interfaces 4:6450–6453. https://doi.org/10.1021/am302268t
Hakimian F, Ghourchian H, Hashemi AS, Arastoo MR, Behnam Rad M (2018) Ultrasensitive optical biosensor for detection of miRNA-155 using positively charged Au nanoparticles. Sci Rep 8:2943. https://doi.org/10.1038/s41598-018-20229-z
Jebelli A, Oroojalian F, Fathi F, Mokhtarzadeh A, Guardia M (2020) Recent advances in surface plasmon resonance biosensors for microRNAs detection. Biosens Bioelectron 169:112599. https://doi.org/10.1016/j.bios.2020.112599
Shabaninejad Z, Yousefi F, Movahedpour A, Ghasemi Y, Dokanehiifard S, Rezaei S, Aryan R, Savardashtaki A, Mirzaei H (2019) Electrochemical-based biosensors for microRNA detection: nanotechnology comes into view. Anal Biochem 581:113349. https://doi.org/10.1016/j.ab.2019.113349
Zhou X, Cao P, Zhu Y, Lu W, Gu N, Mao C (2015) Phage-mediated counting by the naked eye of miRNA molecules at attomolar concentrations in a Petri dish. Nat Mater 14:1058–1064. https://doi.org/10.1038/nmat4377
Persano S, Guevara ML, Wolfram J, Blanco E, Shen H, Ferrari M, Pompa PP (2016) Label-free isothermal amplification assay for specific and highly sensitive colorimetric miRNA detection. ACS Omega 1:448–455. https://doi.org/10.1021/acsomega.6b00109
Mundinamani S (2020) Large area, multilayer graphene films as a flexible electronic material. ACS Omega 5:17479–17485. https://doi.org/10.1021/acsomega.0c01982
Perreault F, Fonseca de Faria A, Elimelech M (2015) Environmental applications of graphene-based nanomaterials. Chem Soc Rev 44:5861–5896. https://doi.org/10.1039/c5cs00021a
Li X, Zhi L (2018) Graphene hybridization for energy storage applications. Chem Soc Rev 47:3189–3216. https://doi.org/10.1039/c7cs00871f
Reina G, González-Domínguez JM, Criado A, Vázquez E, Bianco A, Prato M (2017) Promises, facts and challenges for graphene in biomedical applications. Chem Soc Rev 4400(46):4400–4416. https://doi.org/10.1039/c7cs00363c
Chen D, Feng H, Li J (2012) Graphene oxide: preparation, functionalization, and electrochemical applications. Chem Rev 112:6027–6053. https://doi.org/10.1021/cr300115g
Mouhat F, Coudert FX, Bocquet ML (2020) Structure and chemistry of graphene oxide in liquid water from first principles. Nat Commun 11:1566. https://doi.org/10.1038/s41467-020-15381-y
Paul T, Bera SC, Agnihotri N, Mishra PP (2016) Single-molecule FRET studies of the hybridization mechanism during noncovalent adsorption and desorption of DNA on graphene oxide. J Phys Chem B 120:11628–11636. https://doi.org/10.1021/acs.jpcb.6b06017
Li F, Pei H, Wang L, Lu J, Gao J, Jiang B, Zhao X, Fan C (2013) Nanomaterial-based fluorescent DNA analysis: a comparative study of the quenching effects of graphene oxide, carbon nanotubes, and gold nanoparticles. Adv Func Mat 23:4140. https://doi.org/10.1002/adfm.201203816
Georgakilas V, Perman JA, Tucek J, Zboril R (2015) Broad family of carbon nanoallotropes: classification, chemistry, and applications of fullerenes, carbon dots, nanotubes, graphene, nanodiamonds, and combined superstructures. Chem Rev 115:4744. https://doi.org/10.1021/cr500304f
Hwang HS, Jeong JW, Kim YA, Chang M (2020) Carbon nanomaterials as versatile platforms for biosensing applications. Micromachines 11:814. https://doi.org/10.3390/mi11090814
Lee J, Kim Y-k, Lee S, Yoon S, Kim W-k (2019) Graphene oxide-based NET strategy for enhanced colorimetric sensing of miRNA. Sens Actuators B 282:861–867. https://doi.org/10.1016/j.snb.2018.11.149
Li W, Li Y, Liu Z, Lin B, Yi H, Xu F, Nie Z, Yao S (2016) Insight into G-quadruplex-hemin DNAzyme/RNAzyme: adjacent adenine as the intramolecular species for remarkable enhancement of enzymatic activity. Nucleic Acids Res 44:7373–7384. https://doi.org/10.1093/nar/gkw634
Zhang DY, Winfree E (2009) Control of DNA strand displacement kinetics using toehold exchange. J Am Chem Soc 131:17303–17314. https://doi.org/10.1021/ja906987s
Kim J, Cote LJ, Kim F, Huang J (2010) Visualizing graphene based sheets by fluorescence quenching microscopy. J Am Chem Soc 132:260–267. https://doi.org/10.1021/ja906730d
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2020R1F1A1072921) and the Korea Institute of Toxicology (Grant No. KK-2101–01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J., Na, HK., Lee, S. et al. Advanced graphene oxide-based paper sensor for colorimetric detection of miRNA. Microchim Acta 189, 35 (2022). https://doi.org/10.1007/s00604-021-05140-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-021-05140-1